SPRTN is a mammalian DNA-binding metalloprotease that resolves DNA-protein crosslinks
Figures
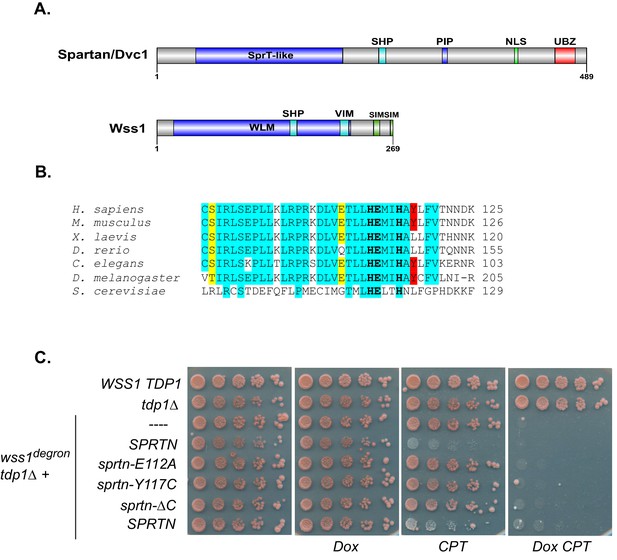
SPRTN fails to complement Wss1 activity in yeast.
(A) Comparison of SPRTN and Wss1 domain organization. (B) Comparison of SprT-like domains from selected SPRTN-containing organisms. Amino acid highlighted in red corresponds to the tyrosine mutated in patients with Ruijs-Aalfs syndrome (SPRTN-Y117C), identical amino acids are in blue and similar amino acids are in yellow. The catalytic glutamic acid and zinc-coordinating histidines are indicated by bold text. (C) Yeast spot assay. Five-fold dilutions of wildtype, tdp1∆, and wss1∆ tdp1∆ cells. wss1∆ tdp1∆ cells harboring empty plasmid (--- = pRS415) or plasmid encoding wild-type SPRTN or corresponding mutants under the endogenous Wss1 promoter. Cells were spotted on CSM-LEU plates with 2 ug/mL doxycycline or 2 ug/mL doxycycline and 40 uM CPT. Doxycycline is used to acutely deplete Wss1 protein levels in tdp1∆ cells to effectively obtain wss1∆ tdp1∆ cells.
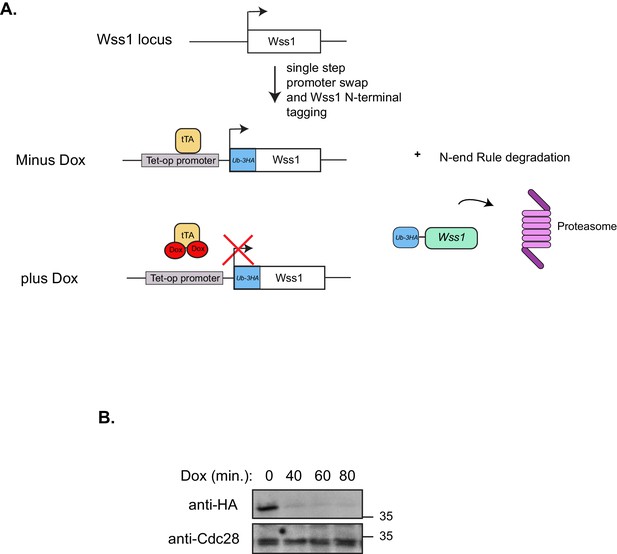
Dual Wss1 protein depletion system.
(A) Schematic illustration describing the dual protein depletion system used to acutely deplete Wss1 from tdp1∆ yeast strains to circumvent the accumulation of cells containing second site suppressors that give resistance to CPT. The endogenous Wss1 promoter and the N-terminal region of Wss1 was replaced, in a single step, with a selection cassette that includes a repressible heterologous promoter and ubiquitin-3HA epitope tag. (B) Western blot analysis confirming Wss1 depletion upon doxycycline [2 ug/mL] addition to yeast strain yKM10.
