Structural basis for the disaggregase activity and regulation of Hsp104
Figures
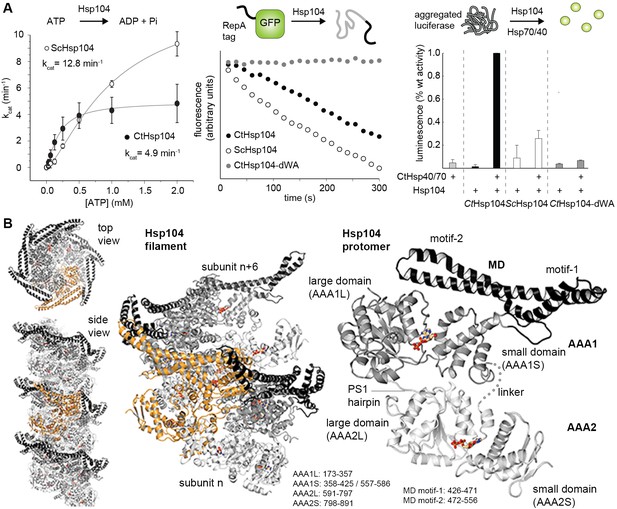
Crystal structure of Hsp104.
(A) Overview of ATPase, GFP unfolding, and luciferase disaggregation assays used to test the functionality of the Hsp104 protein from C. thermophilum (CtHsp104) and its homologue from yeast (ScHsp104). The catalytic inactive double Walker A (WA; K229A/K640A) mutant was used as negative control. For the unfolding and disaggregase assay, the relative activities to the CtHsp104 wild-type protein are shown, with error bars representing standard deviations of 3 independent experiments. Unfoldase activity was measured using a non-physiological ATP/ATPgS mixture required by the assay setup. (B) Ribbon presentation of the crystallized Hsp104 filament (color coded according to domains, with one whole subunit depicted in orange) and of the constituting protomer. ADP molecules are shown in stick representation. See also Figure 1—figure supplements 1 and 2.
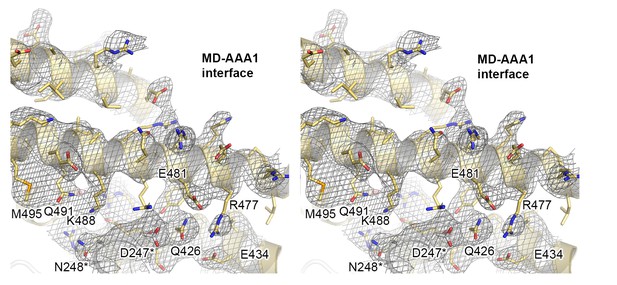
Crystal structure of Hsp104 from C. thermophilum.
Stereo figure showing the central portion of the MD/AAA1 interface (described as tethering site one in Figure 3). Despite medium resolution, the electron density map revealed the overall orientation of most interface residues, as seen in the composite 2FoFc omit map, calculated at 3.7 Å resolution and contoured at 1 σ.

Active sites of AAA1 and AAA2.
Active sites of AAA1 and AAA2 are shown with the partner subunit colored in yellow. The 2FoFc omit density of the bound ADP nucleotide is overlaid on the final structure showing the mechanistically important WA (Walker A), WB (Walker B), S1 (sensor-1), and S2 (sensor-2) residues.
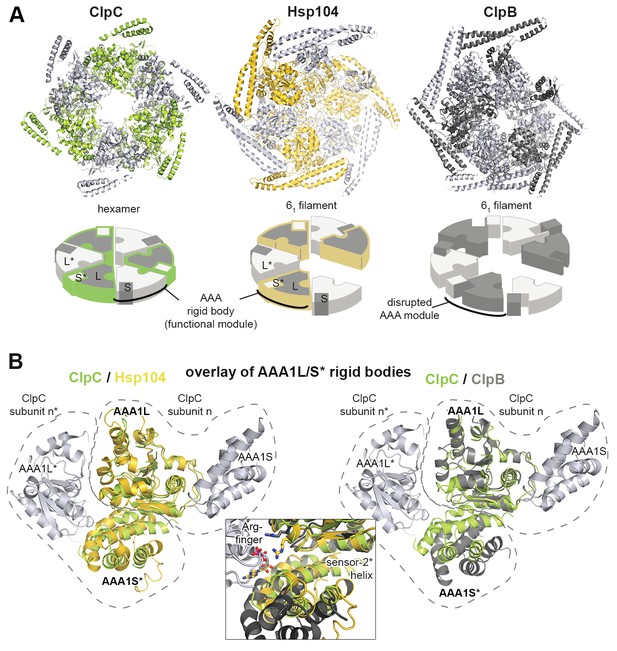
Functional ATPase modules are retained in the Hsp104 filament.
