Single-molecule FRET unveils induced-fit mechanism for substrate selectivity in flap endonuclease 1
Figures
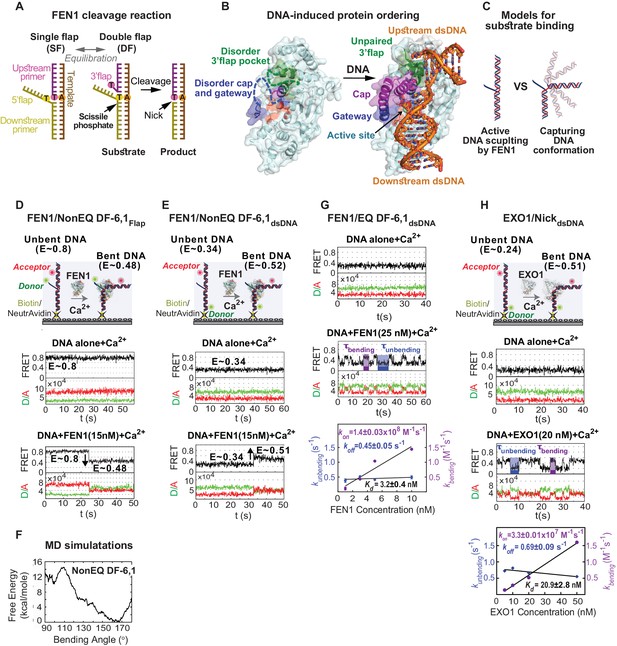
Active junction bending by structure-specific 5’nucleases.
(A) FEN1 cleavage reaction. Schematic showing the equilibration of a flap substrate junction from a single- to a double-flap and its subsequent cleavage by FEN1 to generate a nick that can be sealed by DNA ligase 1. (B) Ordering of FEN1 upon DNA binding. FEN1 alone (1ULI.pdb) (Sakurai et al., 2005) and in complex with bent DNA (3Q8L.pdb) (Tsutakawa et al., 2011), highlighting the various structural features of FEN1 and the regions that undergo through disorder-to-order transitioning upon DNA binding. (C) Active DNA versus DNA conformational capturing models for forming the FEN1 complex with the bent DNA conformer. Monitoring DNA bending of FEN1 and non-equilibrated DF-6,1 using the flap-labeling scheme (NonEQ DF-6,1Flap) (D) and internal labeling-scheme (NonEQ DF-6,1dsDNA) (E). For each labeling, a schematic of the donor and acceptor positions (upper panel) and smFRET time traces of the substrate alone (middle panel) and in presence of FEN1 (lower panel) are shown; change in FRET upon DNA bending in each labeling scheme is highlighted. (F) Analysis of the structure of NonEQ DF-6,1 by MD simulations. The effective free energy profile (PMF) from adaptive biasing force calculations is shown. (G) Bending of equilibrated DF-6,1 (EQ DF-6,1dsDNA) by FEN1. smFRET time traces of EQ DF-6,1dsDNA alone (upper panel) and in the presence of FEN1 (middle panel) and analysis of its DNA bending association rate constant (kon-bending) and dissociation rate constant (koff-unbending) (lower panel) are shown. kbending and kunbending were calculated by fitting an exponential function to the histogram from the population of dwell times of bent (τbending) and unbent (τunbending) conformers, respectively; error bars correspond to the standard deviation of the fit. kon-bending and koff-unbending are calculated from the slope of kbending from a linear regression fit and the mean of kunbending, respectively; the error bars correspond to the standard deviation of the fit. Kd-bending = koff-unbending/kon-bending. (H) Bending of nicked substrate using the internal labeling scheme (NickdsDNA) by EXO1. A schematic of the donor and acceptor positions (upper panel), smFRET time traces of NickdsDNA alone and in the presence of EXO1 (middle panels) and analysis of its kon-bending, koff-unbending and Kd-bending (lower panel) is presented. Donor and acceptor are at identical positions to those in DF-6,1dsDNA in Figure 1E. kon-bending, koff-unbending and Kd-bending were calculated as in 1G. All TIRF-smFRET experiments were acquired at 100 ms.
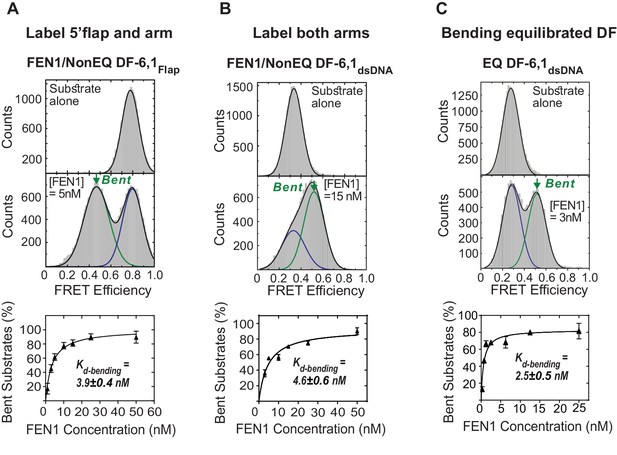
Bending of equilibrated and non-equilibrated DF-6,1 by FEN1.