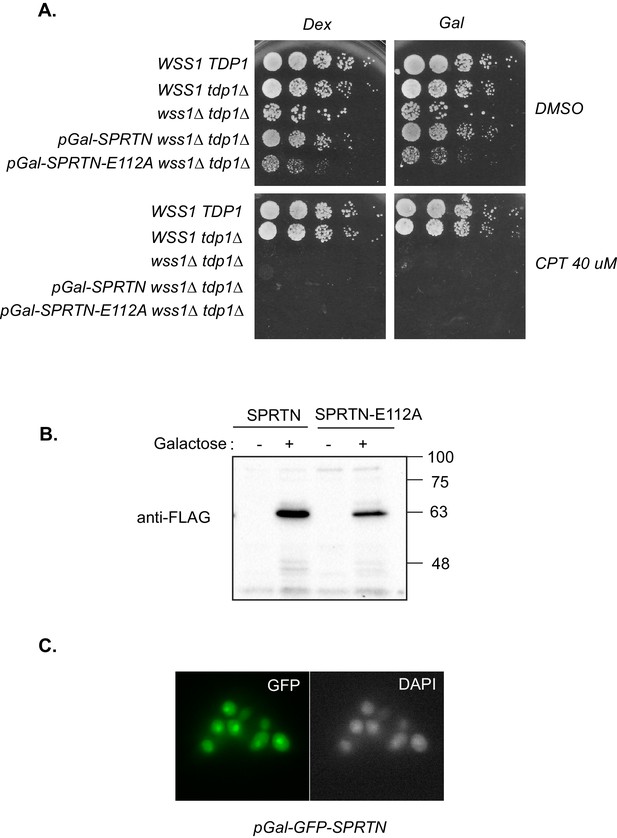
SPRTN complementation in yeast.
(A) Galactose inducible expression of FLAG-SPRTN or corresponding mutants integrated at the URA3 locus of wss1∆ tdp1∆ cells. Five-fold serial dilutions of yeast cells were spotted on rich plates containing either 2% glucose with or without 40 uM CPT or 2% galactose with our without 40 uM CPT. (B) Western blot analysis of FLAG-SPRTN and FLAG-SPRTN-E112A expression in yeast cells. (C) Epi-fluorescent microscopy of fixed cells expressing galactose inducible GFP-SPRTN. Cells were stained with DAPI to locate the nucleus.
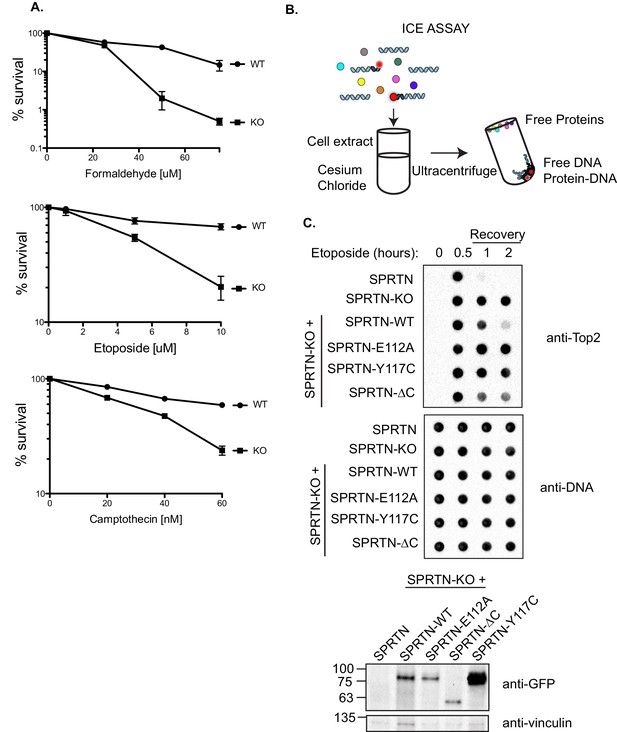
SPRTN is required for DPC removal.
