A nanobody-based toolset to investigate the role of protein localization and dispersal in Drosophila
Figures
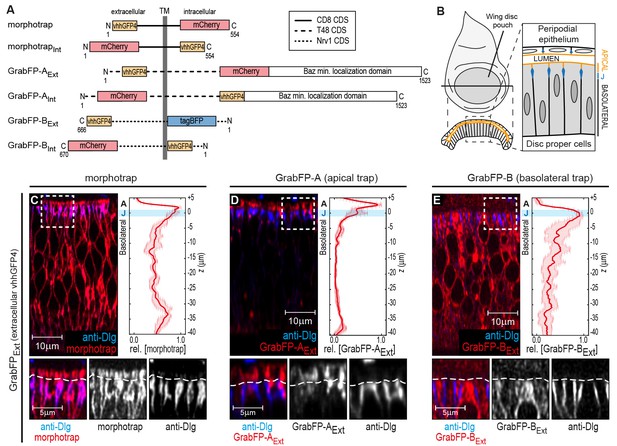
The GrabFP constructs localize to distinct regions along the apical-basal axis.
(A) Linear representation of the six different versions of the GrabFP system; the constructs exist in two topologies with the GFP-nanobody (vhhGFP4) either facing extracellular (Ext) or intracellular (Int). Numbers refer to the amino acid positions from the N-terminus (N) to the C-terminus (C). TM = transmembrane domain, CDS=coding DNA sequence. (B) Schematic representation of wing disc morphology, the junctions (J) are marked in blue. (C–E) Cross-sections of wing discs expressing morphotrap (C), GrabFP-AExt (D) and GrabFP-BExt (E) in the wing pouch (nub::Gal4). The GrabFP tools are shown in red and the junctions are visualized by staining for Dlg (blue). In the magnifications the junctional level is marked by a dashed line. Relative distribution of the GrabFP tools along the A-B axis in respect to the junctions (marked by Dlg) is quantified in the plots to the right (n = 4 for each plot, error bars represent the standard deviation). For details on the quantification see Materials and methods and Figure 1—figure supplement 3.
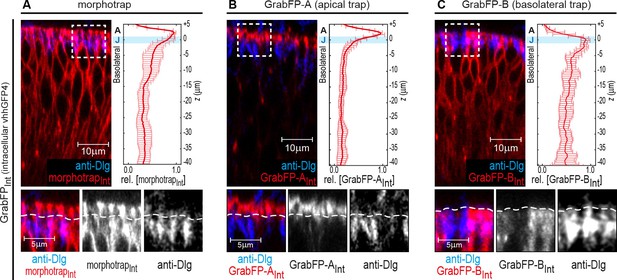
Localization of the GrabFPIntra tools.
(A–C) Optical cross-sections of wing discs expressing the intracellular versions of morphotrapInt (A), GrabFP-AInt (B) and GrabFP-BInt (C) in the disc proper (ptc::Gal4). The GrabFP constructs are shown in red and the junctions are visualized by staining for Dlg (blue). The junctional level is marked by dashed lines in the magnifications (bottom). Quantification of protein localization along the A-B axis is shown in the graphs to the right. (morphotrapInt n = 6, GrabFP-AInt n = 7, GrabFP-BInt n = 6, error bars represent the standard deviation). For details on the quantification see Materials and methods section and Figure 1—figure supplement 3.
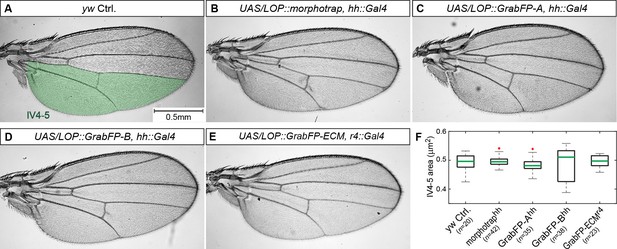
Expression of the GrabFP system allows normal wing development.
(A–E) Male wings of indicated genotypes. Expression of the GrabFPExt tools (using hh::Gal4 (A–D) or r4::Gal4 (E)) does not interfere with wing development and yields viable and fertile flies. Solely expression of GrabFP-AExt in the posterior compartment results in slightly rounder wing shape (compare A to C). (F) Quantification of intervein area between vein 4 and the posterior wing margin (IV4-5), as marked in (A). None of the genotypes showed significantly reduced or increased wing blade area due to the expression of the GrabFP tools. Significance was assessed using a two-sided Student’s t-test with unequal variance (p-values: morphotrap p=0.72, GrabFP-Ahh p = 0.17, GrabFP-Bhh p = 0.61, GrabFP-ECMr4 p=0.55, outliers are marked by a red cross).
-
Figure 1—figure supplement 2—source data 1
Source data for wing area quantification.
Only male wings were used. Intervein area IV4-5 in µm2 as indicated in pannel A.
- https://doi.org/10.7554/eLife.22549.005
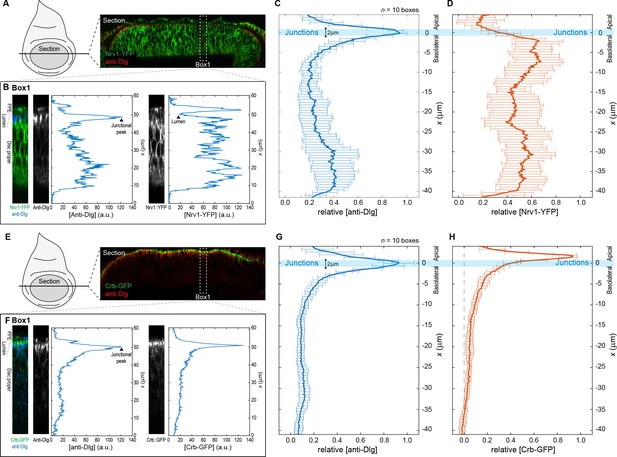
Quantification and analysis of protein distribution along the A-B axis.