(A) Ribbon presentation of ClpC (PDB 3pxg), Hsp104, and ClpB (PDB 4ciu) oligomers highlighting the L/S* rigid bodies (colored). While ClpC was crystallized as a hexamer, Hsp104 and ClpB were crystallized in a helical filament with a six-fold screw axis. The cartoons below depict six L/S* modules (framed) that are formed between adjacent subunits (distinct grey tones). (B) Ribbon presentation of adjacent AAA1 domains of the hexameric ClpC that jointly compose one L/S* rigid body (green). Superimposed are the L/S* modules of the Hsp104 (yellow, left panel) and ClpB (dark grey, right panel) filament. The zoomed-in window emphasizes the distinct orientation of the sensor-2 helix, the central element in linking L and S* sub-domains. ADP and the SRH* arginine finger are shown in stick representation to mark the position of the active site. See also Figure 2—figure supplements 1,2 and 3.
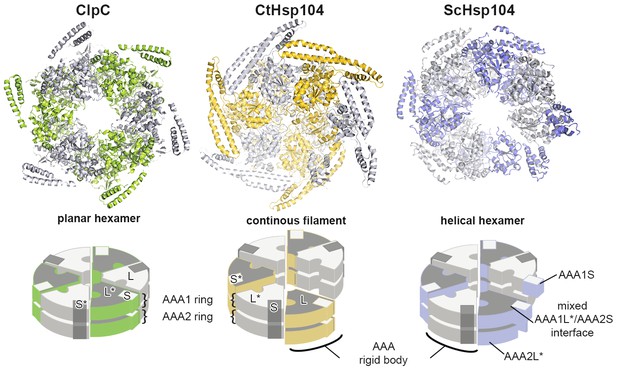
Comparison between planar and helical Hsp100 conformations.
Ribbon presentation of ClpC (PDB 3pxg), Hsp104, and ScHsp104 (PDB 5kne) oligomers highlighting the L/S* rigid bodies (colored). While ClpC was crystallized as a planar hexamer and CtHsp104 as a continuous filament, the helical hexamer structure of ScHsp104 was determined by cryo-electron microscopy. The cartoons below depict the L/S* modules (framed) that are formed between adjacent subunits (distinct grey tones).

Functional ATPase modules are retained in the Hsp104 filament.
Ribbon presentation of two adjacent AAA subunits of ScHsp104 (PDB 5kne (Yokom et al., 2016)) and the ClpC hexamer (PDB 3pxi (Wang et al., 2011)) highlighting the composite L/S* rigid bodies. Individual subunits are encircled. Top: Superimposed are L/S* modules from hexameric ScHsp104 (blue) and ClpC (green) on the AAA1 and AAA2 L/S* modules of the Hsp104 filament (yellow). Bottom: Structural alignment of ClpC with the three L/S* modules present in the filament of ClpB from Thermus thermophilus (ClpB, dark grey; PDB 1qvr [Lee et al., 2003]).

Sequence conservation of HSP100 disaggregases.
The sequence conservation of HSP100 disaggregases is plotted onto the AAA1 and AAA2 domains of one Hsp104 subunit (high sequence conservation is colored pink while low sequence conservation is indicated in green). Neighboring protomers are colored in grey and the most important interfaces are labeled.
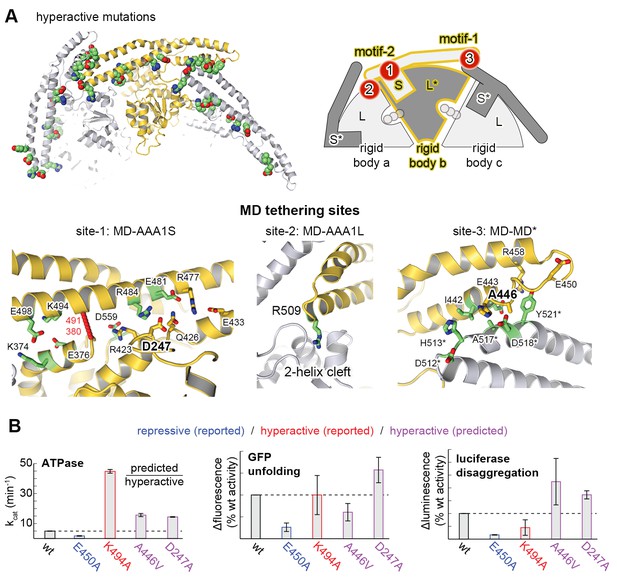
Structural organization of the MD-AAA1 interface.