(A) Clonogenic assay of control SPRTN, and SPRTN-KO MEFs. Control SPRTN MEFs were treated with 4-OHT for 24 hr to induce CRE-mediated excision of SPRTN. Cells were then treated with formaldehyde, Etoposide or CPT at the indicated drug concentrations for 24 hr (72 hr for Etoposide) and subsequently allowed to form colonies for 10 days. Cells were plated in triplicate. Error bars represent S.E.M. (B) In vivo complex of Enzyme (ICE) assay. Schematic illustration of the method used to isolate Etoposide-induced covalent Top2 DNA-protein crosslinks from cells. (C) ICE assay of control SPRTN MEFs, SPRTN-KO MEFs, and conditional SPRTN MEFs reconstituted with SPRTN, SPRTN-E112A, SPRTN-Y117C, or SPRTN-∆C. Cells were either untreated or treated for 30 min with 25 uM Etoposide and immediately processed for ICE assay or processed 1 and 2 hr after washing out Etoposide (recovery). 1 ug of DNA was loaded for each time-point per cell line. Membrane was probed with an antibody against Top2 and subsequently re-probed with anti-thymine dimer antibody after 1 min UV treatment to control for DNA loading. Representative Western blot indicating SPRTN levels after reconstitution of SPRTN-KO MEFs.
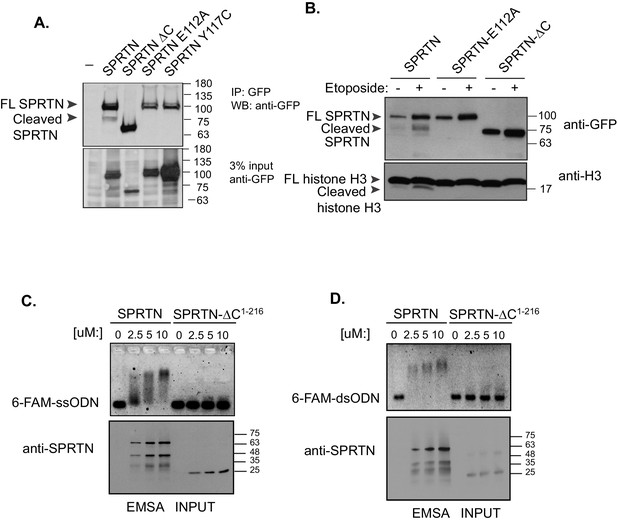
SPRTN is a DNA binding protease.
(A) Western blot analysis of GFP-SPRTN expression. GFP-SPRTN and corresponding mutants were immunoprecipitated after 16 hr of transient transfection in 293T-HEK cells. Arrows indicate full-length (FL) or cleaved fragments. (B) Western blot analysis of histone H3 cleavage in cells overexpressing GFP-SPRTN, GFP-SPRTN-E112A or GFP-SPRTN-∆C. (C) EMSA of full-length recombinant SPRTN or SPRTN-∆C (a.a. 1–216) and 0.25 uM 6-FAM-39-mer single stranded oligodeoxynucleotide (6-FAM-ssODN) or (D) 0.25 uM 6’FAM-39-mer double stranded oligonucleotide (6-FAM-dsODN) .
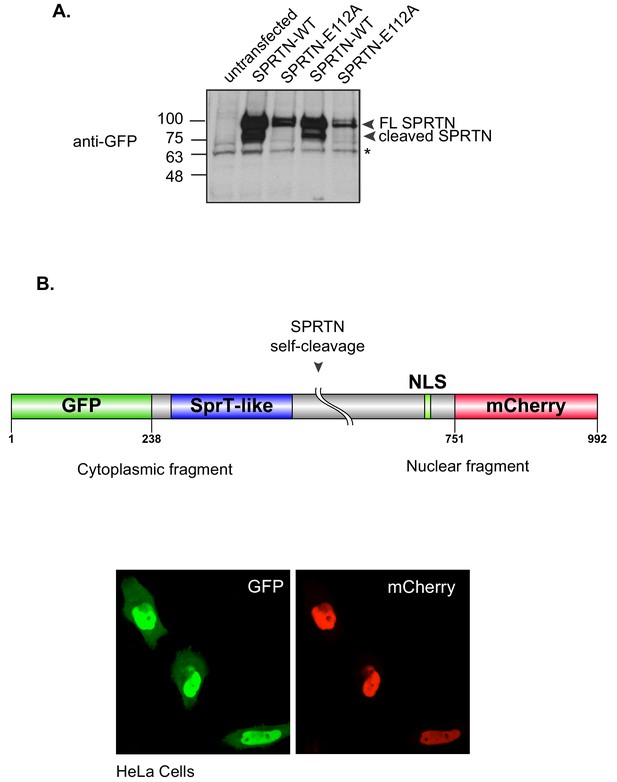
SPRTN self-cleaves in vivo.