(A) TIRF-based single-molecule FRET (smFRET) of FEN1 and NonEQ DF6,1Flap. Histograms of NonEQ DF6,1Flap alone (high FRET) and in the presence of FEN1 (low FRET) (upper panel). Kd-bending (lower panel) is calculated by fitting the percentage of the bent conformer from the FRET histograms at various concentrations of FEN1 with a non-linear least squares regression; the percentages of unbent and bent DNA are calculated by fitting two Gaussians. The uncertainty in calculating the percentage of bent DNA and Kd-bending correspond to the standard deviation of triplicate measurements and the non-linear least squares regression fit, respectively. (B) Kd-bending of FEN1 on NonEQ DF-6,1dsDNA. Kd-bending is calculated as described in Figure 1—figure supplement 1A. (C) Kd-bending of FEN1 on EQ DF-6,1dsDNA. Kd-bending is calculated as described in Figure 1—figure supplement 1A.
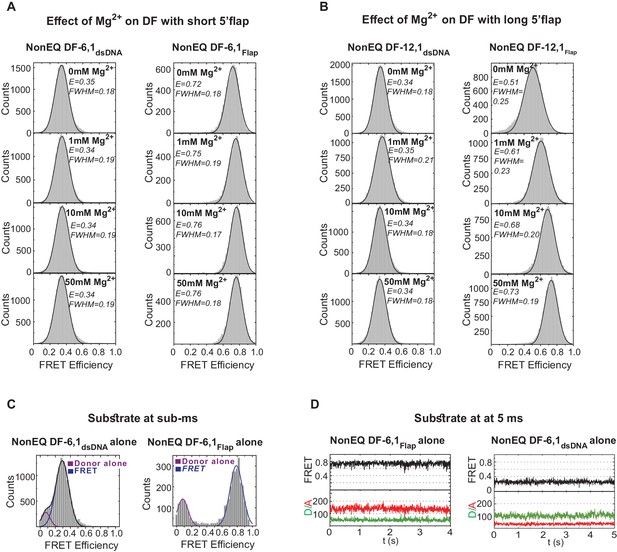
Flap substrates exist as a stable extended conformer.
(A) Effect of various Mg2+ concentration on DF-6,1. TIRF-smFRET histograms of NonEQ DF6,1dsDNA (left panel) and NonEQ DF6,1Flap (right panel) with increasing concentration of MgCl2 (0 mM, 1 mM, 10 mM and 50 mM). The insensitivity of both labeling schemes to increasing divalent metal ion concentrations demonstrates that both labeling schemes report directly on the geometry of the duplex DNA. (B) Effect of various Mg2+ concentration on DF-12,1. TIRF-smFRET histograms of NonEQ DF12,1dsDNA (left panel) and NonEQ DF12,1Flap (right panel) with increasing concentration of MgCl2 (0 mM, 1 mM,10 mM and 50 mM). The sensitivity of the flap-labeling scheme to varying divalent metal ion concentrations but not the internal-labeling scheme demonstrates that the geometry of the duplex DNA is not influenced by the length of the 5’flap and that the flap-labeling scheme is inappropriate to describe the geometry of the duplex DNA only when the 5’flap length exceeds 6 nt. All TIRF-smFRET measurements were acquired at 160 ms temporal resolution. FWHM represents the full width at half maximum of the Gaussian peak. (C) Burst confocal-smFRET histograms from freely diffusing DNA in solution of NonEQ DF6,1dsDNA (0.5 nM) (upper panel) and NonEQ DF6,1Flap (0.5 nM) (lower panel) acquired at sub-ms temporal resolution. No enrichment of other FRET conformers was observed upon increasing the temporal resolution from 160 ms (shown in Figure 1—figure supplement 1A,B) to sub-ms. (D) Confocal-smFRET time traces of surface-immobilized NonEQ DF6,1Flap (upper panel) and NonEQ DF6,1dsDNA (lower panel) with 5 ms temporal resolution.
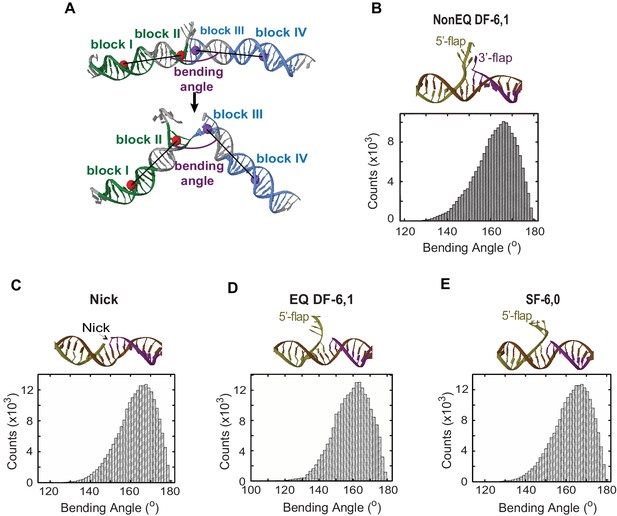
MD simulations of the conformational states and DNA bending energy of nick and various flap structures.
(A) Definition of the DNA bending angle. The bending angle is calculated between two vectors. One vector was defined by using the center of the masses of two blocks of nucleotides: block I (green) and block II (green) and one vector was defined with the center of mass of block III (blue) and block VI (blue). (B, C, D and E) are the averaged structure (upper panel) and histogram of the DNA bending angle (lower panel) for NonEQ DF-6,1, Nick, EQ DF-6,1 and SF-6,0, respectively, taken from the MD simulations. These MD simulations demonstrated that the extended DNA structure was the most energetically favorable conformer in all tested substrates and that these substrates could sample bent conformers up to 140°, which are equivalent to those observed for dsDNA (Sharma et al., 2013).
