Mapping cell type-specific transcriptional enhancers using high affinity, lineage-specific Ep300 bioChIP-seq
Figures
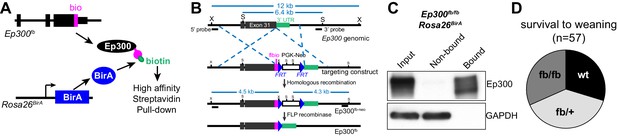
Generation and characterization of Ep300flbio allele.
(A) Experimental strategy for high affinity Ep300 pull down. A flag-bio epitope was knocked onto C-terminus of the endogenous Ep300 gene. The bio peptide sequence is biotinylated by BirA, widely expressed from the Rosa26 locus. (B) Targeting strategy to knock flag-bio epitope into the C-terminus of Ep300. A targeting vector containing homology arms, flag-bio epitope, and Frt-neo-Frt cassette was used to insert the epitope tag into embryonic stem cells by homologous recombination. Chimeric mice were mated with Act:Flpe mice to excise the Frt-neo-Frt cassette in the germline, yielding the Ep300fb allele. (C) Biotinylated Ep300 is quantitatively pulled down by streptavidin beads. Protein extract from Ep300flbio/flbio; Rosa26BirA embryos was incubated with immobilized streptavidin. Input, bound, and unbound fractions were analyzed for Ep300 by immunoblotting. GAPDH was used as an internal control. (D) Ep300flbio/flbio mice from heterozygous intercrosses survived normally to weaning.
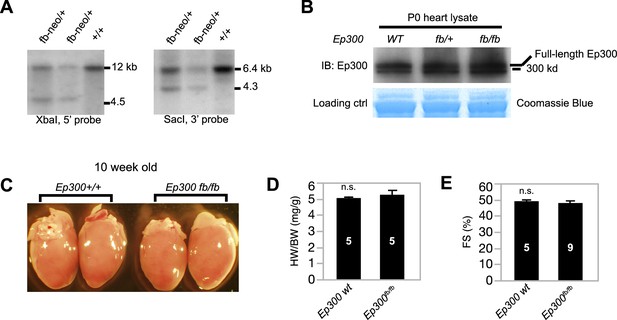
Characterization of Ep300fb allele.
(A) Southern blot genotyping of Ep300fb allele. Probes are marked in Figure 1B. (B) The expression level of tagged Ep300 protein is similar to that of wild-type Ep300 in neonatal mouse hearts. (C, D) The heart weight to body weight ratio of 10-week-old Ep300fb/fb heart was not significantly different from littermates. (E) Echocardiograms of 10-week old Ep300fb/fb mice show that their ventricular function was not significantly different from littermate controls. Sample sizes are shown in bars in D,E. Bars represent mean ± standard deviation. n.s., not significantly different (t-test).
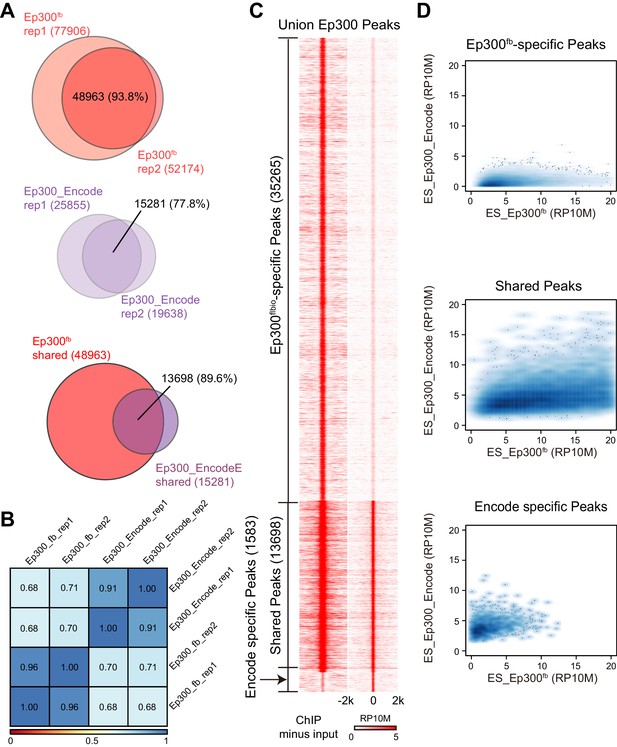
Comparison of Ep300 bioChiP-seq to antibody ChIP-seq for mapping Ep300 chromatin occupancy.
(A–B) Comparison of biological duplicate antibody Ep300 ChIP-seq (Encode) and Ep300 bioChIP-seq (flbio). The Ep300 bioChIP-seq data had greater overlap between replicates and greater intragroup correlation. Most antibody peaks were covered by the bioChIP-seq data. There were 3.3 times more Ep300 regions detected by Ep300 bioChiP-seq. 89.6% of Ep300 regions detected by antibody ChIP-seq were recovered by Ep300 bioChIP. Panel B shows Spearman correlation between samples over the peak regions. (C) Tag heatmap shows input-subtracted Ep300 antibody or bioChIP signal in the union of the Ep300-bound regions detected by each method. (D) Correlation plots show greater Ep300 bioChIP-seq signal compared to antibody bioChIP-seq.
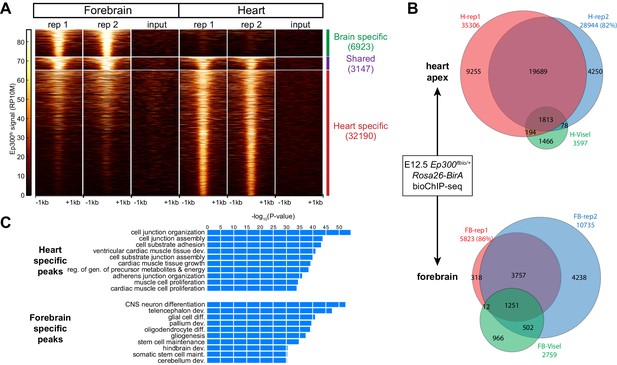
Tissue specific Ep300 bioChiP-seq.
(A) Tag heatmap showing Ep300fb signal in heart and forebrain. Each row represents a region that was bound by Ep300 in heart, forebrain, or both. A minority of Ep300-enriched regions were shared between tissues. (B) Ep300fb pull down from E12.5 heart or forebrain. Ep300-bound regions identified by bioChIP-seq of Ep300fb/fb; Rosa26BirA tissues in biological duplicates are compared to regions identified by antibody-mediated Ep300 ChIP-seq (Visel et al., 2009b). (C) GO biological process terms enriched for genes that neighbor the tissue-specific heart or forebrain peaks. Bars indicate statistical significance.
