Structural basis of protein translocation by the Vps4-Vta1 AAA ATPase
Figures
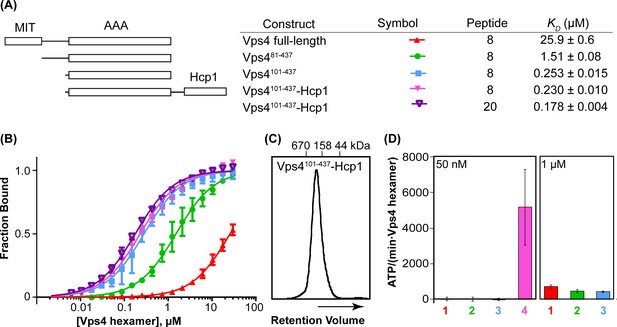
Vps4101-437-Hcp1 is an active hexamer.
(A) Vps4 constructs and peptide-binding affinities assayed by fluorescence polarization. Peptide ‘20’ is a Vps2-derived 20-residue peptide C identified earlier (Han et al., 2015). Peptide ‘8’ is an 8-residue fragment (DEIVNKVL) of peptide ‘20’ that retains essentially full binding affinity. The relatively weak binding of full-length Vps4 reflects autoinhibition mediated by the MIT domains (Han et al., 2015). (B) Fluorescence polarization isotherms corresponding to values in panel A. Means and standard deviations are from four independent experiments. (C) Gel filtration of Vps4101-437-Hcp1 on a Superdex 200 column in 25 mM Tris/HCl pH 7.4, 100 mM NaCl and 1 mM DTT. The protein elutes as a symmetric peak with an apparent molecular mass of 290 kDa, in good agreement with the calculated molecular mass of a hexamer (330 kDa). (D) ATPase activities for Vps4 constructs: 1, Vps4 full-length; 2, Vps481-437; 3, Vps4101-437; 4, Vps4101-437-Hcp1. Vps4 subunit concentrations are indicated. Means and standard deviations from at least three independent measurements.
-
Figure 1—source data 1
Binding of fluorescently labeled ESCRT-III peptides to Vps4, related to Figure 1B.
- https://doi.org/10.7554/eLife.24487.004
-
Figure 1—source data 2
ATPase activity of Vps4 constructs, related to Figure 1D.
- https://doi.org/10.7554/eLife.24487.005
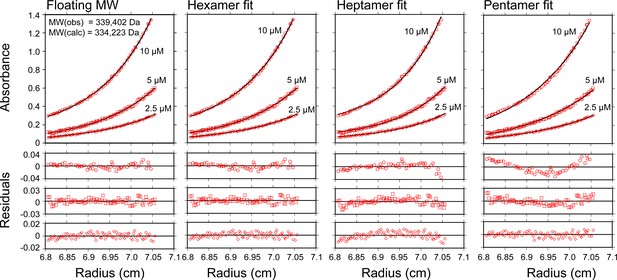
Vps4101-437-Hcp1 is a hexamer.
Equilibrium sedimentation of Vps4101-437-Hcp1. Absorbance is shown as a function of distance from the axis of rotation for three loading concentrations (open circles, 10 µM; open squares, 5 µM; open diamonds, 2.5 µM Vps4 subunits). The data were fit to a single species model where the molecular weight was allowed to float or held constant as indicated (black line), and residuals for each concentration are shown below. The molecular weight was fit to 329,402 Da in good agreement with the expected molecular weight for a hexamer of 334,223 Da. When the molecular weight was held constant, the data agree well with a hexamer model, whereas the residuals are clearly biased (and in opposite directions) when the molecular weight is set to a heptamer or pentamer.