(A) Western blot analysis of SPRTN self-cleavage in cells. 293T-HEK cells were transfected with GFP-SPRTN or GFP-SPRTN-E112A. Experiment was performed in duplicate. Cells were directly lysed in 1X Sample buffer. Asterisk denotes a non-specific cross-reacting band that serves as a loading control. (B) Schematic illustration describing expression system to detect SPRTN self-cleavage in live cells. HeLa cells were transfected with a dual tag GFP (N-terminus) and mCherry (C-terminus) SPRTN expression plasmid.
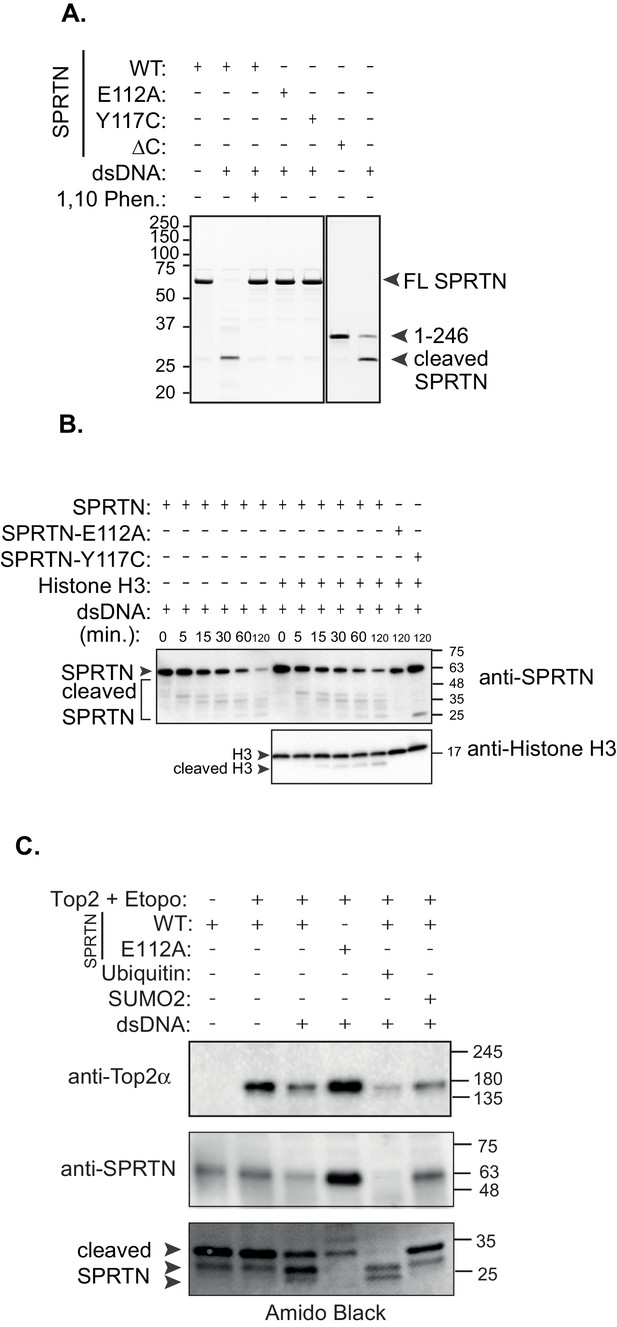
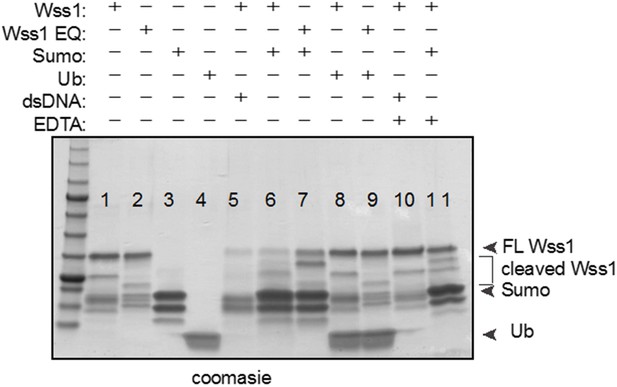
In vitro SPRTN substrate cleavage.
