TGF-β reduces DNA ds-break repair mechanisms to heighten genetic diversity and adaptability of CD44+/CD24− cancer cells
Figures
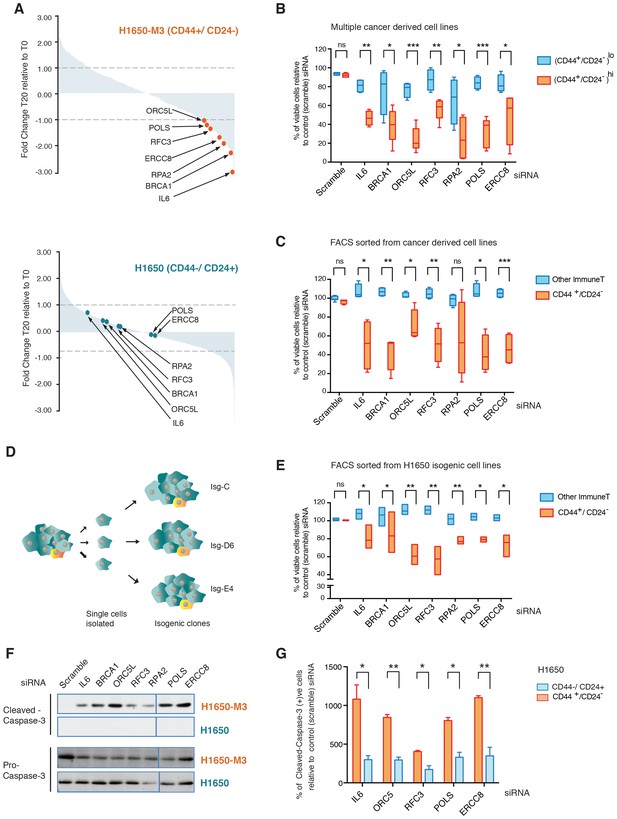
A genome-wide shRNA screen identifies genes involved in DNA damage repair (DDR) that are required for the survival of CD44+/CD24−.
(A) The graph depicts the relative abundance of barcodes recovered from the screen. Each bar represents fold changes of an shRNA expression vector at T20 (i.e., 20 cell passages) compared with T0 (time of infection) in CD44+/CD24− H1650-M3 cells (upper panel) and CD44−/CD24+ H1650 cells (lower panel). Dots indicate unique genes, knockdown of which conferred proliferative disadvantage to CD44+/CD24− (H1650-M3) cells. The data are plotted as the means of three biological replicates in ascending order. A FACS profiling of H1650-M3 and H1650 cells, along with a schematic of the shRNA screen, is provided in Figure 1—figure supplement 1. (B) Validation of shRNA screen hits in tumor-derived cell lines characterized by low CD44+/CD24− cell content (i.e., MCF7, A549 and BT474) compared to cell lines with high CD44+/CD24− content (i.e., NCI-H23, PC9, MDA-MB435S and MDA-MB-231). The box plots show the percentage of viable cells 5 days after transfection with the indicated siRNAs relative to the number of control scramble-siRNA transfected cells. Each box is the mean ± SD of data collected from cell lines with either (CD44+/CD24−)lo or (CD44+/CD24−)hi content, from two independent experiments, each conducted in eight replicates (p-value *<0.05, **<0.005, ***<0.0005, unpaired t-test). FACS profiles for each cell line, relative % of viable cells for each cell line and knockdown efficiency are reported in Figure 1—figure supplement 2. (C) Validation of shRNA screen hits in tumor-derived cell lines FACS-sorted on the basis of the surface expression of CD44 and CD24. The box plots show the percentage of CD44+/CD24− cells and cells of other immune types upon transfection with the indicated siRNA oligonucleotides relative to control (scramble) siRNA. Each box is the mean ± SD of data collected from four cells lines (A549, H1650, PC9 and NCI-H23) upon FACS sorting, each from three replicates from two independent experiments. (p-value *<0.05, **<0.005, ***<0.0005, unpaired t-test). See Figure 1—figure supplement 3 for more details. (D) Schematic of the generation of single cell-derived isogenic cell lines from H1650 cells. See Figure 1—figure supplement 4A for CD44 and CD24 surface marker staining profiles. (E) Validation of shRNA screen hits in the FACS-sorted H1650 single cell-derived isogenic clones—Isg-C, Isg-D6 and Isg-E4. The box plots indicate the percentage of CD44+/CD24− cells and cells of other immune types after transfection with the indicated siRNA oligonucleotides relative to control (scramble) siRNA. Each box is the mean ± SD of data collected from three different isogenic cell lines, each from three replicates from two independent experiments (p-value *<0.05, **<0.005, unpaired t-test). See Figure 1—figure supplement 4B,C for further details. (F) Expression of Pro-caspase three and Cleaved-caspase 3 (i.e., cell death marker) in H1650-M3 (CD44+/ CD24−) and H1650 (CD44−/CD24+) cell lines upon knockdown of indicated gene expression. Samples were collected 3 days post-transfection and protein lysates were immune-blotted with the indicated antibodies. Alpha-tubulin is used as the loading control. See Figure 1—figure supplement 5 for quantification. (G) Percentage of Cleaved-caspase 3-positive cells, normalized to respective scramble controls (set at 100%), in FACS-sorted CD44+/CD24− and CD44−/CD44+ cells in H1650 cell line. Each bar represents mean ± SD of three replicates from two independent experiments(p-value *<0.05, **<0.005, unpaired t-test).
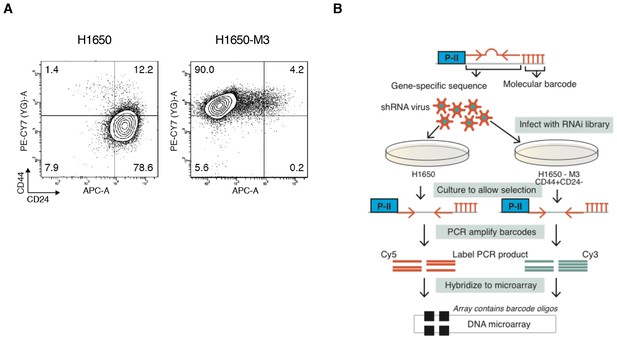
Forward genetic screen performed in H1650 and H1650-M3 cell lines.
(A) FACS analysis of H1650 and H1650-M3 stained with antibodies against cell surface markers CD44 and CD24. (B) A schematic of the RNAi-based, forward genetic screen.
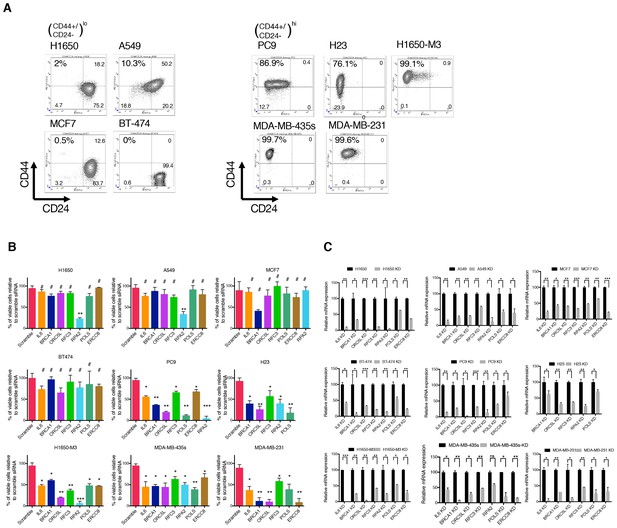
Validation of shRNA screen hits in tumor-derived cell lines characterized by low (H1650, A549, MCF7, and BT-474) and high (PC9, NCI-H23, H1650-M3, MDA-MB-435s and MDA-MB-231) content of CD44+/CD24− cells.