(A) Ribbon presentation of Hsp104 showing the clustering of hyperactive mutations (green, see Supplementary file 1) along the MD-AAA1 and MD-MD* interfaces. The cartoon illustrates the localization of the three major MD tethering sites, which are also shown in structural detail. The 380–491 pair used in cross-linking studies connects motif-2 of the MD with AAA1 (red line). (B) Comparison of mutants predicted to be hyperactive (A446V and D247A should destabilize the MD-AAA1-MD* interface) with reported hyperactive (red, K494A) and repressed (blue, E450A) mutants. Error bars indicate standard deviations.
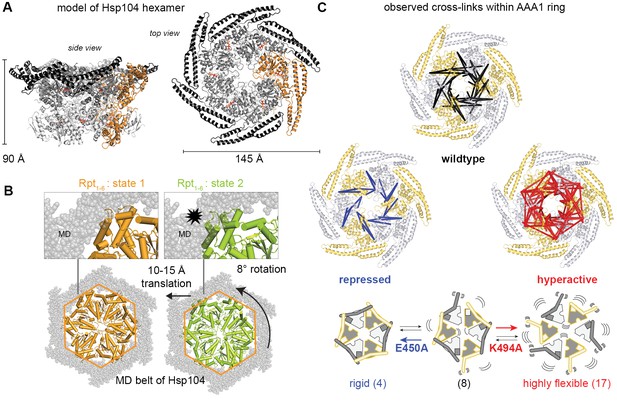
Inhibitory function of the MD.
(A) Ribbon presentation of a modeled Hsp104 hexamer that was constructed from the L/S* rigid body of the crystallized filament. The dimensions of the Hsp104 hexamer are indicated and subunits are colored as in Figure 1B). (B) Ribbon representation of the proteasomal Rpt1-6 present in two alternative conformations (state-1, PDB 4cr2, orange, and state-2, PDB 4cr4, green (Unverdorben et al., 2014)). The two AAA rings are shown together with the MD belt (grey surface) of the superimposed Hsp104 hexamer. Conformational differences between the Rpt1-6 ATPase modules are indicated. The asterisk highlights a hypothetical clash with the MD belt. (C) Ribbon presentation of the modeled Hsp104 hexamer highlighting the L/S* rigid bodies (colored differently). The different numbers of identified Lys-Lys cross-links are represented by intermolecular connections, linking neighboring rigid bodies (wt in black, E450A in blue, K494A in red). The proposed effects of activating and repressing mutations on the dynamics of the Hsp104 hexamer are schematically indicated. See also Figure 4—figure supplements 1 and 2.
-
Figure 4—source data 1
Cross-linked Lys pairs observed by XL-MS.
- https://doi.org/10.7554/eLife.21516.012
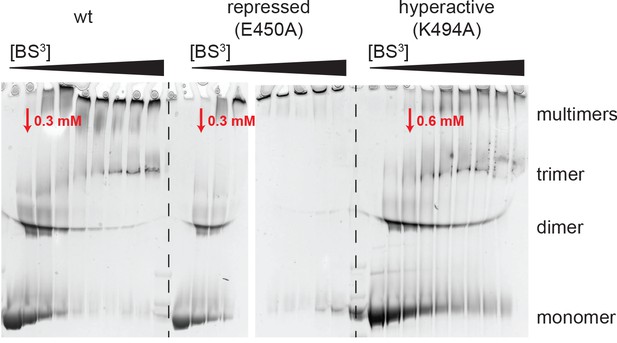
Analysis of BS3 cross-linking efficiency.
The Hsp104 variants (wild-type, repressed (E450), and hyperactive (K494A)) were incubated with increasing BS3 concentrations (from 0 to 35 mM) and analyzed by SDS-PAGE. Although MS experiments characterizing the repressed state of Hsp104 revealed only a small subset of Lys residues being cross-linked, reaction progression was very rapid, as may be expected for a rigid particle. In contrast, cross-linking the highly dynamic hyperactive mutant required larger amounts of the BS3 linker. The BS3 concentrations used in the XL-MS experiment are indicated in red.