(A) In vitro SPRTN self-cleavage reactions. Purified proteins were incubated with or without DNA for 5 hr at 37°C. 1,10 Phe = 1,10 Phenanthroline, a zinc metal chelator and inhibitor of zinc metalloproteases. Proteins were separated by SDS-PAGE and stained with coommasie blue. (B) In vitro histone H3 cleavage. SPRTN was incubated with or without histone H3 (SPRTN:H3 molar ratio of 4:1) in the presence of dsDNA for the indicated time points. SPRTN-E112A or SPRTN-Y117C mutants were incubated for 2 hr. Proteins were separated on an SDS-PAGE gel and transferred to a membrane for Western blot analysis. Histone H3 cleavage as well as SPRTN self-cleavage were monitored by immunobloting with antibodies against histone H3 and SPRTN. (C) In vitro Top2 cleavage. Purified recombinant Top2 was pre-incubated with DNA and Etoposide to irreversibly bind Top2 to DNA. Recombinant SPRTN or SPRTN-E112A was then added alone or in combination with 10-fold molar excess of either ubiquitin or SUMO and incubated for 2 hr at 37°C. Proteins were separated by SDS-PAGE and transferred to a membrane for Western blot analysis. Membrane was stained with amido black to detect SPRTN cleavage fragments.
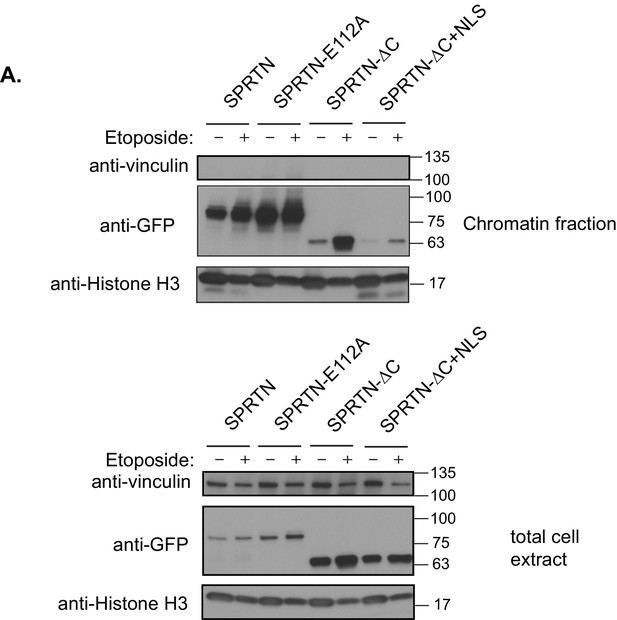
SPRTN recruitment to chromatin.
(A) Chromatin fractionation of 293T-HEK cells expressing GFP-SPRTN, GFP-SPRTN-E112A, GFP-SPRTN-∆C, or GFP-SPRTN-∆C+NLS in the absence or presence of Etoposide. Proteins were separated by SDS-PAGE and transferred to a membrane for Western blot analysis with an antibody against GFP, vinculin or histone H3.