(A) FACS profile of multiple cancer-derived cell lines used in validation of the shRNA screen hits. Cells were stained with antibodies against cell surface markers CD44 and CD24 and analyzed with FACS. (B) The charts represent the percentage of viable cells 5 days after transfection with the indicated siRNA oligonucleotides compared to scramble siRNA=transfected control cells. Each bar represents mean ± SD of two independent experiments each conducted in eight- replicates (n = 16). (C) Efficiency of knockdown upon transfection with the indicated siRNA oligonucleotides and compared to scramble siRNA transfected control cells. Cells were collected 3 days post transfection and analyzed for expression of the indicated mRNA by RT-qPCR. Each bar represents mean ± SD of three replicates. p-value *<0.05, **<0.005, ***<0.0005, unpaired t-test. Non-significant differences are shown by the symbol ‘#’ or ‘ns’.
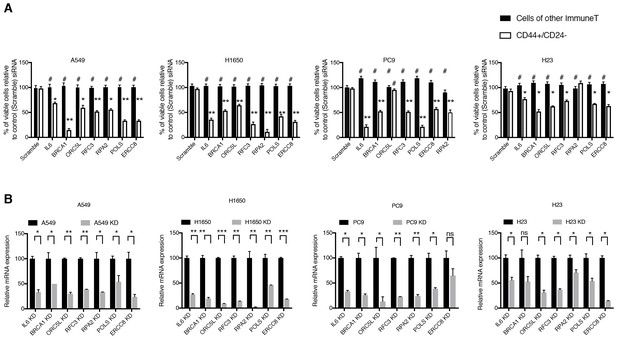
Validation of shRNA screen hits in tumor-derived cell lines FACS sorted on the basis of the surface expression of CD44 and CD24.
( A) The charts represent the percentage of viable CD44+/CD24− cells and cells of other immune types upon transfection with the indicated siRNA oligonucleotides relative to control (scramble siRNA). Each bar represents mean ± SD of three technical replicates from two independent experiments. 10,000 cells were analyzed by FACS for each replicate of each sample. (B) Efficiency of knockdown upon transfection with the indicated siRNA oligonucleotides and compared to control cells transfected with scramble siRNA. Cells were collected 3 days post transfection and analyzed for expression of the indicated mRNA. Each bar represents mean ± SD of three replicates. p-value *<0.05, **<0.005, ***<0.0005, unpaired t-test. Non-significant differences are shown by the symbol ‘#’ or ‘ns’.
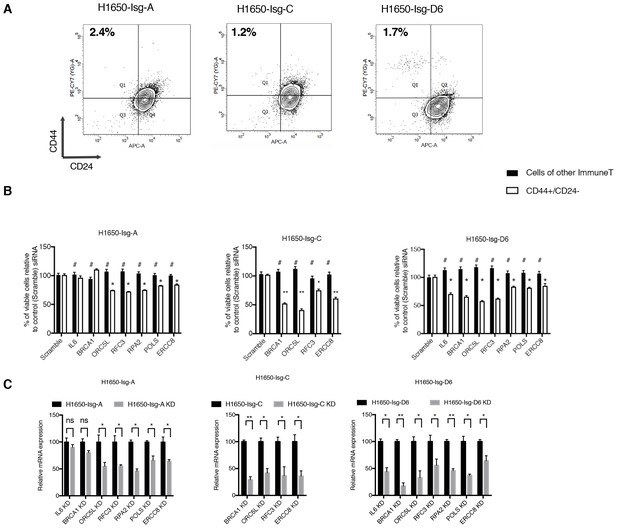
Validation of shRNA screen hits in H1650-derived isogenic clones, FACS sorted on the basis of surface expression of CD44 and CD24.
(A) FACS profile of H1650-derived isogenic cell lines stained with CD44 and CD24 antibodies. The numbers in bold represent the percentage of cells that are CD44+/CD24−. (B) The charts represent the percentage of viable CD44+/ CD24− cells and cells of other immune types upon transfection with the indicated siRNA oligonucleotides relative to control (scramble siRNA). Each bar represents mean ± SD of three technical replicates from two independent experiments. 10,000 cells were analyzed by FACS for each replicate of each sample. (C) Efficiency of knockdown upon transfection with the indicated siRNA oligonucleotides and compared to scramble siRNA transfected control cells. Cells were collected 3 days post transfection and analyzed for expression of the indicated mRNA. Each bar represents mean ± SD of three replicates. p-value *<0.05, **<0.005, ***<0.0005, unpaired t-test. Non-significant differences are shown by the symbol ‘#’ or ‘ns’.
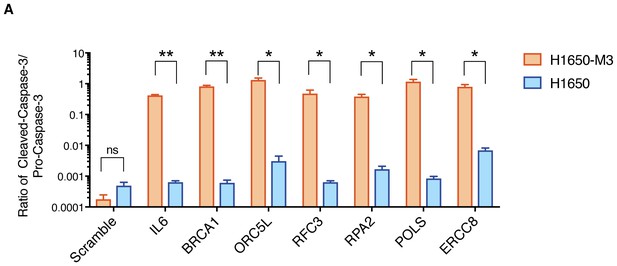
Quantification of apoptosis-mediated increase in lethality in H1650-M3 and H1650.
The chart depicts quantification of relative amounts of Cleaved-caspase-3 between H1650-M3 and H1650 from the immunoblots indicated in Figure 1F. Levels of intensity of each band were quantified using imajeJ32 software and is represented as a ratio to Pro-Caspase-3. The data represent mean ± SD from two independent experiments. p-value *<0.05, **<0.005, unpaired t-test.
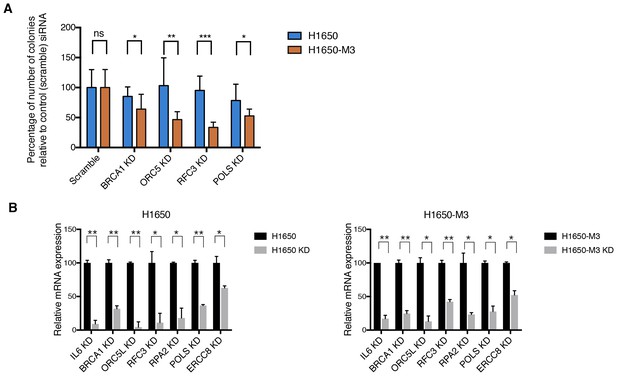
Validation of shRNA screen hits in H1650 and H1650-M3 cells using the clonogenic assay technique.
(A) Clonogenic assay performed on H1650 and H1650-M3 to validate siRNA screen hits for BRCA1, ORC5L, RFC3, and POLS. The charts represent the percentage of viable cells 15 days after transfection with the indicated siRNA oligonucleotides compared to that of scramble-siRNA-transfected control cells. Each bar represents mean ± SD of data collected from ten fields from three different six-well plates (n = 30). (B) Knockdown efficiency for clonogenic assay upon transfection with the indicated siRNA oligonucleotides. Samples were collected 3 days post-transfection and analyzed by immunoblot analysis with antibodies against Total caspase 3, alpha-tubulin and Cleaved-caspase 3. Each bar represents mean ± SD of three replicates. p-value *<0.05, **<0.005, ***<0.0005, unpaired t-test. Non-significant differences are shown by the symbol ‘#’ or ‘ns’.
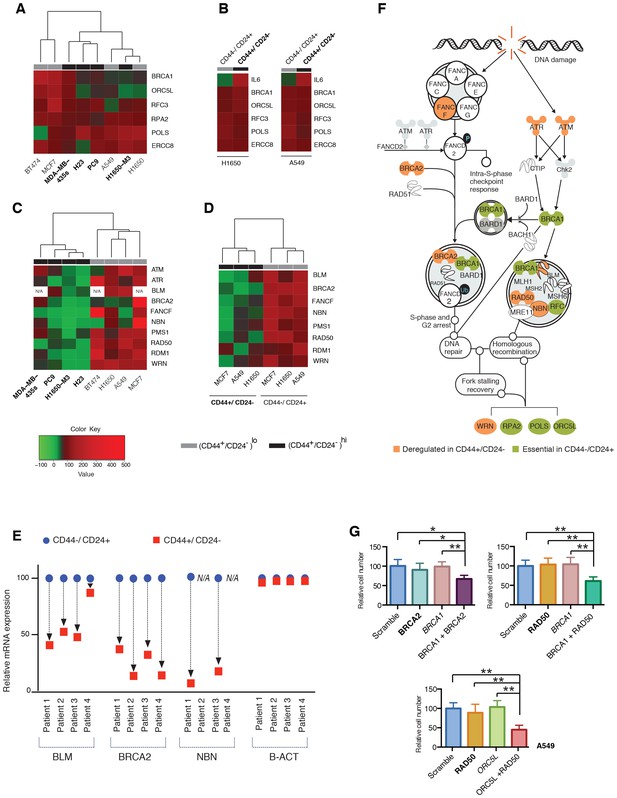
Decreased expression of homology-directed repair (HDR) genes in CD44+/CD24− cells results in synthetic lethal interactions.
