RISC-interacting clearing 3’- 5’ exoribonucleases (RICEs) degrade uridylated cleavage fragments to maintain functional RISC in Arabidopsis thaliana
Figures
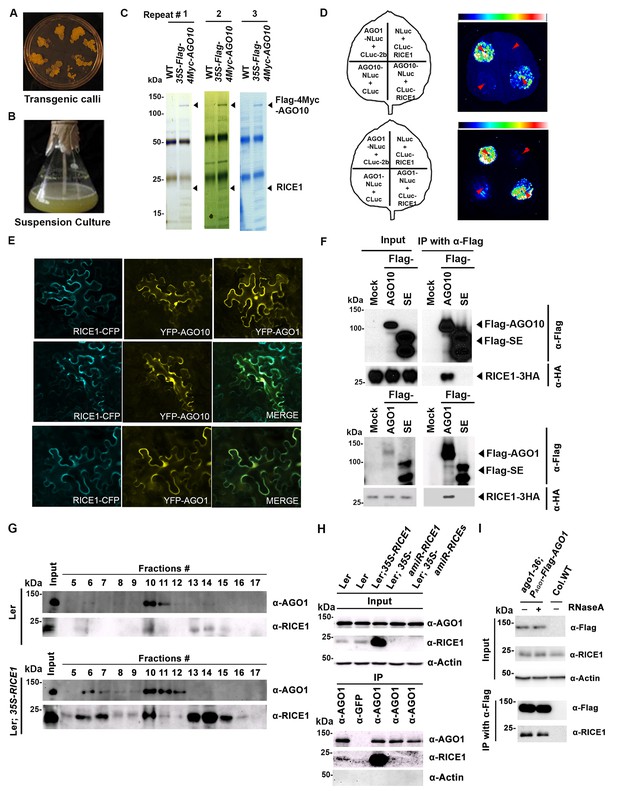
RICE1 is a novel RISC-bound cofactor in Arabidopsis.
(A and B) Generation of transgenic calli and cell lines expressing 35S-Flag-4Myc-AGO10. (C) Gel-code blue stained SDS-PAGE of purified AGO10 complexes for proteomics analysis. (D–F) Specific RICE1-AGO interaction was confirmed in N. benthamiana (N. bentha.) by LCI (D), co-localization (E), and co-IP (F) assays. In (D), the schemes of leaves show different combinations of infiltrated constructs that were fused either to N-terminal (NLuc) and C-terminal (CLuc) regions of luciferase. The LCI photographs on right showed the signals resulting from the protein-protein interaction. The red arrows indicate the infiltration positions. A combination of AGO1 and CMV 2b (Zhang et al., 2006b) serves as a positive control. In (E), RICE1-CFP and YFP-AGO proteins, when expressed alone in the N. bentha, were shown in top panels. The RICE1-CFP and YFP-AGOs, when co-expressed and detected in CFP and YFP channels separately, were shown in middle and bottom panels, respectively. Note: co-localization of RICE1-CFP and YFP-AGOs was observed predominantly in cytoplasm. In (F), IP was done with an anti-Flag antibody. Western blot analyses were conducted using anti-Flag, or -HA antibodies. SE is a negative control. (G–I) Specific interaction between endogenous RICE1 and AGO1 in Arabidopsis was confirmed by SEC (G) and co-IP (H and I) assays. IP was performed with anti-AGO1 (H) or anti-Flag (I) and western blot analyses were detected with anti-AGO1 (or -Flag), -RICE1 and -actin antibodies. Actin is a negative control. In (I), the crude extract was applied with (+) or without not (−) 50 μg/ml RNase A before co-IP.
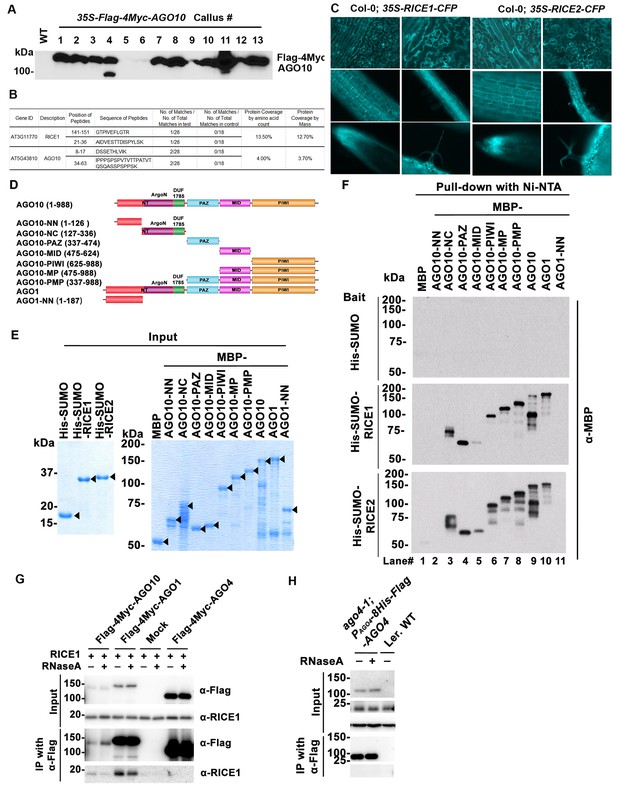
Identification and experimental confirmation of RICEs as novel Arabidopsis RISC-bound cofactors.
