YOD1/TRAF6 association balances p62-dependent IL-1 signaling to NF-κB
Figures
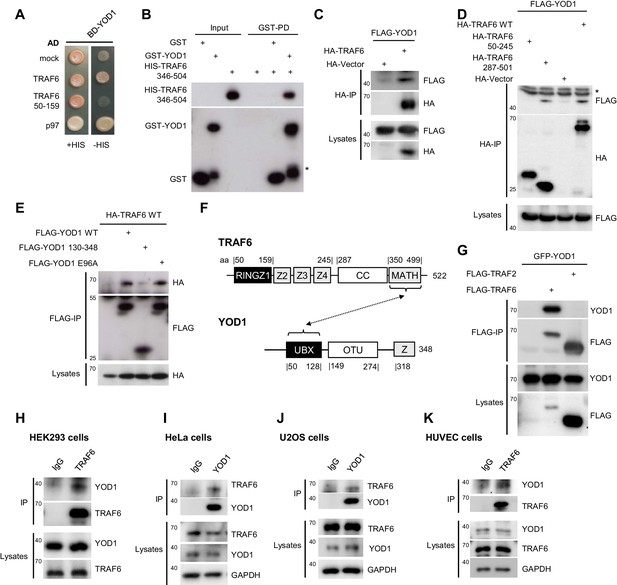
YOD1 interacts with the C-terminal MATH domain of TRAF6.
(A) YOD1 interacts with full length TRAF6 and p97 in a yeast two hybrid assay. Activating domain (AD) and binding domain (BD) fusion constructs were co-transformed as indicated and growth was monitored on -LEU-TRP control (+HIS) and -HIS-LEU-TRP (−HIS) plates. (B) The MATH domain of TRAF6 is sufficient for interaction with YOD1 in vitro. GST-PD were performed with recombinant GST-YOD1 or GST and C-terminal HIS-TRAF6 MATH (346-504) and analyzed by Western Blotting. Asterisk indicates GST-YOD1 truncation product. (C) YOD1 and TRAF6 interact in cells. HEK293 cells were co-transfected with FLAG-YOD1 and HA-TRAF6 or HA-control vector and co-IP was carried out using anti-HA antibodies and analyzed by Western Blot. Asterisk depicts IgGs. (D) YOD1 binds to the C-terminus of TRAF6. YOD1 was co-expressed with TRAF6 deletion or control constructs as indicated. Experiment was performed as in (C). Asterisk depicts IgGs. (E) TRAF6 binds to the UBX domain of YOD1. HA-TRAF6 was co-expressed with FLAG-YOD1, FLAG-YOD1 ΔUBX (130-348) or FLAG-YOD1 E96A. Experiment was performed using anti-FLAG IP as in (C). (F) Schematic summary of the domains required for YOD1/TRAF6 interaction as determined by co-IPs and PDs (compare also Figure 3C). (G) YOD1 does not bind to TRAF2. After transfection of GFP-YOD1 and Flag-TRAF2 or Flag-TRAF6 the experiment was performed using anti-FLAG IP as in C. (H – K) Endogenous interaction of YOD1 and TRAF6. HEK293 (H), HeLa (I), U2OS (J) or HUVEC (K) cells were subjected to TRAF6 (H and K) or YOD1 (I and J) IP as indicated. IgG IP was used as control. Co-precipitation of YOD1 or TRAF6 was analyzed by Western Blotting.
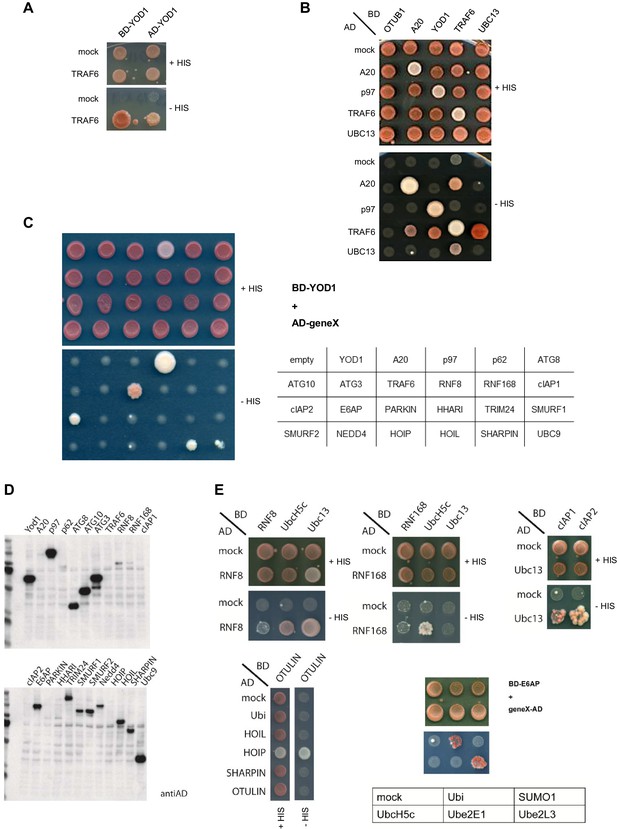

TRAF6/YOD1 interaction in yeast.
(A) AD-YOD1 fusion protein is interacting with BD-TRAF6 and vice versa. YOD1, TRAF6 and mock constructs were co-transformed into the yeast strain PJ69-7A as indicated. AD and BD empty vector were used as control (mock). Co-transformation was confirmed by plating cells on LEU- and TRP- deficient agar plates (+HIS). Interaction of candidate proteins was assessed by plating cells on HIS-, LEU- and TRP- deficient agar plates (-HIS). (B) TRAF6 binds to UBC13, YOD1 and A20 in yeast. Further, TRAF6 and A20 are oligomerizing with themselves and p97 is interacting with YOD1. Experiment was performed as in (A). (C) Specificity test for YOD1 binding in Y2H. A panel of E3 ligases and proteins associated with the ubiquitin system was tested for YOD1 interaction in yeast. TRAF6 and p97 interaction were confirmed and some weaker potential interaction with cIAP2 and SHARPIN was observed. No other interaction was seen. Yeast was co-transformed with BD-YOD1 and the indicated AD constructs. Experiment was performed as in (A). (D) Yeast expression of AD- and BD-constructs used in (C) was verified by Western Blot using anti-AD antibody. (E) Functionality of some selected AD fusion proteins was determined by co-transformation of known interaction partners and interaction was assessed by growth on LEU- and TRP- deficient control plates (+HIS) or HIS-, LEU- and TRP- deficient plates (−HIS) as depicted.