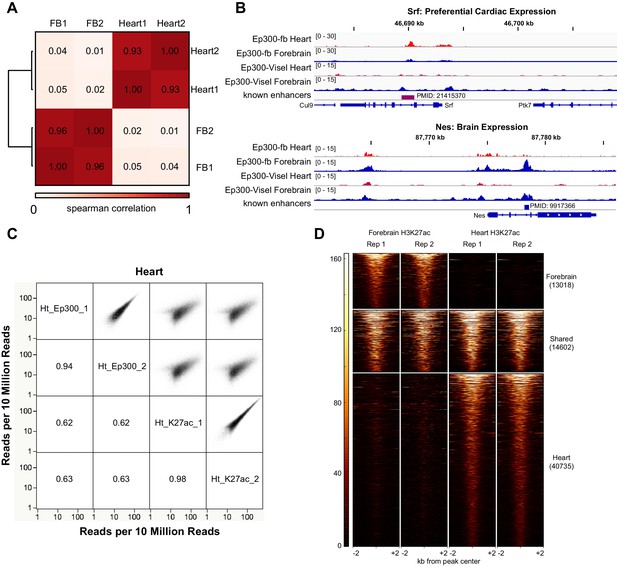
Heart and forebrain Ep300 bioChiP-seq.
(A) Ep300 bioChIP-seq signals within called Ep300 peaks were highly correlated between biological replicates and discordant between heart and forebrain. (B) Genome browser views showing Ep300 bioChIP-seq signal in heart and brain at Srf (preferential cardiac expression) and Nes (preferential Brain expression). (C) Comparison of Ep300 bioChIP-seq and H3K27ac ChIP-seq signals in heart. Numbers indicate correlation coefficients. There was excellent correlation between biological replicates of Ep300 or H3K27ac. Ep300 and H3K27ac also showed highly significant genome-wide correlation. Similar results were observed in forebrain (data not shown). (D) Tag heatmap of H3K27ac-bound regions showing overlap between heart and brain H3K27ac enriched regions from each tissue.
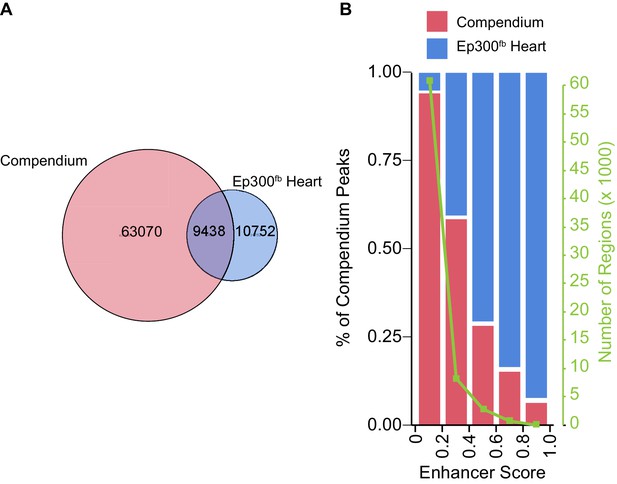
Comparison of heart Ep300fb regions to a compendium of heart enhancers assembled by Dickel et al. (2016).
The Dickel heart enhancer compendium (Dickel et al., 2016) is based on H3K27ac ChIP-seq and Ep300 antibody ChIP-seq. Most compendium regions had low enhancer scores and have low validation frequency in transient transgenic assays. Compendium regions were compared to heart Ep300fb regions using the LiftOver tool to map human genome coordinates to mouse. (A) Venn diagram showing that 13% of prenatal heart compendium regions overlap with heart Ep300 regions. On the other hand, 47% of Ep300fb heart regions overlap with heart compendium regions, and 53% do not. (B) Analysis of the fraction of compendium regions that overlap the Ep300fb heart regions, as a function of enhancer score. Regions with higher enhancer score (and higher likelihood of in vivo validation) corresponded well with heart Ep300 regions.
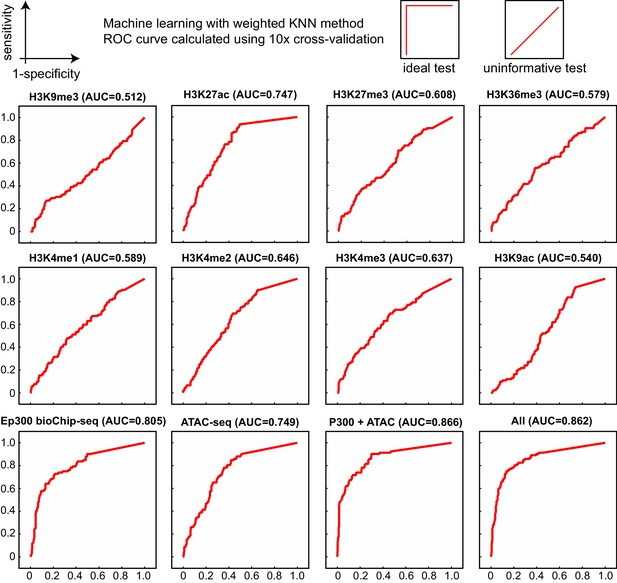
Comparison of enhancer prediction based on indicated chromatin features of E12.5 heart.
Data are from ENCODE except for ATAC-seq (Pu lab, unpublished), and Ep300fb (this manuscript). The KNN classifier was trained on a subset of the VISTA enhancer database and validated on the remaining data, using a 10x cross-validation design. Ep300 was the most accurate single predictor, based on the area under the receiver operating characteristic curve (AUC, indicated in parentheses). All of the features together (All), or just Ep300 plus ATAC-seq together (not shown), gave the best prediction.
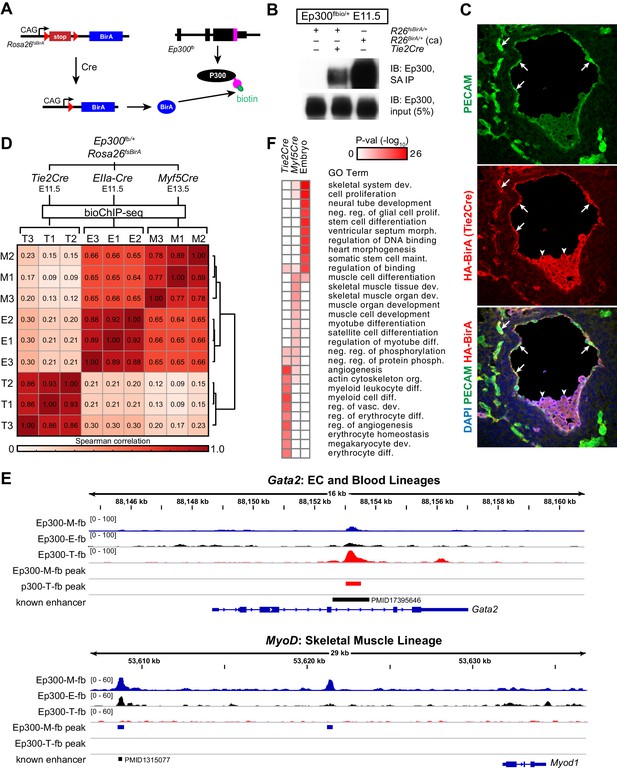
Cre-directed, lineage-selective Ep300 bioChiP-seq.