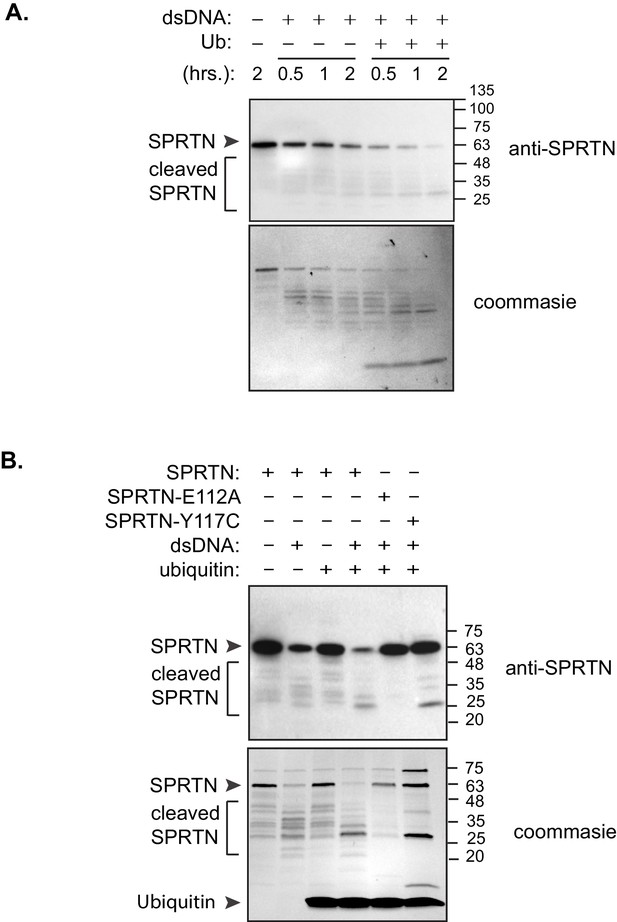
SPRTN activation by ubiquitin.
(A) Ubiquitin enhances SPRTN activity. SPRTN was incubated with histone H3 in the presence or absence of dsDNA and/or ubiquitin for the indicated time points. Proteins were separated by SDS-PAGE and transferred to a membrane for Western blot analysis. SPRTN self-cleavage was monitored by Western blot analysis. (B) Ubiquitin enhances SPRTN self-cleavage. Purified proteins were incubated with dsDNA and/or ubiquitin for 2 hr. Proteins were separated by SDS-PAGE and transferred to a membrane for Western blot analysis. SPRTN self-cleavage was monitored by Western blot analysis.
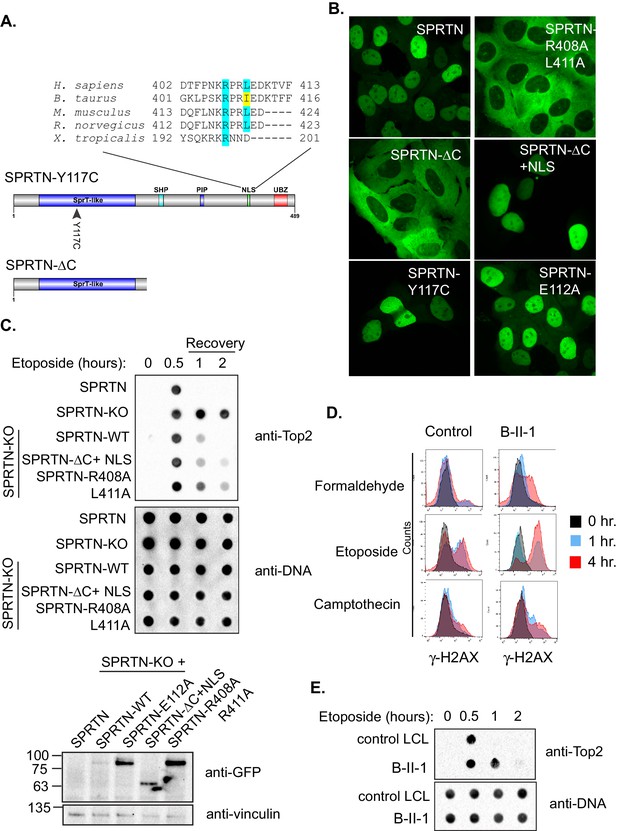
SPRTN mutant proteins from Ruijs-Aalfs syndrome patients are hypomorphic for DPC removal.