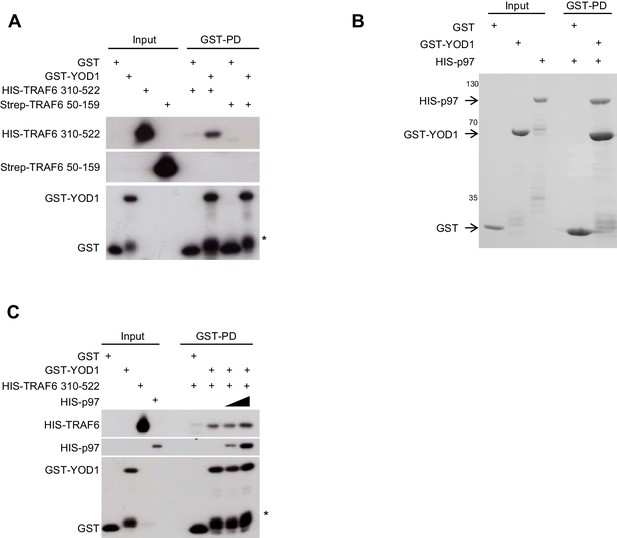
TRAF6/YOD1 interaction is not influenced by p97.
(A) YOD1 is a direct interactor of the TRAF6 C-terminus in vitro. GST-PDs were performed with recombinant GST-YOD1 or GST to test for interaction with N-terminal Strep-TRAF6 RZF1 (50–159) and C-terminal HIS-TRAF6 CC-MATH (310-522). Co-precipitation of interacting proteins was analyzed by Western Blot. (B) Direct interaction of p97 and YOD1 in vitro. GST-PDs were performed with recombinant GST-YOD1 and GST to test for interaction with HIS-p97. Co-precipitation was analyzed by colloidal Coomassie staining of SDS-PAGE. (C) YOD1/TRAF6 interaction is unaffected by YOD1/p97 interaction. GST-PDs were performed with constant amounts of recombinant GST-YOD1 and GST, respectively, and HIS-TRAF6 310–522. Where indicated, HIS-p97 was added in rising amounts. Co-precipitation of interacting proteins was analyzed by Western Blot. Asterisk in (A) and (C) indicates GST-YOD1 truncation product.
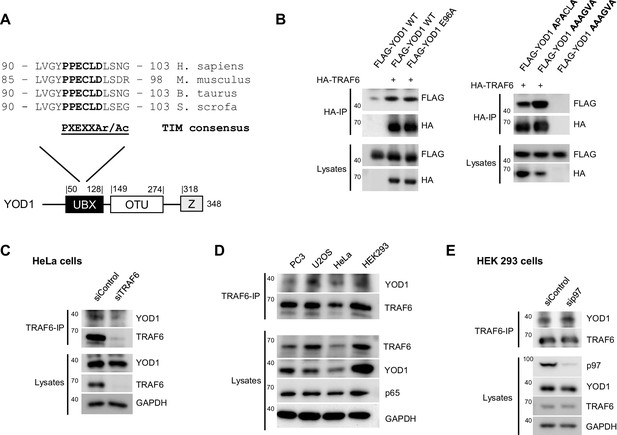
Analysis of YOD1/TRAF6 binding in cells.
(A) One potential TRAF6 interaction motif (TIM) is found in the YOD1 UBX domain. Sequence alignment of YOD1 homologs of different species reveals the existence of a conserved potential TIM in the UBX domain of YOD1. Ar = aromatic, Ac = acidic amino acid, UBX = ubiquitin regulatory X, Z = Zinc Finger (B) TRAF6 binding to YOD1 does not rely on a typical TRAF6 consensus motif in the UBX domain of YOD1. HEK293 cells were co-transfected with HA-TRAF6 and the indicated FLAG-YOD1 constructs with mutations in the putative binding domain. Co-IP was carried out using anti-HA antibodies and analyzed by Western Blot. (C) HeLa cells were transfected with siRNA against TRAF6 (siTRAF6) or control siRNA (siControl) and co-IP was carried out as using anti-TRAF6 antibodies. (D) Comparison of TRAF6 and YOD1 expression and binding in PC3, U2OS, HeLa and HEK293 cells. Protein amounts were adjusted prior to TRAF6-IP. Protein expression in lysates and YOD1 binding after IP was analyzed by Western Blotting. (E) Endogenous YOD1/TRAF6 interaction is independent of p97. HEK293 cells were transfected with siRNA against p97 (sip97) or siControl and after cell lysis, anti-TRAF6 IP was performed and analyzed by Western Blot.
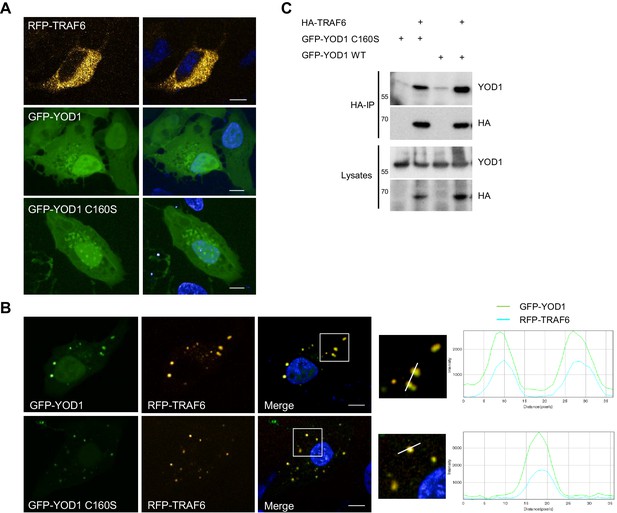
YOD1 co-localizes with TRAF6 in cytosolic speckles.