(A) The heat map represents a hierarchical cluster analysis of BRCA1, ORC5L, RFC3, RPA2, POLS and ERCC8 mRNA expression in the indicated tumor-derived cell lines and (B) in H1650 and A549 cells that were FACS sorted on the basis of their surface expression of CD44 and CD24. mRNA expression was quantified by SYBR-green-based RT-qPCR. Cell lines with high CD44+/CD24− cell content are indicated in bold. The data represent mean ± SD of three replicates from two independent experiments. See Figure 2—figure supplement 1 for details. (C) Hierarchical cluster analysis of the mRNA expression of the indicated HDR genes in multiple tumor-derived cell lines with high (indicated in bold) or low content of CD44+/CD24− cells. mRNA expression was quantified by SYBR-green-based RT-qPCR. The data represent the mean ± SD of three replicates from two independent experiments. See Figure 2—figure supplement 2 for details. (D) Clustering analysis of mRNA expression of the HDR genes in CD44+/CD24− and CD44−/CD24+ cells FACS-sorted from H1650, A549 and MCF7 cells. mRNA expression was quantified by SYBR-green-based RT-qPCR. The data represent the mean ± SD of three replicates from two independent experiments. See Figure 2—figure supplement 3A for details. (E) Expression of BLM, BRCA2 and NBN genes in FACS-sorted CD44−/CD24+ and CD44+/CD24− cells from four human primary NSCLC tumors. mRNA expression was quantified by SYBR-green-based RT-qPCR. Expression of an indicated mRNA in the CD44+/CD24− cells was calculated relative to its expression in CD44−/CD24+ cells from the respective tumor. Each dot represents mean ± SD of three replicates. See Figure 2—figure supplement 3B,C for details. (F) Schematic representation of the functional interactions between genes that we identified in the screen (green) and the HDR genes that we found to be downregulated (orange) in the H1650-M3 (and CD44+/CD24−) cells (see Supplementary file 2 for details). (G) The charts depict the percentage of viable cells 5 days after knockdown of the indicated genes relative to a scramble-siRNA control in the A549 cell line. Each bar represents mean ± SD of eight replicates from two independent experiments. See Figure 2—figure supplement 5 for knockdown efficiency (p-value *<0.05, **<0.005 paired t-test).
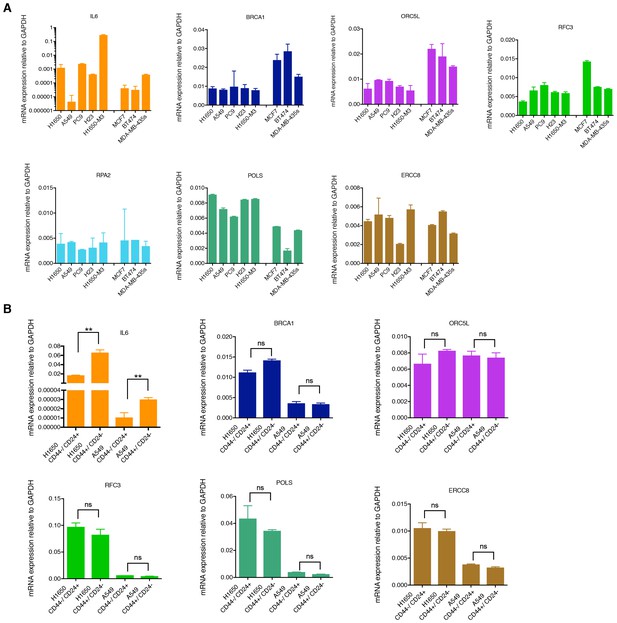
mRNA expression analysis of the shRNA screen hits in tumor-derived cell lines and cells that have been FACS sorted on the basis of their surface expression of CD44 and CD24.
(A) mRNA expression analysis (SYBR-green-based RT-qPCR) of BRCA1, ORC5L, RFC3, RPA2, POLS and ERCC8 in tumor-derived cell lines. The data represent the mean ± SD of three replicates from two independent experiments. (B) mRNA expression analysis (SYBR-green-based RT-qPCR) of BRCA1, ORC5L, RFC3, RPA2, POLS and ERCC8 in H1650 and A549 that have been FACS sorted on the basis of their surface expression of CD44 and CD24. Each bar represents the mean ± SD of three replicates from two independent experiments and represents the mRNA expression of the indicated gene normalized to GAPDH expression of each cell line. p-value **<0.005, unpaired t-test; ns= non-significant.
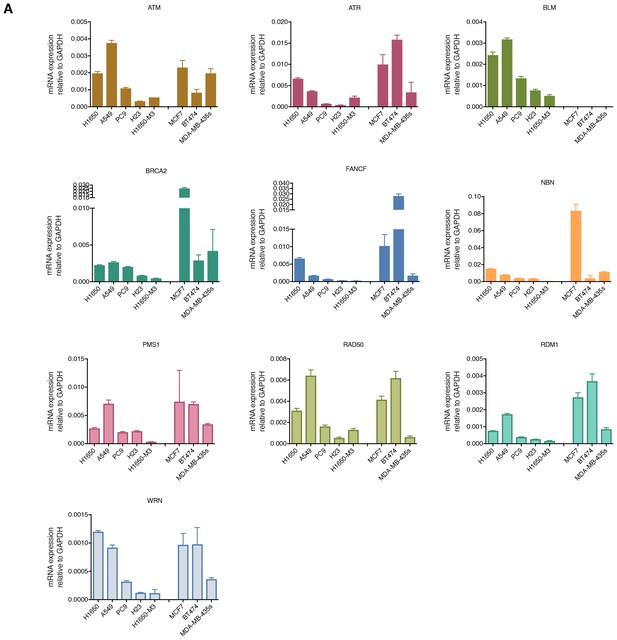
Differential mRNA expression of homology-directed repair (HDR) genes in multiple tumor-derived cell lines.
The chart represents mRNA expression analysis (SYBR-green-based RT-qPCR) of the indicated HDR genes that we observed to be downregulated in the H1650-M3 (CD44+/CD24−) cells across the indicated cell lines. The data represent mean ± SD of three replicates from two independent experiments.
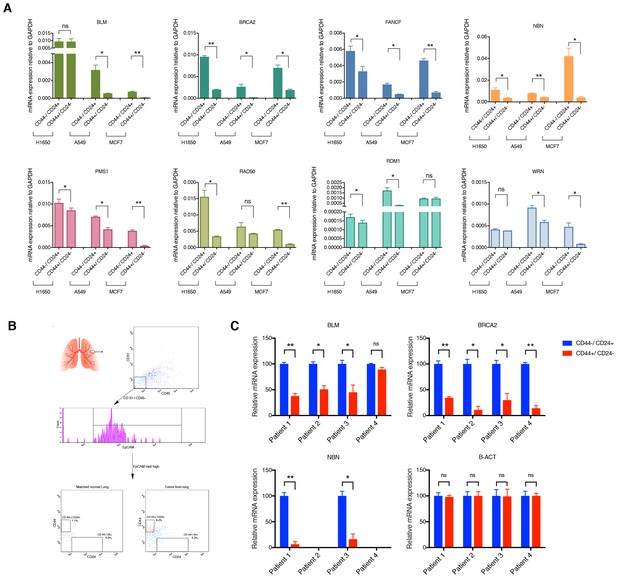
mRNA expression analysis of HDR genes in CD44−/CD24+ and CD44+/CD24− cells FACS sorted from cells lines and patient tumors.