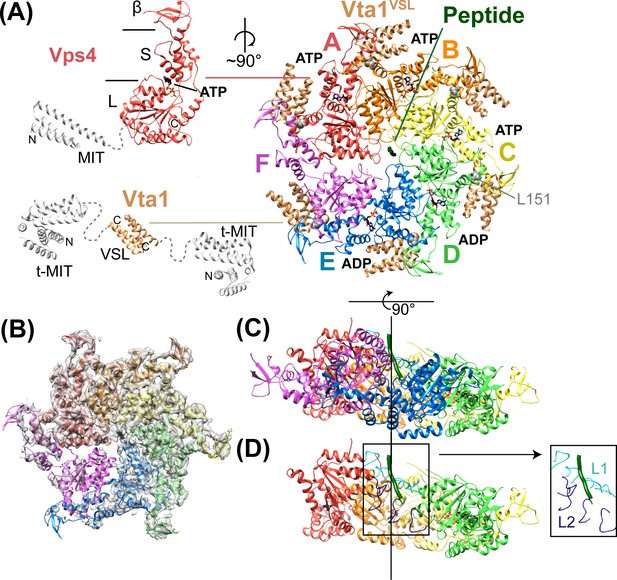
Structure of Vps4101-437:Vta1VSL:ESCRT-IIIpeptide:ADP·BeFx.
(A) Structure of the complex. The Vps4 and Vta1 constructs used for cryo-EM structure determination are shown in color on the left, with excluded segments colored white. MIT, large AAA ATPase (L), small AAA ATPase (S) and β domains of Vps4 are labeled, as are the t-MIT and VSL domains of the Vta1 dimer. L151, a residue critical for hexamerization, is shown in gray spheres. (B) 4.3 Å map with the Vps4 model. (C) Side view of Vps4 hexamer, oriented with the subunit A-D helix axis vertical (black line). (D) Same as panel D but with subunits E and F removed. The inset shows the position of pore loops 1 (L1, residues 203–210, cyan) and pore loops 2 (L2, 240–248, dark blue) relative to the ESCRT-III peptide (dark green).
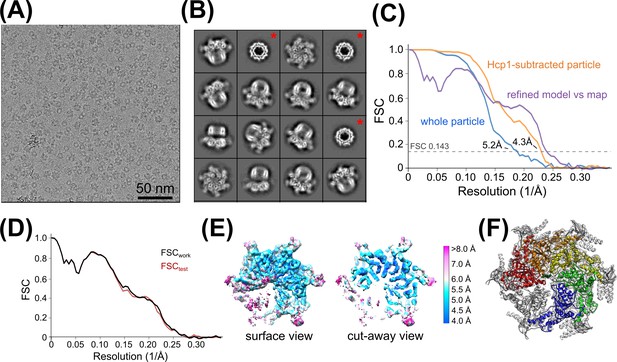
Vps4 3D reconstruction, refinement, and validation.
(A) Representative cryo-EM image of Vps4101-437-Hcp1 particles. (B) Representative 2D class averages of Vps4101-437-Hcp1 particles. Red asterisks indicate classes with disordered Vps4 in which only the Hcp1 template is apparent. (C) ‘Gold standard’ FSC curves generated by RELION before (blue) and after (orange) Hcp1 signal subtraction. The FSC curve of the refined model (comprising the large and small AAA ATPase domains of subunits A-E and substrate peptide) against the final Hcp1-subtracted Vps4 map is shown in purple. (D) Cross-validation of the refined model. The refined model (comprising large and small AAA ATPase domains of subunits A-E and the substrate peptide) was randomly displaced by applying 0.5 Å shifts to all atoms and refined against one of the half maps generated by RELION. FSC curves are shown between the re-refined model against the half map used for re-refinement (FSCwork, black) and between the re-refined model and the other half map (FSCtest, red). The agreement between the two FSC curves is an indicator that the model has not been overfit. (E) Local resolution estimates determined by ResMap. (F) The composite model indicating the refined portions of Vps4 (colored ribbons) and other regions limited to rigid body fitting (Vps4 β domains, subunit F, and Vta1VSL, gray ribbons). Same orientation as panel (E). Note that Vta1 densities are weak prior to 3D classification (see Figure 2—figure supplement 6).