(A) Diffuse localization of TRAF6 and YOD1 upon individual expression. RFP-TRAF6, GFP-YOD1 or GFP-YOD1 C160S were overexpressed in U2OS cells and localization was analyzed by confocal fluorescence microscopy. (B) YOD1 and TRAF6 co-localize in cytosolic speckles upon co-expression. The co-localization is independent of YOD1 catalytic activity. GFP-YOD1 (WT or C160S) and RFP-TRAF6 were co-transfected in U2OS cells and localization was analyzed as in (A). Enlargement of boxed area is shown next to Merge. Plot Profile analysis was conducted to quantify fluorescence intensities and to monitor co-localization along the white line. (C) TRAF6 interacts with YOD1 independent of its catalytic activity. HEK293 cells were co-transfected with HA-TRAF6, GFP-YOD1 WT and GFP-YOD1 C160S constructs as indicated. Co-IP was carried out using anti-HA antibodies and analyzed by Western Blot. Merged pictures include nuclear staining with Hoechst 33342. Scale bars depict 10 µM (A and B).
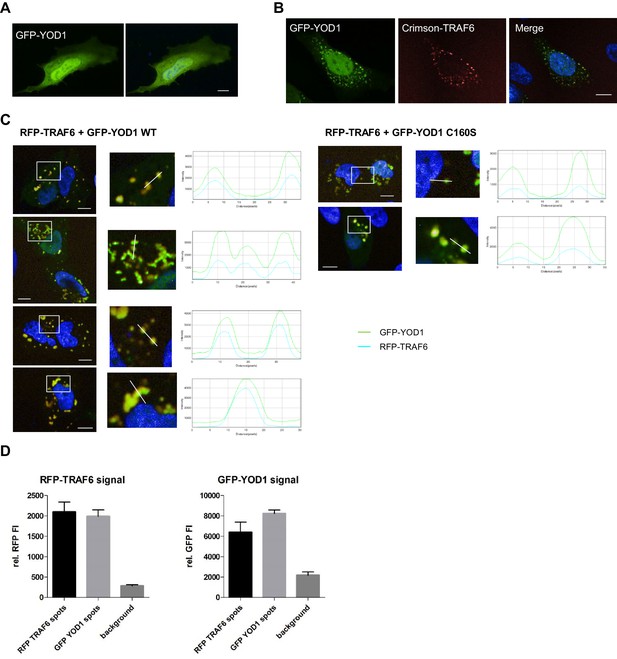
YOD1/TRAF6 co-localization.
(A) YOD1 localizes diffusely in cytoplasm and nucleus of HeLa cells. GFP-YOD1 was transfected in HeLa cells and localization was analyzed by confocal fluorescence microscopy following fixation. (B) YOD1 and TRAF6 co-localize upon overexpression in HeLa cells. GFP-YOD1 and Crimson-TRAF6 were co-transfected in HeLa cells and localization was analyzed as in (A). (C) Example pictures of YOD1 WT (left panel) and C160S (right panel) recruitment to TRAF6 (corresponding to Figure 2B). GFP-YOD1 WT or C160S was expressed together with RFP-TRAF6 in U2OS cells and localization was analyzed by confocal fluorescence microscopy. Only merged images with enlargement of boxed area are shown. Plot Profile analysis was conducted along the white line. Nuclear stainings were performed with Hoechst 33342. Scale bars depict 10 µM. (D) Quantitative measurement of TRAF6-YOD1 co-localization. Co-clustering of fluorescence intensities (FI) was determined by automated fluorescence imaging in >200 co-transfected U2OS cells. For determining RFP or GFP fluorescence background, cells were defined by GFP staining and FI was measured in the whole cell area. RFP signal was enriched in RFP-TRAF6 as well as GFP-YOD1 spots (left) and vice versa GFP signal was enriched in GFP-YOD1 as well as RFP-TRAF6 dots (right), demonstrating co-localization of RFP-TRAF6 and GFP-YOD1. Data depict the mean and standard deviation (SD) of measured mean FI.
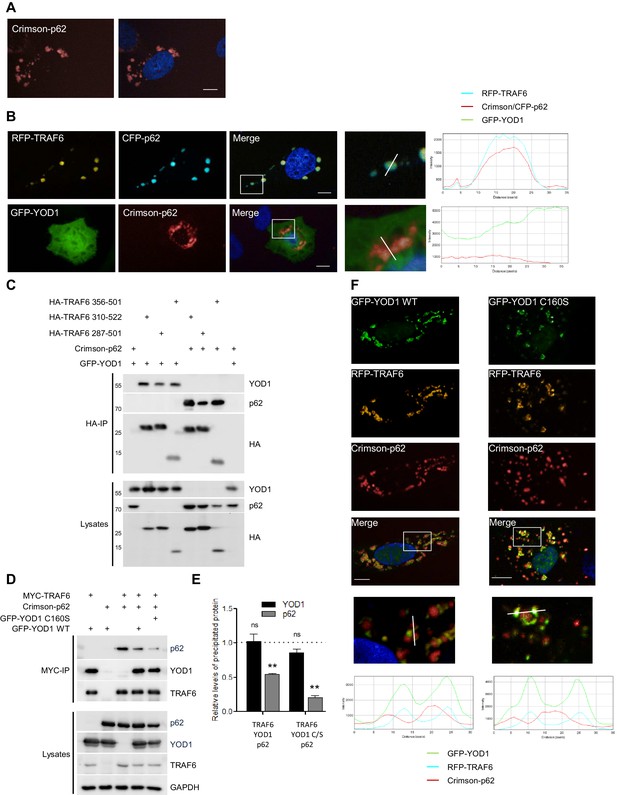
YOD1 competes with p62 for binding to TRAF6 and recruitment to sequestosomes.