(A) Experimental strategy. Lineage-specific Cre recombinase activates expression of BirA (HA-tagged) from Rosa26-flox-stop-BirA (Rosa26fsBirA). This results in Ep300 biotinylation in the progeny of Cre-expressing cells. (B) Cre-dependent Ep300 biotinylation using Rosa26fsBirA. Protein extracts were prepared from E11.5 embryos with the indicated genotypes. R26BirA/+ (ca) and R26fsBirA/+ indicate the constitutively active and Cre-activated alleles, respectively. (C) Immunostaining demonstrating selective Tie2Cre-mediated expression of HA-tagged BirA in ECs of R26fsBirA; Tie2Cre embryos. Arrows and arrowheads indicate endothelial and hematopoietic lineages, respectively. (D) Tissue-selective Ep300 bioChIP-seq. Tie2Cre (T; endothelial and hematopoietic lineages), Myf5Cre (M; skeletal muscle lineages), and EIIaCre (E; germline activation) were used to drive tissue-selective Ep300 bioChIP-seq. Correlation between ChIP-seq signals within peak regions are shown for triplicate biological repeats. Samples within groups were the most closely correlated. (E) Ep300fb bioChIP-seq signal at Gata2 (EC/blood specific) and Myod (muscle specific). Enhancers validated by transient transgenic assay are indicated along with the citation’s Pubmed identifier (PMID). (F) Biological process GO terms illustrate distinct functional groups of genes that neighbor Ep300 bioChIP-seq driven by lineage-specific Cre alleles. The heatmap contains the top 10 terms enriched for genes neighboring each of the three lineage-selective Ep300 regions.
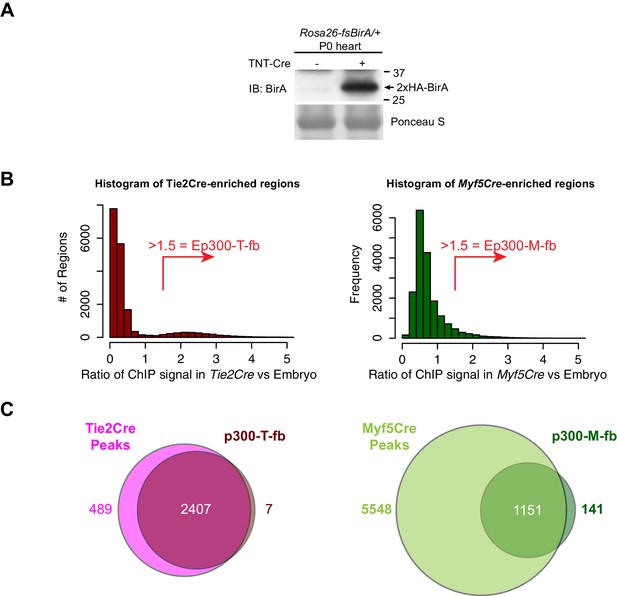
Cre-dependent Ep300 bioChIP-seq.
(A) Cre-dependent expression of BirA. Heart with or without cardiac-specific TNTCre was immunoblotted for expression of BirA from the Rosa26fsBirAallele. (B) Distribution of Ep300fb bioChIP-seq signal when driven by Tie2Cre or Myf5Cre. Regions with Ep300 peaks called in samples from any of the three Cre drivers were scored for their EC- or muscle- enriched signal by calculating the ratio of the region's signal in Tie2Cre or Myf5Cre bioChIP-seq to its signal with ubiquitous Ep300 bioChIP-seq (EIIaCre). The Tie2Cre-enrichment score showed a bimodal distribution with a small peak at values greater than 1.5. A value of greater than 1.5 similarly was used to define Myf5Cre-enriched regions. (C) Relationship of Tie2Cre-enriched (Ep300-T-fb) and Myf5Cre-enriched (Ep300-M-fb) regions to peaks called from Tie2Cre or Myf5Cre samples alone.
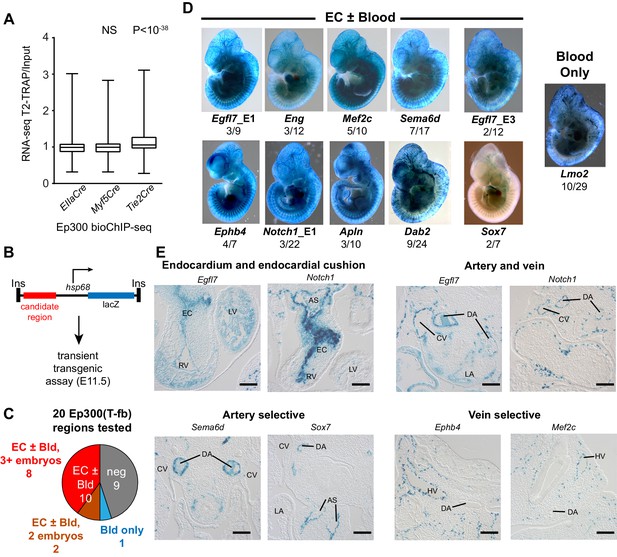
Functional validation of enhancer activity of Ep300-T-fb-bound regions.
(A) Expression of genes neighboring regions bound by Ep300fb in different Cre-marked lineages. Translating ribosome affinity purification (TRAP) was used to enrich for RNAs from the Tie2Cre lineage. Input or Tie2Cre-enriched RNAs were profiled by RNA sequencing. The expression of the nearest gene neighboring Ep300 regions in ECs (Ep300-T-fb regions), but not skeletal muscle (Ep300-M-fb regions), was higher than that of genes neighboring regions bound by Ep300 across the whole embryo (EIIaCre). Box and whiskers show quartiles and 1.5 times the interquartile range. Groups were compared to EIIaCre using the Mann-Whitney U-test. (B) Transient transgenesis assay to measure in vivo activity of Ep300-T-fb regions. Test regions were positioned upstream of an hsp68 minimal promoter and lacZ. Embryos were assayed at E11.5. (C) Summary of transient transgenic assay results. Out of 20 regions tested, nine showed activity in ECs or blood in three or more embryos, and two more showed activity in two embryos. See also Table 2. (D) Representative whole mount Xgal-stained embryos. Enhancers that directed LacZ expression in an EC or blood pattern in two or more embryos are shown. Numbers indicate embryos with LacZ distribution similar to shown image, compared to the total number of PCR positive embryos. (E) Sections of Xgal-stained embryos showing examples of enhancers active in arteries, veins, and endocardium, or selectively active in arteries or veins. AS: aortic sac; CV: cardinal vein; DA: dorsal aorta; EC: endocardial cushion; HV: head vein; LA: left atrium; LV: left ventricle; RV: right ventricle. Scale bars, 100 µm. See also Figure 5—figure supplements 1 and 2 and Table 2.
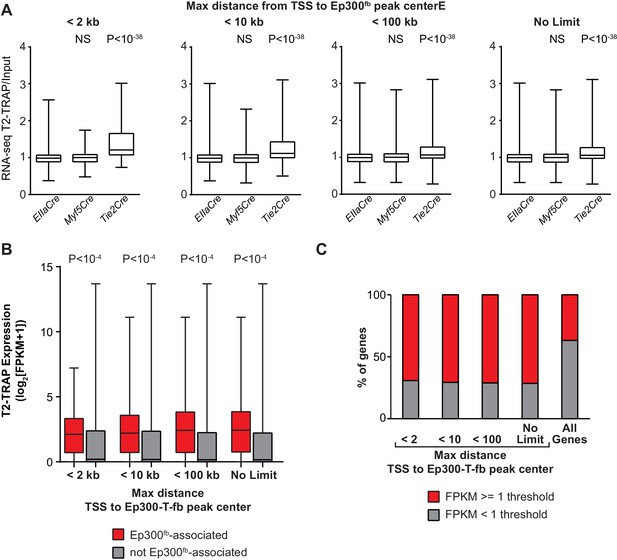
Relationship of EC gene expression to Ep300 regions in ECs, skeletal muscle, and whole embryo.