(A) Comparison of SPRTN-Y117C and SPRTN-∆C mutant proteins found in patients with Ruijs-Aalfs syndrome. The NLS sequence highlights the amino acid conservation in vertebrate SPRTN species. (B) Fluorescent expression of GFP-SPRTN and mutant proteins in live U2OS cells. (C) ICE assay of DNA isolated from SPRTN control, SPRTN-KO cells or SPRTN-KO cells reconstituted with SPRTN-WT, SPRTN-∆C+NLS, or SPRTN-R408A L411A. Cells were either untreated or treated for 30 min with 25 uM Etoposide and immediately subjected to ICE assay or 1 and 2 hr after washing out Etoposide (recovery), as in Figure 2C. Western blot indicates SPRTN levels after reconstitution of SPRTN-KO MEFS. (D) Representative flow cytometry analysis of control LCL cells and patient (B-II-1) LCL cells treated with formaldehyde, Etoposide or CPT for 0, 1 or 4 hr. Cells were collected at indicated time points and processed for γ-H2AX staining. Untreated controls are indicated in black for comparison to treated cells. 10,000 cells were scored for each experiment. (E) ICE assay of DNA isolated from control or B-II-1 cells that were either untreated or treated with 25 uM Etoposide for the indicated times as in Figure 2C and 5C.
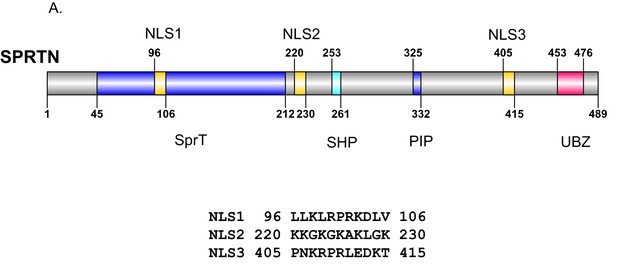
SPRTN nuclear localization signals.
(A) Schematic illustration of SPRTN indicating the position and amino acid sequence of candidate nuclear localization signals identified by bioinformatic analysis. NLS 3 was chosen for further experimentation.
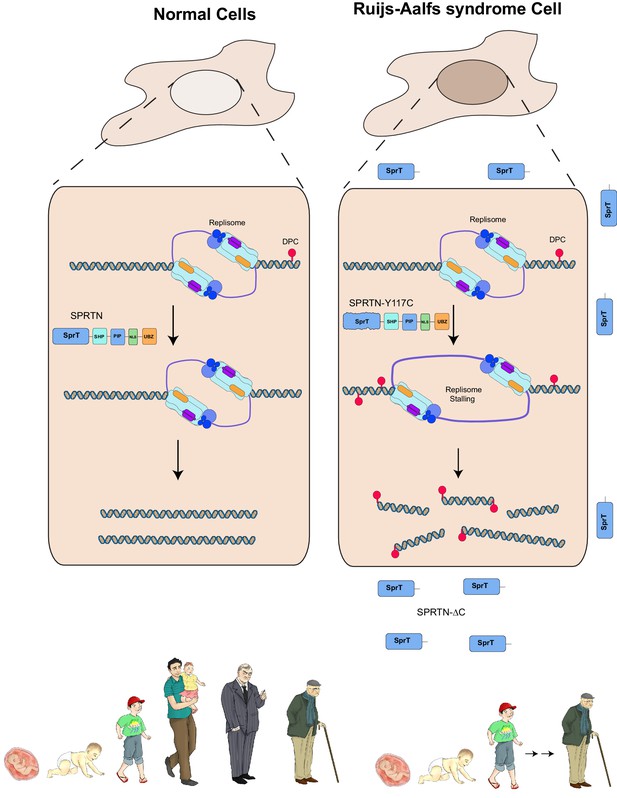
Model of SPRTN removal of DNA-protein crosslinks.
DNA-protein crosslinks may arise stochastically as a consequence of normal metabolic processes occurring at or in the vicinity of DNA or by exogenous sources such as Etoposide. DPCs are resolved by SPRTN to allow for unperturbed and timely DNA replication. When SPRTN is absent or when its activity is compromised, such as in patients with Ruijs-Aalfs syndrome (SPRTN-Y117C or SPRTN-∆C), DPCs can persist and perturb DNA replication and transcription (not shown). DPCs, if not resolved, can lead to genomic instability, tumorigenesis and contribute to accelerated aging.
Additional files
-
Supplemental file 1
Yeast strains used in this study.
- https://doi.org/10.7554/eLife.21491.014