(A) Comparative expression of the indicated HDR genes in FACS-sorted CD44+/ CD24− cells relative to CD44−/ CD24+ cells from H1650, A549 and MCF7. mRNA expression was quantified by RT-qPCR. Each bar is the mean ± SD of three replicates from two different experiments and represents mRNA expression of the indicated gene. p-value *<0.05, **<0.005, unpaired t-test. (B) Schematic of cell sorting from tumors. Tumor-derived single cell suspension was stained with antibodies against CD45, CD31, EpCAM, CD44, and CD24. CD45-; CD31-; EpCAM+ cells were then FACS sorted according to the immune types CD44+/CD24− and CD44−/CD24+. (C) Expression of BLM, BRCA2, NBN and B-ACT genes in FACS sorted CD44−/CD24+ and CD44+/CD24− cells from four NSCL tumors. B-ACT was used as a housekeeping control gene. Each bar represents the mean ± SD of three replicates. p-value *<0.05, ***<0.0005, unpaired t-test.
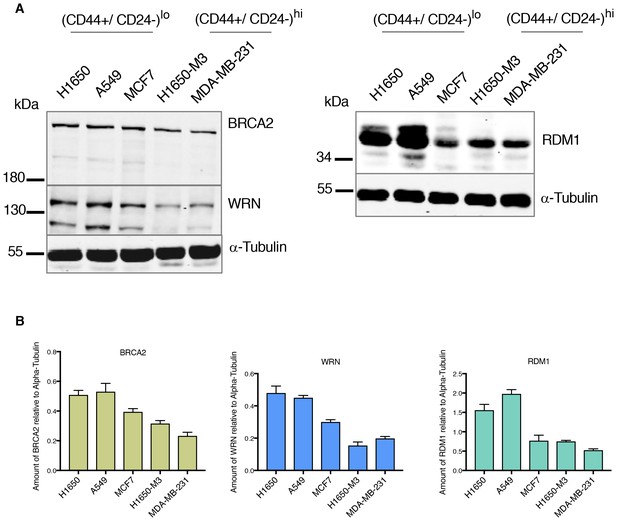
Differential expression of HDR genes in multiple tumor-derived cell lines at protein level.
(A) Expression of BRCA2, WRN and RDM1 in multiple cancer-derived cell lines characterized by different CD44 CD24 immune types. α-Tubulin is used as a loading control. (B) The charts depict the quantification of relative amounts of BRCA2, WRN and RDM1. Levels of intensity of each band were quantified using imajeJ32 software and are represented as a ratio to α-tubulin. Each bar represents the mean ± SD from two independent experiments.
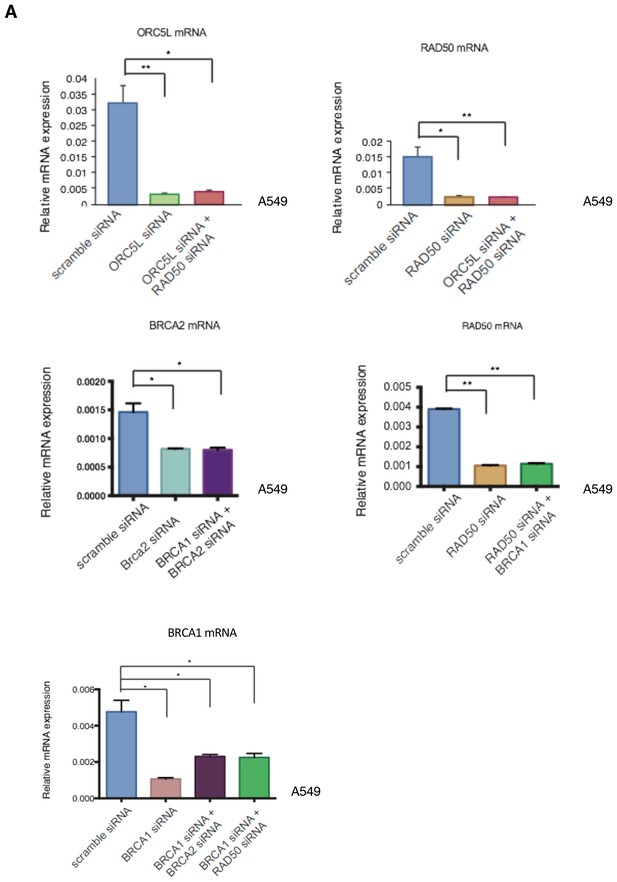
Knockdown efficiencies of the indicated siRNAs in A549 cells.
Knockdown efficiency upon transfection with the indicated siRNA oligonucleotides and compared to scramble siRNA-transfected control cells. Cells were collected 3 days post transfection and analyzed for expression of the indicated mRNA. Each bar represents mean ± SD of three replicates from two independent experiments and represents mRNA expression of the indicated gene normalized to GAPDH expression of each cell line. p-value *<0.05, **<0.005, paired t-test.
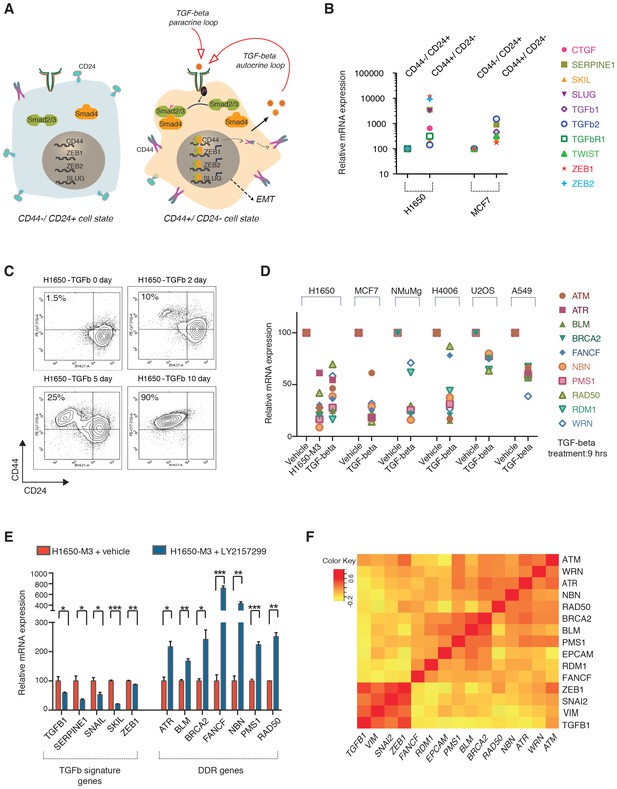
The TGF-β axis controls the expression of the HDR genes that are downregulated in CD44+/CD24− cells.
(A) TGF-β signaling is required for CD44+/CD24− cell state transition. The schematic provided here is based on current literature (Korkaya et al., 2011; Mani et al., 2008). (B) mRNA expression analysis of well-known TGF-β target genes in CD44+/CD24− cells relative to CD44−/ CD24+ cells FACS-sorted from H1650 and MCF7 cell lines. mRNA expression was quantified by SYBR-green-based RT-qPCR. Each dot represents the mean ± SD of three replicates from two independent experiments. See Figure 3—figure supplement 1 for details. (C) FACS analysis of H1650 cells exposed to TGF-β. Cells were treated with TGF-β for the indicated days and stained for the surface expression of CD44 and CD24 and analyzed by FACS. (D) mRNA expression analysis of the indicated HDR genes in TGF-β-treated cells relative to vehicle control across multiple tumor-derived cell lines. Cells were treated for 9 hr with TGF-β1 and TGF-β2 (1 ng/ml each). mRNA expression was quantified by SYBR-green-based RT-qPCR. Each dot represents the mean ± SD of three replicates from two independent experiments. See Figure 3—figure supplement 2B and Figure 3—figure supplement 3 for additional details. (E) The inhibition of TGF-β signaling in the CD44+/CD24− H1650-M3 cells results in an increased expression of HDR genes. TGF-β receptor 1 (TGFBR1) kinase activity was blocked by treatment with 20 μM of LY2157299 (Selleckchem) for 72–96 hr. Expression of TGF-β signature genes were used as a control for the efficacy of LY2157299 treatment. mRNA expression was quantified through SYBR-green-based RT-qPCR. Each bar represents the mean ± SD of three replicates from three independent experiments (p-value *<0.05, **<0.005 paired t-test). (F) Meta-analysis of human breast tumor dataset (BRCA) generated by the TCGA Research Network: http://cancergenome.nih.gov/. Average Pearson correlation coefficients (PCCs) between every pair of genes displayed were calculated and the matrix was generated. Pearson correlation coefficients ranged from −1 to +1. The matrix indicates an inverse co-regulation of the TGF-β1 and the HDR genes that we found to be downregulated in CD44+/CD24− cells.