(A) p62 localizes to sequestosomes. Crimson-p62 was transfected in U2OS cells and localization was analyzed by confocal fluorescence microscopy. (B) TRAF6, but not YOD1, is recruited to p62-containing aggregates. RFP-TRAF6 and CFP-p62 or GFP-YOD1 and Crimson-p62 were co-transfected in U2OS cells and localization was analyzed as in (A). Enlargement of boxed area is shown next to Merge. Plot Profile analysis was conducted to quantify fluorescence intensities and monitor co-localization along the white line. (C) YOD1 and p62 bind to the C-terminal MATH domain of TRAF6. HEK293 cells were co-transfected with the indicated constructs. Co-IP was carried out using anti-HA antibodies and analyzed by Western Blot. (D and E) YOD1 impedes p62/TRAF6 interaction. (D) HEK293 cells were co-transfected with GFP-YOD1 WT, GFP-YOD1 C160S, Crimson-p62 and MYC-TRAF6 constructs as indicated. co-IP was carried out using anti-MYC antibodies and analyzed by Western Blot. (E) For quantification of Figure 3D and two additional experiments, amounts of YOD1 or p62 bound to TRAF6 in double transfected cells were set to 1. Changes in binding upon co-expression of all three proteins were measured using LabImage 1D software. Data depict the mean and standard error of the mean (SEM) of three independent experiments. Significance for the decrease p62 and YOD1 versus control was evaluated using Student’s t-test (**p<0,01; ns = not significant). (F) YOD1 WT and C160S diminish recruitment of TRAF6 to p62 aggregates. GFP-YOD1 WT or C160S, respectively, RFP-TRAF6 and Crimson-p62 were co-expressed in U2OS cells and localization was analyzed as in (A). Enlargement of boxed area is shown below Merge. Plot Profile analysis was conducted along the white line. Merged pictures include nuclear staining with Hoechst 33342. Scale bars depict 10 µM (A, B and F).
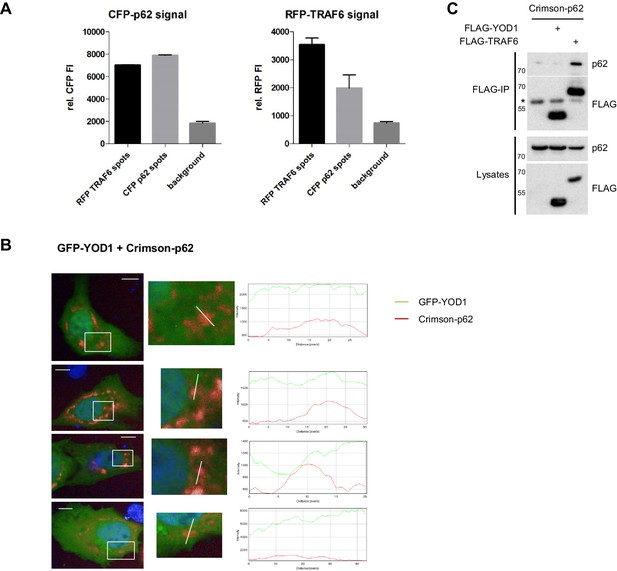
TRAF6, but not YOD1, is interacting with p62.
(A) Quantitative measurement of TRAF6-p62 co-localization. Co-clustering of FI was determined by automated fluorescence imaging in >150 co-transfected U2OS cells. For determining RFP or CFP fluorescence background, nuclei were defined by the Hoechst33342 staining and FI was measured around this area. The CFP signal is enriched in RFP-TRAF6 as well as CFP-p62 spots (left) and vice versa RFP signal is enriched in CFP-p62 as well as RFP-TRAF6 dots (right), showing co-localization of RFP-TRAF6 and CFP-p62. Data depict the mean and standard deviation (SD) of measured mean FI. (B) Example pictures showing that YOD1 and p62 are not specifically co-localizing (corresponding to Figure 3B). GFP-YOD1 WT was expressed together with Crimson-p62 in U2OS cells and localization was analyzed by confocal fluorescence microscopy. Only merged images with enlargement of boxed area are shown. Plot Profile analysis was conducted along the white line. Nuclear stainings were performed with Hoechst 33342. Scale bars depict 10 µM. (C) p62 interacts with TRAF6, but not with YOD1. HEK293 cells were transfected as indicated and co-IP was carried out using anti-FLAG antibodies. Asterisk indicates migration of IgGs.
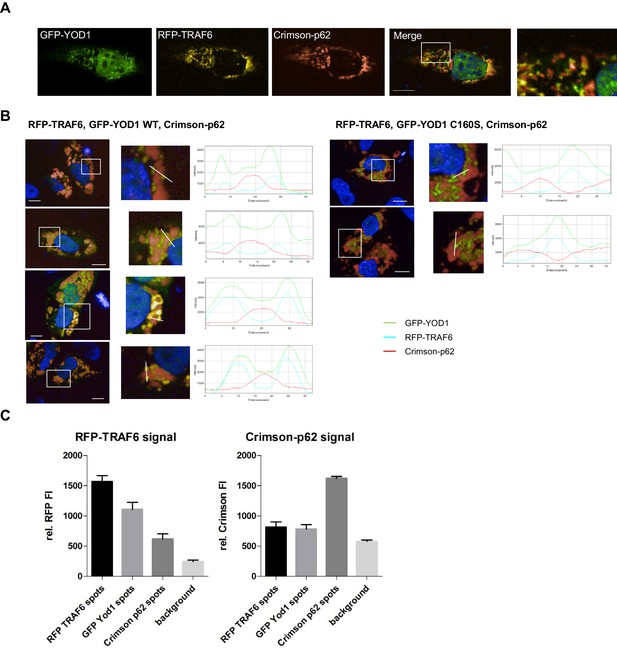
YOD1/p62 competition for TRAF6 binding.