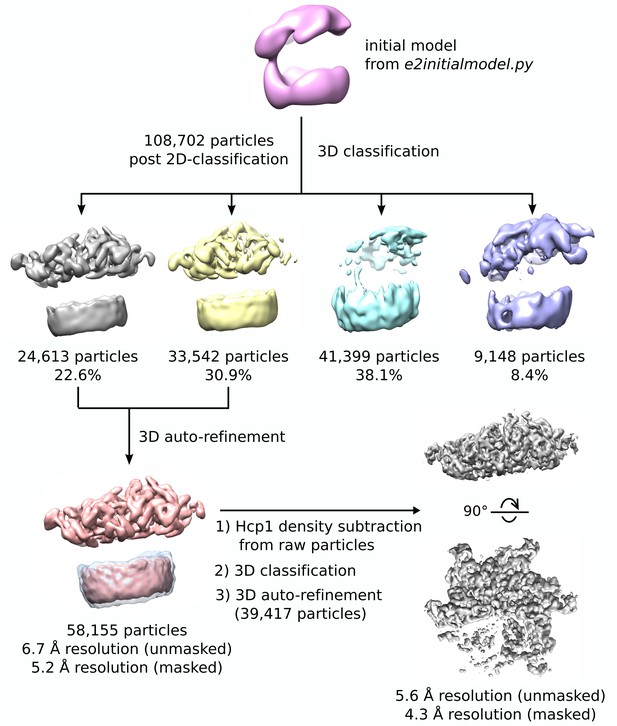
3D reconstruction workflow.
Flow chart depicting classification and refinement of Vps4 particles. An initial model was generated from a gallery of non-CTF corrected 2D class averages, which was then used as a starting point for 3D classification. Particles from two classes showed ordered Vps4 features, which were then used to compute a 6.7 Å resolution consensus structure. Hcp1 densities were subtracted from raw images, followed by an additional round of RELION 3D classification and auto-refinement, which produced the final 4.3 Å resolution reconstruction.
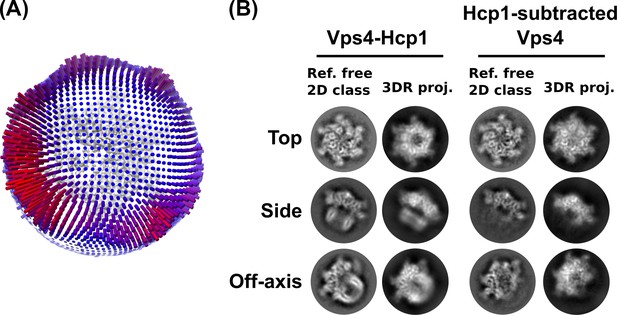
Additional validation of the 3D reconstruction.
(A) Angular distribution plot based on RELION assignments and visualized in UCSF Chimera. (B) Comparison between reference-free 2D class averages and re-projections of the 3D reconstructions of the Vps4101-437-Hcp1 particle and Hcp1-subtracted Vps4 particle.
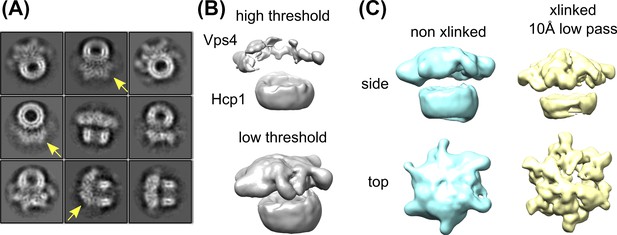
Glutaraldehyde crosslinking improves the Vps4 density without distorting the structure.
(A) Reference-free 2D class averages of non-crosslinked Vps4101-437-Hcp1 particles. Note that the Vps4 features are weaker and smeared out (yellow arrows) relative to the Hcp1 template and relative to the crosslinked sample (Figure 2—figure supplement 1). (B) 3D reconstruction of non-crosslinked Vps4101-437-Hcp1 particles reveals the Vps4 hexamer only at very low thresholds. (C) Comparison between non-crosslinked (cyan) versus crosslinked particles (yellow) reveal consistent features.
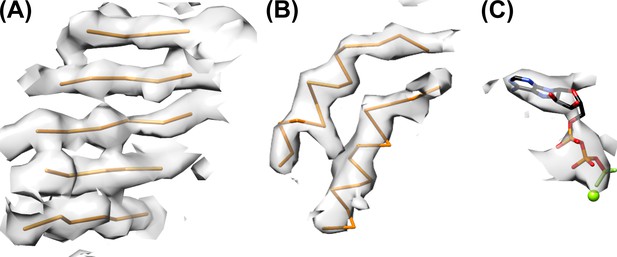
Refined model and representative density.