(A) EC enrichment of actively translating genes that neighbored Ep300 regions in skeletal muscle (Ep300-M-fb), whole embryo (Ep300-E-fb), and endothelial cell/blood (Ep300-T-fb), using several different thresholds for the maximum distance allowed to associate genes to regions. The relationship between Ep300-T-fb regions and gene expression was not dependent upon the threshold used. The rightmost panel (no limit to maximum distance) is the same as in Figure 5A. Mann-Whitney U-test. (B) EC expression of actively translating genes that neighbored Ep300-T-fb, compared to those that did not, using several different thresholds for the maximum distance used associate genes to regions. Ep300-T-fb-associated genes were more highly expressed regardless of the distance threshold used. Ep300-T-fb genes were significantly more highly expressed (Mann-Whitney U-test). (C) Fraction of genes expressed or not expressed, for Ep300-T-fb-associated genes compared to all genes. Ep300-T-fb associated genes were more frequently expressed than non-associated genes. However, not all genes associated with Ep300-T-fb regions were expressed, and some genes without Ep300-T-fb regions were expressed. This may reflect additional mechanisms of gene regulation as well as limitations of the gene-to-region association rule additional rule.
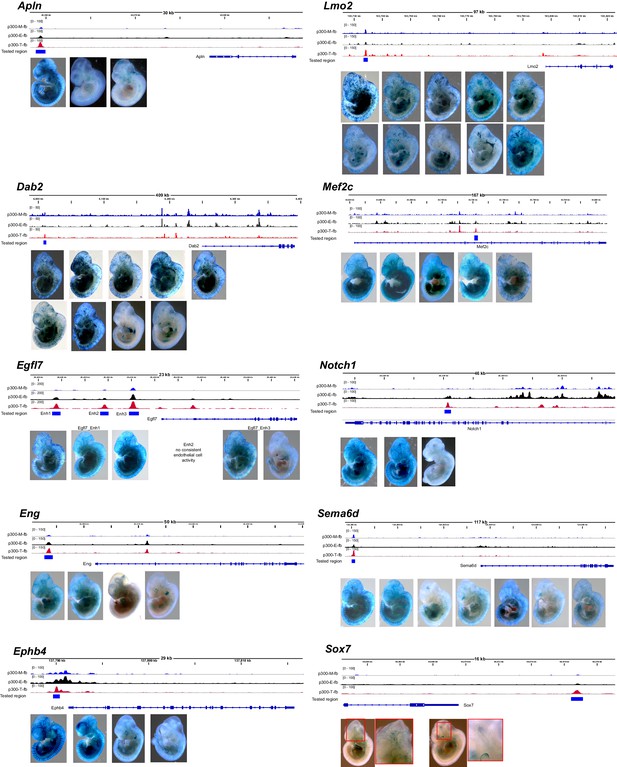
Transient transgenic assays to measure in vivo activity of candidate endothelial cell/blood enhancers.
For each tested enhancer with positive endothelial cell or blood activity, we show the candidate enhancer’s Ep300 signal in skeletal muscle (Ep300-M-fb), whole embryo (Ep300-E-fb), and endothelial cell/blood (Ep300-T-fb), with respect to the gene both. The whole mount images show each of the positive Xgal-stained embryos.
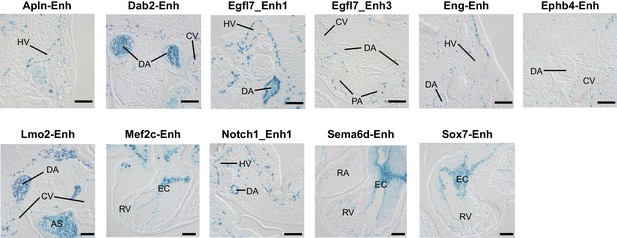
Histological sections of transient transgenic embryos.
Histological sections of embryos containing the indicated enhancer-driven lacZ transgene. AS: aortic sac; CV: cardinal vein; DA: dorsal aorta; EC: endocardial cushion; HV: head vein; PA: pulmonary artery; RA: right atrium; RV: right ventricle. Bar, 100 µm.
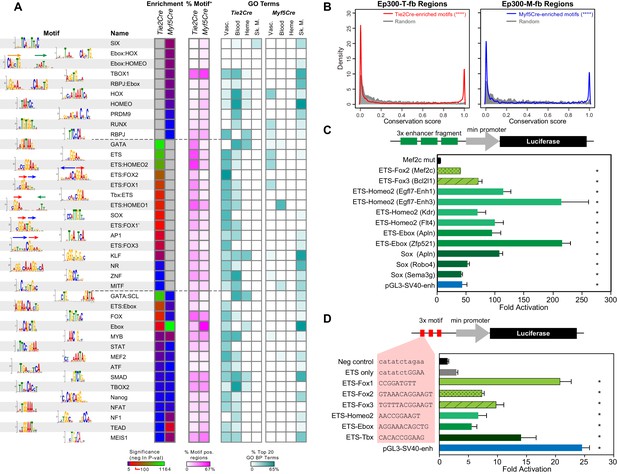
Motifs enriched in Ep300-T-fb and Ep300-M-fb regions.
(A) Motifs enriched in Ep300-T-fb or Ep300-M-fb regions. 1445 motifs were tested for enrichment in Ep300 bound regions compared to randomly permuted control regions. Significantly enriched motifs (neg ln p-value>15) were clustered and the displayed non-redundant motifs were manually selected. Heatmaps show statistical enrichment (left), fraction of regions that contain the motif (center), and GO terms associated with genes neighboring motif-containing, Ep300-bound regions (fraction of the top 20 GO biological process terms). Grey indicates that the motif was not significantly enriched (neg ln p-value≤15). (B) Conservation of sequences matching Tie2Cre- or Myf5Cre-enriched motifs within 100 bp of the summit of Ep300-T-fb or Ep300-M-fb regions, compared to randomly selected 12 bp sequences from the same regions. PhastCons conservation scores across 30 vertebrate species were used. ****p<0.0001, Kolmogorov–Smirnov test. (C) Luciferase assay of activity of enhancers containing indicated motifs. Three repeats of 20–30 bp regions from Ep300-bound enhancers linked to the indicated gene and centered on the indicated motif were cloned upstream of a minimal promoter and luciferase. The constructs were transfected into human umbilical vein endothelial cells. Luciferase activity was expressed as fold activation above that driven by the enhancerless, minimal promoter-luciferase construct. *p<0.05 compared to Mef2c-enhancer with mutated ETS:FOX2 motif (Mef2c mut). n = 3. (D) Luciferase assay of indicated motifs repeated three times within a consistent DNA context. Assay was performed as in C. *p<0.05 compared to negative control sequencing lacking predicted motif. n = 3. Error bars in C and D indicate standard error of the mean.
