Collagen IV and basement membrane at the evolutionary dawn of metazoan tissues
Figures
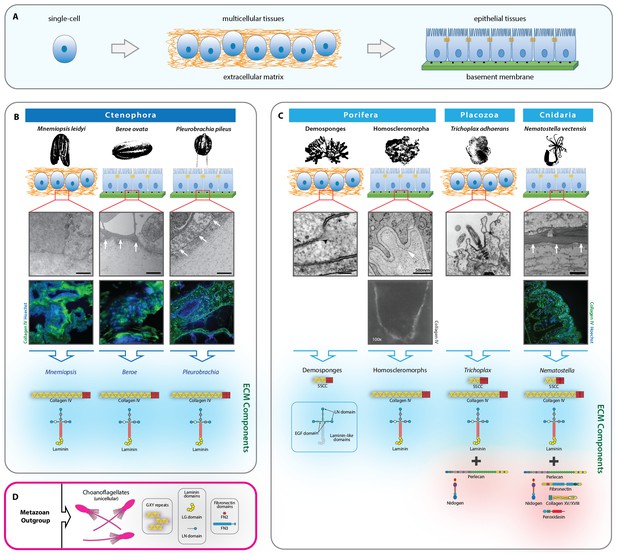
Extracellular matrix of the non-bilaterian animal phyla.
(A) The transition from single-cell organisms to complex multicellular animals was enabled by an extracellular matrix. (B) Electron microscopy (EM) and immunohistochemistry (IHC) of the Ctenophora species, Mnemiopsis (IHC: 20X magnification), Pleurobrachia (IHC: 20X magnification), and Beroe (IHC: 40X magnification) and ECM components of Ctenophora. (C) Electron microscopy (EM) and immunohistochemistry (IHC) of the non-bilaterian animal phyla, Cnidaria (Nematostella; 20X magnification), Placozoa (Trichoplax), and Porifera (Homoscleromorpha and Demosponges) and ECM components of Porifera, Placozoa, and Cnidaria. Demosponge EM reproduced from Figure 1E of Adams, et al., Freshwater Sponges Have Functional, Sealing Epithelia with High Transepithelial Resistance and Negative Transepithelial Potential, PLoS ONE, 2010, volume 5; Homoscleromorph EM reproduced from Figure 3B, Leys et al., Epithelia and integration in sponges, Integrative and Comparative Biology, 2009, volume 49 with permission from Oxford University Press; Homoscleromorph IHC reproduced from Boute et al., Type IV collagen in sponges, the missing link in basement membrane ubiquity, Biology of the Cell, 1996, volume 88 with permission from Wiley; Trichoplax EM reproduced from Ruthmann et al., The ventral epithelium of Trichoplax adhaerens (Placozoa): Cytoskeletal structures, cell contacts and endocytosis, Zoomorphology, 1986, volume 106 with permission from Springer. (D) ECM components in choanoflagellates, the unicellular sister-group to metazoa. All scale bars 500 nm, unless otherwise noted.
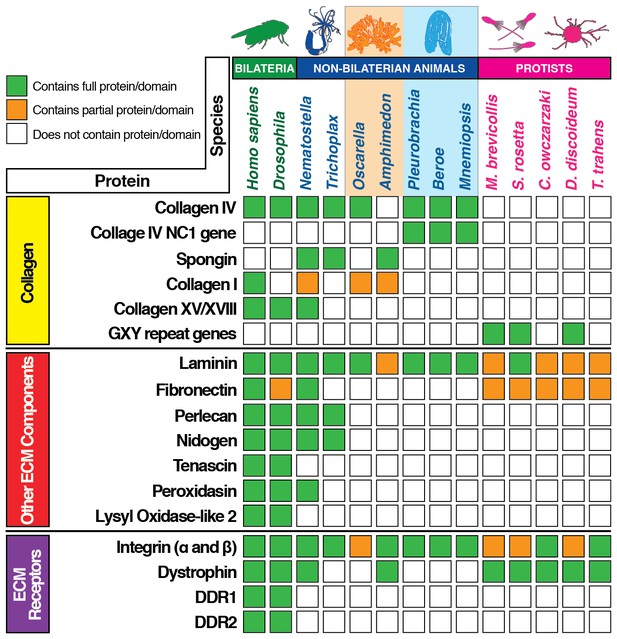
Extracellular matrix gene content across bilaterian, non-bilaterian animal, and unicellular protist phyla.
Protein BLAST searches using the human ortholog of each protein as bait was conducted for ECM gene content analysis. Where possible (with exception of ctenophore species), we performed a search by protein name across each database. The databases used were Ensembl (http://protists.ensembl.org), NeuroBase (http://neurobase.rc.ufl.edu), AmoebaDB (http://amoebadb.org) and NCBI’s Blast (https://blast.ncbi.nlm.nih.gov/Blast.cgi). Complete hits are denoted in green, while partial protein or domain sequences are denoted in orange. White boxes indicate absence of that protein/domain.