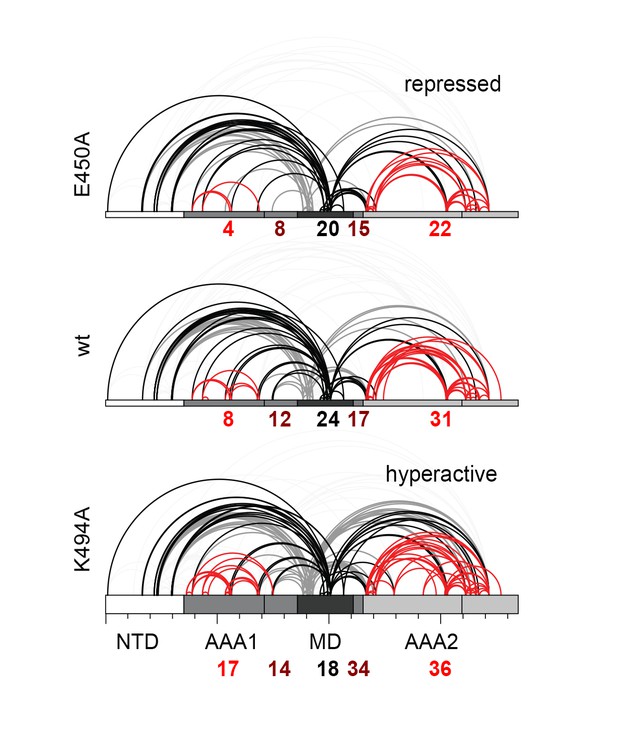
Distribution of identified XL-MS cross-links.
Identified XL-MS cross-links are visualized by lines connecting corresponding Lys residues. The number of identified Lys-Lys pairs (Figure 4—source data 1) within the AAA rings (red), between residues of the MD's (black), and between the MD and the AAA rings (dark red) are given.
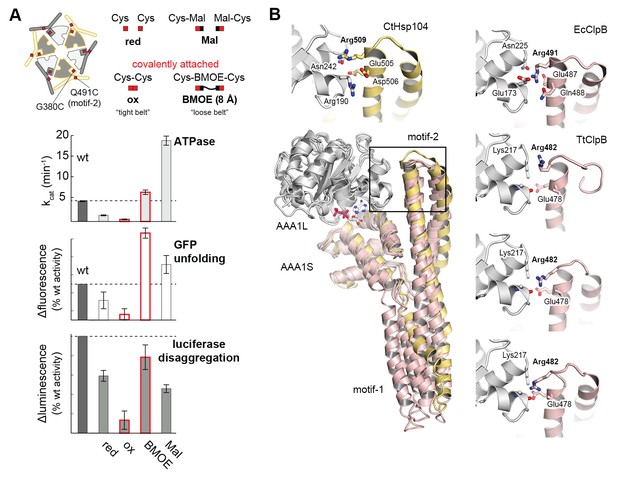
Regulatory role of motif-2.
(A) As schematically shown, the G380C/Q491C mutant allows to covalently connect MD and AAA1 by a ‘tight’ (Cys-Cys linkage) or a ‘loose’ belt (Cys-BMOE-Cys). ATPase, unfolding and disaggregase activity of the G380C/Q491C mutant under reducing (red) and oxidizing conditions (ox) and upon reaction with the chemical cross-linker (BMOE) or free maleimide (Mal) are shown in comparison to the wild-type enzyme. Error bars represent standard deviations. See also Figure 5—figure supplement 1. (B) Interaction between MD and AAA1 domain, observed in Hsp104 and ClpB crystal structures (EcClpB PDB 4ciu, TtClpB PDB 1qvr). As shown in the superposition (Hsp104 MD in yellow; ClpB MD conformations in red) and the zoomed-in windows, motif-2 binds via a conserved arginine to the same two-helix cleft in AAA1L. Adjacent polar contacts should stabilize this interaction.

Identification of a Cys-Cys pair to covalently fix the MD belt.
Monitoring the cross-linking specificity of different Cys-Cys pairs by SDS-PAGE. Prior to loading to the gel, samples were either incubated at reducing (red) and oxidizing conditions (ox) or treated with a chemical cross-linking reagent (BMOE).
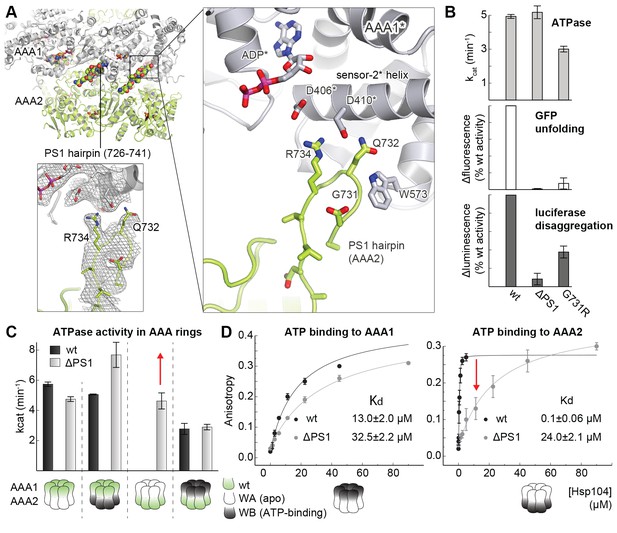
Functional coupling of the two AAA rings of Hsp104.