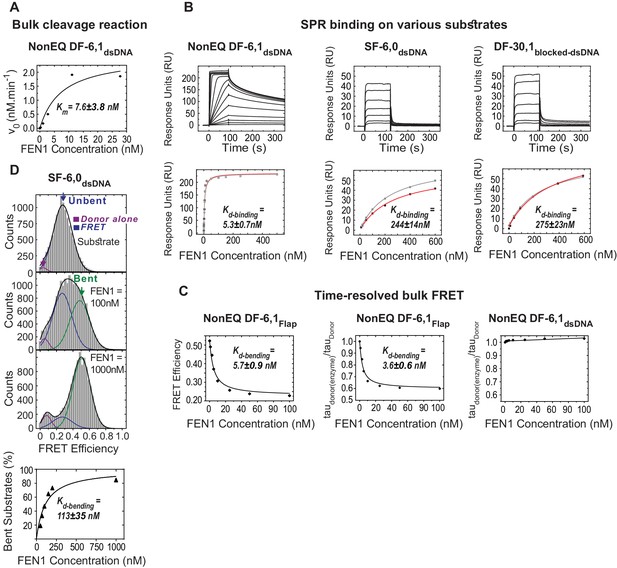
Bulk cleavage, SPR binding and time-resolved bulk FRET of selected substrates.
(A) Cleavage of NonEQ DF-6,1dsDNA (0.5 nM) by bulk assays. A plot of initial rates (v0, nM.min−1) in relation to the FEN1 concentration fitted with a generalized non-linear least-squares regression using a Michaelis-Menten model. The values for v0 were estimated using bulk cleavage assays with different time intervals as described in the Materials and methods. This plot was used to determine the steady-state Km. Uncertainty in Km corresponds to the standard error of the fit. (B) SPR binding studies of FEN1 and NonEQ DF-6,1dsDNA (left panel), SF-6,0dsDNA (middle panel) and DF-30,1blocked-dsDNA (right panel). The sensorgram of binding of increasing concentrations of FEN1 is shown (left panel). The maximum response units (RU) reached at each FEN1 concentration were fitted using the steady-state affinity model to obtain the equilibrium dissociation binding constant (Kd-binding) (lower panel). The uncertainty corresponds to the standard deviation of N = 2 runs. 170 RU, 70 RU and 90 RU of NonEQ DF-6,1dsDNA, SF-6,0dsDNA and DF-30,1blocked-dsDNA were immobilized on the surface, respectively. (C) Bulk-FRET measurements of NonEQ DF-6,1. The FRET efficiency of the donor-acceptor pair in NonEQ DF-6,1Flap at different FEN1 concentrations (left panel) and relative fluorescence lifetimes (taudonor(enzyme)/taudonor) at different FEN1 concentrations without acceptors (middle panel) are shown. The results show that FEN1 binding influenced the fluorescence intensity of the donor in the flap-labeling scheme. Kd-bending after correcting for the effect of the donor was similar to that obtained from uncorrected apparent FRET (Figure 1—figure supplement 1C); Kd-bending was calculated using a standard quadratic equation for the simple bimolecular association as described in Materials and methods and the uncertainty corresponds to the standard error of the fit. The uncertainty corresponds to the standard error of the fit. Bulk-FRET measurements of NonEQ DF-6,1dsDNA (right panel). The relative fluorescence lifetime (taudonor(enzyme)/taudonor) OF DF-6,1dsDNA at different FEN1 concentrations without acceptors (right panel), showing no effect on the donor fluorescence intensity upon FEN1 binding in the internal-labeling scheme. (D) Burst confocal-smFRET histograms from freely diffusing substrate missing the 3’flap but containing the 5’flap (SF6,0dsDNA) (0.5 nM) in solution acquired at sub-ms temporal resolution (upper panel). Kd-bending (lower panel) calculated as described in Figure 1—figure supplement 1A.
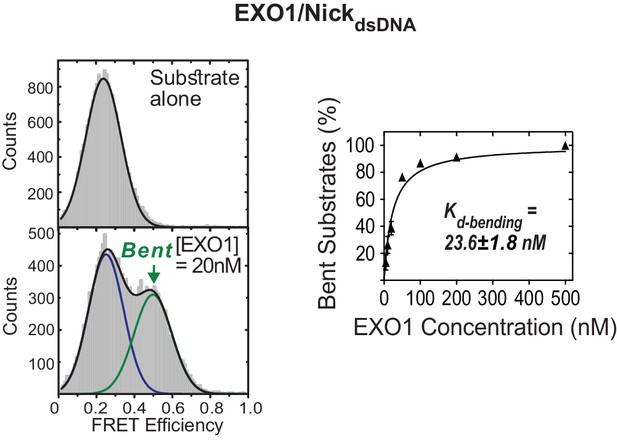
Active bending of nicked DNA by EXO1.
TIRF-smFRET histograms of NickdsDNA alone and in the presence of EXO1 (left panel) and Kd-bending (right panel). Kd-bending is calculated as described in Figure 1—figure supplement 1A. The donor and acceptor in NickdsDNA are at identical positions to those in NonEQ DF6,1dsDNA.