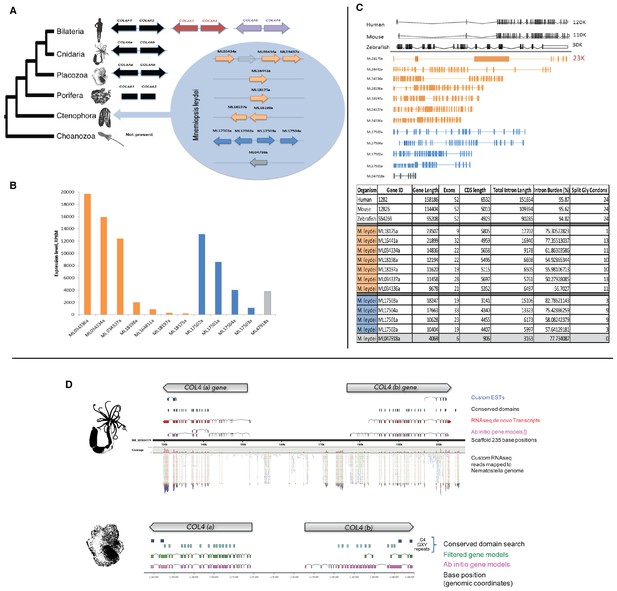
Mnemiopsis reveals multiple duplications of collagen IV genes, and non-bilaterian animal phyla collagen IV organization is similar to Bilateria.
(A) Collagen IV Gene Orientation. Mnemiopsis collagen IV genes were separated into two groups base on genomic orientation. Group I genes are found on the same scaffold (colored in blue). Group II genes (colored in orange) are spread across four different scaffolds and do not have head-to-head orientation. (B) Transcriptome analysis of Mnemiopsis confirmed a total of 11 collagen IV genes and one NC1 proto-domain gene (colored in grey). (C) Human COL4A1 spans 150 kb, contains 52 exons and has an intron composition of 95%. Mnemiopsis collagen IV genes are approximately one sixth the length of human collagen IV genes ranging in length from 1 to 23 kb with an intronic composition of 50–82%. (D) In Nematostella and Trichoplax, two collagen IV genes are located in head-to-head orientation on one genomic scaffold (Nematostella, scaffold 14; Trichoplax, scaffold 235), which indicates that they share one chromosome. The arrows at the top of each species indicate gene orientation: either minus or plus strands. The search for conserved domains revealed multiple collagens repeats (PF01391, light blue boxes) and C4 domains (PF01413, dark blue) further support that these genes belongs to collagen IV gene family. Pfam domains were identified using HMM against the genomic sequence. Mapping RNAseq reads to the genome strongly supports the proposed collagen IV genes model.
-
Figure 3—source data 1
Mnemiopsis collagen IV gene expression by RNA-Seq (RPKM).
- https://doi.org/10.7554/eLife.24176.006
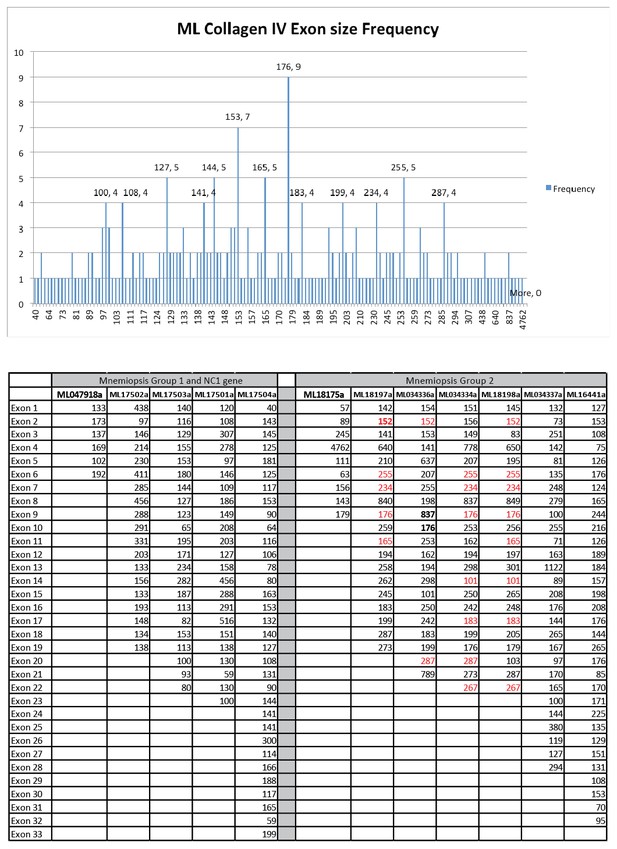
In an effort to detect conservation of exon size between ML collagen IV genes a database composed of all the exons from each gene was compiled.
Calculation of exon size frequency revealed a total of 75 different exon lengths (ranging from 57 to 837 bp), which were repeated at least once. Only six of these exon sizes were repeated more than four times throughout the database (upper graph). Conservation of exon length and position (marked in red) is present among some of the group II genes (lower table).
-
Figure 3—figure supplement 1—source data 1
Frequency of >exon lengths in Mnemiopsis collagen IV genes.
- https://doi.org/10.7554/eLife.24176.008
-
Figure 3—figure supplement 1—source data 2
Conservation of exon length and position is present among some of the groups.
Conserved exon lengths and positions are marked in red.