Procedure for obtaining relative concentration profiles along the A-B axis of DP cells for the basolateral marked Nrv1-YFP (A–D) and the apical marked Crb-GFP (E–H). (A) Optical cross-section of a wing disc expressing Nrv1-YFP (green) and stained for Dlg (blue) as obtained when using the reslice function in ImageJ (NIH). (B) Single fluorescence intensity profiles of anti-Dlg and Nrv1-YFP fluorescence extracted from a rectangular area of 16 µm width (e.g. Box1) using the plot profile function of ImageJ. (C–D) Individual profiles as extracted in (B) were aligned according to the position of the junctional peak of the Dlg signal and merged to average concentration profiles. Average profiles of n = 10 sections are shown for Dlg (C) and Nrv1-YFP (D). The junctions are defined as the region 1 µm above and below the average Dlg peak (light blue bar). Nrv1 localization is restricted to the basolateral compartment, and indeed our quantifications show that Nrv1-YFP levels are high along the basolateral compartment but drop at the junctions. (E–H) Similar steps as in (A–D) for the extraction of average Crb-GFP profiles. Crb is a determinant of apical compartment identity and exclusively localizes to the apical compartment as visualized by the quantification.
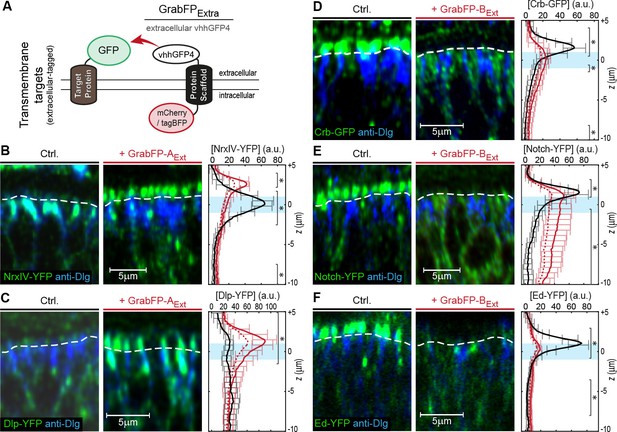
Mislocalization of transmembrane proteins using the GrabFPExt system.
(A) In the GrabFPExt system, the GFP-nanobody (vhhGFP4) faces the extracellular space and can interact with extracellular-tagged transmembrane proteins. (B–C) Optical cross-section of wing disc cells expressing either NrxIV-YFP (B) or Dlp-YFP (C) alone (Ctrl., left) or together with GrabFP-AExt (middle). The junctional level is marked by a dashed line. Quantification of absolute target protein localization (right) along the A-B axis in the absence (black) or in the presence of GrabFP-AExt (continuous red line). Dashed lines represent profiles corrected for increased GFP/YFP fluorescence due to nanobody binding. The position of the junctions is marked by a blue bar. (Error bars show the standard deviation). (D–F) Optical cross sections showing the localization of Crb-GFP (D), Notch-YFP (E) or Ed-YFP (F) in the absence (left) or in the presence of GrabFP-BExt (middle). Quantifications are shown to the right. (Sample numbers for plots in B-F: NrxIV n = 10, Dlp n ≥ 8, Notch n ≥ 8, Crb n = 8, Ed n ≥ 6, significance was assessed comparing wild type with corrected profiles using a two-sided Student’s t-test with unequal variance, *p<0.05).
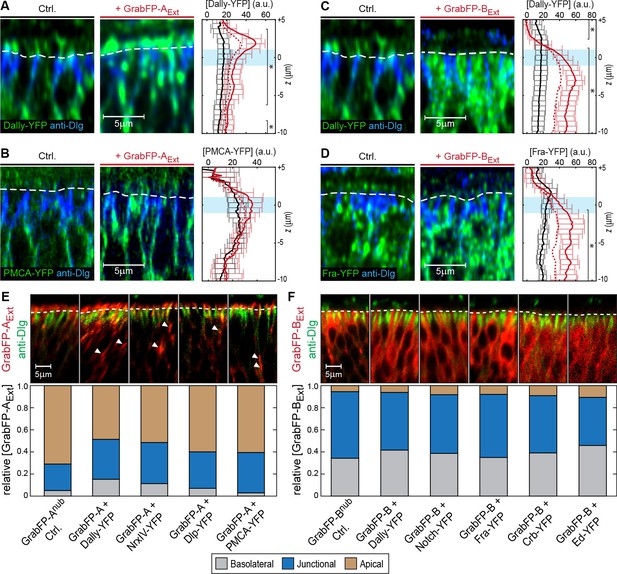
Examples of target protein mislocalization using the GrabFPExtra system.