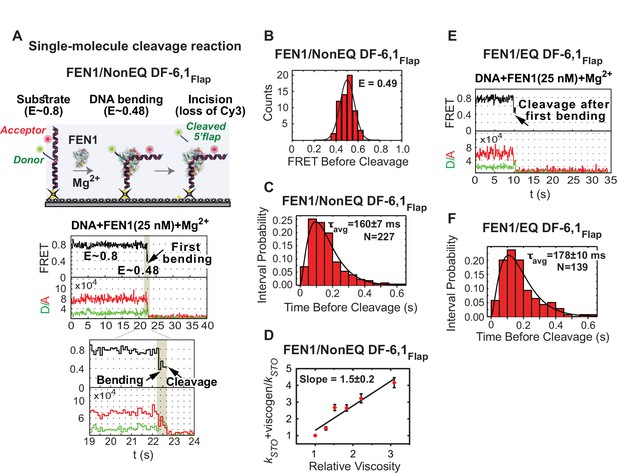
Cleavage of cognate substrate by FEN1.
(A) Cleavage of NonEQ DF-6,1Flap. Schematic showing the simultaneous monitoring of DNA bending and 5’flap cleavage at the single-molecule level (upper panel). A representative smFRET time trace with a zoomed-in view showing the cleavage of NonEQ DF-6,1Flap in which FEN1 never misses the opportunity to bend the DNA and cleave it (lower panel). (B) FRET of the bent state before cleavage of NonEQ DF-6,1Flap fitted with a Gaussian distribution from multiple cleavage events. (C) Dwell times of the bent state prior to cleavage of NonEQ DF-6,1Flap fitted with a gamma distribution to calculate average dwell time (τavg) from the number of independent experiment N = 3; the uncertainty corresponds to the standard error of the mean. Single turnover kcat (kSTO) is = 1/τavg. Cleavage was performed at 50 ms temporal resolution. (D) Effect of low molecular weight viscogen (glycerol) on disorder-to-order transitioning in FEN1. Graph showing relative kSTO of NonEQ DF6,1Flap cleavage upon addition of glycerol at increasing relative viscosity from N = 2–3 fitted with a linear regression to calculate the slope of the curve; the error corresponds to the standard error of the mean. kSTO was determined as in Figure 2C. (E) A representative smFRET time trace showing the cleavage of EQ DF-6,1Flap in which FEN1 never misses the opportunity to bend the DNA and cleave it. (F) A histogram showing the distribution of dwell times of the bent state prior to cleavage of EQ DF6,1Flap. τavg from N = 3 is calculated as in Figure 2C at a temporal resolution of 50 ms.
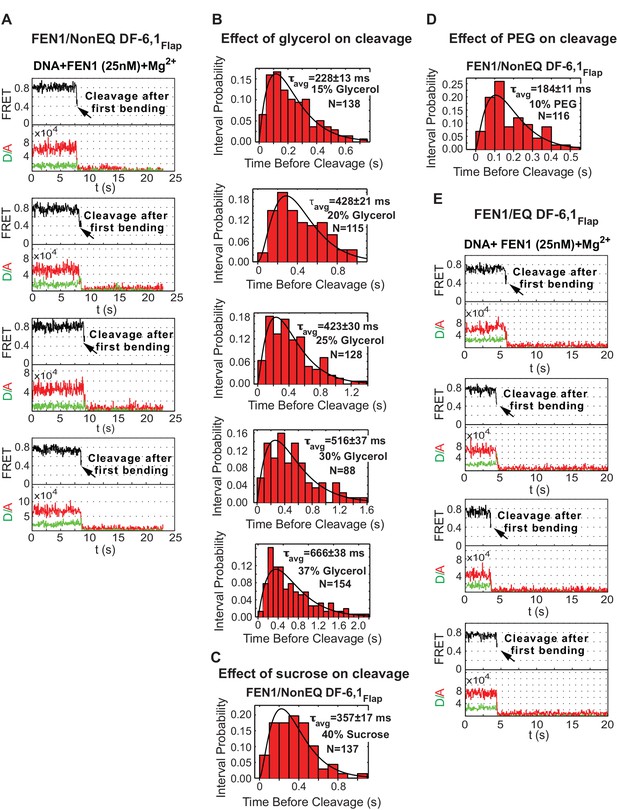
Single-molecule cleavage of cognate substrates and probing conformational changes to FEN1 by viscogens.
(A) Representative TIRF-smFRET time traces showing the cleavage of NonEQ DF6,1Flap in which FEN1 always bends and cleaves the DNA. (B) Histograms showing the distribution of dwell times of bent state prior to the cleavage of NonEQ DF6,1Flap by FEN1 in the presence of varying concentrations of glycerol. For each concentration, the histogram was fitted and τavg was calculated from N = 4 as described in Figure 2C. (C and D) Histograms showing the distribution of dwell times of the bent state prior to the cleavage of NonEQ DF6,1Flap by FEN1 in the presence of 40% sucrose or 10% PEG-8000, respectively. The histogram was fitted and τavg was calculated from N = 4 as described in Figure 2C. (E) Representative TIRF-smFRET time traces showing the cleavage of NonEQ DF6,1Flap in which FEN1 always bends and cleaves the DNA.
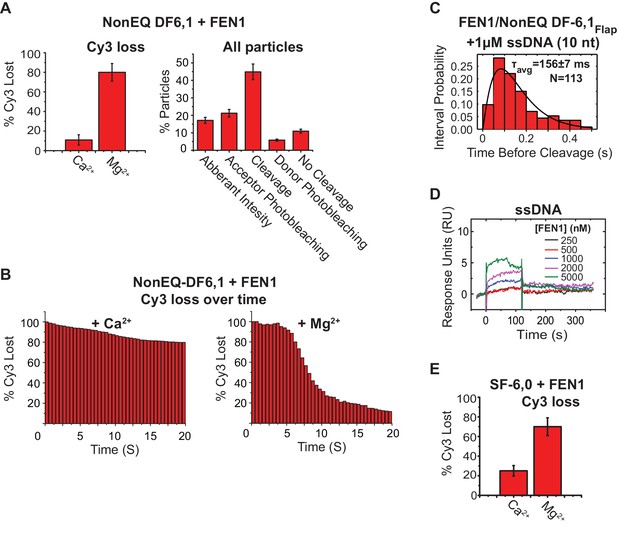
Controls for assigning donor loss to the cleavage and immediate departure of the 5’flap.