- https://doi.org/10.7554/eLife.24176.009
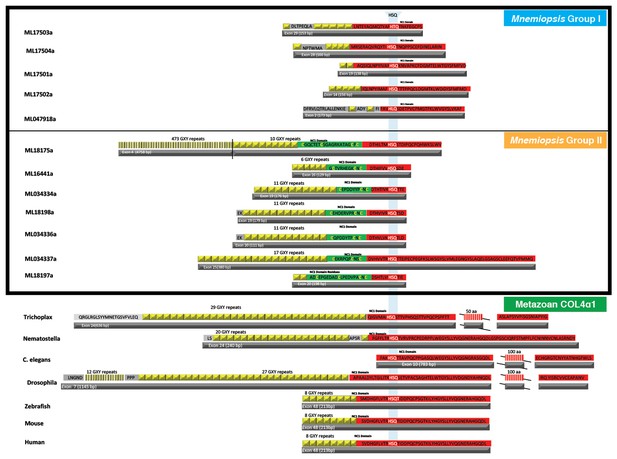
The first NC1 domain (red) coding exon of collagen IV has several features conserved throughout the animal kingdom, including the collagenous domain (yellow), which is a stretch of interrupted Gly-X-Y repeats at the 5’ end, and the presence of an HSQ coding region (white text in red).
In addition to these conserved features, Mnemiopsis Group II collagen IV genes encode for a cysteine loop region (green) on the collagenous domain/ NC1 domain junction exon.
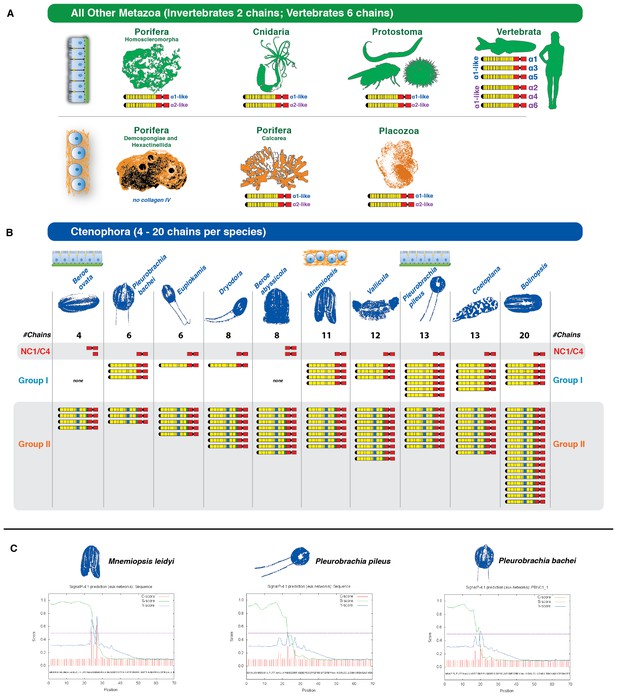
Collagen IV in Ctenophora underwent numerous gene duplication events resulting in an unprecedented diversity.
(A) Collagen IV chain distribution across non-bilaterian animal phyla and Bilateria. Two collagen IV chains are found across invertebrates, and six chains in chordate/vertebrate lineages. The poriferan class of Demosponges lacks collagen IV and BM. (B) Ctenophora collagen IV chains range from four to twenty distinct chains across species, indicating a variable number of gene duplication events. Ctenophora chains can be split into Group I, Group II, and NC1/C4 subgroupings. All ctenophore species contain Group I, II, and NC1 genes except for the two Beroe species, which lack Group I chains. (C) NC1 genes identified across Ctenophora were analyzed for signal peptide presence to determine whether sequences were truncated, or represented standalone NC1 proteins. Putative signal peptides were detected in at least three ctenophore NC1 genes, Mnemiopsis (ml047918a), Pleurobrachia pileus (pp_COL4_i), and Pleurobrachia bachei (PBNC1_1) based on SignalP prediction (http://www.cbs.dtu.dk/services/SignalP/).
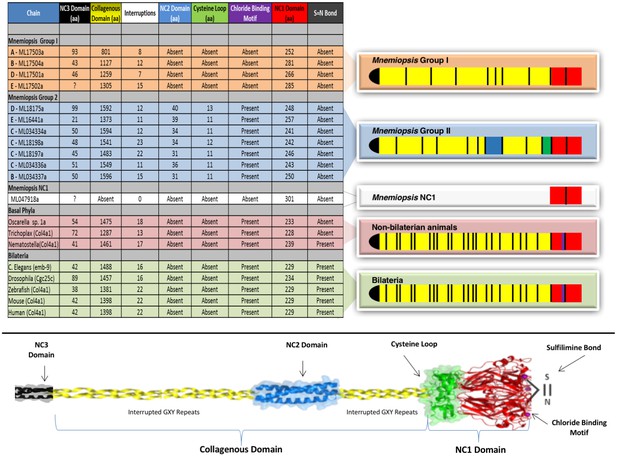
Collagen IV structural features are conserved across metazoa, and Ctenophora exhibits novel domains.
Signature features of collagen IV are found in each of the identified in Mnemiopsis chains. The collagenous region (yellow) of each chain contains characteristic interruptions (black lines) of the Gly-X-Y motif repeats. Group II chains also possess a NC2 domain (blue), which interrupts the collagenous region, and a cysteine loop (green) that is an extension of canonical NC1 domain (red). The NC1 domain of each chain is composed of two C4 domains. Group II chains possess the chloride-binding motif (purple) within the NC1 domain. While conservation of most Mnemiopsis collagen IV features can be found throughout metazoan species the NC2 domain and cysteine loop are structural innovations restricted to Ctenophora.