(A) YOD1 competes with p62 for TRAF6 co-localization in HeLa cells. GFP-YOD1, RFP-TRAF6 and Crimson-p62 were co-expressed in HeLa cells and localization was analyzed by confocal fluorescence microscopy. Enlargement of boxed area is shown next to Merge. Merge pictures show nuclear staining using Hoechst 33342. Scale bar depicts 10 µM. (B) Example pictures of YOD1 WT (left panel) and C160S (right panel) recruitment to TRAF6 and prevention of TRAF6/p62 aggregates (corresponding to Figure 3F). GFP-YOD1 WT or C160S (right) were expressed together with RFP-TRAF6 and Crimson-p62 in U2OS cells and localization was analyzed by confocal fluorescence microscopy. Only merged images with enlargement of boxed area are shown. Plot Profile analysis was conducted along the white line. Nuclear stainings were performed with Hoechst 33342. Scale bars depict 10 µM. (C) Quantitative analyses of TRAF6/YOD1 and TRAF6/p62 co-localization in triple transfected U2OS cells. Co-clustering of FI was determined by automated fluorescence imaging in >100 triple transfected U2OS cells (PerkinElmer Imaging). For determining RFP or Crimson fluorescence background, cells were defined by GFP staining and FI was measured in this area. The RFP signal is enriched in RFP-TRAF6 as well as GFP-YOD1 and to a lesser extent in Crimson-p62 spots (left). Crimson-p62 is enriched neither in RFP-TRAF6 nor in GFP-YOD1 spots compared to background levels. Data depict the mean and standard deviation (SD) of measured mean FI.
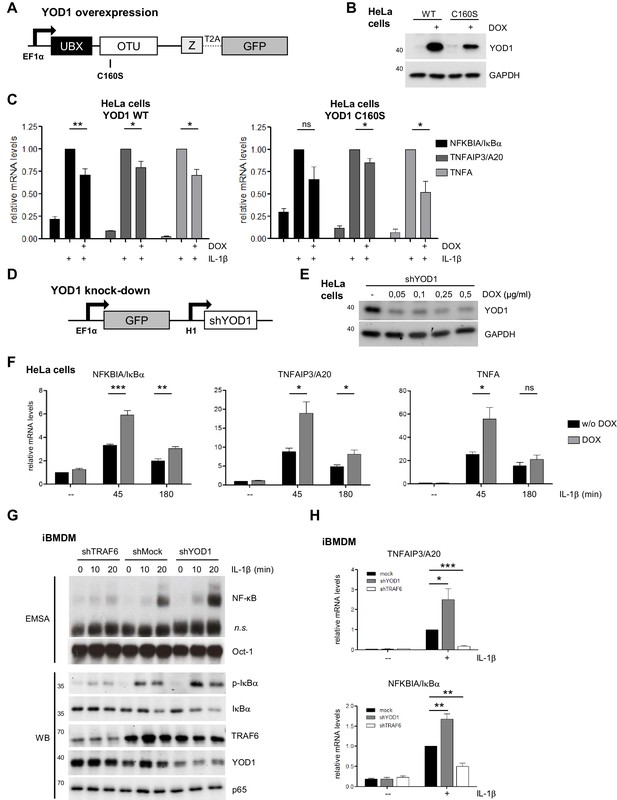
YOD1 is a negative regulator of IL-1β-induced NF-κB signaling.
(A) Schematic representation of YOD1 overexpression constructs. YOD1 WT or C160S and GFP were co-expressed using T2A site under the control of EF1α promoter, which in turn is DOX/tTR-KRAB-controlled. (B) YOD1 WT and YOD1 C160S are overexpressed upon doxycycline (DOX) treatment of lentivirally transduced HeLa cells. Transduced cells were grown in DOX containing medium for 72 hr and after cell lysis subjected to Western Blotting. (C) YOD1 WT (left panel) or C160S (right panel) overexpression diminishes NF-κB target gene expression. Infected HeLa cells were treated with DOX for 72 hr and stimulated with IL-1β for 60 min. Expression of indicated transcripts was analyzed by qRT-PCR. Bars show mean and standard error of the mean (SEM) of five independent experiments. (D) Schematic representation of YOD1 shRNA construct. GFP and shYOD1 were expressed under control of EF1α and H1 promoter, respectively. Both promoters are DOX/tTR-KRAB-controlled. (E) YOD1 protein levels are reduced in shYOD1 cells. Cells were treated for 72 hr with 0,05–0,5 µg/ml DOX as indicated and YOD1 knock-down was analyzed by Western Blot. (F) YOD1 knock-down results in enhanced NF-κB target gene expression. shYOD1-infected HeLa cells were treated with DOX for 72 hr and stimulated with IL-1β for the indicated time points. RNA was isolated and transcripts were analyzed by qRT-PCR as indicated. Bars show mean and SEM of four independent experiments. (G) TRAF6 and YOD1 exert opposing effects on NF-κB signaling and activation in iBMDM. iBMDM transduced with control shMock, shTRAF6 or shYOD1 were stimulated with IL-1β as indicated. NF-κB and Oct-1 (control) DNA binding was assessed by EMSA (n.s. = non-specific band). IκBα phosphorylation, degradation and knock-down efficiencies were analyzed by Western Blotting. (H) YOD1 knock-down promotes, while TRAF6 depletion impairs NF-κB target gene expression in iBMDM. iBMDM transduced as in (G) were stimulated with IL-1β for 45 min. Transcript levels were analyzed by qRT-PCR as indicated. Bars show mean and SEM of seven independent experiments. Significance was evaluated using Student’s t-test (*p<0,05; **p<0,01; ***p<0001; ns = not significant).
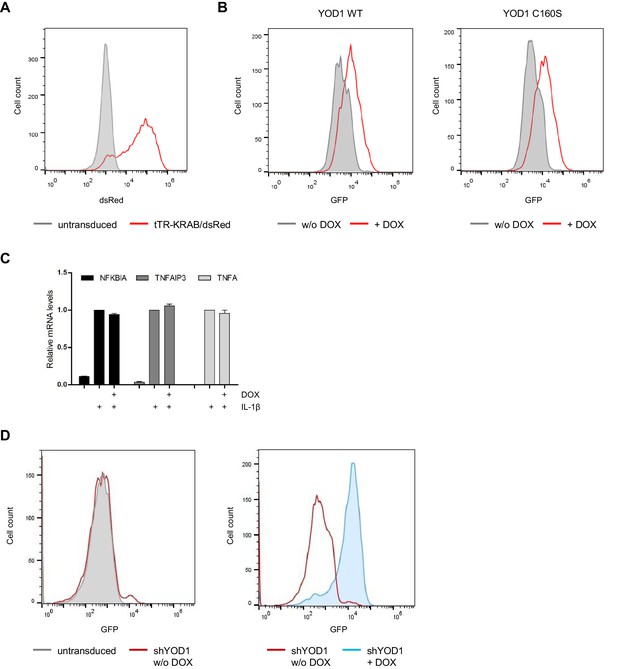
Lentiviral transduction and DOX control treatment of HeLa cells.