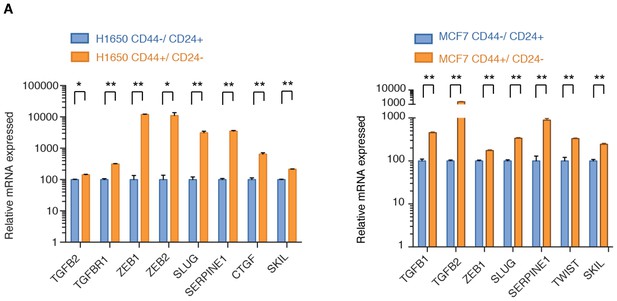
TGF-β signaling is upregulated in FACS-sorted CD44+/CD24− cells.
Expression of well-known TGF-β target genes in CD44+/ CD24− cells relative to CD44−/ CD24+ cells sorted from H1650 and MCF7. mRNA expression was quantified by SYBR-green-based RT-qPCR. Each bar represents the mean ± SD of three replicates from two independent experiments and represents mRNA expression of the indicated gene normalized to GAPDH expression of each cell line (n = 6). p-value *<0.05, **<0.005 unpaired t-test.
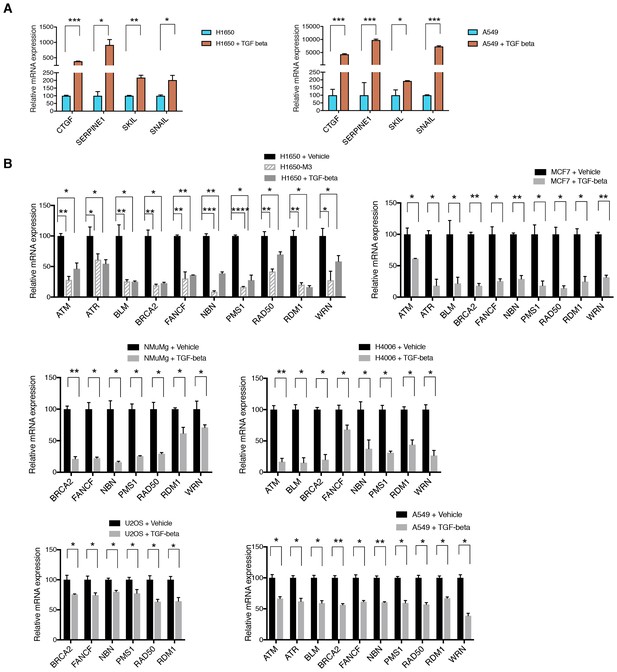
Active TGF-β signaling control the expression of multiple HDR genes.
(A) TGF-β signature target genes measured by SYBR-green-based RT-qPCR in H1650 and A549 upon treating the cells with TGF-β for 9 hr. mRNA expression was quantified by RT-qPCR. Each bar represents the mean ± SD of three replicates from three independent experiments and represents mRNA expression of the indicated gene. p-value *<0.05, **<0.005 , ***<0.0005, paired t-test. (B) mRNA expression analysis of indicated HDR genes in TGF-β-treated cells relative to vehicle control in the indicated cell lines. mRNA expression was quantified by SYBR-green-based RT-qPCR. Each bar represents the mean ± SD of three replicates from two independent experiments (n = 6), normalized to respective GAPDH mRNA expression. Cells were treated for 9 hr with TGF-β (1 ng/ml each of TGF-β1 and TGF-β2). p-value *<0.05, **<0.005 ***<0.0005, paired t-test.
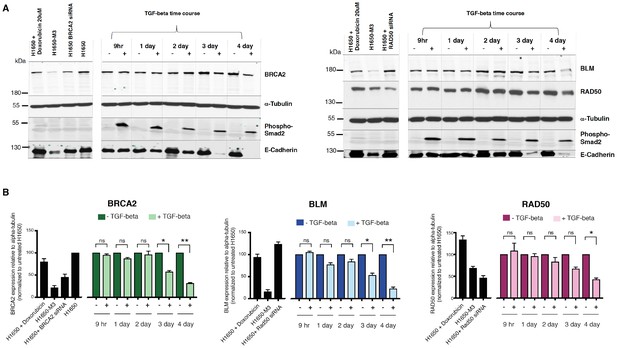
TGF-β signaling controls the expression of multiple HDR genes at the protein level.
(A) Western blot analysis of the expression of BRCA2, BLM and RAD50 upon vehicle or TGF-β treatment (9 hr, 1 day, 2 day, 3 day or 4 day) in H1650 cells. Phospho-Smad2 and E-cadherin are used as a marker of active TGF-β signaling and MET, respectively. α-Tubulin is used as a loading control. (B) The charts depict the quantification of the relative amounts of BRCA2, BLM and RAD50. Levels of intensity of each band were quantified using imajeJ32 software, represented as a ratio of the protein of interest to α-tubulin, and normalized to levels detected in respective TGF-β untreated (–TGF-β) samples. The data represent the mean ± SD from two independent experiments. p-value *<0.005, ***<0.001, unpaired t-test with Welch’s correction.
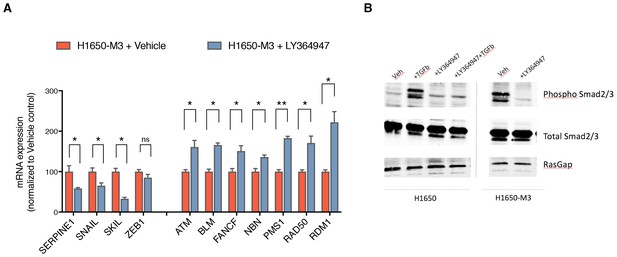
Inhibition of TGF-β signaling in the CD44+/CD24− H1650-M3 cells results in an increased expression of HDR genes.
(A) TGF-β receptor 1 (TGFBR1) kinase activity was blocked by treatment with 1 uM LY364947 (Selleckchem) for 48–72 hr. Expression of TGF-β signature genes were used as a control for the efficacy of LY364947 treatment. mRNA expression was quantified through SYBR-green-based RT-qPCR. Each bar represents the mean ± SD of three replicates from two independent experiments (p-value *<0.05, **<0.005 paired t-test). (B) Western blot analysis of H1650 and H1650-M3 cells treated with LY364947 (1 uM) shows that the inhibitor specifically inhibits TGF-β signaling, as phosphorylation of Smad2 and Smad3 are repressed after 48 hr of treatment.
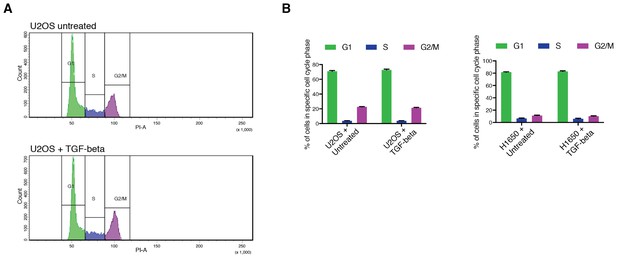
Exposure to TGF-β does not alter the cell cycle distribution in cancer cell lines.
(A) TGF-β treatment does not alter the cell cycle distribution in U2OS cells. The histograms represent U2OS cell cycle distribution without or with TGF-β (1 ng/ml of each of TGF-β1 and TGF-β2, 9 hr). 10,000 cells were analyzed by FACS for each replicate of each sample. (B) Quantification of cell cycle distribution from U2OS and H1650 cells upon 9 hr treatment with TGF-β (1 ng/ml of each of TGF-β1 and TGF-β2). Each bar represents the mean ± SD of three replicates from two independent experiments.