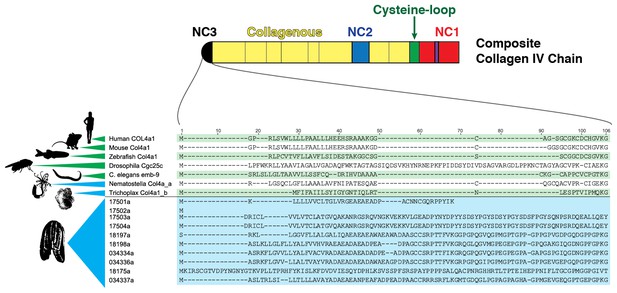
Multiple sequence alignment of the NC3 domain from various metazoan species reveals a high degree of conservation in this region among Mnemiopsis sequences, which is not seen in other metazoan sequences.
https://doi.org/10.7554/eLife.24176.013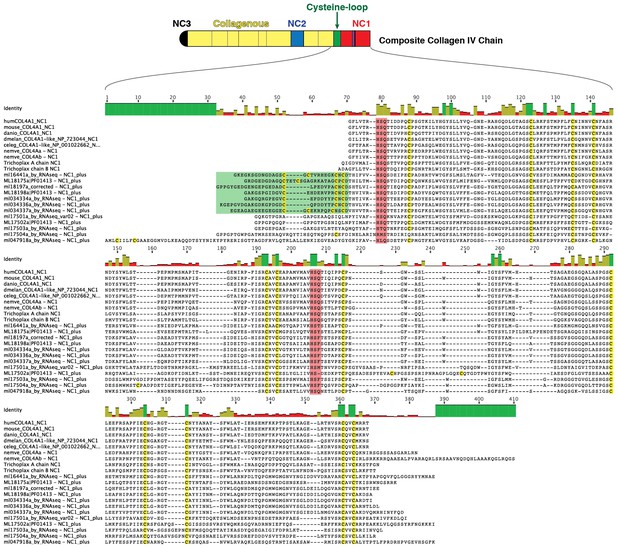
Multiple-sequence alignment of collagen IV NC1 domain sequences across human, mouse, zebrafish, fly, C. elegans, Nematostella, and Trichoplax, compared to the ctenophore representative, Mnemiopsis leidyi (MLXXXXXX).
Several hallmarks of collagen IV domains are present in Mnemiopsis: conservation of 12 cysteine residues throughout the NC1 (highlighted in yellow), and conservation of an HSQ- motif at the N-terminal side of both C4 domains comprising the whole NC1 domain (highlighted in red).
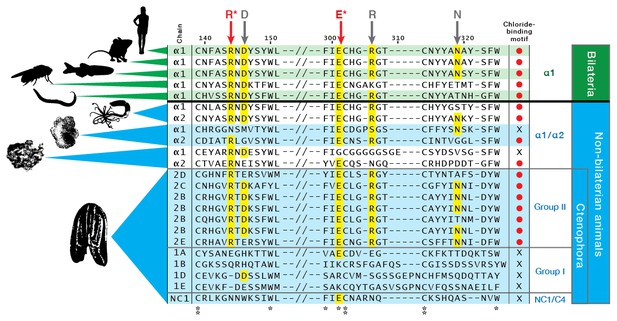
The chloride-motif has been identified previously in humans to Trichoplax.
The chloride-motif is highly conserved in Mnemiopsis Group II chains but is absent in Group I chains, and the standalone NC1 gene.
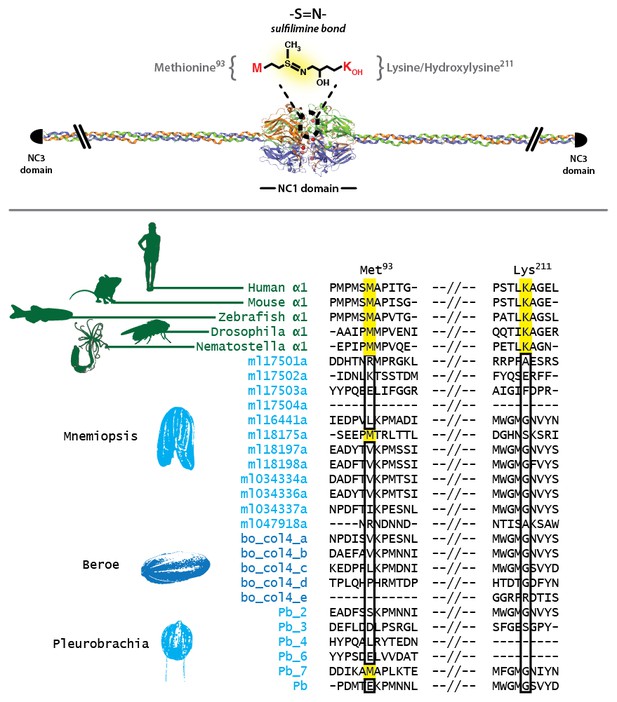
Sulfilimine bond crosslinking of collagen IV occurs between Methionine-93 and Lysine/Hydroxylysine-211 residues between adjoining NC1 domain interfaces.