(A) HeLa cells are efficiently transduced with tTR-KRAB-dsRed constructs. After the first infection with tTR-KRAB-T2A-dsRed, cells were analyzed for dsRed expression by FACS. (B) YOD1-T2A-GFP transduction in HeLa cells. Following tTR-KRAB-T2A-dsRed infection, cells were transduced with YOD1 (WT or C160S)-T2A-GFP containing vectors. Cells were analyzed by FACS and sorted for GFP expression. GFP expression was induced by treatment with DOX for 72 hr. (C) DOX treatment does not affect NF-κB target gene expression in HeLa parental cells. HeLa cells were treated with DOX for 72 hr and stimulated with IL-1β for 60 min. Expression of indicated transcripts was analyzed by qRT-PCR. Bars show mean and standard error of the mean (SEM) of four independent experiments. (D) HeLa cells are efficiently transduced with shYOD1. tTR-KRAB-T2A-dsRed expressing cells were transduced with shYOD1 containing lentivirus. Cells show almost no leakiness (-DOX, left panel). shYOD1 and GFP expression is efficiently induced by DOX-treatment for 72 hr (right panel).
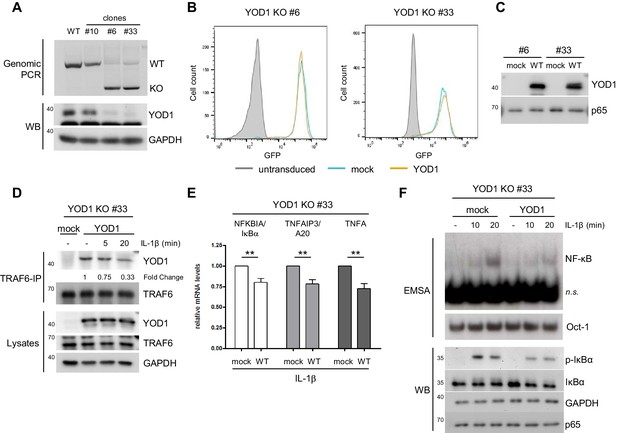
Reconstitution of YOD1-deficient HeLa cells impairs IL-1β-induced NF-κB signaling.
(A) Validation of YOD1 KO HeLa cell clones. YOD1 genomic DNA and protein levels in parental HeLa cells and in cell clones generated by CRISPR/Cas9 gene editing were checked by PCR and Western Blot. (B) YOD1 deficient HeLa clones #6 and #33 are efficiently transduced with empty vector (mock) and YOD1 WT. Cells were transduced and homogenous populations of GFP expressing cells were sorted by FACS. FACS of GFP expression after sorting is shown. (C) Reconstitution of YOD1-deficient cell clones #6 and #33 with YOD1 WT. YOD1-deficient HeLa cells were transduced with YOD1 WT or mock constructs and YOD1 expression was analyzed by Western Blot. (D) YOD1/TRAF6 interaction in reconstituted YOD1-deficient HeLa clone #33 is decreasing upon IL-1R engagement. Cells were stimulated with IL-1β for the indicated time points. Anti-TRAF6 IPs were conducted and interaction of YOD1 was analyzed by Western Blot. Quantification of YOD1 bound to TRAF6 is shown. Numbers indicate the fold change after IL-1β stimulation (unstimulated set to 1). (E) Reconstitution of YOD1-deficient HeLa clone #33 with YOD1 WT diminishes NF-κB target gene expression. Cells were stimulated with IL-1β for 40 min. RNA was isolated and transcripts were analyzed by qRT-PCR as indicated. Bars show mean and SEM of seven independent experiments. Significance was evaluated using Student’s t-test (*p<0,05; **p<0,01; ***p<0001; ns = not significant). (F) YOD1 re-expression in YOD1-deficient HeLa clone #33 diminishes NF-κB activation and IκBα phosphorylation and degradation. Cells were stimulated with IL-1β for the indicated time points and NF-κB DNA binding was assessed by EMSA (n.s. = non-specific band). Oct-1 EMSA served as loading control. IκBα phosphorylation and degradation was analyzed by Western Blot.
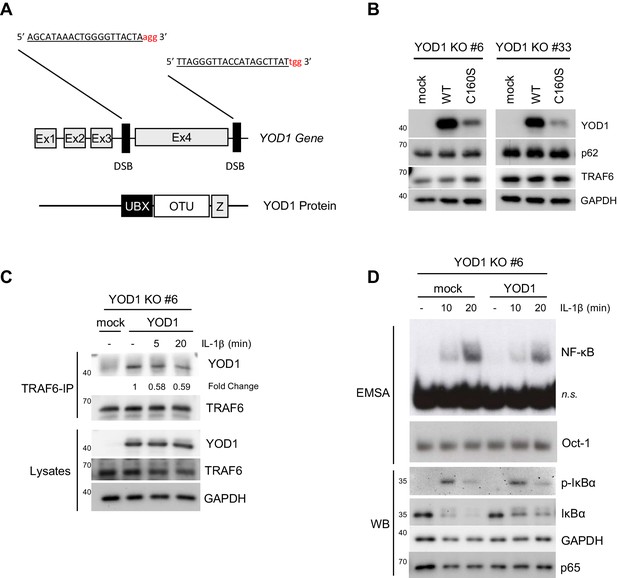
Generation, reconstitution and analyses of YOD1-deficient HeLa cells.