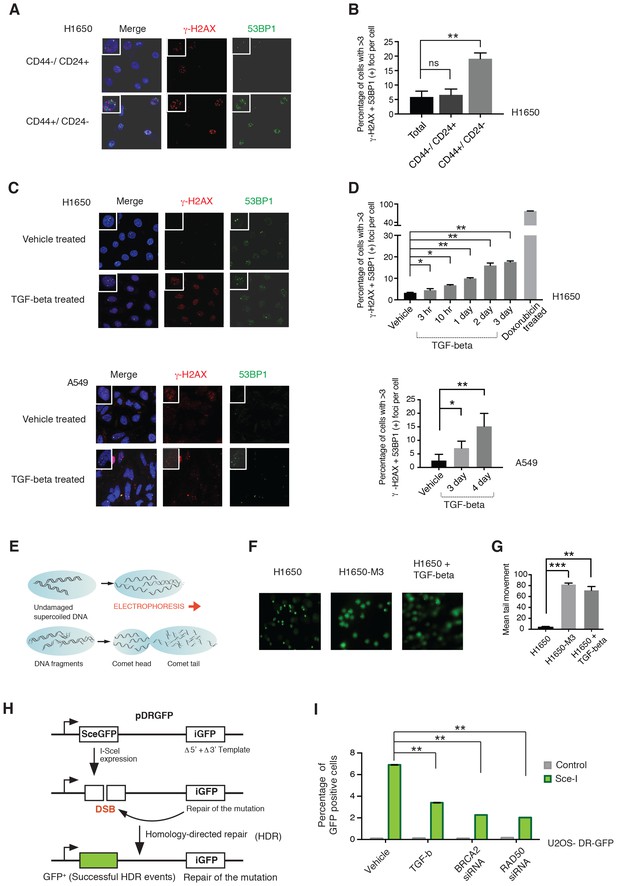
CD44+/CD24− cells are characterized by defect in DNA damage repair.
(A) CD44+/CD24− cells are characterized by increased DNA DSBs compared with CD44−/CD24+ cells. CD44+/CD24− and CD44−/CD24+ cells sorted from H1650 were stained with antibodies against γ-H2AX (red) and 53BP1 (green). DAPI (blue) was used as a counter-stain. Insets in the left upper corner show a representative nucleus. (B) The chart represents quantification of the experiment depicted in (A). Each bar represents the mean ± SD of the percentage of cells with more than three γ-H2AX and 53BP1 double-positive foci per field in CD44−/CD24+ cells and CD44+/CD24− cells, FACS-sorted from the H1650 cell line. Approximately ten fields were counted, for a total of 100 cells (n = 100) (p-value **<0.005, unpaired t-test). (C) H1650 and A549 cells treated with vehicle (DMSO) or TGF-β (1 ng/ml of each of TGF-β1 and -β2, for 4 days) were stained with antibodies against γ-H2AX (red) and 53BP1 (green). DAPI (blue) was used as a counter-stain. Insets in the left upper corner show a representative nucleus staining; analysis of an additional cell line is provided in Figure 4—figure supplement 1. (D) The chart represents quantification of the experiment depicted in (C). Each bar represents the mean ± SD of the percentage of cells with more than three γ-H2AX and 53BP1 double-positive foci per field in vehicle or TGF-β treated cells. Approximately ten fields were counted, for a total of 100 cells (n = 100). (p-value *<0.05, **<0.005, paired t-test). Doxorubicin treatment (10 μM for 24 hr) was used as a positive control. (E) Schematic of Comet assay. (F) Comet assay in H1650, H1650-M3 (CD44+/CD24−) and TGF-β-treated H1650 cells (treatment for 5 days), showing increased DNA strand breaks. (G) The chart represents quantification of the mean ± SD of tail movement of the samples depicted in (F), conducted in three replicates in two independent experiments. (p-value **<0.005, ***<0.0005, unpaired t-test). (H) Schematic of the DR-GFP assay. (I) The chart indicates the percentage of GFP-positive cells upon transfection with pCBASce-I (expressing Sce-I endonuclease) compared with control cells (untransfected cells). siRNA-mediated knock-down of RAD50 and BRCA2 was used as a homologous recombination (HR) efficiency control. Each bar represents mean ± SD of three replicates from two independent experiments (n = 6) (p-value **<0.0005, paired t-test). See Figure 4—figure supplement 3 for knockdown efficiencies with the indicated siRNAs.
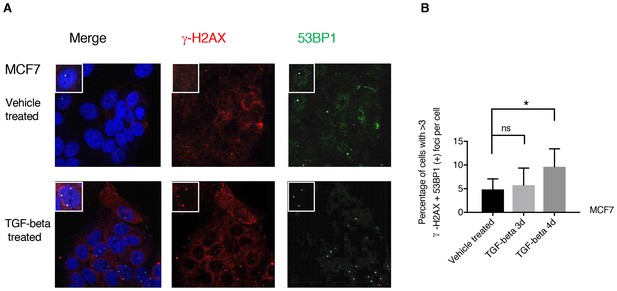
TGF-β treatment increases the number of 53BP1/γ-H2AX double-positive foci in MCF7 cells.
(A) MCF7 cells treated with TGF-β (1 ng/ml each of TGF-β1 and TGF-β2) or DMSO for 3 days and 4 days were stained with antibodies against γ-H2AX (red) and 53BP1 (green). DAPI (blue) was used as a counter-stain. Insets in the left upper corner show a representative nucleus. (B) The chart represents quantification of the experiment depicted in (A). Each bar represents the mean ± SD of the percentage of cells with more than threeγ-H2AX and 53BP1 double-positive foci per field in vehicle or TGF-β-treated cells. Approximately ten fields were counted, for a total of 100 cells (n = 100). (p-value *<0.005, paired t-test).
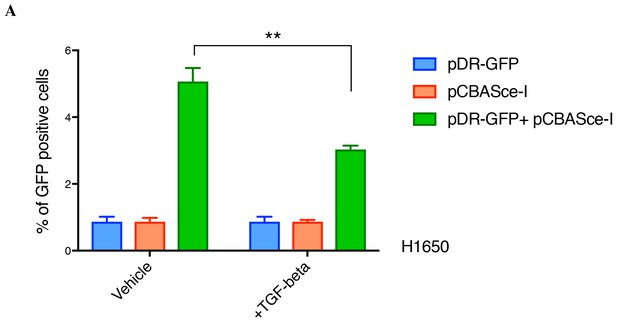
TGF-β reduces the efficiency of homologous recombination in a DR-GFP assay in H1650 cells.
The chart represents the percentage of GFP-positive cells upon transfection with pDR-GFP or pCBASce-I or both, under vehicle or TGF-β treatment. Each bar represents mean ± SD of three replicates from two independent experiments. (p-value **<0.005, unpaired t-test with desired FDR 1%.)
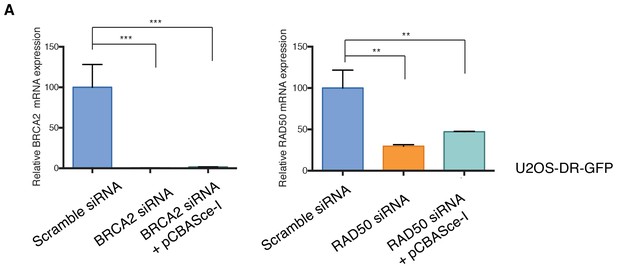
Knockdown efficiencies of the indicated siRNAs in U2OS-DR-GFP cells.
Efficiency of knockdown upon transfection with the indicated siRNA oligonucleotides, without or with pCBASce-I (plasmid expressing Sce-I endonuclease), and compared to scramble-siRNA-transfected control cells. Cells were collected 3 days post-transfection and analyzed for expression of the indicated mRNA. Each bar represents the mean ± SD of three replicates from two independent experiments. p-value **<0.005, ***<0.0005, paired t-test.