(A) (Left panel) bar chart comparing the Cy3 donor signal lost in cognate substrate due to photobleaching (+Ca2+) or to both photobleaching and incision (+Mg2+) in presence of FEN1. The donor loss increases significantly in the presence of Mg2+ as compared to what would be expected for its loss in the presence of Ca2+. (Right panel) The detailed classification of all single-molecule time traces in the field of view for cognate substrate in the presence of Mg2+ as explained in Materials and methods (right panel). This shows that most of the traces exhibited donor loss instantaneously only upon FEN1 bending, demonstrating that donor loss in the presence of Mg2+ is due 5’flap cleavage by FEN1. The uncertainty corresponds to the standard deviation between multiple movies in the presence of either CaCl2 or MgCl2. (B) Time-dependent loss of Cy3 molecules under binding (Ca2+) or cleavage (Mg2+) conditions, showing that sudden loss of Cy3 coincides with FEN1 introduction into the focal volume in the case of Mg2+, while Cy3 loss in the case of Ca2+ is minimal and not abrupt. (C) Histograms showing the distribution of dwell times of the bent state prior to the cleavage of NonEQ DF6,1Flap by FEN1 in the presence of excess 10 nt ssDNA as a competitor, showing no effect on the cleavage dwell time when compared to cleavage of NonEQ DF6,1Flap by FEN1 (Figure 2C). The histogram was fitted and τavg was calculated from N = 2 as described in Figure 2C. (D) SPR binding of FEN1 to 22 nt ssDNA, showing only residual binding of FEN1 at extreme concentrations of FEN1. (E) Similar analysis to that described in Figure 2—figure supplement 2A (left panel), showing that most of the loss of the donor molecules in the noncognate substrate SF-6,0 resulted from cleavage by FEN1 in the presence of Mg2.
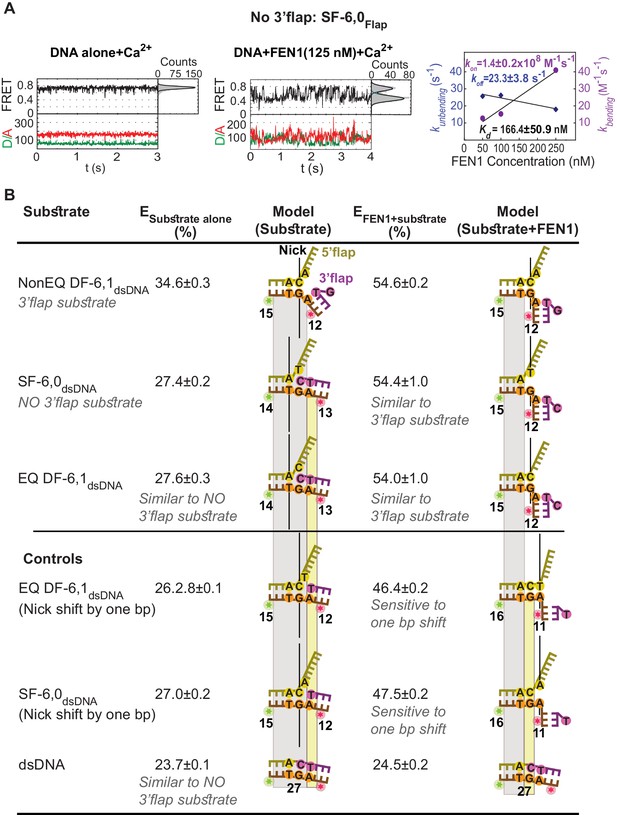
Active sculpting of the 3’end of a nick junction creates a 3’flap and drive protein ordering.
(A) Confocal-based smFRET time traces of surface-immobilized SF6,0Flap alone (left panel) and in the presence of FEN1 (middle panel) acquired at 5 ms, showing rapid transitions from high to low FRET upon DNA bending. kon-bending and koff-unbending of SF6,0Flap by FEN1 (right panel) calculated as in Figure 1G. (B) FEN1 actively creates a 3’flap at the nick junction of cognate and noncognate substrates. Determining the status of the 3’flap in equilibrated DF and SF junctions by comparing the FRET states of various nick-junction positions in NonEQ DF6,1dsDNA, EQ DF6,1dsDNA and SF6,0dsDNA in the absence or presence of FEN1. 0.5 nM DNA and saturating concentrations of FEN1 were used (1000 nM for SF6,0dsDNA and its one base pair shift construct and 200 nM for the remaining constructs). FRET values were determined by fitting the burst confocal-smFRET histograms from freely diffusing DNA at sub-ms temporal resolution with a Gaussian. FRET is reported as a percentage, and the uncertainty corresponds to the standard deviation of N = 3.