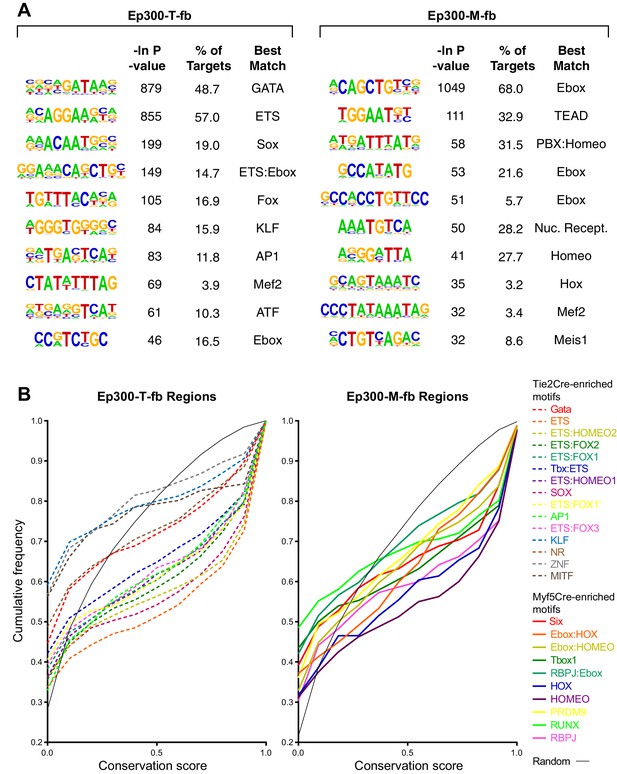
Motifs with enriched Ep300 occupancy in Tie2Cre or Myf5Cre embryo samples.
(A) De novo motif discovery. The top 10 discovered motifs are shown, with the significance P-value, percent of regions containing the motif, and the best matching known motif class. (B) Conservation of Tie2Cre-enriched motifs (dashed lines) or Myf5Cre-enriched motifs (solid lines) within 100 bp of Ep300-T-fb or Ep300-M-fb peak summits. The distribution of conservation scores for each indicated motif is shown as a cumulative frequency plot. Random indicates randomly sampled 12 nt regions from within the Ep300-T-fb or Ep300-M-fb regions.
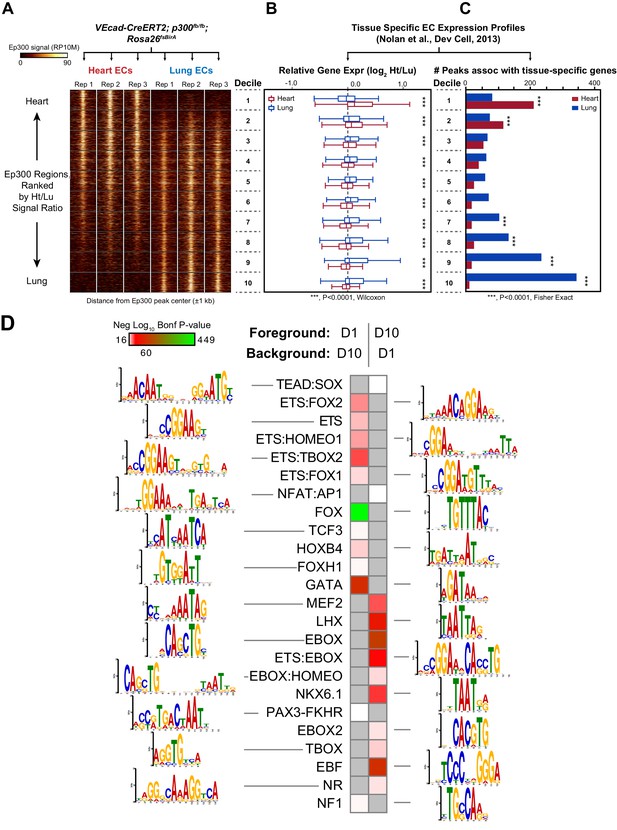
Enhancers in adult heart and lung ECs.
VEcad-CreERT2 and neonatal tamoxifen pulse was used to drive BirA expression in ECs. Ep300 regions in adult heart or lung ECs were identified by bioChIP-seq in biological triplicate. Ep300 regions were ranked into deciles by the ratio of the Ep300-VE-fb signal in heart to lung. (A) Tag heatmap shows that the top and bottom deciles have selective Ep300 occupancy in heart and lung ECs, respectively. (B) Expression of genes in heart ECs (red) or lung ECs (blue) neighboring Ep300 regions, divided into deciles by Ep300-VE-fb signal ratio in heart and lung. ***p<0.0001, Wilcoxon test. Expression values were obtained from (Nolan et al., 2013). Box plots indicate quartiles, and whiskers indicate 1.5 times the interquartile range. (C) Genes with selective expression in heart or lung ECs were identified by K-means clustering (see Materials and methods and Figure 7—figure supplement 1). The number of Ep300-VE-fb regions in heart (red) or lung (blue) neighboring these genes was determined, stratified by decile. ***p<0.0001, Fisher's exact test. (D) Enrichment of selected motifs in Ep300-VE-fb regions from heart (decile 1) compared to lung (decile 10) or vice versa. Grey indicates no significant enrichment. Displayed non-redundant motifs were selected from all significant motifs by manual curation of clustered motifs.
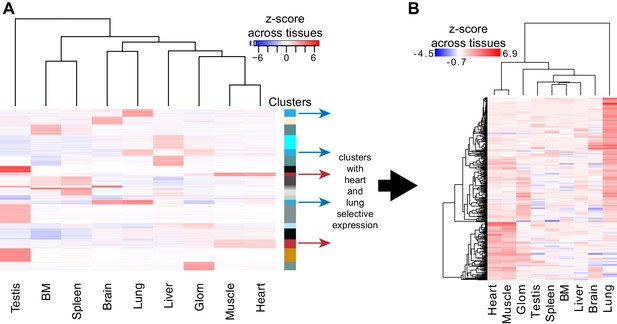
Organ-specific EC gene expression.
(A) Genes with at least four-fold differential mean expression between highest and lowest EC samples were identified from microarray data (Nolan et al., 2013). These genes were grouped by k-means clustering. Genes in clusters containing selective heart or lung expression were analyzed further. Expression of these heart or lung EC genes is displayed in panel (B) using hierarchical clustering.
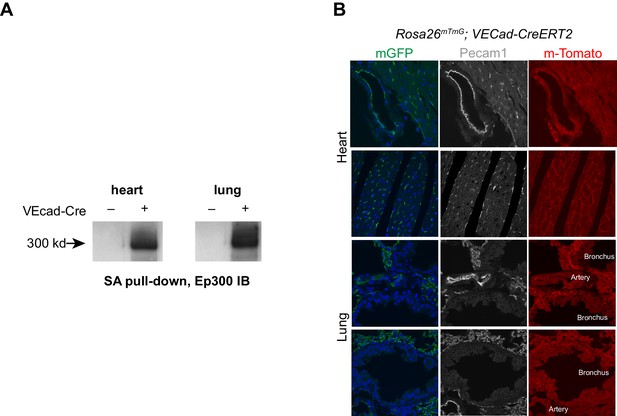
VEcad-CreERT2 activation of Ep300 biotinylation.
(A) VEcad-CreERT2 activated Ep300 biotinylation by Rosa26fsBirA, permitting its pulldown on streptavidin (SA) beads. (B) EC-selective activation of the Rosa26mTmG Cre reporter allele by VEcad-CreERT2. Tamoxifen was administered at P1 and P2, and tissues were analyzed at six weeks of age.