(A) Central β-sheet of subunit B with density. (B) Helices of subunit B. (C) Nucleotide density with ADP·BeFx and magnesium ion bound to subunit B.
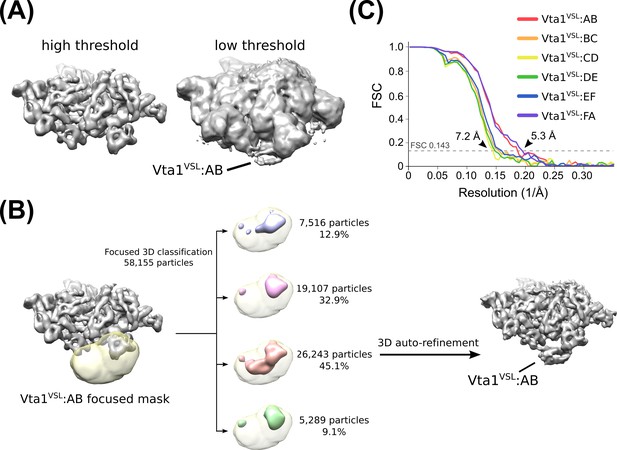
Identification and classification of Vta1 density.
(A) Vta1VSL densities are visible at low threshold levels in an overall Hcp1-subtracted map. This observation prompted us to perform focused 3D classification with a mask over the expected Vta1 binding site. (B) Flow chart depicting 3D classification of the consensus structure with a focused mask (yellow) at the interface of subunits A and B. Classification revealed one distinct class with robust Vta1 features and the particles were isolated and subjected to an additional round of RELION auto-refinement (light red). The same strategy was employed for each interface. (C) ‘Gold standard’ FSC plots of each of the six Vta1VSL datasets derived from focused 3D classification.
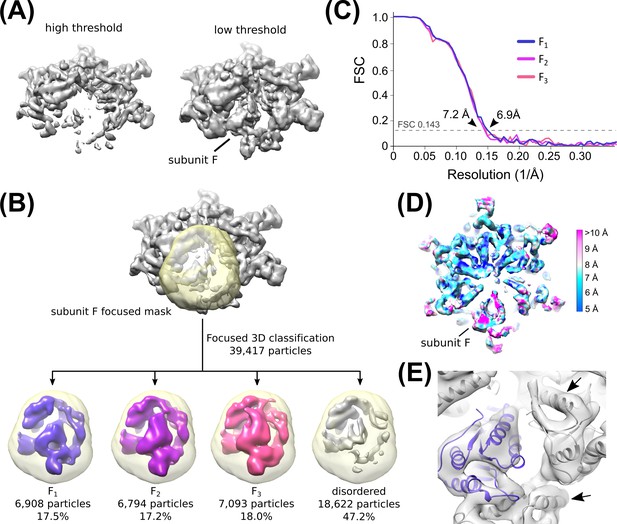
Classification of subunit F Density.
(A) Subunit F is poorly resolved but visible at low threshold levels in an overall Hcp1-subtracted map. This prompted us to perform focused 3D classification with a mask over subunit F. (B) Flow chart depicting 3D classification of the consensus structure with a focused mask (yellow) over subunit F. Classification revealed three distinct classes that could accommodate a rigid-body fit of the Vps4 crystal structure. See Figure 2—figure supplement 6 and Methods for details. (C) ‘Gold standard’ FSC plots of the three subunit F datasets derived from focused 3D classification. (D) Cut-away view depicting the local resolution of the F1 map determined by ResMap. Note that despite the overall ~7 Å resolution of the map, subunit F itself is less well resolved. (E) Fitting of Vps4 coordinates into the F1 structure confirms that our map resolves individual helices for Vps4 subunits A-E (arrows), despite the lower resolution density for subunit F (blue).