Sulfilimine crosslinking confers structural integrity to collagen IV networks. Sulfilimine bond crosslinking residues are conserved throughout Bilateria to Cnidaria. Ctenophora has no conservation of Met-93 and Lys/Hyl-211 residues, indicating lack of sulfilimine bond crosslinking within the phylum.
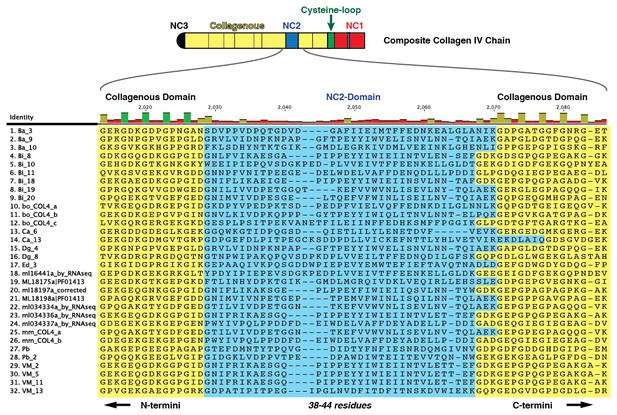
Multiple sequence alignment of 32 partial ctenophore collagen IV sequences spanning 10 species reveals the NC2 domain (highlighted in blue), spanning 38–44 amino acids.
However, very little sequence homology was observed across species for this region.
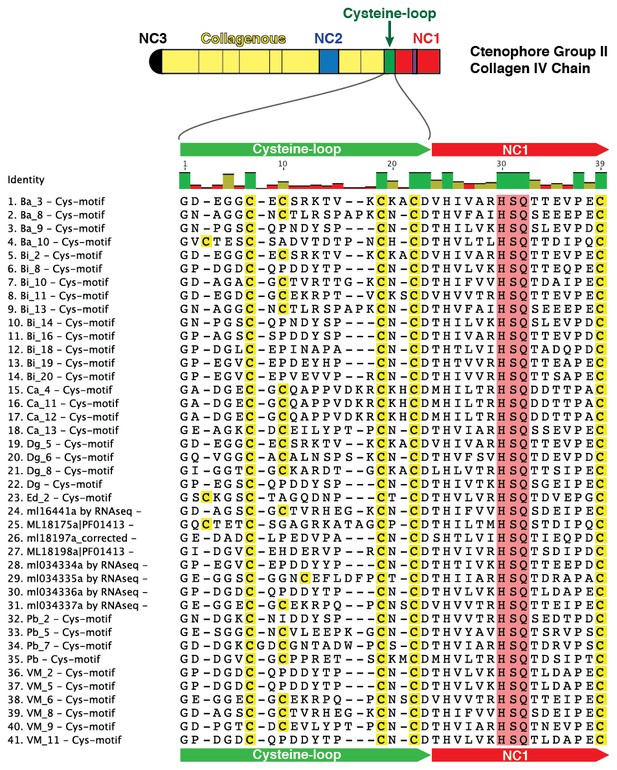
Multiple sequence alignment of partial sequences from 41 collagen IV genes exhibiting the cysteine-loop region of NC1 domains in Group II chains across Ctenophora.
Cysteine-loop domains across species contain either three or four conserved cysteine residues. HSQ motif and preceding seven residues demarcate the classical start of the NC1 domain.
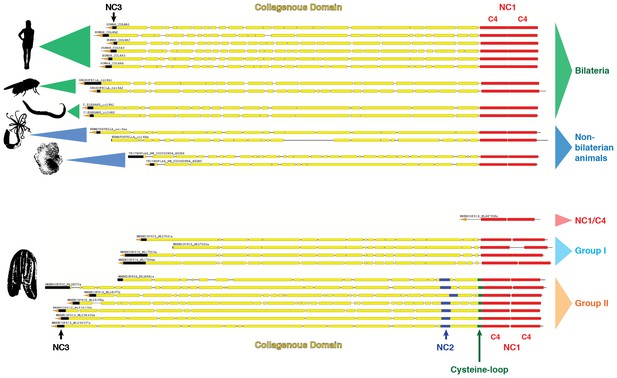
Collagen IV in Ctenophora and the non-bilaterian animal phyla are structurally homologous to bilateria.
Representative collagen IV sequences from human (UniProt entries P02462, P08572, Q01955, P53420, P29400, Q14031), Drosophila (UniProt entries P08120, O18407), C. elegans (UniProt entries P17139, P17140), Nematostella, Trichoplax, and Mnemiopsis genomes. The following regions are depicted: (based on prediction from http://www.cbs.dtu.dk/services/SignalP/; shown as orange arrow), NC3 domain (black box), uninterrupted triple helical segments with at least three GXY repeats (yellow boxes), NC2 domain (blue box), cysteine loop (green box), C4 domains (based on conserved domain search at http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi; shown as red boxes).