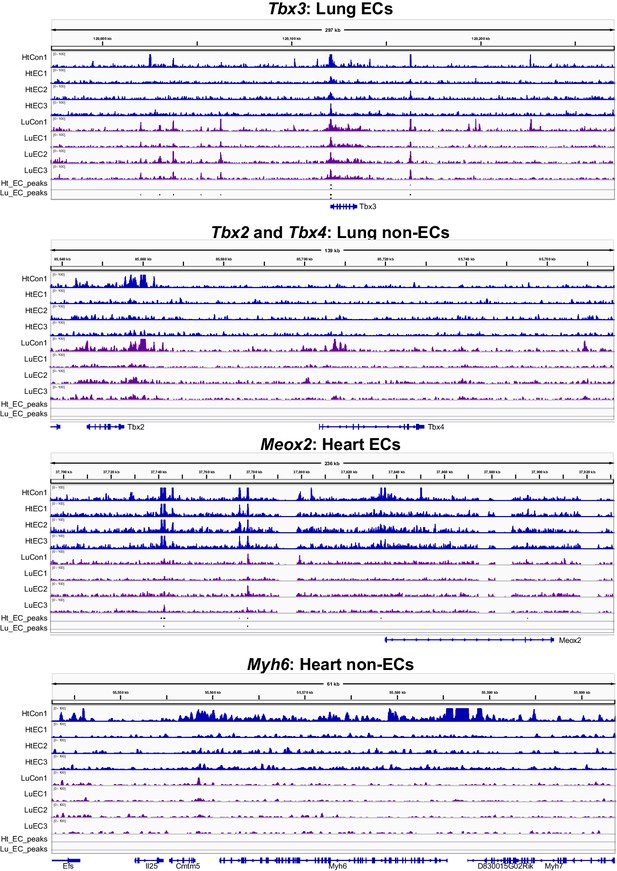
Ep300fb bioChIP-seq signal in whole organ or ECs.
Ep300fb bioChIP-seq signal in whole organ (‘Con’; EIIaCre) or ECs ('EC’; VEcad-CreERT2) at genes with EC or non-EC expression in lung or heart.
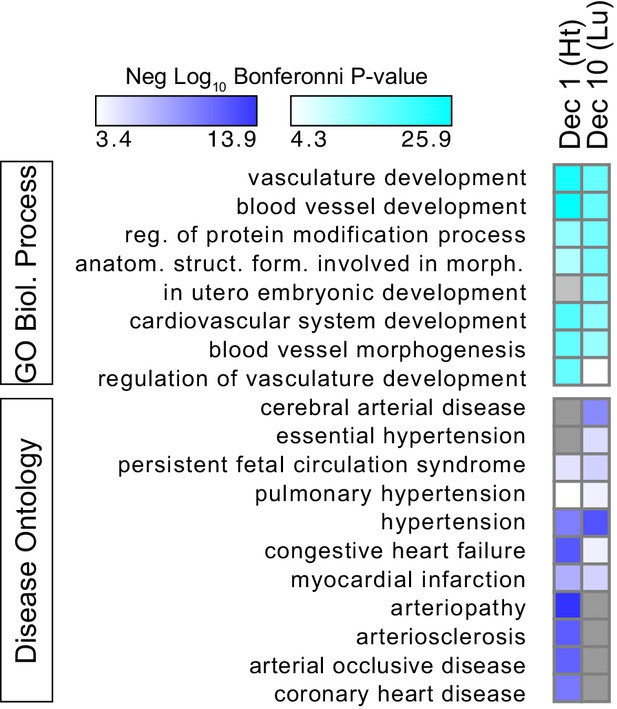
Gene functional terms enriched in decile 1 or decile 10 of Ep300-VE-fb regions.
GO biological process (top) or disease ontology (bottom) terms enriched in decile 1 or decile 10 of Ep300-VE-fb regions. Grey indicates no significant enrichment. The top five biological process terms in deciles 1 and 10 and selected relevant disease ontology terms were are displayed.
Tables
mm9 genome coordinates of regions with EC activity as determined by transient transgenic assay. Vista_XXX indicates that the region was obtained from the VISTA enhancer database. Lifeover indicates that the region was inferred by liftover from the human genome. For enhancers obtained from the literature, Pubmed was searched for ‘endothelial cell enhancer’. The resulting references were manually curated for transient transgenic testing of candidate endothelial cell enhancer regions.
| Chr | Start | End | Note |
|---|---|---|---|
| chr9 | 37206631 | 37209631 | Robo4;PMID17495228;bloodVessels |
| chr4 | 94412022 | 94413640 | Tek;PMID9096345;bloodVessels |
| chr8 | 106625634 | 106626053 | Cdh5;PMID15746076;bloodVessels |
| chr11 | 49445663 | 49446520 | Flt4;posEC;liftoverFromPMID19070576;FoxETS |
| chr6 | 99338223 | 99339713 | FoxP1;posEC;liftoverFromPMID19070576;FoxETS |
| chr8 | 130910720 | 130911740 | Nrp1;posEC;liftoverFromPMID19070576;FoxETS |
| chr18 | 61219017 | 61219690 | Pdgfrb;posEC;liftoverFromPMID19070576;FoxETS |
| chr4 | 137475841 | 137476446 | Ece1;posEC;liftoverFromPMID19070576;FoxETS;artery |
| chr4 | 114743698 | 114748936 | Tal1;PMID14966269;endocardium;bloodVessels |
| chr2 | 119152861 | 119153661 | Dll4;PMID23830865;arterial |
| chr13 | 83721919 | 83721962 | Mef2c;PMID19070576;FoxETS |
| chr13 | 83711086 | 83711527 | Mefec;PMID15501228;panEC |
| chr2 | 32493213 | 32493467 | Eng;liftoverFromPMID16484587;bloodVessels |
| chr2 | 32517844 | 32518197 | Eng;liftoverFromPMID18805961;bloodVessels;blood |
| chr6 | 88152598 | 88153791 | Gata2;PMID17395646;PMID17347142;bloodVessels |
| chr9 | 32337302 | 32337549 | Fli1;PMID15649946;bloodVessels |
| chr5 | 76370571 | 76372841 | KDR;PMID10361126;bloodVessels |
| chr6 | 125502138 | 125502981 | VWF;PMID20980682;smallbloodVessels |
| chr5 | 148537311 | 148538294 | Flt1;liftoverFromPMID19822898 |
| chr17 | 34701455 | 34702277 | Notch4;liftoverFromPMID15684396 |
| chr2 | 155568649 | 155569132 | Procr;liftoverfromPMID16627757;bloodVessels[7/17] |
| chr9 | 63916890 | 63917347 | Smad6;liftoverfromPMID17213321;bloodVessels |
| chr19 | 37510161 | 37510394 | Hhex;liftoverfromPMID15649946;bloodVessels;blood |
| chr5 | 76357892 | 76358715 | Flk1;PMID27079877;bloodVessesl;artery |
| chr11 | 32145270 | 32146411 | vista_101;heart[9/12];bloodVessels[7/12] |
| chr13 | 28809515 | 28811310 | vista_265;neural tube[7/8];bloodVessels[3/8] |
| chr17 | 12982583 | 12985936 | vista_89;heart[6/10];bloodVessels[8/10] |
| chr4 | 131631431 | 131635142 | vista_80;bloodVessels[10/10] |
| chr4 | 57858433 | 57860639 | vista_261;bloodVessels[5/8] |
| chr5 | 93426293 | 93427320 | vista_397;limb[12/13];bloodVessels[12/13] |
| chr8 | 28216799 | 28218903 | vista_136;heart[7/10];other[6/10];bloodVessels[4/10] |
| chr3 | 87786601 | 87790798 | liftoverFromvista_1891;somite[7/7];midbrain;(mesencephalon)[6/7];limb[7/7];branchial;arch[7/7];eye[7/7];heart[7/7];ear[7/7];bloodVessels[6/7] |
| chr6 | 116309006 | 116309768 | liftoverFromvista_2065;bloodVessels[9/9] |
| chr19 | 37566512 | 37571839 | liftoverFromvista_1866;bloodVessels[5/5] |
| chr7 | 116256647 | 116260817 | liftoverFromvista_1859;neuraltube[8/8];hindbrain;(rhombencephalon)[5/8];midbrain;(mesencephalon)[8/8];forebrain[8/8];heart[7/8];bloodVessels[5/8];liver[4/8] |
| chr18 | 14006285 | 14007917 | liftoverFromvista_1653;bloodVessels[5/8] |
| chr2 | 152613921 | 152616703 | liftoverFromvista_2050;bloodVessels[5/5] |
| chr14 | 32045520 | 32047900 | liftoverFromvista_2179;bloodVessels[5/7] |
| chr15 | 73570041 | 73577576 | liftoverFromvista_1882;bloodVessels[8/8] |
| chr8 | 28216817 | 28218896 | liftoverFromvista_1665;heart[5/7];bloodVessels[7/7] |
Summary of transient transgenic validation of candidate EC enhancers.