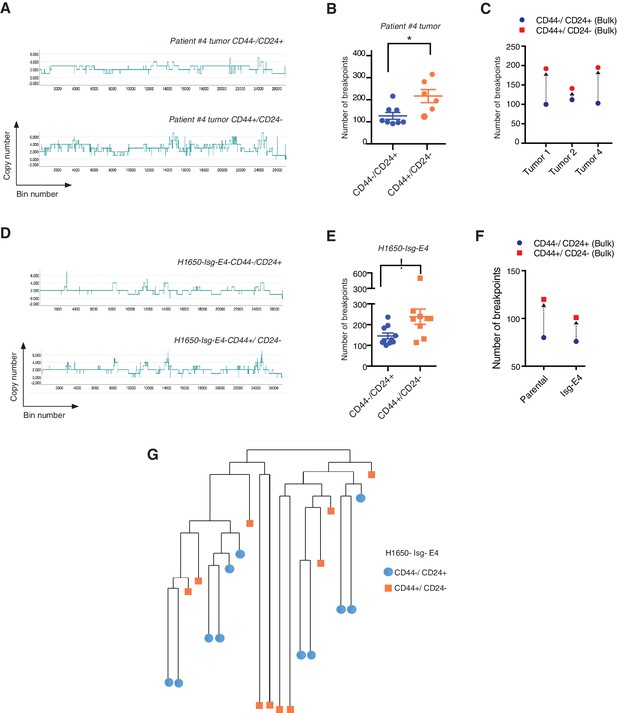
CD44+/CD24− cells have higher copy number alterations and increased genetic diversity.
(A) The graph illustrates a representative copy number profile of CD44−/CD24+ cells and CD44+/CD24− cells sorted from an NSCLC patient (Patient #4). The x-axis corresponds to bins across the genome space from chr1 on the left to the sex chromosomes on the right. The y-axis corresponds to the copy number value at each bin. (B) The chart represents the number of DNA joint points in FACS-sorted CD44−/CD24+ cells (blue circle) and CD44+/CD24− cells (orange circle) from the indicated human NSCLC tumor. Each dot represents the analysis of a single cell. The breakpoint matrix (utilized to calculate DNA joint points), cluster dendogram and heat-map of normalized read counts (Figure 5—figure supplement 1A) were generated using Ginkgo, an open-source web platform for interactive analyses of CNA. (Garvin et al., 2015) (http://qb.cshl.edu/ginkgo). A variable bin size of 175 kb is used. p-value * < 0.05, unpaired t-test with Welch’s correction. Error bars indicate standard deviation. (C) The chart represents number of DNA joint points in CD44−/CD24+ (blue circle) and CD44+/CD24− (red circle) cells FACS-sorted from the indicated primary human NSCLC. Each dot represents the analysis of a cell type collected and sequenced in bulk. (D) The graph illustrates a representative copy number profile of one CD44−/CD24+ and one CD44+/CD24− FACS-sorted cell from the H1650-derived isogenic cell line H1650-Isg-E4. (E) The chart represents the number of DNA joint points in FACS-sorted CD44−/CD24+ cells (blue circle) and CD44+/CD24− cells (orange squares) from the H1650-Isg-E4 cell line. Each dot represents the analysis of a single cell. The breakpoint matrix (utilized to calculate DNA joint points) is generated along with the cluster dendogram and heat-map of normalized read-counts (Figure 5—figure supplement 1B) using Ginkgo. A variable bin size of 175 kb is used. p-value *<0.05, unpaired t-test with Welch’s correction. Error bars indicate standard deviation. (F) The chart depicts the number of DNA joint points in CD44−/CD24+ (blue circle) and CD44+/CD24− (red square) cells FACS-sorted from the H1650 (parental) and H1650 isogenic Isg-E4 cell lines. Each dot represents the analysis of a cell type collected and sequenced in bulk. (G) Cluster dendogram of normalized read-counts across segment breakpoints (using Euclidian distance and the ward-clustering method) of CD44−/CD24+ cells (blue circle) and CD44+/CD24− cells (orange squares) FACS-sorted from the H1650-Isg-E4 cell line. Each dot represents a single cell. The cluster dendogram is generated with Ginkgo.
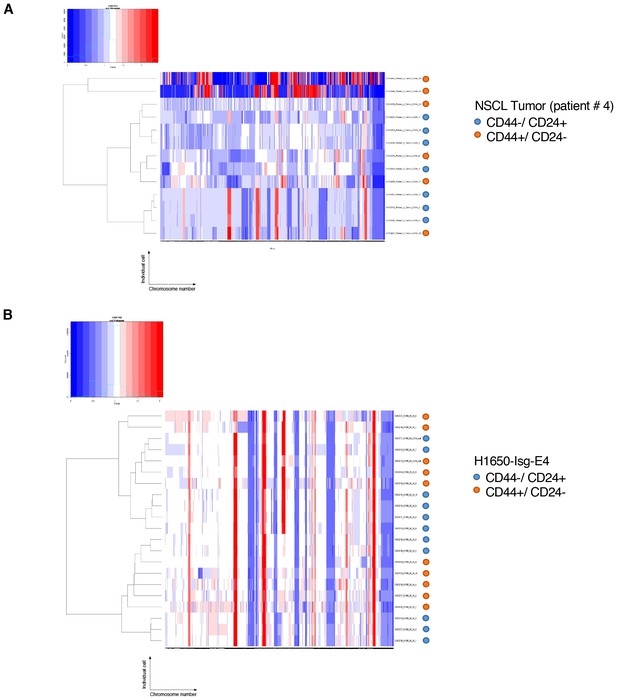
Heat-map of normalized read counts of FACS-sorted cells based on CD44 and CD24 surface markers from patient tumor and from the H1650-derived isogenic cell line H1650-Isg-E4.
Heat-maps of normalized read counts across segment breakpoints (using Euclidian distance and ward clustering method) of CD44+/CD24− cells and CD44−/CD24+ cells sorted from (A) an NSCL tumor (patient #4) and (B) the H1650-derived isogenic cell line (H1650-isg-E4) are displayed. CD44−/ CD24+ cells are marked with a blue circle and CD44+/CD24− cells are marked with an orange circle. The cluster dendogram and heat-map of normalized read counts were generated using Ginkgo, an open-source web platform for interactive analyses of CNA (Garvin et al., 2015) (http://qb.cshl.edu/ginkgo).
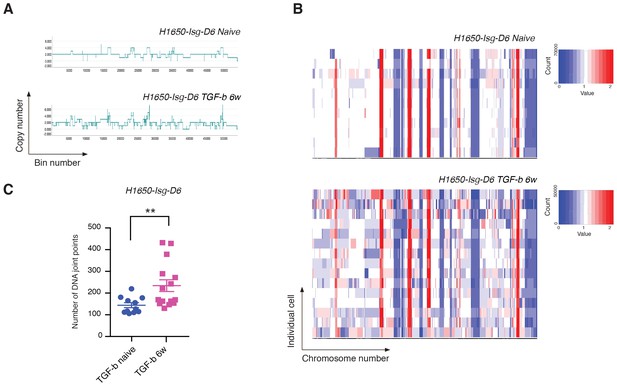
Exposure to TGF-β is sufficient to increase CNA and genetic diversity of the cell population.
(A) The graph illustrates a representative copy-number profile of one TGF-β-naïve cell and one TGF-β-treated (TGF-b 6w) cells from the H1650 isogenic/single cell-derived cell line Isg-D6. H1650-Isg-D6 was treated with vehicle (DMSO) or TGF-β (1 ng/ml of each of TGF-β1 and -β2) for six weeks. Single cell CNA analysis was performed upon TGF-β withdrawal. The x-axis corresponds to bins across the genome space from chr1 on the left to the sex chromosomes on the right. The y-axis corresponds to the copy number value at each bin. (B) Heat map of normalized read counts of TGF-β-naïve and TGF-β-treated (TGF-b 6w) cells from the H1650-Isg-D6 cell line, across segment breakpoints with a variable bin size of 50 kb (using Euclidian distance and the ward clustering method). Each horizontal line across the y-axis represents an individual cell, whereas the x-axis annotates the CNA across chromosomes from chr1 on the left to the sex chromosomes on the right. (C) The chart represents the number of DNA joint points in TGF-β-naïve (blue circle) and TGF-b 6w (purple squares) cells from the H1650-Isg-D6 cell line. Each dot represents the analysis of a single cell. The breakpoint matrix (utilized to calculate DNA joint points) is generated along with the cluster dendogram and heat-map of normalized read-counts (as shown in (B)) using Ginkgo with a variable bin size of 50 kb (http://qb.cshl.edu/ginkgo). p-value **=0.0064, unpaired t-test with Welch’s correction.