(A–B) Effects of GrabFP-AExt expression on the localization of Dally-YFP (A) and PMCA-YFP (B). Optical cross-sections of target proteins alone (Ctrl., left) or co-expressed with GrabFP-AExt (middle). Quantification of target-protein levels (left) in the absence (black) or the presence of GrabFP-AExt (continuous red line). Corrected profiles are indicated by a dashed red line. (C–D) Localization of Dally-YFP (C) and Fra-YFP (D) in control conditions (left) and when co-expressed with GrabFP-BExt (middle), quantification is shown to the left. (E–F) Representative cross-sections (top) and quantification (bottom) of relative GrabFP-AExt (E) and GrabFP-BExt (F) localization when expressed alone (Ctrl.) or when co-expressed with YFP/GFP-tagged target protein. Target proteins of basolateral localization tend to mislocalize GrabFP-AExt (E) toward the junctional/basolateral compartment (see reduction in relative apical localization [brown]). In contrast, GrabFP-BExt (F) is more resistant to mislocalization by apically localizing target proteins and shows only slight increases in apical localization. Sample numbers for shown quantifications in A-F: GrabFP-AExt n = 4, +Dally n = 10, +NrxIV n = 6, +Dlp n = 8, +PMCA n = 5, GrabFP-BExt n = 4, +Dally n = 10, +Notch n = 8, +Fra n = 9, +Crb n = 9, +Ed n = 6. Statistical significance was assessed comparing control with corrected profiles using a two-sided Student’s t-test with unequal variance (*p<0.05).
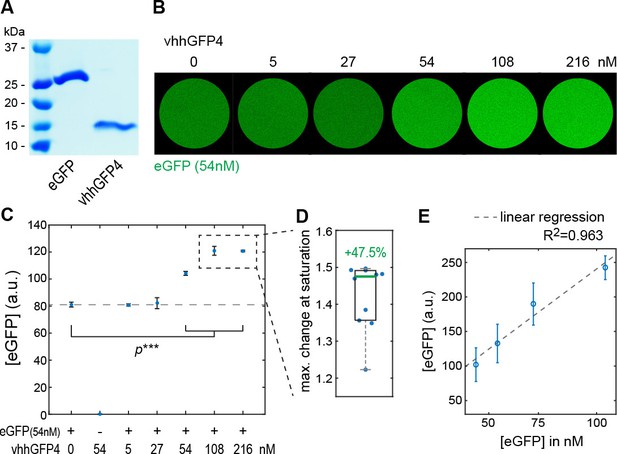
Modulation of EGFP fluorescent properties by vhhGFP4 binding in vitro.
(A) Coomassie staining of SDS-PAGE loaded with purified eGFP (25 kDa) and vhhGFP4 (14 kDa). See Materials and methods for details. (B) Fluorescence in vitro essay to estimate changes in eGFP fluorescence upon binding to vhhGFP4. Titration of defined amounts of purified vhhGFP4 on 54 nM purified eGFP. (C) Quantification of eGFP fluorescence as shown in (B). vhhGFP4 concentrations of 54 nM and higher resulted in a significant increase in fluorescence levels (two-sided t-test with equal variance, ***p≤0.0005). Plot shows average values from five independent replicas; error bars represent the standard deviation. (D) At saturating vhhGFP4 concentrations (108 nM and 216 nM vhhGFP4), we observed a mean increase of 47.5% (1.475-folds) in eGFP fluorescence levels. Individual data points from the 108 nM and 216 nM experiments from all five replicas are plotted and were included in the calculations; the green line represents the median. (E) To test if fluorescent levels reflect changes in protein concentration, defined amounts of eGFP were imaged (43.5 nM, 54.4 nM, 72.5 and 108.8 nM) under the imaging conditions used for the experiments shown in (B–D). Indeed, the obtained fluorescence levels increased proportionally to the respective eGFP concentrations. Hence, our imaging conditions were within the linear range of the fluorophores and suitable to detect changes in eGFP fluorescence.
-
Figure 2—figure supplement 2—source data 1
Source data pannels C-E.
- https://doi.org/10.7554/eLife.22549.010
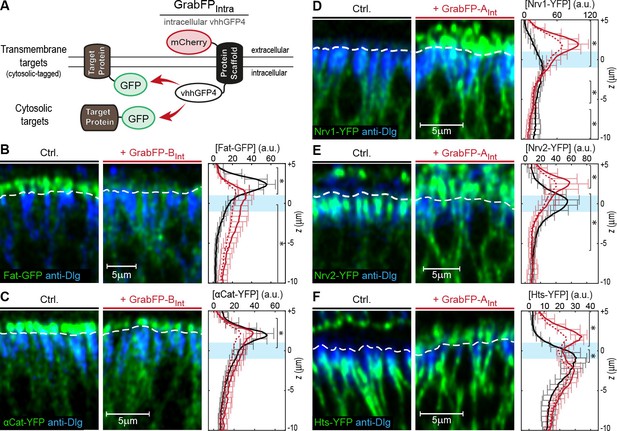
Mislocalization of GFP/YFP-tagged proteins using the GrabFPInt system.
(A) With the GFP-nanobody facing the cytosol, the GrabFPInt system can interact with cytosolic proteins and transmembrane proteins tagged along their cytosolic portion. (B–C) Optical cross-sections of wing disc cells expressing either Fat-GFP (B) or αCat-YFP (C) alone (Ctrl., left) or together with GrabFP-BInt (middle). A dashed line marks the junctional level. Quantification of relative target-protein localization (right) along the A-B axis in the absence (black) or in the presence of GrabFP-BInt (continuous red line). Profiles corrected for fluorescence increase due to nanobody binding are depicted by a dashed red line. The position of the junctions is marked by a blue bar. (Error bars show the standard deviation). (D–F) Optical cross-sections showing the localization of Nrv1-YFP (D), Nrv2-YFP (E) or Hts-YFP (F) in the absence (left) or in the presence of GrabFP-AInt (middle). Quantifications are shown to the right. (Sample numbers for plots in B-F: Fat n = 10, αCat n = 9, Nrv1 n = 10, Nrv2 n = 10, HTS n = 10, significance was assessed comparing control with the corrected profiles using a two-sided Student’s t-test with unequal variance, *p<0.05).