| Neighboring gene | Region (mm9) | Size (bp) | Location w/r gene | Distance to TSS | Whole mount | Sections | Ref. (PMID) | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| #PCR pos | #LacZ pos | # EC or blood pos | Endo | Art | Vein | Blood cells | ||||||
| Apln | chrX:45358891–45359918 | 1028 | 3'_Distal | 28,624 | 10 | 3 | 3 | + | + | + | − | − |
| Dab2 | chr15:6009504–6010497 | 994 | 5'_Distal | −239,788 | 24 | 10 | 9 | + | ++ | + | +++ | − |
| Egfl7_enh1 | chr2:26427040–26428029 | 990 | 5'_Distal | −9,041 | 9 | 3 | 3 | +++ | +++ | +++ | + | − |
| Eng | chr2:32493216–32494019 | 804 | 5'_Distal | −8,497 | 12 | 4 | 3 | ++ | ++ | ++ | − | 16484587 |
| Ephb4 | chr5:137789649–137790412 | 764 | 5'_Proximal | −1,306 | 7 | 5 | 4 | ++ | − | +++ | − | − |
| Lmo2 | chr2:103733621–103734378 | 758 | 5'_Distal | −64,152 | 29 | 14 | 10 | − | − | − | +++ | − |
| Mef2c | chr13:83721522–83722451 | 930 | Intragenic | 78,954 | 10 | 6 | 5 | + | − | +++ | + | 19070576 |
| Notch1_enh1 | chr2:26330255–26331184 | 930 | Intragenic | −28,622 | 22 | 3 | 3 | +++ | ++ | ++ | − | − |
| Sema6d | chr2:124380522–124381285 | 764 | 5'_Distal | −55,128 | 17 | 10 | 7 | ++ | +++ | − | + | − |
| Egfl7_enh3 | chr2:26433680–26434642 | 963 | 5'_Distal | −2,415 | 12 | 2 | 2 | + | ++ | +++ | − | − |
| Sox7 | chr14:64576382–64577118 | 737 | 3'_Distal | 14,207 | 7 | 4 | 2 | + | ++ | − | − | − |
| Aplnr | chr2:85003436–85004412 | 977 | 3'_Distal | 27,407 | 12 | 3 | 0 | NA | NA | NA | NA | − |
| Egfl7_enh2 | chr2:26431273–26431995 | 723 | 5'_Proximal | −4,942 | 9 | 3 | 1 | + | − | − | − | − |
| Emcn | chr3:136984933–136986103 | 1171 | 5'_Distal | −18,524 | 15 | 1 | 1 | − | − | − | + | − |
| Ets1 | chr9:32481485–32482133 | 649 | Intragenic | −21,818 | 11 | 1 | 0 | NA | NA | NA | NA | − |
| Foxc1 | chr13:31921976–31922827 | 852 | 3'_Distal | 23,887 | 13 | 3 | 1* | NA | NA | NA | NA | − |
| Gata2 | chr6:88101907–88102696 | 790 | 5'_Distal | −46,356 | 8 | 0 | 0 | NA | NA | NA | NA | − |
| Lyve1 | chr7:118020264–118021043 | 780 | 5'_Distal | −3,128,028 | 7 | 1 | 1 | + | + | − | − | − |
| Notch1_enh2 | chr2:26345973–26347118 | 1146 | Intragenic | 12,796 | 9 | 0 | 0 | NA | NA | NA | NA | − |
| Sox18 | chr2:181397552–181398335 | 784 | 3'_Proximal | 8401 | 13 | 2 | 1 | − | ++ | − | − | − |
-
*EC/blood pattern on whole mount not validated in histological sections.
Oligonucleotides used in this study.