(A) The PS1-hairpin of AAA2, which was well-defined by electron density (inset: omit density map contoured at 1.0 σ), forms specific contacts within the AAA1* active site. Bound ADP and interacting residues are shown in stick mode. (B) Characterization of the PS1 deletion (∆PS1) and the G731R mutant showing that the PS1-hairpin is essential for unfoldase and disaggregase activity, but not for ATPase activity. (C and D) ATPase and mant-ATP binding assays reveal the role of the PS1-hairpin in adjusting the activities and nucleotide binding affinities of AAA1 and AAA2 to each other. Strongest effects of the ∆PS1 mutation are highlighted (red arrow). The used AAA variants (WA/WB combined with wildtype) are indicated. Error bars represent standard deviations.

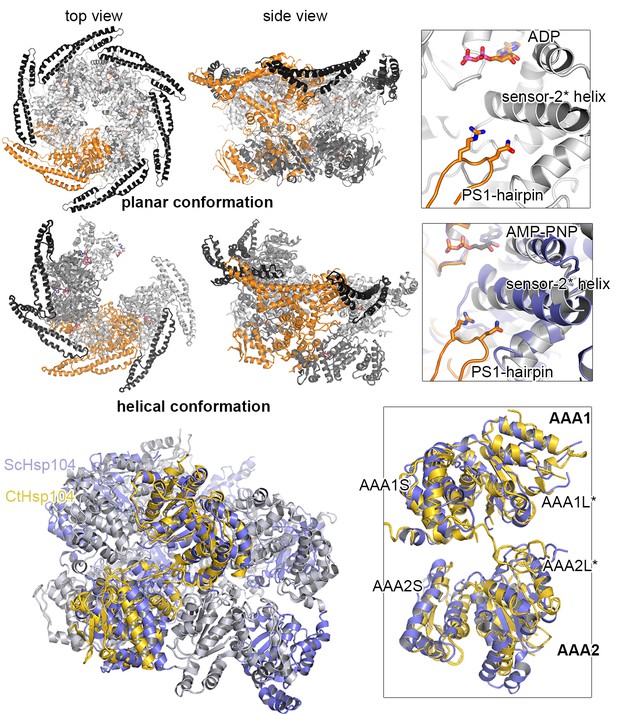
Position of the PS1-hairpin in hexameric Hsp104.
Top and middle row: Ribbon presentation of molecular models showing CtHsp104 in the planar and helical conformation. Models were constructed by overlying the AAA1 part of the crystallized CtHsp104 L/S* rigid bodies onto the respective AAA1 modules of the hexameric ClpC (top, PDB 3pxi [Wang et al., 2011]) or ScHsp104 (middle, PDB 5kne [Yokom et al., 2016]). Of note, the subunit that closes the hexamer by forming an uncanonical AAA1S-AAA2L* interface is left out from the alignment. The zoomed in windows illustrate the position of the PS1-haripin with nucleotides and functional residues shown in stick mode. Models are colored according to Figure 1B and ScHsp104 is shown in blue. Bottom row: Ribbon presentation showing the alignment of one Ctsp104 rigid body (yellow) onto the ScHsp104 hexamer (L/S* modules colored in blue and grey, respectively) with one functional unit magnified. As the MD is only partially resolved in the ScHsp104 EM structure, this domain was omitted from the presentation.
Effect of PS1-hairpin deletion on ScHsp104 activity.
ATPase and GFP unfolding activity of wt and ΔPS1 mutants of Ct and ScHsp104 (Ct: C. thermophilium, Sc: S. cerevisiae). Error bars represent standard deviations and arrows indicate the decrease in unfolding activity.
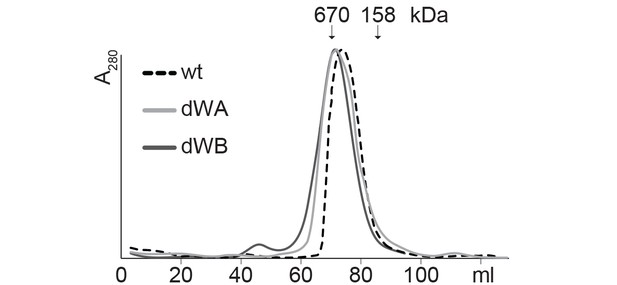
SEC profiles of CtHsp104 dWA and dWB mutants.