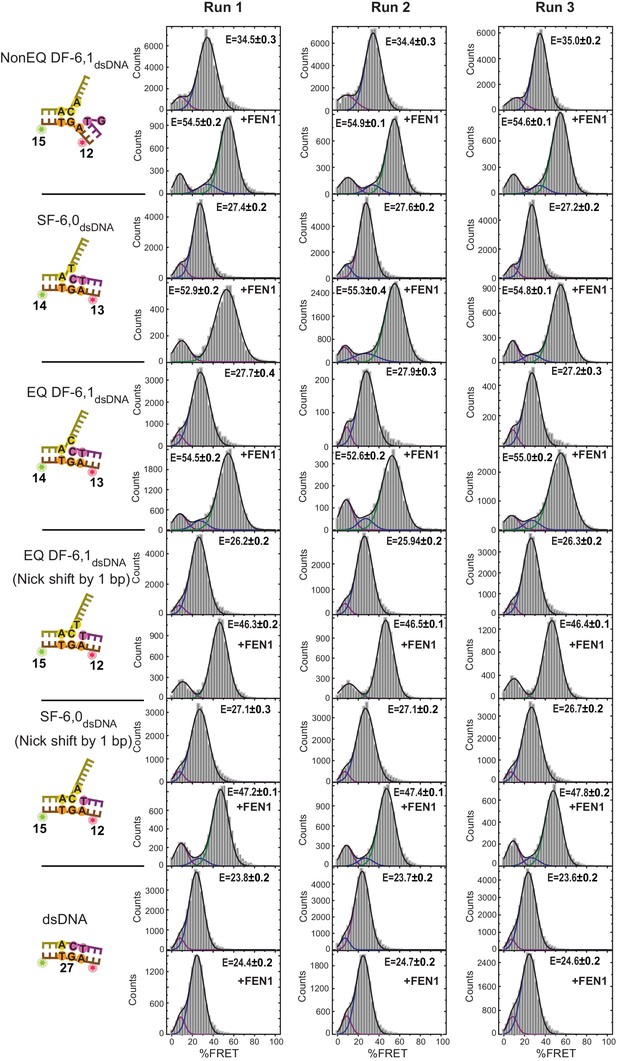
All histograms corresponding to the data shown in Figure 3B.
Histograms were fit with Gaussian distribution to get the mean of the distribution. Uncertainties correspond to the error of the fit.
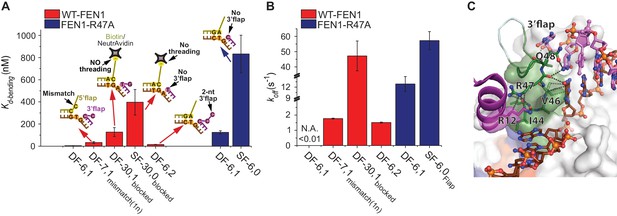
Verification of the bent DNA conformer by the 3’flap-induced protein ordering.
(A) Bar chart comparing Kd-bending for FEN1-WT or FEN1-R47A on various non-equilibrating flap substrates using the internal labeling scheme. Used noncognate substrates include SF-6,0, DF containing 1 nt mismatch at the nick junction (DF-7,1mismatch(1nt)), DF containing biotin-NeutrAvidin on the 5’flap to block 5’flap threading (DF-30,1blocked) and its SF version (SF-30,0blocked), and DF containing 2 nt 3’flap (DF-6,2). (B) Bar chart comparing koff-unbending for FEN1-WT or FEN1-R47A on various non-equilibrating flap substrates using the internal labeling scheme. The lower estimate of koff-unbending for FEN1-WT on DF-6,1 corresponds to the 60 s acquisition time where transitions were rarely detected. Kd-bending and koff-unbending are calculated as in Figure 1—figure supplement 1A and Figure 1G, respectively. koff-unbending was determined from multiple FEN1 concentrations except for FEN1-R47A on SF-6,0 and FEN1 on DF-7,1mismatch(1nt), which were determined from two and one concentration, respectively. The smFRET technique and temporal resolutions used in Figure 4A,B are described in Figure 4—figure supplement 1. (C) R47 acts as a sensor that couples structuring of the 3’flap-binding pocket and the cap-helical gateway. R47 in the hydrophobic wedge mediates multiple interactions, where it stacks against the first base pair on the 3’flap side of the junction while its side chain C-caps the α2 in the gateway (highlighted in green) and stacks with K128 on α5 in the cap (highlighted in purple) (3Q8L.pdb) (Tsutakawa et al., 2011).
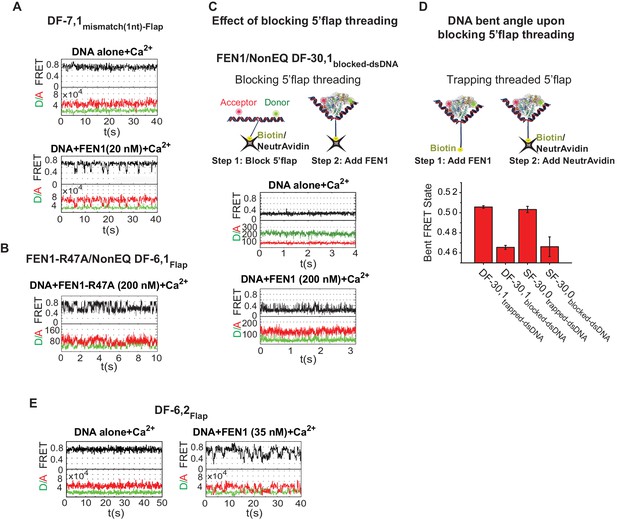
Bending kinetics of various noncognate substrates.