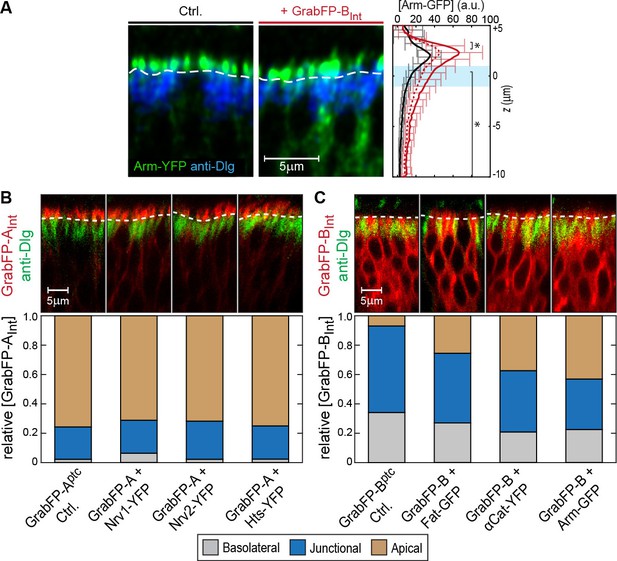
Examples of GFP/YFP-target protein mislocalization using the GrabFPIntra system.
(A) Optical cross-section of DP cells expressing Arm-YFP (Ctrl., left) and Arm-YFP together with GrabFP-BInt (middle). Average Arm-YFP protein distribution along the A-B axis in the absence (black) and in the presence of GrabFP-BInt (original data: continuous red line, corrected data: dashed line) is plotted to the right. (B–C) Representative cross-sections (top) and quantification (bottom) of relative GrabFP-AInt (B) and GrabFP-BInt (C) localization when expressed alone (Ctrl.) or when co-expressed with YFP/GFP-tagged target proteins. While GrabFP-AInt is robust to mislocalization by target proteins, GrabFP-BInt tends to be mislocalized to the apical compartment in all three conditions tested. (GrabFP-AInt Ctrl. n = 7, Nrv1 n = 10, Nrv2 n = 10, Hts n = 10, GrabFP-BInt Ctrl. n = 6, Fat n = 10, αCat n = 9, Arm n = 8).
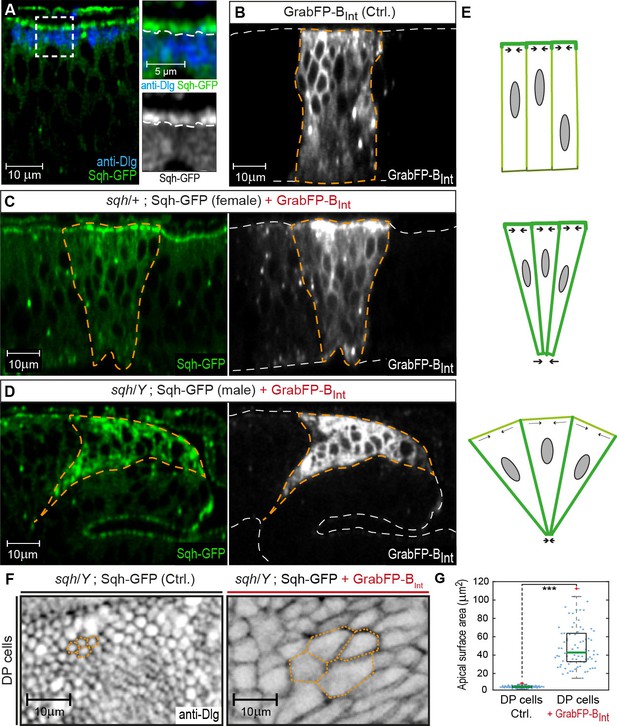
GrabFP-BInt-mediated Sqh-GFP mislocalization results in changes of DP cell shape.
(A) Optical cross-section of a wing disc expressing Sqh-GFP (green), stained for Dlg (blue). In the magnifications, the junctional level is marked by a dashed line. (B–D) Optical cross-sections of wing discs expressing GrabFP-BInt (grey) in the patched domain (marked by dashed orange line, ptc::Gal4) either alone (Ctrl., (B) or together with Sqh-GFP (green) in heterozygous sqh females (C) and hemizygous sqh males (D). Sqh-GFP mislocalization causes a drastic increase of basolateral Sqh-GFP (C–D). Mislocalization of Sqh-GFP causes cell shape alterations resulting in a triangular shape of the ptc domain (C–D), compared to the rectangular shape of the ptc domain in control discs (B). The white dashed line marks the apical (top) and basal (bottom) surface of DP cells. (E) Schematic representation of the effect of Sqh-GFP mislocalization. Tension is higher in the apical cortex of columnar cells due to polarization of myosin II activity (top). Mislocalization of Sqh-GFP causes increased basolateral tension, leading to constriction of the basolateral cell area (middle). In sqh hemizygous conditions, the apical surface expands due to decreased apical myosin II activity (bottom). (F) Projections of the junctional level of the DP columnar epithelium of the genotype shown in (D) either in the absence of GrabFP-BInt (left, normal Sqh::GFP localization) or in the presence of GrabFP-BInt (right, mislocalized Sqh::GFP). (G) Quantification of apical surface area as marked in (F). The green line marks the median, statistical significance was assessed using a two-sided Students t-test (***p<0.0005), n ≥ 77.
-
Figure 4—source data 1
Source data for apical surface area.