SEC profiles confirm the formation of stable hexamers for CtHsp104 (wt) and the catalytic inactive double Walker A (dWA: K230A/K641A) and double Walker B (dWB: D295A/E296A) mutants. Elution volumes of marker proteins are indicated.
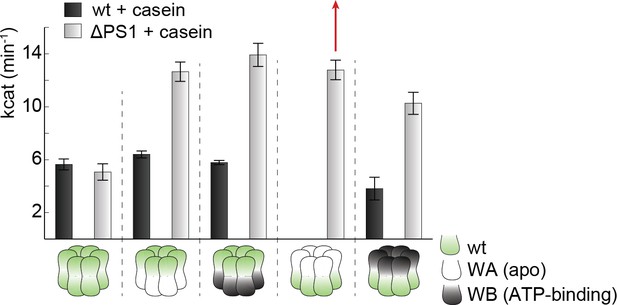
Effect of casein on ATPase activity.
ATPase activity assays confirm the importance of the PSI-hairpin in coordinating activities between AAA1 and AAA2 in the presence of casein. The used AAA variants (WA/WB combined with wildtype) are schematically indicated. The strongest effect of the ∆PSI mutation is highlighted (red arrow). Error bars represent standard deviation.
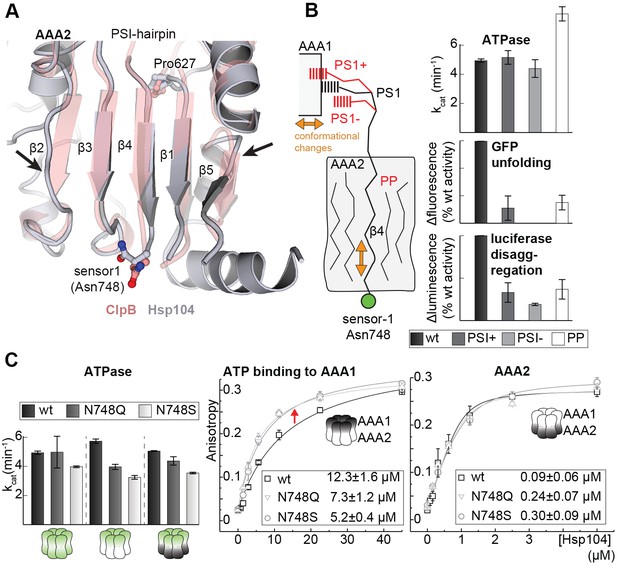
Signal path between AAA1 and AAA2.
(A) Superposition of the AAA2 domain of the Hsp104 filament (grey) with the AAA2 domain of ClpB that was crystallized as isolated domain (salmon; PDB 4lj5). The ribbon model illustrates the distortion of strands β2 and β5 (arrows) and the shorter strands β1, β3, and β4 observed in the Hsp104 structure. See also Figure 7—figure supplement 1. (B) Model of how conformational changes of an AAA1 ATPase module or reorientations of strand β4 and the sensor-1 residue are communicated by the PSI-hairpin. Mutants predicted to decouple signaling (PS1+, PS1- and PP) were analyzed for their ATPase, unfolding, and disaggregation activities. See also Figure 7—figure supplements 2 and 3. (C) ATPase activity assay and mant-ATP binding data of sensor-1 mutants (N748S and N748Q). Strongest effects compared to the respective wildtype control are highlighted (red arrow). The used AAA variants (WA/WB combined with wildtype) are indicated. Error bars represent standard deviations.
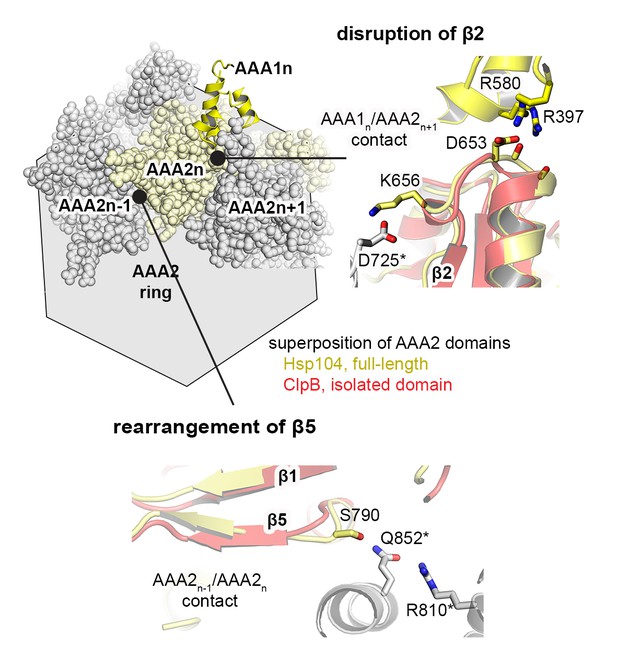
Signal path between AAA1 and AAA2.