(A) Western blot analysis of transgenic Arabidopsis calli expressing Flag-4Myc-tagged AGO10 using an anti-Myc antibody. (B) Peptides recovered from proteomics analysis of AGO10 complex. (C) Cytoplasmic distribution of RICE1 and RICE2 in various tissues of Arabidopsis. Microscopic images were taken of stably transgenic Arabidopsis plants expressing 35S-RICE1 (or −2)-CFP at the different growth stages. (D) Schematic diagram of full-length and truncated forms of AGO1 and AGO10. The numbers refer to the amino acid residues in AGO proteins. Locations of the N-terminal (including the conserved PFAM domain ArgoN and DUF1785), PAZ, MID, and PIWI domains are shown. (E and F) In vitro pull-down assays of MBP-tagged target proteins with the indicated His-tagged bait proteins using nickel bound nitrilotriacetate (Ni-NTA) resins. (E) Input of bait proteins (His-SUMO and His-SUMO-RICEs) and target proteins (MBP-tagged full-length or truncated AGOs). Coomassie brilliant blue R250 staining of the proteins shows their mobility. The major bands representing the target proteins are indicated with arrowheads. (F) Output of in vitro pull-down assays of MBP-tagged target proteins. Western blots were analyzed with a monoclonal anti-MBP antibody. All His-SUMO-tagged bait proteins and MBP-tagged target proteins were purified from E. coli using nitrilotriacetate (Ni-NTA) and amylose resins, respectively. In all assays, 2 μg of target proteins were pulled down with the indicated bait proteins (2 μg each) using Ni-NTA resins.
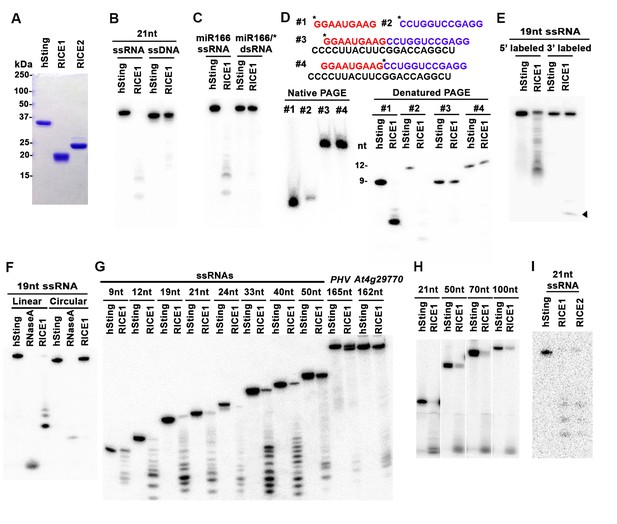
RICEs are 3’-to-5’ exoribonucleases preferring ss RNA.
(A) Coomassie brilliant blue staining of purified RICEs. hSting protein serves as a control throughout experiments. (B) RICE1 favored ss RNA over ss DNA as a substrate. (C) RICE1 did not degrade ds RNA. miR166/*, the 32P-labeled miR166 was annealed with unlabeled miR166*. (D) RICE1 did not degrade nicked but not released miRNA/*. Top panel shows positions of labeled nucleotides (*) and schematic diagram of nicked but still annealed miRNA/*. The nucleotides in red are the 5’ 9-nt fragment of miR166* and the nucleotides in blue are the 3’ 12-nt fragment of miR166*. (E) RICE1 was a 3’-to-5’ ribonuclease. Released single nucleotide is shown with an arrowhead. (F) RICE1 was an exoribonuclease. The circular substrates were generated by the self-ligation of 5’ labeled linear RNAs with T4 RNA ligase. (G) RICE1 efficiently degraded ss sRNA but not long RNA with secondary structures. Partial mRNAs of PHV and At4g29770 (165 nt and 162 nt in length) serves as long ss RNA substrates. (H) RICE1 efficiently degraded both sRNA and long RNA homopolymer (poly-As in 21-, 50-, 70-, and 100-nt length). (I) RICE2, similar to RICE1, is a 3’-to-5’ exoribonuclease. All the substrates in the enzymatic assay were 5’ 32P-labeled except the ones in (E) as indicated.
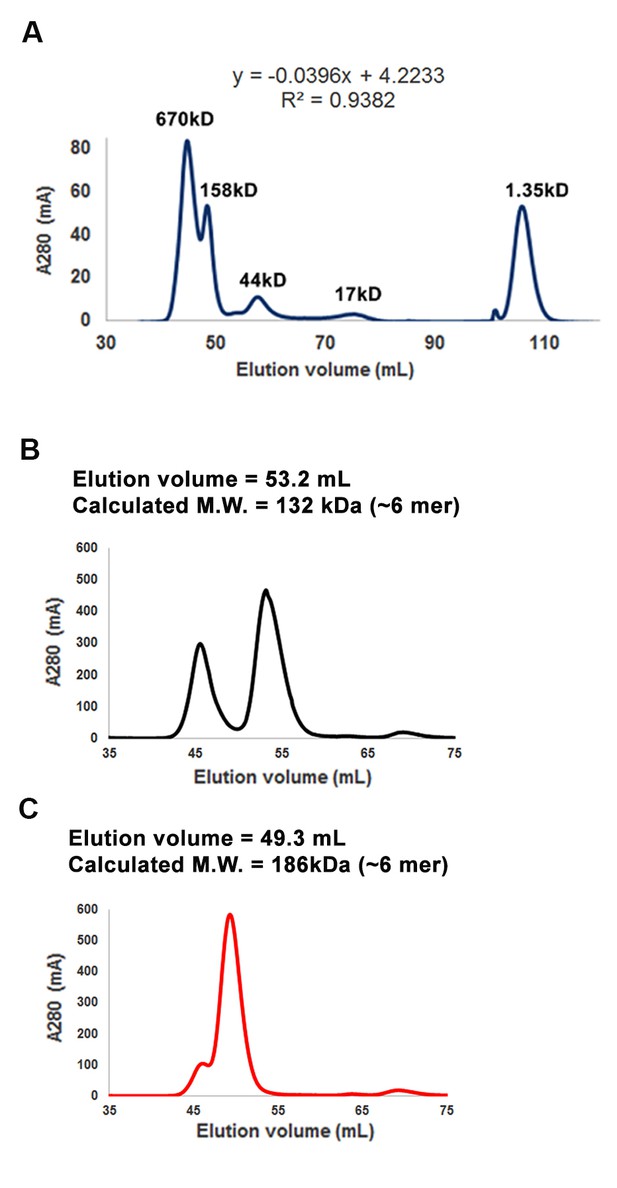
Purification of RICE1 and RICE2 proteins.