(A) Schematic representation of the Cas9/sgRNA-targeting sites in the YOD1 gene. sgRNA-targeted sequences are underlined and the protospacer-adjacent motif (PAM) is labeled in red. Induced double-strand breaks (DSB) are marked. Ex = Exon. (B) Expression of YOD1 WT and YOD1 C160S after reconstitution of YOD1 deficient HeLa clones #6 and #33. YOD1 expression was analyzed by Western Blotting. (C) YOD1-TRAF6 interaction in reconstituted YOD1-deficient cell clone #6 is partially lost upon IL-1R engagement. Cells were stimulated with IL-1β for the indicated time points. Anti-TRAF6 co-IPs were conducted and co-precipitation of YOD1 was analyzed by Western Blot. Quantification of YOD1 bound to TRAF6 is shown. Numbers indicate the fold change after IL-1β stimulation (unstimulated set to 1). (D) Reconstitution of YOD1-deficient cell clone #6 with YOD1 WT diminishes IκBα degradation and NF-κB activation. Cells were stimulated with IL-1β for the indicated time points and NF-κB DNA binding was assessed by EMSA (n.s. = non-specific band). Oct-1 EMSA served as loading control. IκBα phosphorylation and degradation was analyzed by Western Blot.
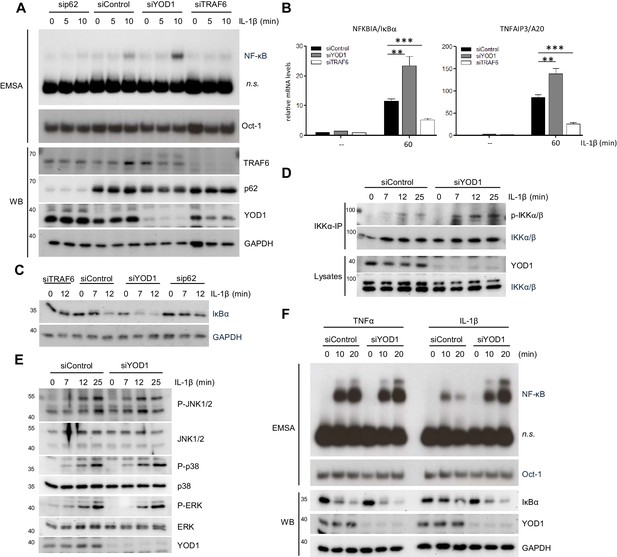
YOD1 and TRAF6/p62 exert opposing effects on IL-1β-induced NF-κB activation.
(A) YOD1 knock-down promotes while TRAF6 and p62 knock-down impair NF-κB activation. HeLa cells were transfected with control siRNA or siRNA targeting YOD1, TRAF6 or p62 and stimulated with IL-1β as indicated. NF-κB and Oct-1 (control) DNA binding was assessed by EMSA (n.s. = non-specific band). Knock-down efficiency was confirmed by Western Blotting. (B) YOD1 knock-down promotes, while TRAF6 depletion impairs NF-κB target gene expression. HeLa cells were transfected with siRNA as indicated and subsequently stimulated with IL-1β for 60 min. Transcript levels were analyzed by qRT-PCR as indicated. Bars show mean and SEM of four to five independent experiments. Significance was evaluated using Student’s t-test (**p<0,01; ***p<0001). (C) IκBα degradation is enhanced after YOD1 knock-down, but inhibited after TRAF6 or p62 knock-down. HeLa cells were transfected with siRNAs and stimulated with IL-1β as indicated. IκBα degradation was analyzed by Western Blot. (D) YOD1 is a negative regulator of IKK T-loop phosphorylation upon IL-1R engagement. HeLa cells were transfected with siControl or siYOD1 and stimulated with IL-1β as indicated. Anti-IKKα-IP was carried out to precipitate IKKα and IKKβ, and IKKα/β phosphorylation was analyzed by Western Blot. (E) YOD1 knock-down does not affect MAPK activation. HeLa cells were transfected with siRNA as in (D) and stimulated with IL-1β for the indicated time points. After cell lysis, MAPK activation was determined by Western Blotting using phospho-specific antibodies. (F) YOD1 specifically regulates IL-1β-, but not TNFα-induced NF-κB signaling. HeLa cells were transfected with siRNA as in (D) and stimulated with IL-1β or TNFα as indicated. NF-κB and Oct-1 (control) DNA binding was assessed by EMSA (n.s. = non-specific band). IκBα degradation and knock-down efficiency was confirmed by Western Blot.
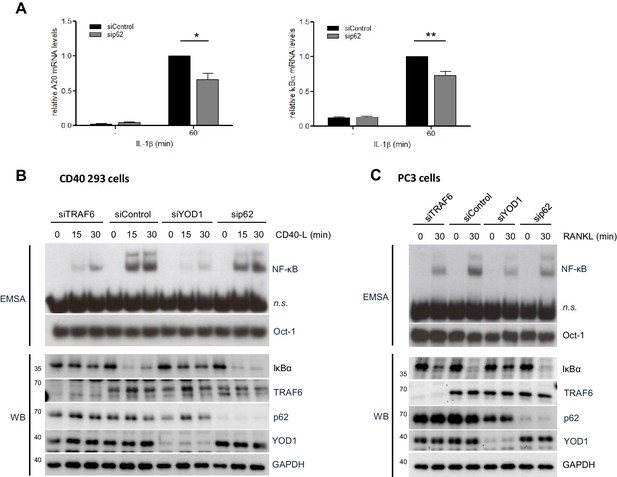
Functional impact of TRAF6, p62 and YOD1 on CD40 and RANK stimulation.
(A) p62 is required for NF-κB target gene induction upon IL-1β stimulation. HeLa cells were transfected with siRNA against p62 and stimulated with IL-1β for 60 min. Transcript levels were analyzed by qRT-PCR as indicated. Bars show means and SEM of seven independent experiments. Significance was evaluated using Student’s t-test (*p<0,05; **p<0,01). (B) YOD1, but not p62, is required for TRAF6-dependent CD40 stimulation in CD40 expressing 293 cells. CD40 293 cells were transfected with control siRNA or siRNA targeting YOD1, TRAF6 or p62 and stimulated with CD40 ligand as indicated. NF-κB and Oct-1 (control) DNA binding was assessed by EMSA (n.s. = non-specific band). IκBα degradation and knock-down efficiency was confirmed by Western Blotting. (C) YOD1, but not p62, is required for TRAF6-dependent RANK stimulation in PC3 cells. PC3 cells were transfected with siRNA and after over-night serum starvation stimulated with RANK-L for the indicated time points. Extracts were analyzed as in (B).