The structural presentation shows three AAA2 domains of the Hsp104 filament (space filling model, colored by subunits) together with a motif protruding from a juxtaposed AAA1 domain (yellow ribbon). The zoomed-in windows illustrate structural details of how contacts with AAA1 and neighboring subunits may destabilize the central β-sheet of AAA2. For comparison, the structure of Hsp104 (yellow) is aligned with the AAA2 domain of ClpB, which was crystallized in isolated form (red, PDB 4lj5 [Zeymer et al., 2014]).
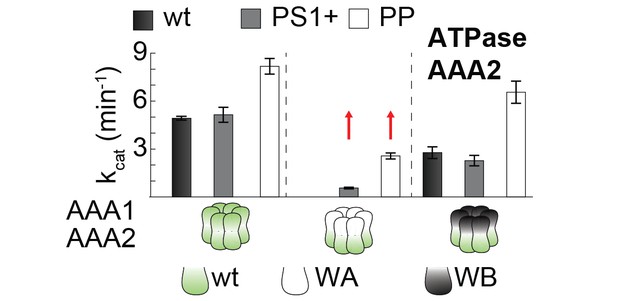
PS1+ and PP mutations partially uncouple the ATPase activity of AAA1 and AAA2.
ATPase activity assay of wt, PS1+ and PP mutants. WA and WB mutations were introduced as indicated. Error bars represent standard deviations.
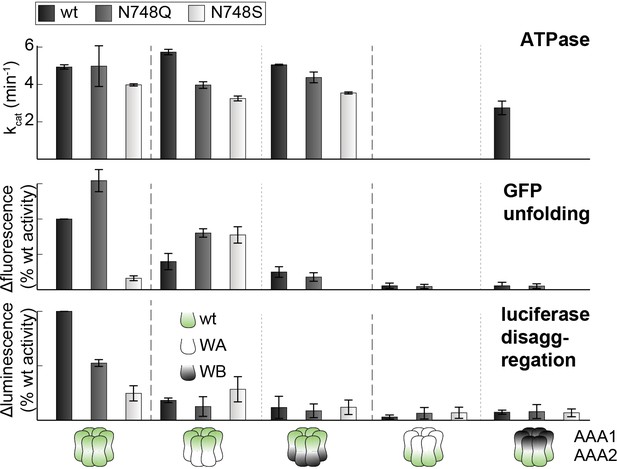
Activities of sensor-1 mutations.
Activity assays of wt and sensor-1 mutants (N748S and N748Q). WA and WB mutations were introduced as indicated. Error bars represent standard deviations.
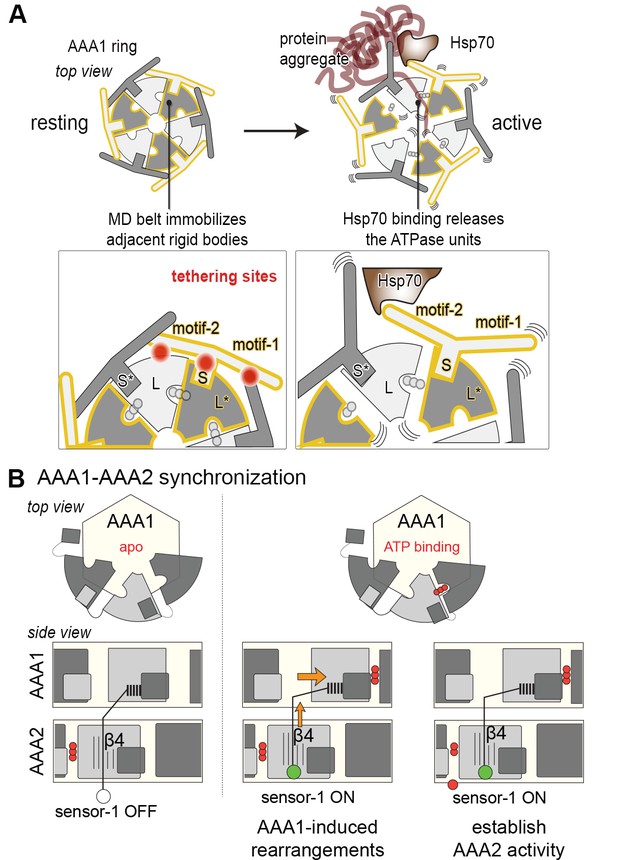
Novel mechanistic features of the Hsp104 disaggregase.