- https://doi.org/10.7554/eLife.22549.014
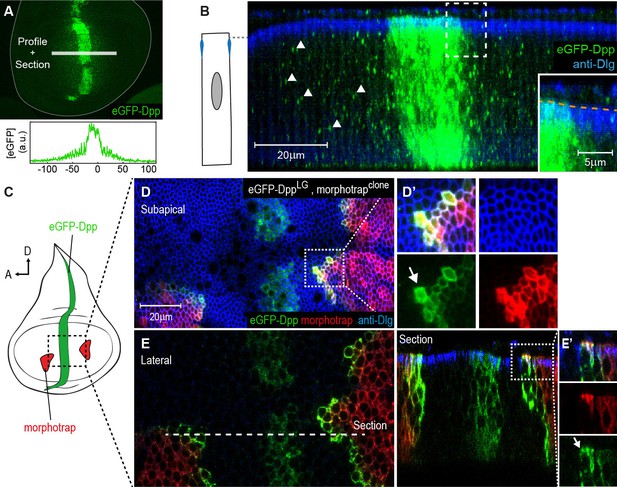
The Dpp morphogen spreads in the apical and basolateral compartment.
(A) Wing disc expressing eGFP-Dpp in the central Dpp stripe and eGFP fluorescence profile (bottom). (B) Optical cross-section of a wing disc as shown in (A) additionally stained for Dlg (blue). eGFP-Dpp is prominently observed in spots (arrowheads) along the lateral axis of the disc but not in the wing disc lumen (see magnified insert). (C) Scheme of morphotrap expression in clones and eGFP-Dpp in the central dpp stripe. (D) Subapical projection of a wing disc expressing eGFP-Dpp in the dpp stripe and two lateral morphotrap clones. Magnifications to the right show apical eGFP-Dpp immobilization on the proximal surface of morphotrap clones. (E) Lateral projection of the wing disc shown in (D). An optical cross-section to the right shows low level apical (also see arrow in magnification in [E’]) and high level basolateral immobilization of eGFP-Dpp.
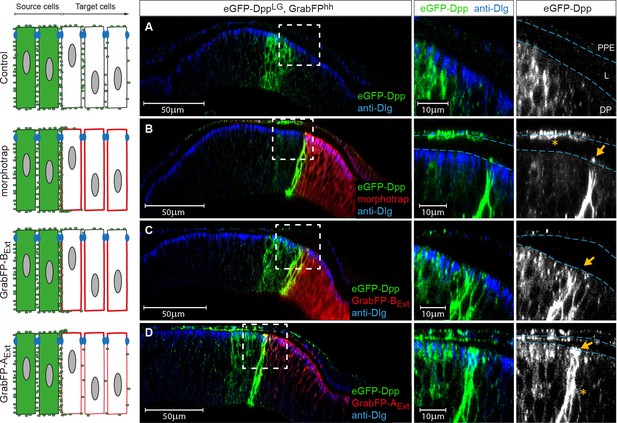
The GrabFPExt system can interfere with specific sub-fractions of the Dpp morphogen gradient.
Optical cross sections of wing discs either expressing eGFP-Dpp (green) in the stripe source (A) or eGFP-Dpp in the stripe and the different versions of the GrabFP system (red, B–D) in the posterior compartment of disc proper and PPE cells (hh::Gal4). (A) When expressed alone (Ctrl.), eGFP-Dpp is mainly observed in the lateral plane of the DP epithelium. Peripodial epithelium (PPE), lumen (L) and disc proper epithelium (DP). (B) Posterior expression of morphotrap results in strong eGFP-Dpp immobilization along the basolateral domain and low or no apical immobilization (see arrow in the magnification to the right). eGFP-Dpp is also immobilized on the apical surface of PPE cells overlaying the Dpp DP source (see asterisk in magnification). (C) Posterior expression of GrabFP-BExt results in exclusive immobilization of eGFP-Dpp in the basolateral domain. No apical immobilization is observed, neither in DP (see arrow) nor in PPE cells. (D) Expression of GrabFP-AExt in the posterior compartment results in strong basolateral (asterisk) and apical (arrow) immobilization of eGFP-Dpp.
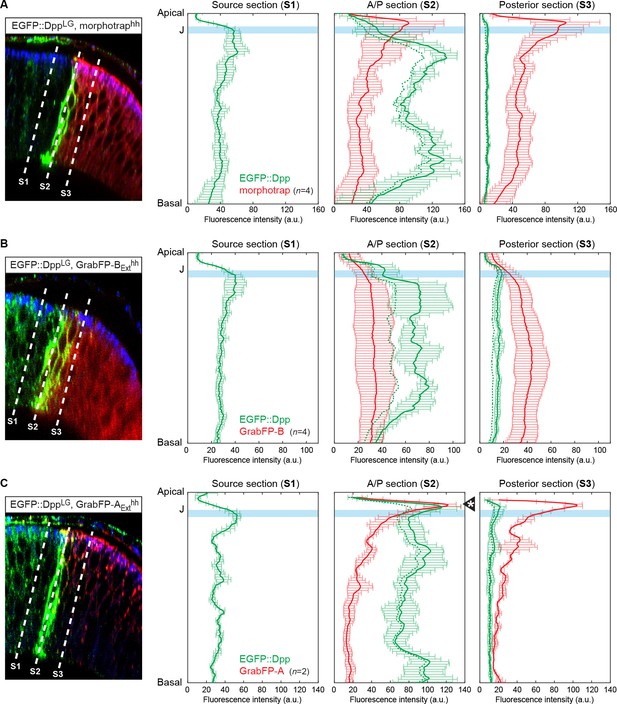
Quantification of differential eGFP-Dpp accumulation by morphotrap, GrabFP-BExt and GrabFP-AExt.