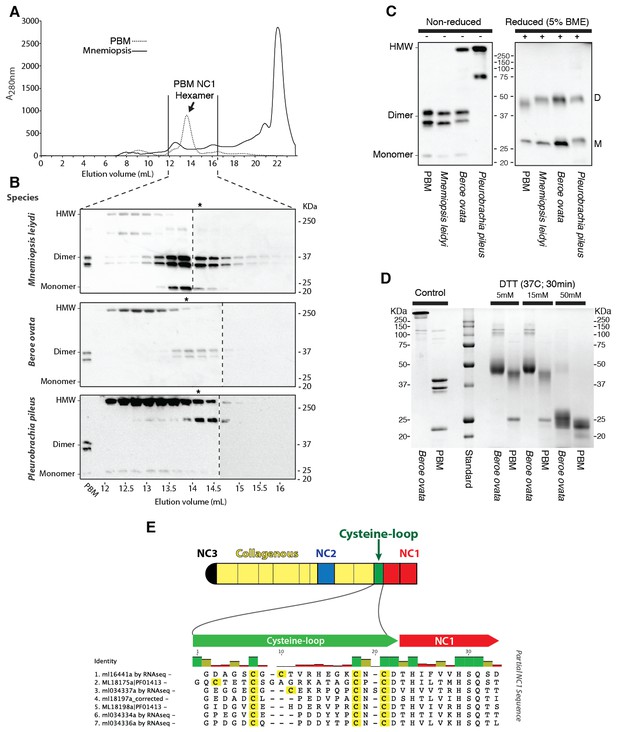
Ctenophora exhibits a novel collagen IV cross-linking mechanism.
(A) Gel filtration chromatography elution profile of Mnemiopsis collagenase digest (black) and native, purified placental basement membrane NC1 hexamer (dashed) run successively. Three ctenophore species were digested with bacterial collagenase to solubilize NC1 hexamer for analysis of collagen IV crosslinking. (B) Western blot of gel filtration fractions encompassing elution of NC1 hexamer (12 mL to 16.4 mL) from Mnemiopsis, Pleurobrachia, and Beroe, developed with NC1-specific monoclonal antibodies. HMW=high-molecular-weight complex. (C) Western blot of ctenophore NC1 hexamer separated by SDS-PAGE under reducing (+) and non-reducing (-) conditions (5% β-mercaptoethanol). (D) Reduction of the high-molecular-weight complex from Beroe (first lane, >250 kDa) following by alkylation results in formation of dimers at low DTT concentration, and complete reduction to monomers at high DTT concentration. (E) Structure of Ctenophora collagen IV group II chain, highlighting cysteine-loop region of the NC1, and multiple-sequence alignment of cysteine-loop region of Group II chains of Mnemiopsis (NC1 domain is partial sequence).
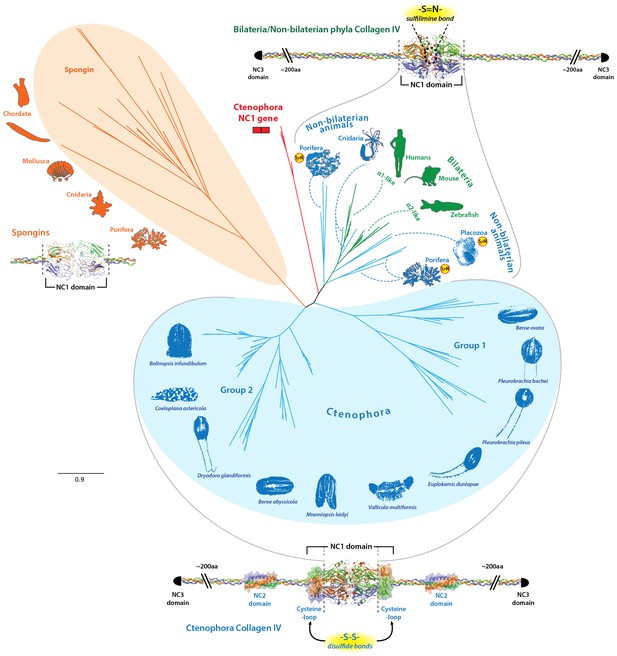
Evolutionary relationships of collagen IV and spongin NC1 domains across metazoa compared to Ctenophora.
Metazoan collagen IV chains feature a sulfilimine bond cross-linked collagen IV network, with the exception of the cnidarian, Hydra, and the non-bilaterian animal phyla Porifera and Placozoa. However, the structural domains across bilaterians and the non-bilaterian animal phyla are homologous; however, Ctenophora also contains novel domains. Unrooted maximum likelihood tree of collagen IV NC1 domains in human, mouse, zebrafish, Trichoplax (Placozoa), Pseudocorticium jarrei (Porifera), and Oscarella sp. (Porifera), in comparison with 10 ctenophore species. All analyses were based off amino acid sequence alignments of the NC1 domain, omitting the cysteine-loop region of Ctenophora NC1 domains.