(A) A restraint mask composed of the MD keeps the disaggregase inactive by immobilizing ATPase modules of the AAA1 ring. Binding of Hsp70 to the MD opens the safety belt and activates Hsp104 to act on the presented protein aggregate. The zoomed-in windows illustrate the multiple contact sites established by the MD that result in a physical tethering of adjacent rigid bodies (neighboring subunits are colored in different grey tones and rigid bodies are framed). (B) Allosteric coupling of the two AAA rings by the PS1 motif of AAA2. Nucleotide-dependent movements of an AAA1 module are transduced via the PS1-hairpin over a large distance leading to the repositioning of the catalytic sensor-1 residue of AAA2. As indicated, the functional switch relies on the reorientation of strand β4 in the AAA2 domain.
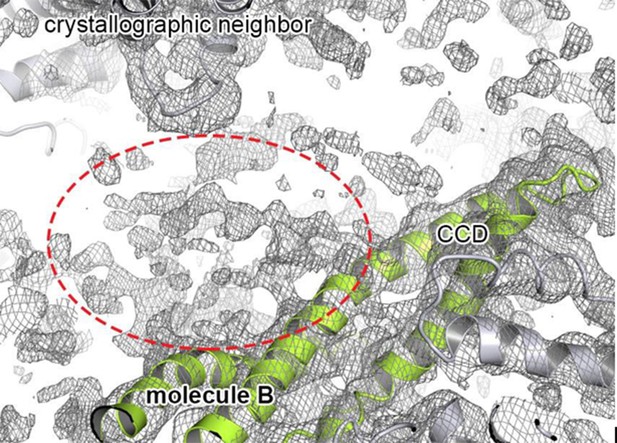
Extra density that may account for the N-terminal domain.
Cartoon representation showing the CCD of molecule B colored in green, together with the 2FoFc electron density contoured at 0.8 σ. The observed extra density for molecule B that could not be used for model building is highlighted. The neighboring subunits (molecule C and crystallographic neighbors) are colored in grey.
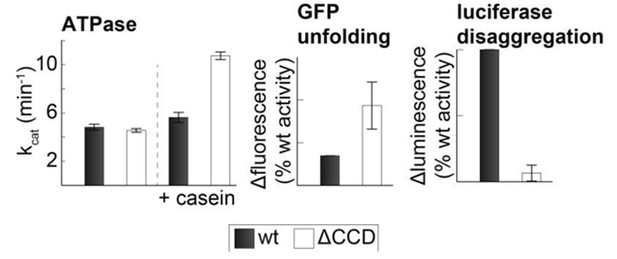
Activity of the CCD deletion mutant.
Substrate-induced ATPase activity of CtHsp104ΔP427-G553 (ΔCCD) is higher than for wild-type Hsp104. Moreover, the ΔCCD mutant showed an increased GFP-unfolding activity while the luciferase disaggregation activity of the corresponding Hsp104/Hsp70 system was reduced.
Tables
Data collection and refinement statistics.
5d4w | |
|---|---|
Data collection | |
Space group | P21 |
Cell dimensions | |
a, b, c (Å) | 144.9, 93.2, 144.4 |
α, β, γ (°) | 90, 119.7, 90 |
Resolution (Å) | 47.2–3.7 (3.8–3.7)1 |
Rmerge | 0.192 (1.624) |
Rpim | 0.130 (1.121) |
I/σI | 5.3 (0.8) |
Completeness (%) | 98.3 (97.9) |
Redundancy | 3.1 (2.9) |
Refinement2 | |
Resolution (Å) | 47.6–3.7 |
No. reflections | 35560 |
Rwork/Rfree | 0.237/0.277 |
No. atoms (all) | 16272 |
Protein | 16110 |
ADP | 162 |
B-factors (all) | 210.7 |
Protein | 210.9 |
ADP | 188.6 |
R.m.s deviations | |
Bond lengths (Å) | 0.006 |
Bond angles (°) | 1.082 |
-
1Highest resolution shell is shown in parenthesis.
-
2One crystal was used for measurement and the stereochemistry of the model was validated with Molprobity (Chen et al., 2010).
Additional files
-
Supplementary file 1
Published Hsp104 and ClpB mutants with repressive or hyperactive phenotype.
- https://doi.org/10.7554/eLife.21516.027
-
Supplementary file 2
Sequence alignment of Hsp104 and related HSP100 disaggregases.
- https://doi.org/10.7554/eLife.21516.028