eGFP-Dpp immobilization pattern along the A-B axis in the posterior compartment when either morphotrap (A), GrabFP-BExt (B) or GrabFP-AExt (C) are expressed in posterior cells (hh::Gal4). Left column: Optical cross-sections as shown in Figure 5. Positions at which eGFP-Dpp and GrabFPExt localization was measured are indicated by dashed lines (S1–S3). Middle-right column: Plots of average eGFP-Dpp (green line, corrected profile is shown as dashed line) and GrabFPExt (red) levels along the A-B axis as positions indicated in the left column. eGFP-Dpp levels are strongly increased along the A/P boundary (S2 section) in all conditions. Importantly, neither morphotrap (A, middle-right) nor GrabFP-BExt (B, middle-right) immobilize eGFP-Dpp in the apical compartment above the junctions (thick blue line). In contrast, GrabFP-AExt shows strong eGFP-Dpp immobilization in the apical compartment (arrowhead in C, middle-right). This might be due to GrabFP-AExt mediated mislocalization of basolateral eGFP-Dpp to the apical compartment. In all conditions eGFP-Dpp levels drop after the first 2–3 cell rows (green in S3) suggesting that indeed posterior GrabFPExt expression results in impaired posterior Dpp dispersal. In the plots, thick lines represent average fluorescence values and error bars show the standard deviation.
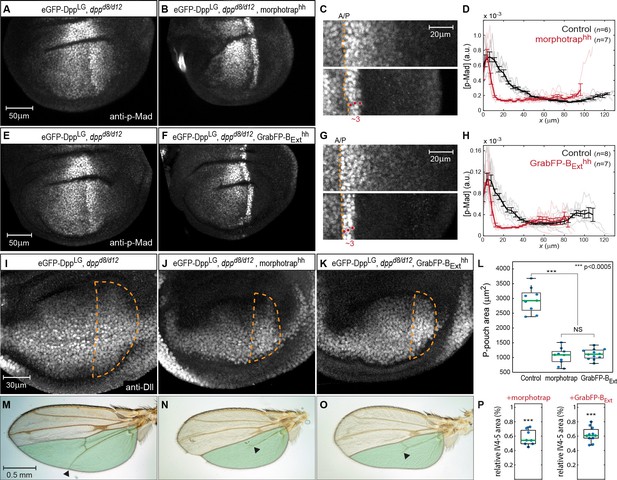
Basolateral Dpp spreading is required for patterning and size control.
(A–B) p-Mad staining in representative dppd8/d12 mutant wing disc rescued by eGFP-Dpp (A) and in dppd8/d12 wing disc rescued by eGFP-Dpp expressing morphotrap in the posterior compartment (hh::Gal4, B). (C) Magnifications of the posterior, dorsal pouch region of the images shown in (A–B). The A/P boundary is marked by a dashed yellow line. (D) Average posterior p-Mad profiles of 98–100 hr AEL old dppd8/d12 wing disc rescued by eGFP-Dpp (black) and dppd8/d12 wing disc rescued by eGFP-Dpp expressing morphotrap (red). (E–H) Representative wing discs and quantification of p-Mad levels in dppd8/d12 wing disc rescued by eGFP-Dpp (E, black in H) and dppd8/d12 wing disc rescued by eGFP-Dpp expressing GrabFP-BExt in the posterior compartment (F, red in H). (I–K) Representative 98–100 hr AEL old wing discs of the indicated genotypes stained for Distal-less (Dll) as a marked for pouch size. The posterior wing pouch is outlined by a dashed yellow line. (L) Quantification of posterior wing pouch area as shown in (I–K). (Control n = 9, morphotrap n = 10, GrabFP-BExt n = 12) (M–O), Female wings of the genotypes indicated. The area posterior to vein 4 (IV4-5) is marked in green. Block of apical and basolateral, as well as block of basolateral Dpp dispersal results in a loss of the distal parts of wing vein 5 and hence patterning (see arrowheads). (P) Quantification of relative IV4-5 area as indicated in (M–O). (***p>0.0005, Control n = 10, morphotrap n = 8, GrabFP-BExt n = 11).
-
Figure 7—source data 1
Wing pouch area and IV4-5 area.
Only male wing discs and wings were included in the quantification.
- https://doi.org/10.7554/eLife.22549.019
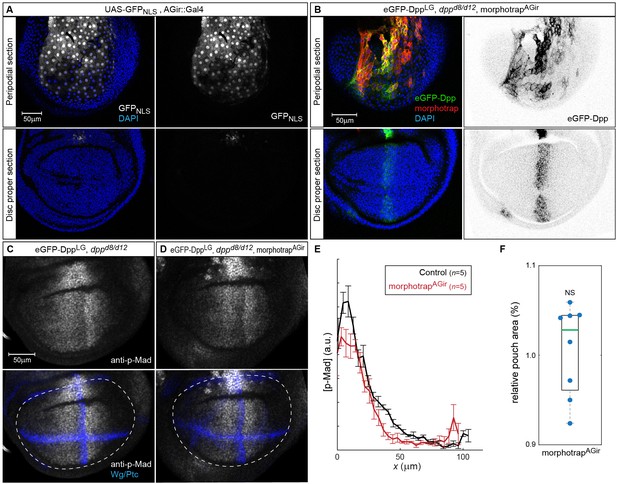
morphotrap expression in PPE cells interferes with luminal Dpp spreading.
(A) Expression of UAS-GFPNLS under control of the AGir::Gal4 driver line in a third instar wing imaginal disc. The activity of AGir::Gal4 is restricted to PPE cells and a small group of cells in the dorsal hinge region of the DP. (B) dppd8/d12 mutant wing disc rescued by eGFP-Dpp (green) expressing morphotrap (red) in the PPE (AGir::Gal4). A projection of the PPE plain shows that high amounts of luminal eGFP-Dpp are immobilized along PPE cells expressing morphotrap (top), while eGFP-Dpp dispersal is undisturbed in DP cells (bottom). (C–F) Patterning and growth are largely unaffected by morphotrap mediated immobilization of eGFP-Dpp along PPE cells. Peak levels of Dpp signaling activity, visualized by p-Mad (grey), are slightly reduced in the morphotrap condition (D) compared to controls (C, 98–100 hr AEL). Quantification is shown in (E). Pouch size (white dashed line in C–D), as defined by the inner Wg ring (blue in C–D), is also not affected by modifying luminal Dpp dispersal (F).