(A) Gel filtration of protein standards using Hiload 16/600 Superdex 75. (B) Size exclusion chromatogram shows that RICE1 forms hexamer in vitro. (C) Size exclusion chromatogram of RICE2 shows that RICE2 oligomerizes in vitro.
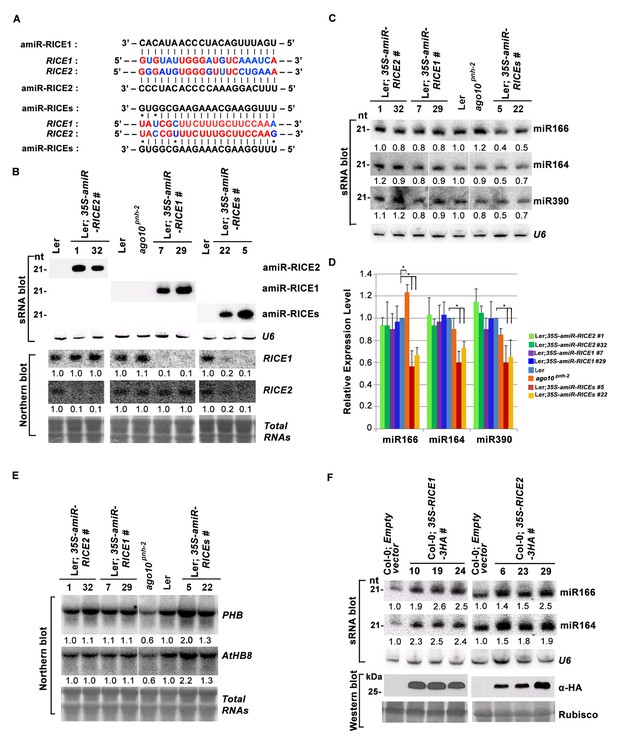
RICE1 and RICE2 are positive regulators for miRNA accumulation in vivo.
(A) Sequence alignment of artificial miRNAs and their targets. The nucleotides in red are perfect matched region between RICE1 and RICE2 target sites, while the nucleotides in blue are mismatched region between RICE1 and RICE2 target sites. (B) Generation of artificial miRNA transgenic lines targeting specific RICE1, RICE2, or both transcripts. (C–E) Concurrent knockdown of RICE1 and RICE2 consistently reduced miRNA abundance (C and D) and upregulated the targeted transcripts (E). (F) Ectopic expression of RICE1-3HA in wild-type Ler significantly enhanced accumulation of tested miRNAs. sRNA and RNA gel blot analyses were done with the indicated 32P-labelled probes. U6 and total RNAs stained with methylene blue serves as loading controls. For (C), additional two biological replicates were shown in Figure 3—figure supplement 2C. Quantification of three biological replicates was statistically analyzed and the significant difference (Student t-test, p<0.05) shown by the asterisk (*) in (D). Also see in Supplementary file 1A. Western blot analyses were performed with anti-HA antibody and Rubisco stained with Ponceau S serves a loading control. The amount of miRNAs or mRNAs in wild-type plants or transgenic plants harboring empty vectors was arbitrarily designated as 1.0, and the relative amount in the other lines was normalized to that of control plants.
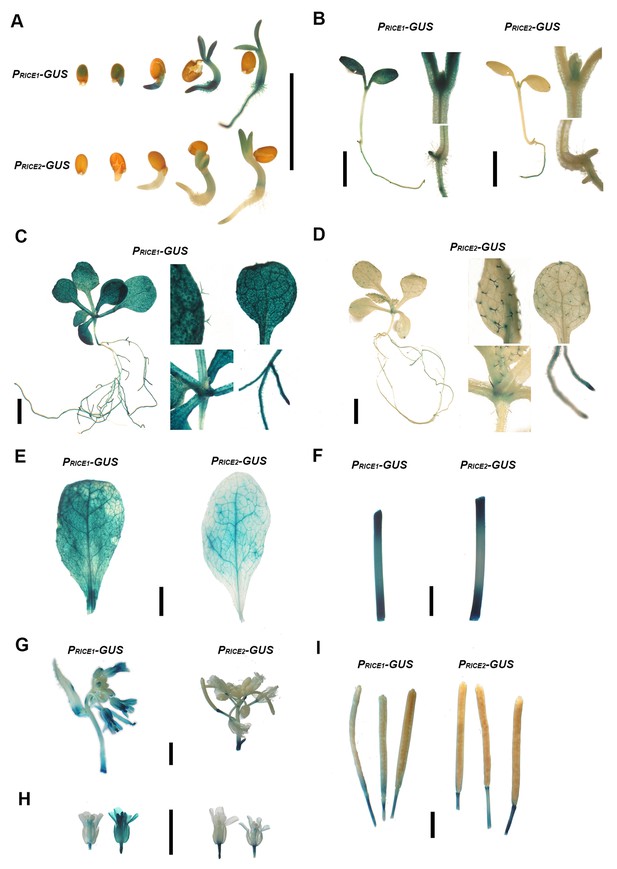

RICE1 is ubiquitously expressed in various organs and tissues throughout different developmental stages whereas RICE2 expression is more restricted and less abundant.
Note: the expression patterns of RICE1 and RICE2 are inferred from GUS gene regulated through their native promoters. Expression of RICE1 and RICE2 in germinating seeds (A), 3 day seedlings (B), 10 day seedlings (C and D), mature leaves (E), stem (F), inflorescence (G), flowers (H), and matured siliques (I). Black scale bars represent 0.5 cm in length.