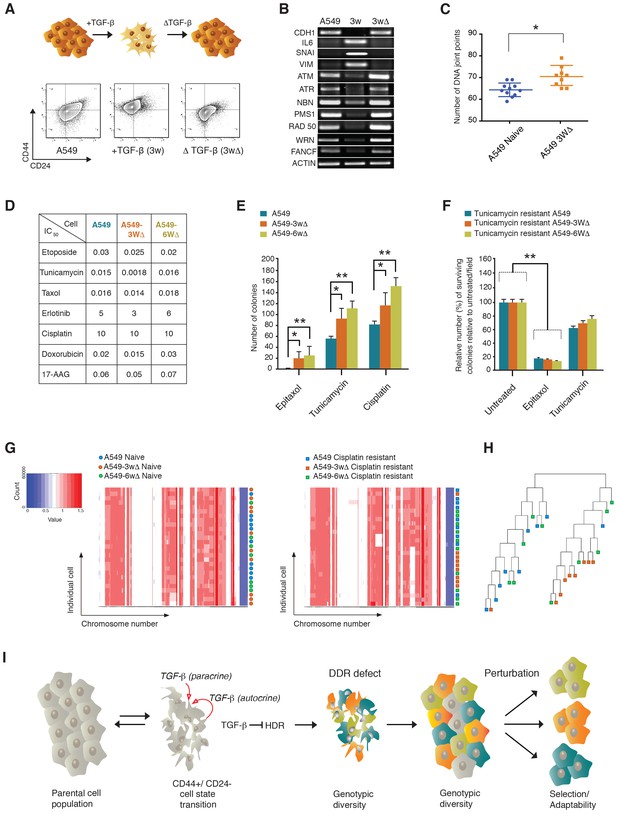
The TGF-β-induced CD44+/CD24− cell state increases the adaptability of cell populations.
(A and B) A549 cells exposed to TGF-β acquire phenotypic and molecular changes characteristic of a CD44+/CD24− cell state. Upon TGF-β withdrawal, the cells return to their original cell state, as indicated by the FACS analysis in (A) and the RT-PCR analysis in (B). See Figure 7—figure supplement 1 for quantification of FACS analysis. (C) The chart indicates the number of DNA joint points in TGF-β-naïve and TGF-β-treated A549 cells. Each dot represents the analysis of a single cell. The breakpoint matrix (utilized to calculate DNA joint points) is generated using Ginkgo (http://qb.cshl.edu/ginkgo). A variable bin size of 175 kb is used. p-value *<0.05, unpaired t-test. (D) The table depicts the IC50 values of A549, A549-3W△ and A549-6W△ cells in the context of treatment with the indicated drugs. (E and F) TGF-β treatment increased the adaptability of cells. Cells that were transiently exposed to TGF-β for 3 or 6 weeks (A549-3W△, A549-6W△) were then treated with the indicated drugs upon TGF-β withdrawl. (E) The number of colonies (mean ± SD) that have survived epitaxol (1.6 μM), tunicamycin (3.2 μM) and cisplatin (1 mM) treatment. Notably, the concentration of drugs used in this experiment corresponds to approximately >100X the IC50. Two independent experiments, each with three replicates, were carried out and approximately 50 fields were counted for each sample. (p-value *<0.05, **<0.005, paired t-test.) (F) A549 tunicamycin-resistant clones were grown in regular/drug-free medium for a week and then retested for sensitivity to tunicamycin (3.2 μM) or epitaxol (1.6 μM). The plot represents mean ± SD number of colonies surviving 5 days after treatment compared with untreated cells, from two independent experiments each with three replicates (p-value **<0.005, paired t-test). (G) Heat map of normalized read counts across segment breakpoints (using Euclidian distance and the ward-clustering method) of the indicated cells. Each horizontal line across the y-axis represents an individual cell, whereas the x-axis annotates the CNA across chromosomes from chr1 on the left to the sex chromosomes on the right. A heat map of cisplatin-naïve cells is shown on the left and of cisplatin-resistant cells on the right. (H) Cluster dendogram of normalized read counts across segment breakpoints (using Euclidian distance and the ward-clustering method) of cisplatin-resistant-A549, cisplatin-resistant-A549-3W△ and cisplatin-resistant-A549-6w△. The cluster dendogram and heat-map of normalized read counts were generated using Ginkgo. (I) Schematic of proposed model. When cells transit into a CD44+/CD24− state, they acquire mesenchymal-like features and autocrine secretion of TGF-β that leads to the downregulation of HDR genes. This process results in a hyper-mutable phenotype that spurs genetic diversity and intra-tumor clonal heterogeneity. Consequently, following a Darwinian model of cancer evolution, the transition of cancer cells into a CD44+/CD24− state or exposure to TGF-β leads to an increased adaptability to any given perturbation.
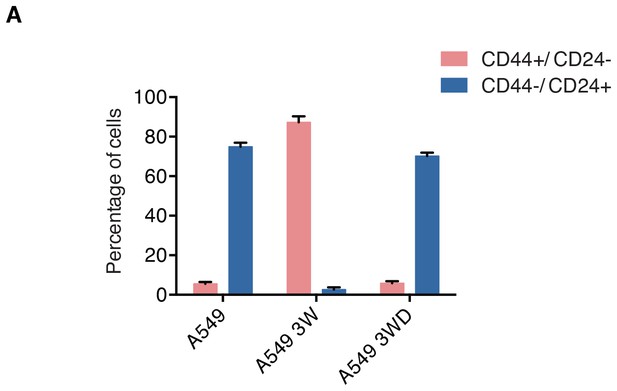
A549 cells undergo changes in the surface expression of CD44 and CD24 before, during and after exposure to TGF-β.
The chart depicts the percentage of of A549 cells in the CD44+/CD24− state and the CD44−/ CD24+ state, before, during and post exposure to TGF-β. A549 cells before, during and post exposure to TGF-β were stained for surface expression of CD44 and CD24 and analyzed by FACS. Each bar represents the mean ± SD of three replicates from three independent experiments. 10,000 cells were analyzed for each replicate of each sample.
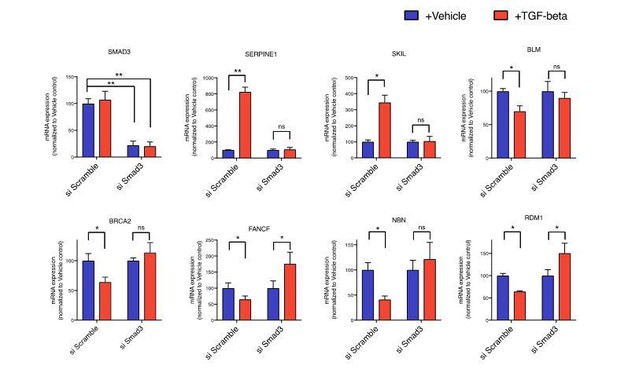
mRNA expression analysis of SMAD3, SERPINE1, SKIL and the indicated HDR genes in vehicle or TGF-β-treated cells relative to vehicle control upon siRNA mediated SMAD3 knock down.
si Scramble was used as an experimental control. mRNA expression was quantified by SYBR-green-based RT-qPCR. Each bar represents mean ± SD of 3 replicates from 2 independent experiments. (p-value * < 0.05, **< 0.005 paired t-test).
Additional files
-
Supplementary file 1
Mutation analysis of the following cells lines (used in our study) from
COSMIC (Catalogue Of Somatic Mutations In Cancer). http://cancer.sanger.ac.uk/cosmic
- https://doi.org/10.7554/eLife.21615.031
-
Supplementary file 2
Relative abundance of DNA-damage/-repair pathway components from gene expression profiles of H1650 and H1650-M3 cells.
- https://doi.org/10.7554/eLife.21615.032
-
Supplementary file 3
Sequences of primers used in quantitative-RT-PCR in this study.
- https://doi.org/10.7554/eLife.21615.033