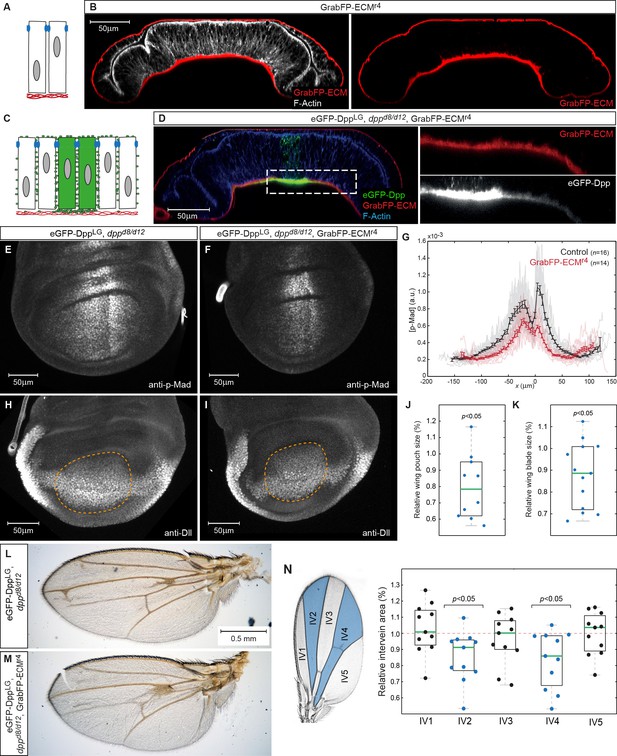
Basal Dpp is required to control wing size.
(A) Schematic representation of GrabFP-ECM localization when expressed in the larval fat body. (B) Wing disc optical cross-section of an animal expressing GrabFP-ECM in the fat body, stained for mCherry (GrabFP-ECM, red) and F-Actin (Phalloidin, white). (C) Schematic of eGFP-Dpp immobilization in the ECM by GrabFP-ECM. (D) Optical cross-section of a dppd8/d12 mutant wing disc rescued by eGFP-Dpp (green) and GrabFP-ECM (red) localizing to the basal lamina. Tissue outlines are visualized by F-Actin staining (blue). Magnification to the right shows strong eGFP-Dpp accumulation below Dpp source cells. (E–F) 98–100 hr AEL old wing discs of the indicated genotype stained for p-Mad. (G) Average p-Mad gradient at 98–100 hr AEL. (H–I) 98–100 hr AEL old wing discs of the above indicated genotypes stained for Dll. The wing pouch is outlined by a dashed yellow line and quantified in (J). (J) Relative wing pouch area of dppd8/d12 mutant wing disc rescued by eGFP-Dpp and GrabFP-ECM localizing to the basal lamina (n = 11). (K) Relative wing blade area of dppd8/d12 mutant wing disc rescued by eGFP-Dpp and GrabFP-ECM localizing to the basal lamina (n = 14). (L–M) Representative female wings of the genotypes indicated. (N) Quantification of intervein area in GrabFP-ECM flies relative to control wings (n = 11).
-
Figure 8—source data 1
Wing pouch and blade areas.
Only male wing discs and wings were included in the quantification.
- https://doi.org/10.7554/eLife.22549.022
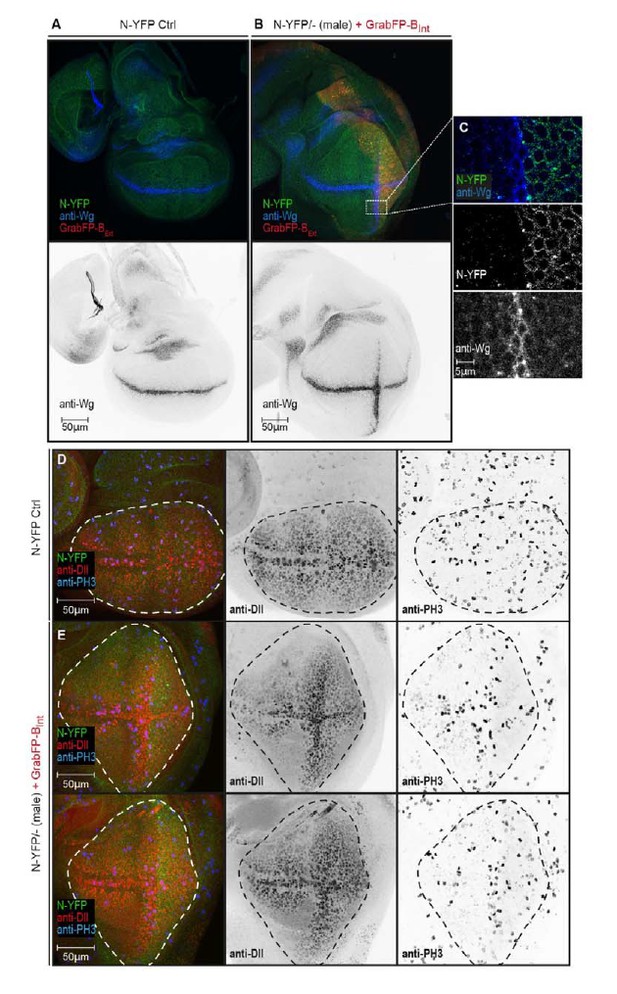
N-YFP mislocalization causes the induction of an ectopic Wg stripe.