(A) TIRF-smFRET time traces of DF-7,1mismatch(1nt)-Flap alone and in the presence of FEN1. (B) Surface-immobilized confocal-smFRET time traces of NonEQ DF-6,1Flap in the presence of FEN1-R47A at 5 ms temporal resolution. (C) The effect of blocking 5’flap threading on DNA bending by FEN1. Schematic showing the strategy used to block 5’flap threading into the cap-helical gateway by introducing NeutrAvidin/biotin linkage at the 5’end of the 5’flap of DF-30,1 (termed of DF-30,1blocked-dsDNA) prior to the addition of FEN1 (upper panel). Surface-immobilized confocal-smFRET time traces of DF-30,1blocked-dsDNA alone and in the presence of FEN1 at 5 ms temporal resolution (lower panel). The substrate was immobilized by surface-coated-NeutrAvidin via the biotin group on the 5’flap. (D) A bar chart comparing final bent FRET states of DF-30,1trapped-dsNDA, DF-30,1blocked-dsDNA, SF-30,0trapped-dsDNA and SF-30,0blocked-dsDNA using burst confocal-smFRET histograms from freely diffusing substrates acquired at sub-ms temporal resolution. FEN1 concentrations were 5000 nM for SF-30,0internal-blocked, 1000 nM for DF-30,1internal-blocked,1000 nM for SF-30,0internal-trapped and 200 nM for DF-30,1internal-trapped. To trap a threaded 5’flap, FEN1 was first pre-incubated with the substrate before NeutrAvidin was added to bind the biotin on the 5’flap (as shown in the schematic in the upper panel). The FRET value in each condition represents the average of N = 3 and the uncertainty corresponds to the standard error of their fits. (E) TIRF-smFRET time traces of DF6,2Flap alone and in the presence of FEN1.
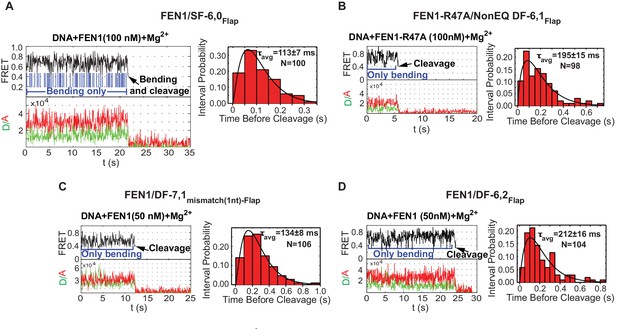
Cleavage of noncognate substrates by FEN1.
(A, B, C and D) Representative smFRET time traces showing the cleavage of FEN1 on SF6,0Flap, FEN1-R47A on NonEQ DF6,1Flap, FEN1 on DF-7,1mismatch(1nt)-Flap and FEN1 on DF-6,2Flap, respectively and the distribution of dwell times of the bent state prior to cleavage (τavg). Cleavage occurs after a random number of missed cleavage opportunities from a bent conformer as illustrated in blue arrows in Figure 5A. τavg is calculated from N = 4 as in Figure 2C at a temporal resolution of 50 ms.
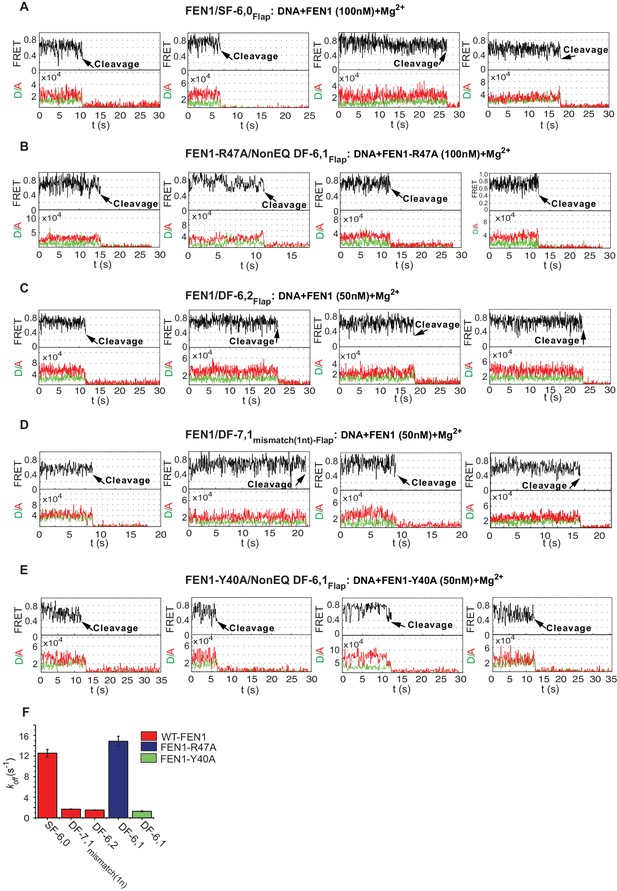
FEN1 cleavage of various noncognate substrates.