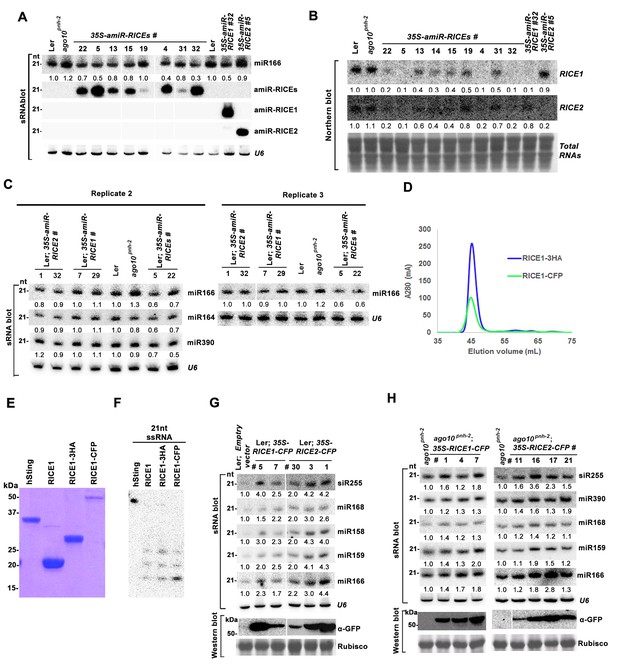
RICEs positively regulate miRNA accumulation in vivo.
(A) miR166 abundance in additional transgenic lines expressing artificial miRNAs specifically targeting RICE1, RICE2 or both. sRNA blot analyses were done with the indicated 32P-labelled probe. U6 serves as a loading control. (B) RNA gel blot analyses of RICE1 and RICE2 transcripts in the transgenic lines expressing various artificial miRNAs. Methylene blue staining of total RNAs in blots shows loading controls. (C) Additional biological replicates for the miRNA abundance in artificial miRNA knockdown lines targeting RICE1, RICE2 or both. (D) Size exclusion chromatogram of purified RICE1-3HA/-CFP proteins show that the proteins form oligomers in vitro. (E) Coomassie brilliant blue R250 staining of SDS-PAGE shows purity and mobility of RICE1-3HA/-CFP. (F) Both RICE1-3HA and RICE1-CFP are catalytically-active exoribonucleases in vitro. (G and H) Ectopic expression of RICE1- or RICE2-CFP in wild-type Ler (G) and ago10pnh-2 background (H) significantly enhanced accumulation of tested miRNAs. U6 serves as a loading control. Western blot analyses were performed with anti-GFP antibody and Rubisco stained with Ponceau S shows the loading control. The numbers listed on the bottom of panels represented the relative ratios of miRNAs in each line compared with that of wild-type plants or transgenic plants harboring empty vectors where the ratio was arbitrarily assigned a value of 1.0 after normalization with the loading control.
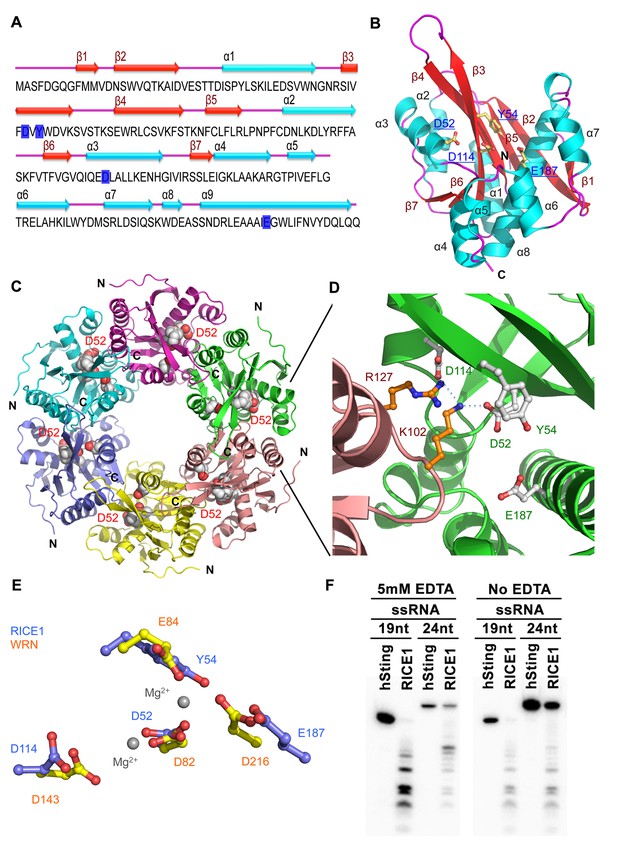
Crystal structure of RICE1.
(A) The amino acid sequence and secondary structure of RICE1. (B–D) Structures of RICE1 monomer and hexamer. In (B) α-helices, β-strands, and loops are colored in cyan, green, and pink, respectively. In (A and B), residues (D52, Y54, D114 and E187) in the active site are highlighted in blue. In (C), each monomer is shown in different colors. Active site residues are shown by the sphere representations, with active site residues D52 of each monomer labeled. In (D), the active sites of RICE1 are located at the interface between RICE monomers. Key residues (D52, Y54, D114 and E187) in the active site are shown as ball-and-stick models. Salt bridges are denoted by blue dotted lines. (E) Overlay of active sites residues of RICE1 and WRN exonuclease. (F) Enzyme assays showed that EDTA did not affect the catalytic activity of RICE1.
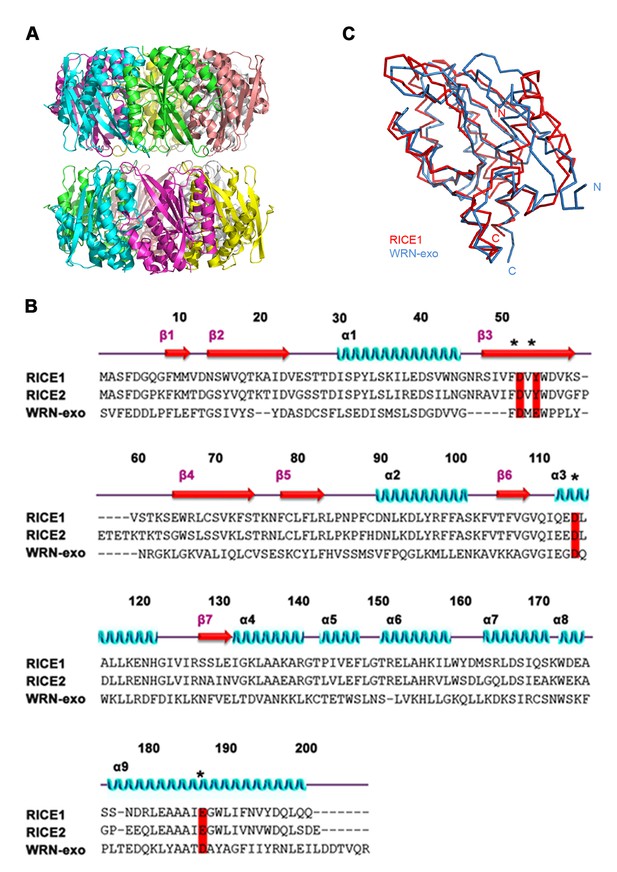
Structural features of RICE1.