| Genotyping primers | |||
|---|---|---|---|
| Name | Sequence (5'- > 3') | Comments | |
| Ep300fb-f | AATGCTTTCACAGCTCGC | 0.28 kb for wild-type, 0.43 kb for Ep300fb knockin | |
| Ep300fb-r | AAACCATAAATGGCTACTGC | ||
| Forward common | CTCTGCTGCCTCCTGGCTTCT | Rosa26-fs-BirA, 0.33 kb for wildtype, 0.25 kb for knockin | |
| Wild type reverse | CGAGGCGGATCACAAGCAATA | ||
| CAG reverse | TCAATGGGCGGGGGTCGTT | ||
| LacZ-f | CAATGCTGTCAGGTGCTCTCACTACC | 0.42 kb, genotyping of transient transgenic | |
| LacZ-r | GCCACTTCTTGATGCTCCACTTGG | ||
| Primers to amplify Ep300 peak regions for transient transgenic assay. 4 nucleotides CACC have been added to all the forward primers for TOPO Cloning. | |||
| Name | Sequence (5'- > 3') | ||
| Apln_f | CACCGGAGGCTGAGCAATGAATAG | ||
| Apln_r | TTGGCTGGGGAAGAGTAAGC | ||
| Aplnr_f | CACCTCTCTCTCTGGCTTCG | ||
| Aplnr_r | CCTCAGAATGTTTTCATGG | ||
| Dab2_f | CACCGTGGAAATCATAGCAC | ||
| Dab2_r | GGTTGGAATAAAAGAGC | ||
| Egfl7_Enh1_f | CACCGCCTACCCAGTGCTGTTCC | ||
| Egfl7_Enh1_r | CTGGAGTGGAGTGTCACG | ||
| Egfl7_Enh2_f | CACCGCTAGGGGCTTCTAGTTC | ||
| Egfl7_Enh2_r | AGGTCTCTTCTGTGTCG | ||
| Egfl7_Enh3_f | CACCTGTTAGTGGTGCTCCC | ||
| Egfl7_Enh3_r | TCCAAGGTCACAAAGC | ||
| Emcn_f | CACCAGCACACCTCGTAAAATGG | ||
| Emcn_r | GAGTGAAGTAAGACATCGTCC | ||
| Eng_f | CACCAAACTAATTAAAAAACAAAGCAGGT | ||
| Eng_r | CATATGTACATTAGAACCATCCA | ||
| Ephb4_f | CACCTGGGTCTCATCAACCGAAC | ||
| Ephb4_r | CCTATCTACATCAGGGCACTG | ||
| Ets1_f | CACCTTCGTCAGAAATGATCTTGCCA | ||
| Ets1_r | TAGCAAGAGAGCCTGGTCAG | ||
| Foxc1_f | CACCTCTCTGCTTCAAGGCACCTT | ||
| Foxc1_r | TGGATAGCATGCAGAGGACA | ||
| Gata2_f | CACCTTCTCTTGGGCCACACAGA | ||
| Gata2_r | ATCTGCTCCACTCTCCGTCA | ||
| Lmo2_f | CACCTGGTTTTGCTTGCTAC | ||
| Lmo2_r | CATTTCTAAGTCTCCAC | ||
| Lyve1_f | CACCTACTGCCATGGAGGACTG | ||
| Lyve1_r | AGACACCTGGCTGCCTGATA | ||
| Mef2c_f | CACCGGAGGATTAAAAATTCCCC | ||
| Mef2c_r | CCTCTTAAATGTACGTG | ||
| Notch1_Enh1_f | CACCTCCCAAATGCTCCACGATG | ||
| Notch1_Enh1_r | GAGGAATGGCGAGAAATAGAC | ||
| Notch1_Enh2_f | CACCGAAGGCAGGCAGGAATAAC | ||
| Notch1_Enh2_r | TGGACAGGTGCTTTGTTG | ||
| Sema6d_f | CACCTCTTAACCACTATCTCC | ||
| Sema6d_r | ACTTCCTACACAGTTC | ||
| Sox18_f | CACCTTGGGGGGAAAGAGTG | ||
| Sox18_r | GACTTCATCCCATCTC | ||
| Sox7_f | CACCACAGAGCCCCTGCATATGT | ||
| Sox7_r | GCATGGTTTCTGAAGCCCAAAT | ||
| 3x repeated enhancer regions for luciferase assay. Core motifs of interest are highlighted in red. | |||
| Name | Sequence (5'- > 3') | ||
| ETS-FOX2 (Mef2c) | CAGGAAGCACATTTGTCTACGCTTTCCTGTGAGCAGGAA GCACATTTG TCTACGCTTTCCTGTGAGCAGGAAGCAC ATTTGTCTACGCTTTCCTGTGAG | ||
| Mef2c-mut | CAAGAAGCACATTTGTCTACGCTTTCCTGTGAGCAAGAAGCACATTTG TCTACGCTTTCC TGTGAGCAAGAAGCACATTTGTCTACGCTTTCCTGTGAG | ||
| ETS-FOX3 (Bcl2l1) | CAGTAGATCAGTAGATCAGTAGAT | ||
| ETS-HOMEO2 (Egfl7_Enh1) | GACAGGCGGGACAGGCGGGACAGGCGG | ||
| ETS-HOMEO2 (Egfl7_Enh3) | ACACATTCCTGACACATTCCTGACACATTCCTG | ||
| ETS-HOMEO2 (Flt4) | ACAGTCTTTACAGTCTTTACAGTCTTT | ||
| ETS-HOMEO2 (Kdr) | CAACAGGACAACAGGACAACAGGA | ||
| Sox (Apln) | CAGTTCCTCGCAGTTCCTCGCAGTTCCTCG | ||
| Sox (Robo4) | GCCAGAACAATGAAGCCTGCACGGCCAGAACAATGAAGCCTGCAC GGCCAGAACAATGAAGCCTGCACG | ||
| Sox (Sema3g) | CGAATGGAAAGGGAGGGGAGAACGAATGGAAAGGGAGGGGAGA ACGAATGGAAAGGGAGGGGAGAA | ||
| ETS-Ebox (Apln) | AGGCGGATAGGCGGATAGGCGGAT | ||
| ETS-Ebox (Zfp521) | TTATCCACAAGGATCGTTATCCACAAGGATCGTTATC CACAAGGATCG | ||
| 3x repeated motifs within a similar DNA context. Motifs are indicated in red. | |||
| Neg. control | TGTGAGTGTGAGTGTGAG | ||
| ETS_alone | TGTGAGTGTGAGTGTGAG | ||
| ETS-FOX1 | TGTGAGTGTGAGTGTGAG | ||
| ETS-FOX2 | TGTGAGTGTGAGTGTGAG | ||
| ETS-FOX3 | TGTGAGTGTGAGTGTGAG | ||
| ETS_Ebox | TGTGAGTGTGAGTGTGAG | ||
| ETS-HOMEO2 | TGTGAGTGTGAGTGTGAG | ||
| Tbx-ETS | TGTGAGTGTGAGTGTGAG | ||
Additional files
-
Supplementary file 1
Tissue-specific Ep300-bound regions identified in this study.
Each tab of the excel spreadsheet contains the Ep300-bound regions in the following conditions: Each tab of this spreadsheet shows the Ep300-bound regions in the following samples. H1.narrowPeak: E12.5 heart, replicate 1. H2.narrowPeak: E12.5 heart, replicate 2. FB1.narrowPeak: E12.5 forebrain, replicate 1. FB2.narrowPeak: E12.5 forebrain, replicate 2. Ep300-T-fb: Tie2Cre-enriched regions. Average Ep300 signal in Tie2Cre (T2), Myf5Cre (M5), and EIIaCre (E) is shown in reads per million. T2/E and M5/E show the ratio of signals. Tie2Cre-enriched regions were defined as peak regions with T2/E ratio > 1.5. Ep300-M-fb: Myf5Cre-enriched regions. Average Ep300 signal in Tie2Cre (T2), Myf5Cre (M5), and EIIaCre (E) is shown in reads per million. T2/E and M5/E show the ratio of signals. Myf5Cre-enriched regions were defined as peak regions with M5/E ratio > 1.5. Ep300-E-fb: Ep300-bound peaks called from whole embryo (ubiquitous BirA expression). Ep300-VE-fb: Merged VEcad-CreERT2-driven Ep300 peaks in adult heart and lung. The peaks were ranked by the ratio of Ep300 signal in heart ECs compared to lung ECs and then grouped into deciles.
- https://doi.org/10.7554/eLife.22039.025
-
Supplementary file 2
Transcription factors expressed in embryonic ECs.
E10.5 embryo T2-TRAP and input RNA-seq data were analyzed to identify DNA-binding transcriptional regulators with detectable expression level in ECs (T2-TRAP log2 (fpkm +1)>0.8) and preferential EC expression (T2-TRAP/input >1). Genes with DNA binding domains belonging to the indicated families were annotated based on literature searches and previously publshed catalogs of transcriptional regulators (Fulton et al., 2009; Kanamori et al., 2004).
- https://doi.org/10.7554/eLife.22039.026