(A) Representative TIRF-smFRET time traces showing the cleavage of SF6,0Flap in which FEN1 cleaves the DNA after multiple unsuccessful bending events. (B) Representative TIRF-smFRET time traces showing the cleavage of NonEQ DF6,1Flap by FEN1-R47A in which FEN1-R47A cleaves the DNA after multiple unsuccessful bending events. (C) Representative TIRF-smFRET time traces showing the cleavage of DF6,2Flap in which FEN1 cleaves the DNA after multiple unsuccessful bending events. (D) Representative TIRF-smFRET time traces showing the cleavage of DF7,1mismatch(1nt)-Flap in which FEN1 cleaves the DNA after multiple unsuccessful bending events. (E) Representative TIRF-smFRET time traces showing the cleavage of NonEQ DF6,1Flap by FEN1-Y40A in which FEN1-Y40A cleaves the DNA after multiple unsuccessful bending events. (F) Bar chart comparing koff-unbending of the DNA bending events that did not lead to 5’flap cleavage in presence of Mg2+ in the cases of FEN1-WT on noncognate substrates and FEN1-Y40A and FEN1-R47A on NonEQ DF-6,1. The koff-unbending values were similar to those reported under no cleavage condition in the presence of Ca2+ that are presented in Figure 4B. The slight decrease in rates in the cases of FEN1-WT on SF-6,0 and FEN1-R47A on NonEQ DF-6,1 in the presence of Mg2+ relative to Ca2+ is a result of averaging due to the lower temporal resolution in TIRF-based smFRET (Mg2+ case) versus immobilized confocal-based smFRET (Ca2+ case).
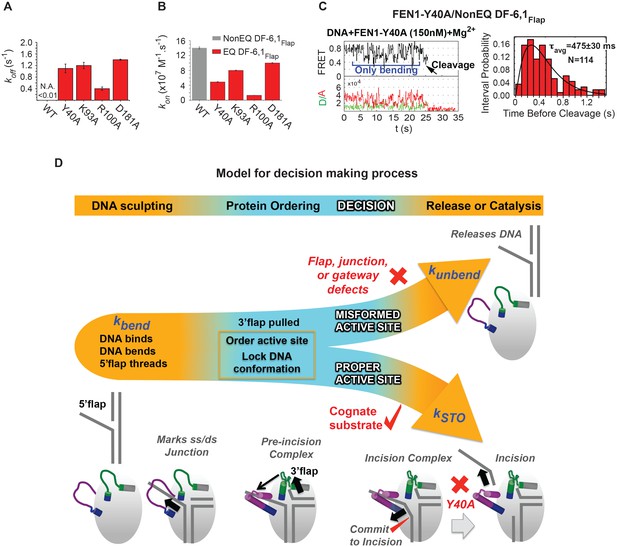
Role of active-site residues in active positioning of the 5’flap and the junction.
(A) Bar chart comparing koff-unbending of WT-FEN1 and the FEN1 active-site mutants Y40A, K93A, R100A and D181A on NonEQ DF6,1Flap. koff-unbending is calculated as described in 1G. (B) Bar chart comparing kon-bending of wild-type FEN1 on EQ DF6,1Flap and the FEN1 mutants Y40A, K93A, R100A and D181A on NonEQ DF6,1Flap. kon-bending is calculated as described in Figure 1G. (C) A representative smFRET time trace showing the cleavage of NonEQ DF6,1Flap by FEN1-Y40A after multiple trials of DNA bending (left panel) and a histogram showing the distribution of dwell times of the bent state prior to cleavage (right panel). τavg from N = 4 is calculated as in Figure 2C at a temporal resolution of 50 ms. (D) Model for control of catalytic selectivity by the DNA mutual-induced fit mechanism in FEN1. DNA sculpting: FEN1 actively bend a variety of structures to verify the key features of its cognate DF substrates of fully paired ss/dsDNA nick junction, threaded 5’flap into the cap-helical gateway and 3’flap. Protein ordering: FEN1 actively pulls the 3’end of the nick junction to create a 3’flap and drive protein ordering, which in turn orders the active site and locks the DNA conformation. Decision: the active site and locked DNA conformer are always in catalytically competent form in cognate substrate, while they are primarily in catalytically incompetent form in noncognate substrates (no 5’flap threading, no 3’flap, mispair junctions) and FEN1 mutants (R47A, K93A and R100A). DNA release or catalysis: the DNA will shift or unpair to move the scissile phosphate into the active site for cleavage as probed by the flap/junction positioning-residue Y40, while in noncognate substrates FEN1 promotes DNA dissociation.
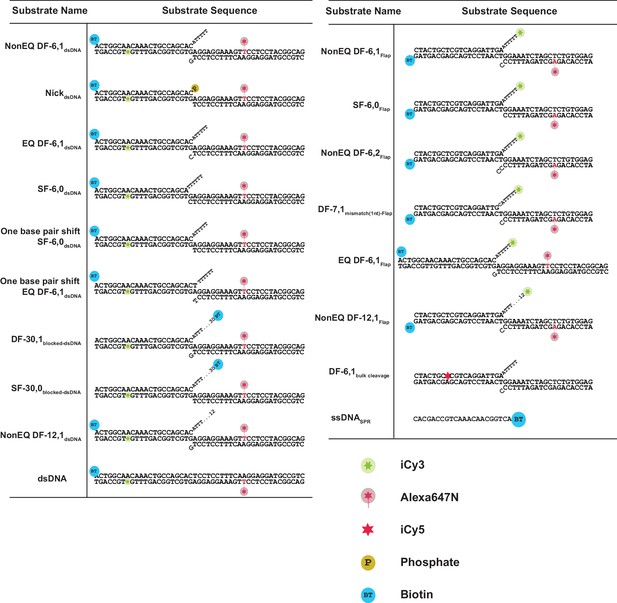
Sequences of DNA constructs used in this study.
Schematic of the substrates with internal-labeling and flap-labeling schemes used in this study including their sequences. A legend of the symbols used is shown.