(A) Double donut assembly of RICE1 hexamers in the crystal. (B) Sequence alignment with RICE1 and its Arabidopsis paralog, RICE2, as well as structural homolog, WRN-exo. (C) Overlaid structures of RICE1 and WRN-exo.
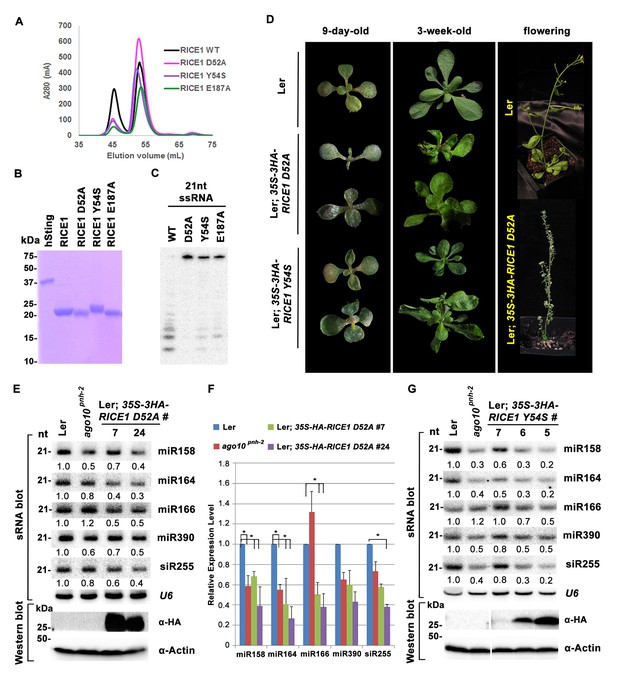
Mutations in catalytic sites impair RICE1 function in vivo.
(A) Chromatograph of representative RICE1 variants with mutations in catalytic sites. (B) SDS-PAGE of purified RICE1 mutants and control protein hSting. (C) Representative RICE1 mutants were compromised exoribonucleases compared to wild type (WT). (D) Phenotypic anomaly of transgenic plants ectopically expressing catalytically-inactive RICE1 variants in different stages. Note: the severe sterility of 35S-3HA-RICE D52A transgenic plants. (E–G) Ectopic expression of RICE1 variants caused dominant-negative effect on miRNA accumulation in vivo. In (E) and (G), sRNA blot analyses were done with the indicated 32P-labelled probes. The amount of miRNAs in wild-type plants was arbitrarily designated as 1.0, and the relative amount in the other lines was normalized to that of control plants. In (F), quantification of three biological replicates was statistically analyzed and the significant difference (Student t-test, p<0.05) shown by the asterisk (*). Additional biological replicates were shown in Figure 5—figure supplement 1C and Supplementary file 1B. Western blot analyses were performed with anti-HA or anti-actin antibodies.
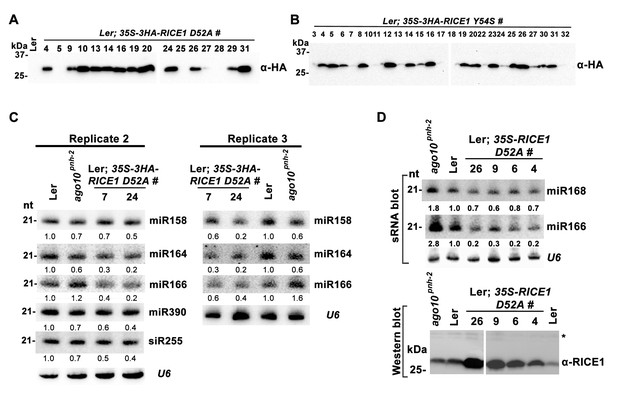
Catalytic inactive RICE1 impairs the accumulation of miRNAs in vivo.
(A and B) Western blot analyses of transgenic plants expressing RICE1 D52A and Y54S using anti-HA antibody. (C and D) Ectopic expression of HA-tagged (C) and untagged (D) RICE1 D52A caused dominant-negative effect on miRNA accumulation in vivo. sRNA blot analyses were done with the indicated 32P-labelled probes. The amount of miRNAs in wild-type plants was arbitrarily designated as 1.0, and the relative amount in the other lines was normalized to that of control plants. The right column chart showed the quantification of sRNA blots and asterisks (*) indicated statistically significant differences between wild type and transgenic plants (Student t-test, p<0.05).