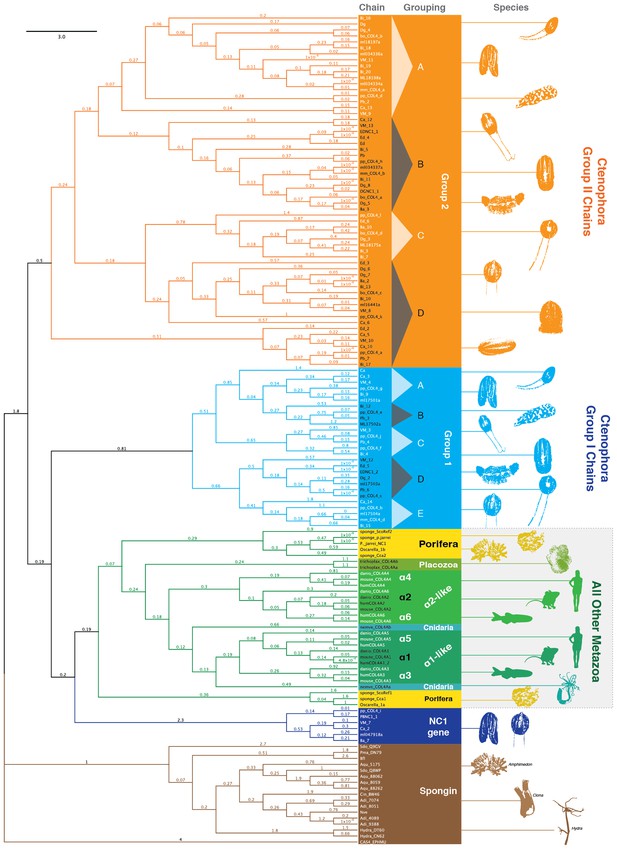
NC1 domain phylogeny across metazoa.
Phylogenetic analysis was conducted using maximum likelihood (using the RAxML software with the PROTGAMMAAUTO option) on NC1 domains spanning human, mouse, zebrafish, Nematostella, Trichoplax, sponges, and 10 ctenophore species. Ctenophore collagen IV separates into two major groups apart from the other non-bilaterian animal phyla and Bilateria. These two groups are consistent with genomic orientation groupings of Group I and Group II and can be further subdivided into subgroups Group I (IA-IE) and Group II (IIA-IID) based on phylogenetic affinity.
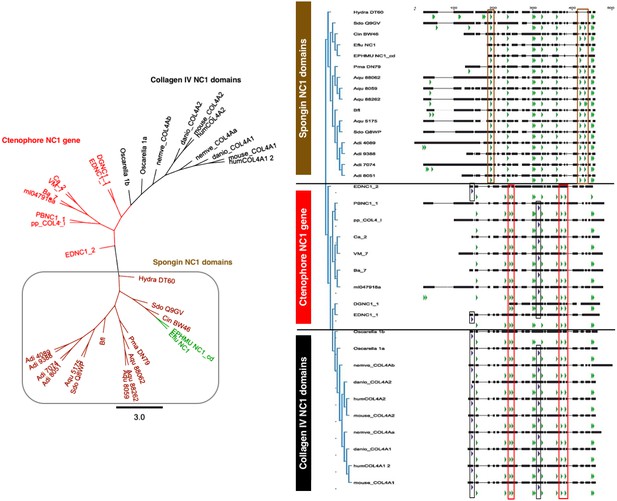
Spongins show conservation of key primary structure features within the NC1 region as compared to collagen IV.
Multiple sequence alignment reveals the conservation of cysteine residues (green arrows) across all four families of collagen IV. Seven cysteines are common to all sequences, while spongins share three unique cysteine residues (brown box). Likewise, the ctenophore NC1 protein and the NC1 domain of bilaterian collagen IV show conservation of four cysteine residues not found in spongin sequences (red box). The ctenophore sequences also show conservation of the second HSQ motif (purple arrows) found within the bilaterian NC1 domain (black box). No HSQ motifs were detected in the spongin sequences.
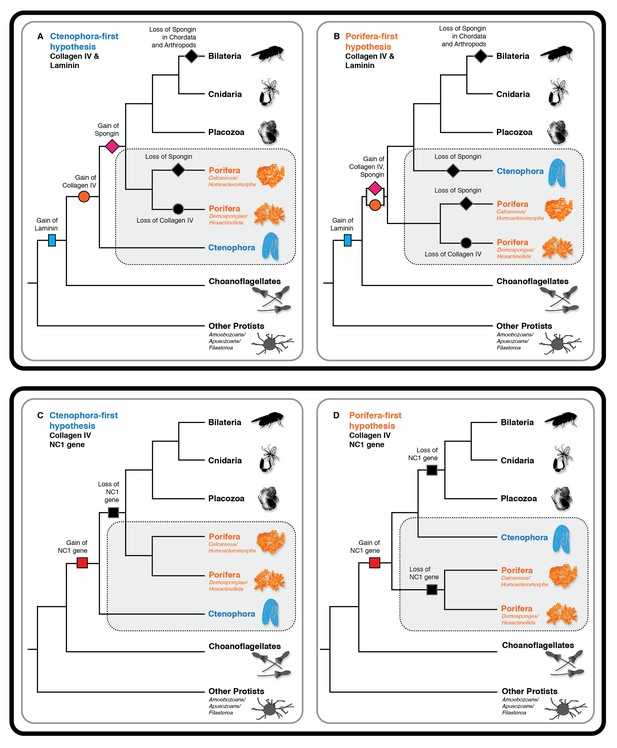
Collagen IV and Laminin gene evolution under Ctenophora or Porifera-first hypotheses.
(A and B) Comparison of collagen IV, spongin, and laminin gene evolution gain and loss evolutionary events in Ctenophora-first and Porifera-first hypotheses. (C and D) Comparison of NC1 gene evolution gain and loss evolutionary events in Ctenophora-first and Porifera first hypotheses.