(A) Representative wing disc of a N-YFP (green) hemizygous male (control) stained for Wg (upper panel, blue; lower panel, black). Wg is expressed along the D/V boundary. (B) Basolateral mislocalization of N-YFP (green) in hemizygous male N-YFP/Y flies in the posterior compartment by GrabFP-BExt (red) leads to the induction of a secondary Wg stripe along the A/P boundary (upper panel, blu; lower panel, black). (C) Blow-up of region marked in (B), showing that Wg is only induced in the row of cells adjacent to the posterior compartment. D, Wing disc of a male control fly stained for the Notch target Distal-less (Dll, red) and the mitotic marker phosphor-Histone H3 (PH3, blue). The pouch outline is marked by a dashed line. Distal-less staining marks the oval pouch and the observed proliferation pattern is fairly uniform. E, Wing discs of two N-YFP hemizygous males expressing GrabFP-BExt in the posterior compartment. Ectopic Wg induction results in expansion of the pouch along the dorsal-ventral axis as shown by the expansion of the Dll staining. This change in pouch shape might be a reason of a non-uniform proliferation pattern in this background, with increased cell division along the ectopic Wg stripe.
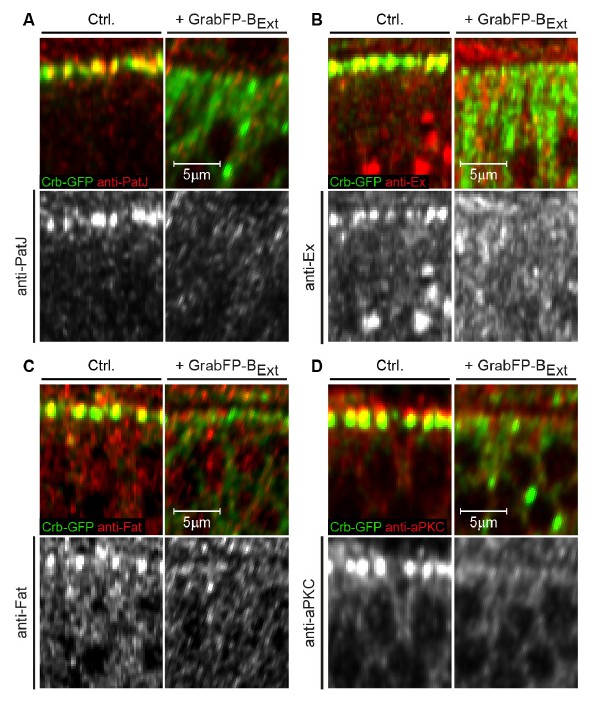
Mislocalization of Crb-GFP affects the localization of apical Crb interaction partners.
(A-D) Optical cross-sections of Crb-GFP (green) homozygous GFP-tagged wing discs showing the anterior control compartment (Ctrl., left) and the posterior compartment expressing GrabFP-BExt (right). Discs were stained for the Crb interacting proteins PatJ (A), Expanded (Ex, B), Fat (C) and aPKC (D). In all cases, loss of apical Crbs due to mislocalization resulted in a striking reduction of the apical levels of all Crb interactors tested. Interestingly, no basolateral increase in PatJ, Ex, Fat or aPKC is observed, suggesting that there are control mechanisms excluding these proteins from the basolateral compartment.
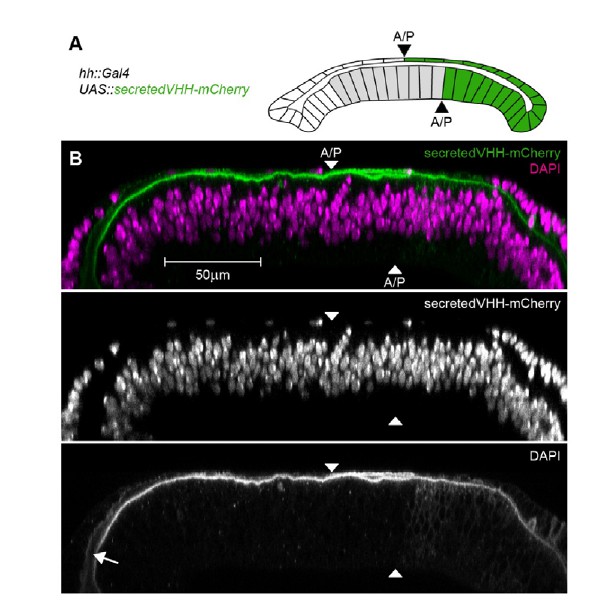
Detection of a secreted VHH-mCherry fusion in the wing disc lumen.
(A) Schematic representation of the experimental setup. We expressed a fusion of the mouse CD8 signal peptide, the vhhGFP4 GFP-nanobody and mCherry (secretedVHH-mCherry) under the control of hh::Gal4 in the posterior compartment of otherwise wild type wing discs. Importantly, the anterior/posterior compartment boundary (A/P) in the peripodial epithelium (top layer) is shifted anteriorly compared to the disc proper (bottom layer). However, hh::Gal4 is exclusively active in posterior cells. (B) Optical cross-sections of wing discs expressing secretedVHH-mCherry (green) in the posterior compartment (hh::Gal4). The wing disc shown was permeabilized by several washes with PBST and subsequently stained for DAPI (magenta). Despite the permeabilization, secretedVHH-mCherry was not washed out and can even be detected in the anterior most parts of the lumen (arrow).