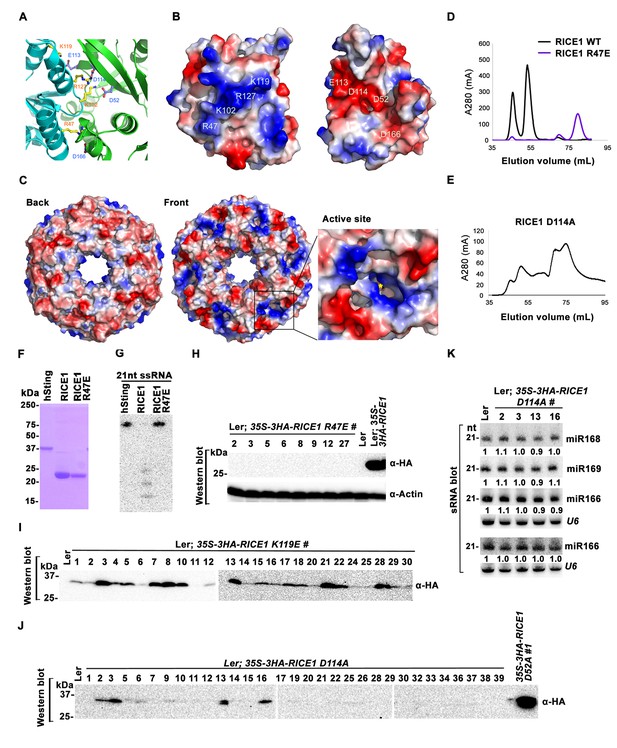
Mutations at the dimer interface impair RICE1 function and protein stability in vivo.
(A) Interface between RICE1 monomers. Amino acid residues involved in the interaction are shown in ball-and-stick models. (B and C) The electrostatic potential at the surfaces of RICE1 monomers and hexamer. Positively and negatively charged surfaces are colored blue and red, respectively. In (C), the active site of RICE1 (indicated by the yellow asterisk) is located at the bottom of a deep groove between RICE1 monomers. (D and E) SEC assays of RICE1 R47E and D114A mutants defective in oligomerization. Note: SEC assay was conducted using Hiload 16/600 Superdex 200 in (E) instead of routinely used Superdex 75. (F) SDS-PAGE of purified wild type, mutated RICE1, and control protein hSting. (G) Oligomerization is essential for RICE1 catalytic activity. (H) RICE1 R47E mutant proteins failing in oligomerization are unstable in vivo. (J) RICE1 D114A, unlike D52A, Y54S, and K119E mutants, was unstable and barely detectable in vivo. (K) Ectopic expression of 3HA-RICE1 D114A had no significant effect on miRNA accumulation in vivo. Western blot analyses were performed with anti-HA antibody. sRNA blot analyses were done with the indicated 32P-labelled probes. The amount of miRNAs in wild-type plants was arbitrarily designated as 1.0, and the relative amount in the other lines was normalized to that of control plants.
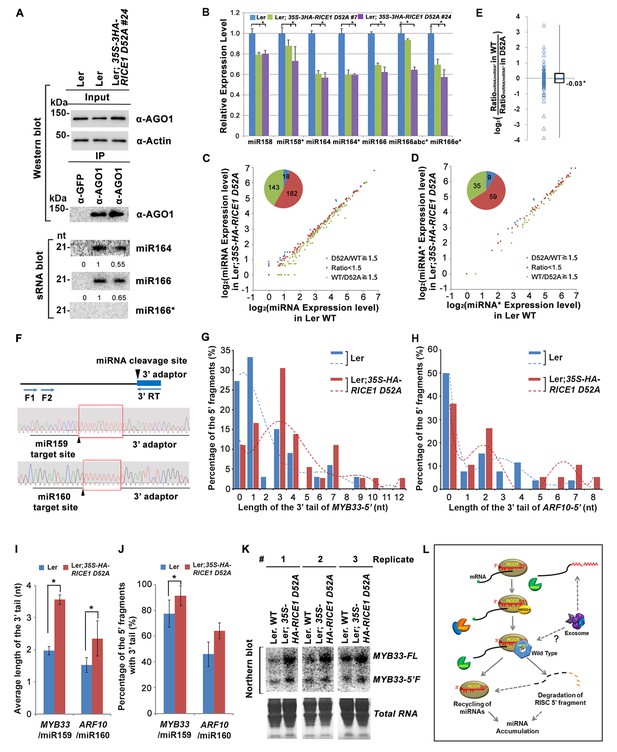
Catalytically-inactive RICE1 extends uridylated 5’ cleavage fragments from RISC activity whereas decreases the abundance of RISC-harbored miRNA in vivo.
(A) Ectopic expression of catalytically-inactive RICE1 mutants impaired the accumulation of miRNAs in RISC. Western blot analyses were done with anti-AGO1 and anti-Actin antibodies. Actin serves as a loading control. sRNA blot analyses of sRNA recovered from AGO1 IP were conducted with 32P-labeled probe against miR166. The ratio of miR166/AGO1 in wild type was arbitrarily designated as 1.0, and the relative amount in each sample was normalized to that of control plants. Additional biological replicates were shown in Figure 7—figure supplement 1A,B. (B) Concurrent reduction of miRNA and miRNA* abundance in the transgenic plants expressing dominant-negative RICE1 mutant. The relative level of miRNA or miRNA* was normalized to that in Ler plants where the amount was arbitrarily designated as 1.0 with ±SD from at least three experiments, asterisks (*) indicate statistically significant differences (Student t-test, p<0.05). See also in Supplementary file 1C. (C and D) sRNA-seq analyses of miRNA or miRNA* expression in wild type and transgenic plants expressing dominant-negative RICE1 mutant. X and Y axes indicate the logarithm base two for the miRNA or miRNA* expression level in wild type and 35S-HA-RICE1 D52A transgenic plants, respectively. The labels in the bottom right of the chart indicated the categories of miRNA or miRNA*s: D52A/WT ≥1.5, upregulated in 35S-HA-RICE1 D52A transgenic plants for more than 1.5 fold; WT/D52A ≥1.5, downregulated in transgenic plants for more than 1.5 fold; Ratio <1.5, the rest with fold changes less than 1.5 fold. The pie in the top left of the chart indicated the numbers of different categories of miRNA or miRNA*s. See also in Supplementary file 1D and 1E. (E) Comparison of the ratio of miRNA vs miRNA* in wild type and transgenic plants expressing dominant-negative RICE1 mutant. The empty triangle indicates the logarithm base two for the ratios of individual pairs of RatiomiRNA/miRNA* in wild type relative to the ones in 35S-HA-RICE1 D52A transgenic plants. RatiomiRNA/miRNA* = miRNA expression level/miRNA* expression level. *, the median value of the logarithm values mentioned above in the whole populations of tested miRNA/miRNA* pairs. See also in Supplementary file 1F. (F) Schematic diagram of 3’ al-RACE to detect the uridine addition (red rectangle) in the 3’ end of RISC-cleaved 5’ fragment. The miR159- and miR160-cleavage sites were shown with black triangle. (G–I) Uridylation of 5’ fragments in Ler and Ler; 35S-HA-RICE1 D52A. The representative length distribution patterns of uridylated MYB33-5’ (G) and ARF10-5’ (H), and the fitting curves in dotted lines indicate the tendency of the length distribution, respectively; the mean length of the 3’ tails and the mean percentage of uridylated clones in total clones were shown in (I) and (J), respectively; significant difference (Student t-test, p<0.05) was indicated with the asterisk (*). See the details for three biological replicates in Figure 7—figure supplement 2A–D and Supplementary file 3A and 3B. (K) The abundance of MYB33-5’ in Ler and Ler; 35S-HA-RICE1 D52A. MYB33-5’ RNAs were detected by Northern blot in three biological replicates, using probes for MYB33-5’ from miR159-AGO1-mediated cleavage. FL, full-length MYB33 transcripts; 5’F, 5’ fragment. Methylene blue staining of total RNAs shows the loading controls. (L) A proposed model for RICE functions in miRNA-RISC. RICEs facilitate the clearance of the uridylated tail of 5’ RISC-cleaved fragments and promote RISC recycling and miRNA stability.