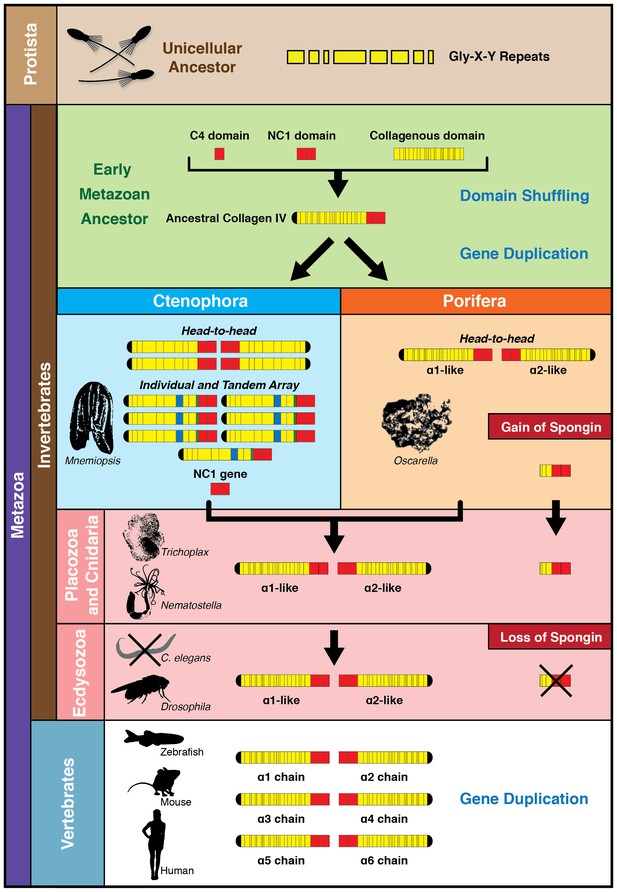
The non-bilaterian animal phyla reveal an evolutionary model for collagen IV.
Based off our phylogenetic analysis: (1) the presence of the ancestral NC1 domain in Ctenophora may have resulted from tandem duplication of the ancestral C4 or conservation of the ancestral NC1 domain. The last common ancestor of ctenophore and the non-bilaterian animal phyla may have expressed both the ancestral C4 and the ancestral NC1 domain. (2) Intergenic duplication of the ancestral collagen IV resulted in the head-to-head orientation. Errors in gene duplication may have given rise to spongins as they lack the domain-swapping region of the NC1 domain (a determined by predicative modeling) and have truncated collagenous tails. (3) The presence of six collagen IV genes arranged in a head-to-head orientation in vertebrates likely resulted from the two rounds of genome duplication that occurred in the vertebrate lineage.
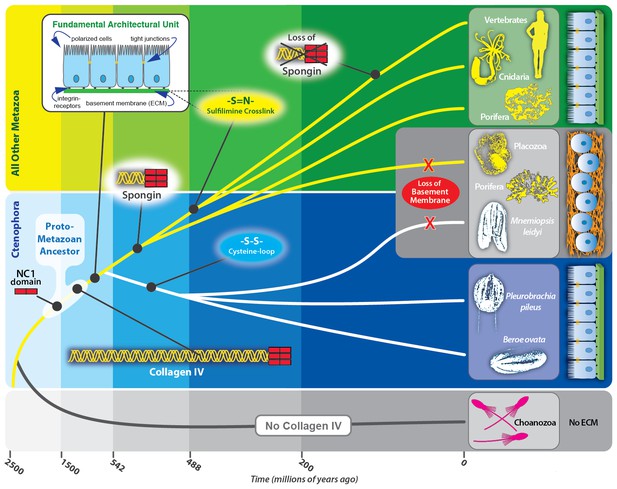
Collagen IV enabled the transition to multicellularity and the evolution of epithelial tissues in metazoa.
Collagen IV was a primordial innovation in early metazoan evolution, providing the architectural foundation for ECM formation. Choanoflagellates exist as singular or in colonies, yet do not have an ECM. Spongins are similar in domain structure and phylogeny to NC1 domains across metazoan collagen IV, and are variants of collagen IV, arising during the divergence of demosponges. Collagen IV, as a member of the basement membrane toolkit, enabled the evolution of multicellularity. Basement membranes juxtaposed to plasma membrane underlying a layer of polarized cells are a fundamental architectural unit of epithelial tissues. A layer of apical/basal-polarized cells that are laterally connected by tight junctions between plasma membranes, and basally anchored via integrin receptors embedded in plasma membranes to a basement membrane suprastructure is a fundamental architectural unit.
Additional files
-
Supplementary file 1
Intron/exon boundaries and split-glycine codons in Mnemiopsis.
(A) The human collagen IV one gene consists of 52 exons. The number of exons in the Mnemiopsis collagen IV genes ranges from 6 to33. A characteristic feature of collagen IV is the presence of split glycine codons. Eleven of the collagen IV genes possess split glycine codons. (B) Intronic regions are in lowercase and exon regions are in bolded UPPERCASE, underlined nucleotides represent partial codons, Green ‘G’ in the residue column denote glycines encoded by a split codon.
- https://doi.org/10.7554/eLife.24176.027