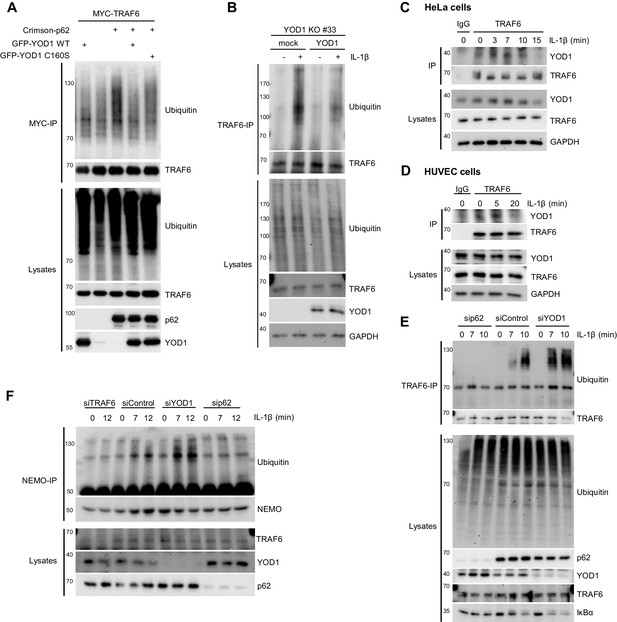
YOD1 counteracts TRAF6/p62-triggered ubiquitination.
(A) YOD1 prevents augmented TRAF6 ubiquitination upon p62 binding. MYC-TRAF6, Crimson-p62, GFP-YOD1 WT and GFP-YOD1 C160S were co-transfected in HEK293 cells as indicated. After cell lysis under denaturing conditions (1% SDS), anti-MYC IP was conducted. TRAF6 ubiquitination was analyzed by Western Blot. (B) Reconstitution of YOD1-deficient HeLa cells diminishes TRAF6 ubiquitination. Mock or YOD1 reconstituted HeLa clone#33 was stimulated with IL-1β for 8 min and TRAF6 ubiquitination was analyzed as in (A), using anti-TRAF6 antibodies for IP. (C and D) Endogenous YOD1/TRAF6 interaction is lost upon IL-1β stimulation. HeLa cells (C) and HUVEC (D) were stimulated with IL-1β for the indicated time points and co-IPs were conducted using anti-TRAF6 or IgG antibodies. Co-IP of YOD1 was analyzed by Western Blot. (E) YOD1 knock-down promotes while p62 depletion inhibits IL-1-induced TRAF6 ubiquitination. HeLa cells were transfected with siRNAs and stimulated with IL-1β as indicated. TRAF6 ubiquitination was analyzed as in (B). (F) YOD1 knock-down promotes while TRAF6 and p62 knock-down impede NEMO ubiquitination. Experiment was essentially conducted as in (E), using anti-NEMO antibodies for IP.
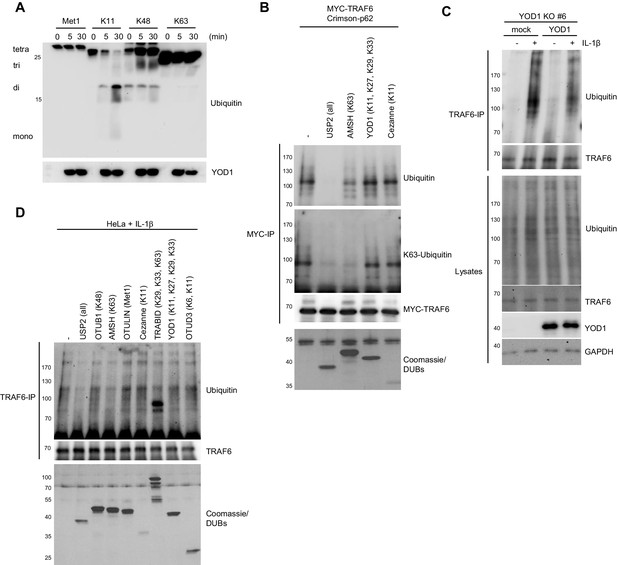
TRAF6 poly-ubiquitination mainly consists of YOD1-resistant K63 linkages.
(A) YOD1 readily cleaves K11-, but not Met1-, K48- and K63-linked ubiquitin chains. Recombinant GST-YOD1 was incubated with the respective tetra-ubiquitin chains and after the indicated time points, the reaction was stopped and samples were analyzed by Western Blotting. (B) TRAF6 poly-ubiquitination upon co-expression with p62 mainly consist of K63 linkages. MYC-TRAF6 and Crimson-p62 were co-transfected in HEK293 cells. After anti-MYC-IP, sample was split and incubated with the recombinant DUBs USP2, AMSH, YOD1 and Cezanne as indicated. Chain restriction was analyzed by Western Blotting, while presence of DUBs was confirmed by Coomassie-staining of SDS-PAGE. (C) Re-introduction of YOD1 in YOD1-deficient HeLa cell clone #6 diminishes TRAF6 ubiquitination. Cells were stimulated with IL-1β for 8 min and after cell lysis under denaturing conditions (1% SDS), anti-TRAF6 IP was conducted and TRAF6 ubiquitination was analyzed by Western Blot. (D) Ubiquitin chains conjugated to TRAF6 upon IL-1β stimulation constitute K63-linkages. HeLa cells were stimulated as in (C) and endogenous TRAF6 was precipitated by IP. After extensive washing, sample was split to incubate with the indicated DUBs. Chain restriction was analyzed by Western Blotting and Coomassie staining as in B.