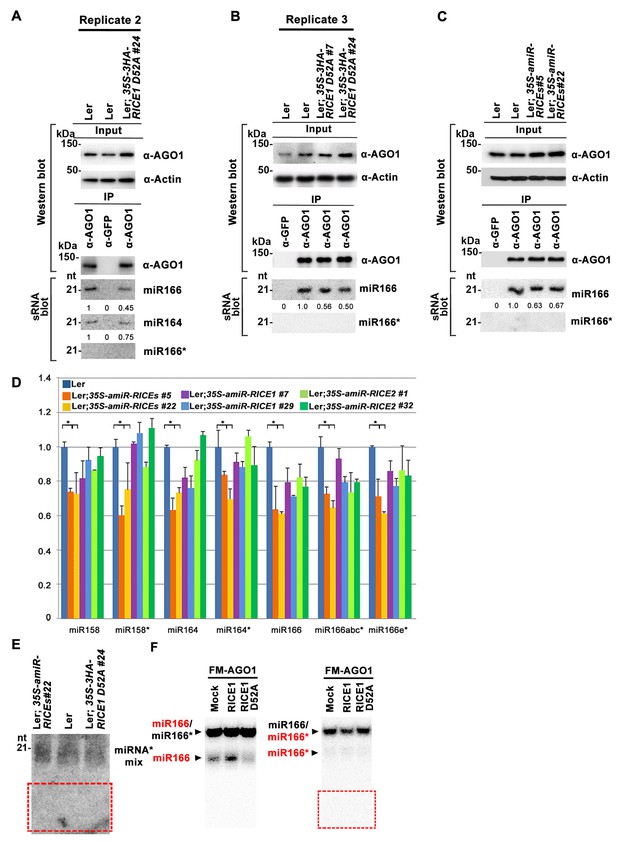
RICEs affect the accumulation of miRNAs and miRNA*.
(A–C) Ectopic expression of catalytically-inactive RICE1 mutants (A and B) and downregulation of RICEs (C) decreased miRNA accumulation in RISC. Western blot analyses were done with the crude extract (Input) and the IP products using anti-AGO1 antibody (top panels). Actin serves as a loading control. sRNA blot analyses of sRNA recovered from AGO1 IP were conducted with 32P-labelled probe against miR166. The ratio of miR166/AGO1 in wild-type Ler was arbitrarily designated as 1.0, and the relative amount in each sample was normalized to that of control plants. (D) Quantitative PCR showed the concurrent reduction of miRNA and miRNA* abundance in the transgenic plants expressing artificial miRNAs specifically targeting RICE1, RICE2 or both. The relative level of miRNA and miRNA* was normalized to that in Ler plants where the amount was arbitrarily assigned a value of 1 with ±SD from at least three experiments, asterisks indicate statistically significant differences between wild type and transgenic plants. (E) sRNA blot analysis of highly-enriched low-molecular-weight RNAs prepared from wild-type and RICE-compromised transgenic lines. Numerous 21-nt and truncated oligo probes complementary to miR165 (a–b)*, miR166 (a–g)*, miR159b*, miR160c*, miR162b*, miR164b*, miR167b*, miR168a*, and miR171a* were pooled for sRNA blot. Note: The region marked in red box is the place where 9-nt and 12-nt miRNA* fragments would have been detected, if present. (F) In vitro RISC assembly assays to test the effect of the D52A mutation in RICE1 on removal of miRNA*s and nicked miRNA* in wheat germ. Flag-4Myc-AGO1, RICE1 wild type and D52A mutant protein were synthesized in wheat germ extract from TNT T7 Quick Coupled transcription/translation system before application of miR166/166* duplexes containing either 32P-labelled miR166 or miR166* (indicated by RED). The reaction mixtures were finally extracted with equal volumes of TE-saturated phenol. The RNA in the aqueous phase was recovered, analyzed with native 15–18% PAGE. The 32P signals were detected after exposure to a phosphorImager plate (Molecular Dynamics). Negative control reaction using mock-translated wheat germ was performed in parallel. Note: 9-nt and 12-nt miRNA* fragments were not detected in the red box region.
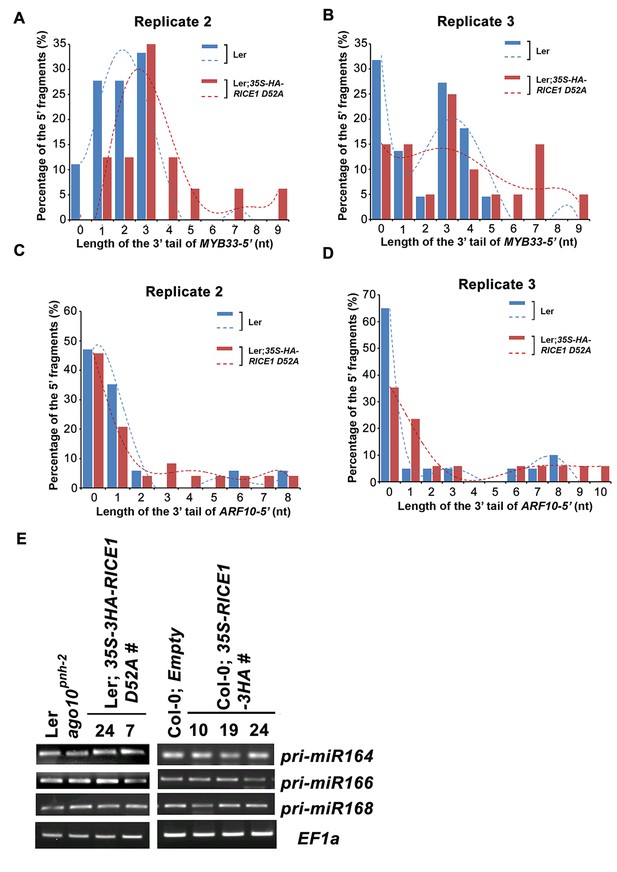
RICEs affect the accumulation of the uridylated 5’ RISC cleavage fragments.
(A–D) The length distribution pattern of uridylated MYB33-5’ (A and B) and ARF10-5’ (C and D), and the fitting curves in dotted lines indicate the tendency of the length distribution pattern, respectively. (E) semi RT-qPCR analysis of selected pri-miRNAs show that RICEs do not affect miRNA biogenesis in Arabidopsis.
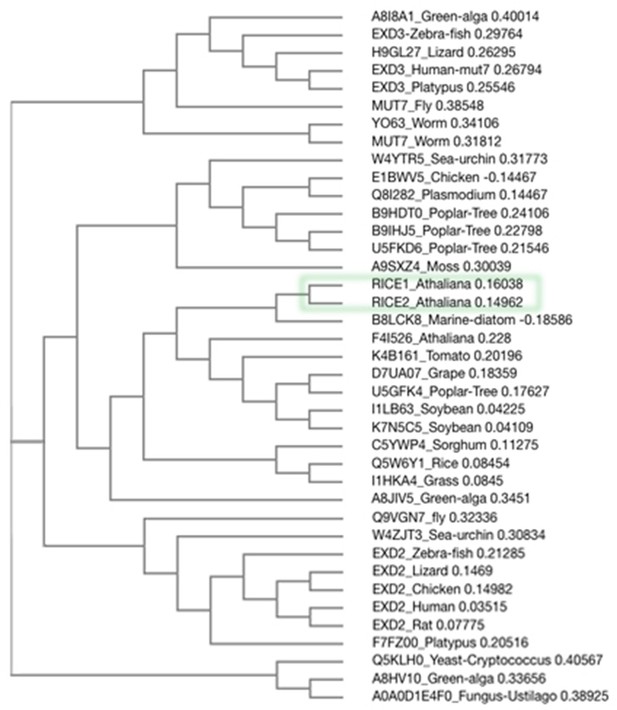

Phylogenetic analysis of RICE proteins and their orthologs in eukaryotes.
Alignment of RICE proteins and their orthologs in eukaryotes was done with Clustal Omega. The evolutionary relationships were inferred.
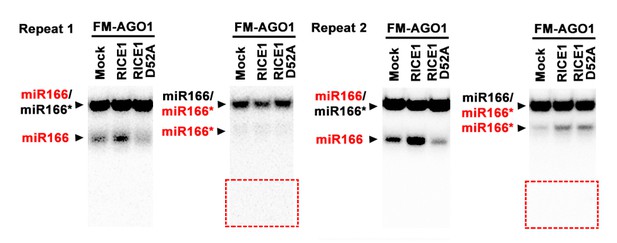
Effect of the D52A mutation in RICE1 on removal of miRNA*s in wheat germ.
Note: (1) The residue miRNA*s in the repeat #2 might be due to incomplete annealing (that is, more molarity of miRNA* than miRNA; or dissociation of miRNA/miRNA* complexes during sample processing); (2) The red box regions are supposed to be locations for 9-nt and 12-nt miRNA* fragments if present. (3) We have done this experiment for dozens of times: results related to accumulation of miRNA and miRNA*s were not very stable, however, the failure to detect 9-nt and 12-nt was always consistent.
Additional files
-
Supplementary file 1
Quantitative examination of miRNA and miRNA expression in wild-type and RICE compromised mutants.
(A) Quantification of miRNA expression in artificial miRNA knockdown lines. (B) Quantification of miRNA expression in catalytically-inactive mutants. (C) Quantification of miRNA/miRNA* expression in catalytically-inactive mutants. (D) Expression pattern of miRNAs in wild type and 35S-HA-RICE1 D52A transgenic plants. (E) Expression pattern of miRNA* in wild type and 35S-HA-RICE1 D52A transgenic plants. (F) Ratio of miRNA/miRNA* in wild type and 35S-HA-RICE1 D52A transgenic plants. (G) Primers and oligos used in this study.
- https://doi.org/10.7554/eLife.24466.019
-
Supplementary file 2
Statistics of crystallographic analysis for RICE1.
- https://doi.org/10.7554/eLife.24466.020
-
Supplementary file 3
3’ RACE for 5’ uridylated cleavage fragments from RISC activity in wild type and the transgenic plant overexpressing catalytic inactive RICE1.
(A) Detailed sequences and ratios of 3’ uridylated tails of 5’ RISC cleavage fragments. (B) Statistic analysis of the average length of 3’ uridylated tail (nt) and percentage of the 5’ fragments with such tails.
- https://doi.org/10.7554/eLife.24466.021