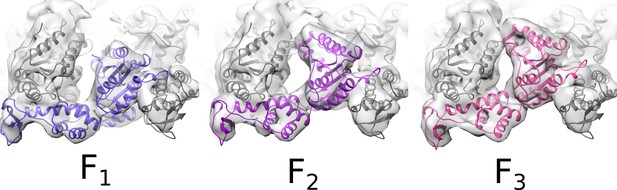
Rigid-body fitting of Vps4 subunit F.
Rigid-body fitting of Vps4 subunit F (colored ribbon) into three different density maps from focused 3D classification. The three models are related by pivoting of up to 16° about a point near the contact with Vta1VSL, close to the small AAA ATPase and β domains.
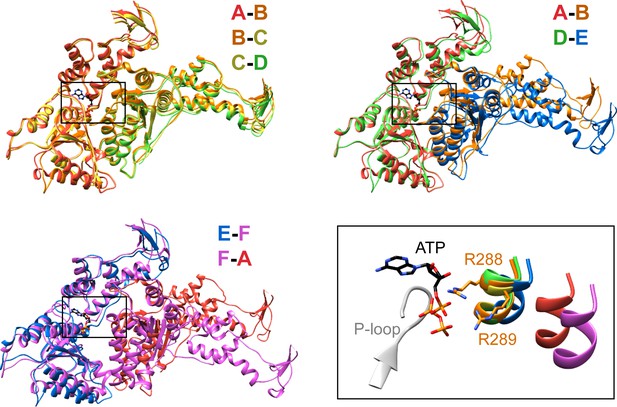
Interfaces in the asymmetric Vps4 hexamer.
Vps4 subunit pairs superimposed on the large AAA ATPase domain of the first subunit, as indicated. A-B, B-C and C-D interfaces are equivalent. The nucleotide-binding site is slightly expanded at the D-E interface due to a 15° relative rotation of subunit E. The E-F and F-A sites are open for nucleotide exchange. Inset (black rectangle), Close-up on the nucleotide binding site showing the nucleotide and coordinating P-loop for the first subunit, with the R288/R289-containing helix of the second subunits in color. These arginine finger Cα atoms shift by 2 Å at the D-E interface relative to the A-B, B-C, and C-D subunits. The E-F and F-A interfaces are shifted by 8 Å and 16 Å, respectively.
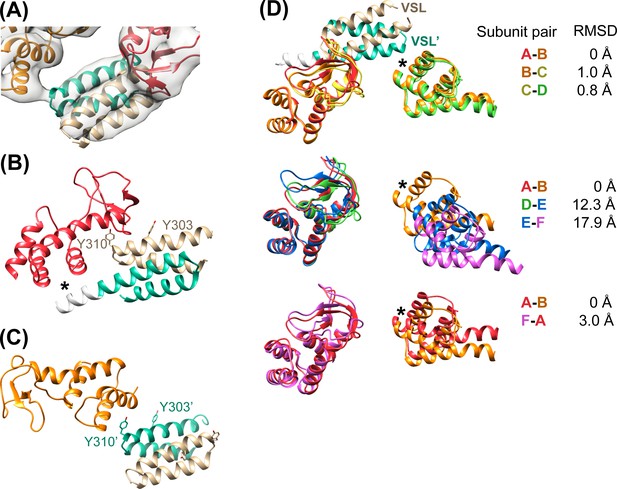
Vta1VSL contacts with Vps4.
(A) Density for the most clearly defined Vta1VSL (bound to the Vps4 subunit A β domain). The Vta1VSL subunits are colored tan and teal. (B) Vta1VSL interaction with the first Vps4 subunit. This interface is modeled identically to a crystal structure of Vta1VSL in complex with a truncated Vps4 construct (Yang and Hurley, 2010). Additional N-terminal residues in the longer Vta1 construct used in this study are shown in white and their interaction with the small AAA ATPase domain of Vps4 is indicated with an asterisk. (C) Vta1VSL interaction with the second Vps4 subunit. Y303’ and Y310’ are labeled. (D) Overlap of subunit pairs on the small AAA ATPase domain of the first Vps4 (residues 301–349 and 403–411). Consequent RMSD values are shown for residues 300–311 and 320–331 of the second Vps4 subunit at the second Vta1 interface (asterisk).
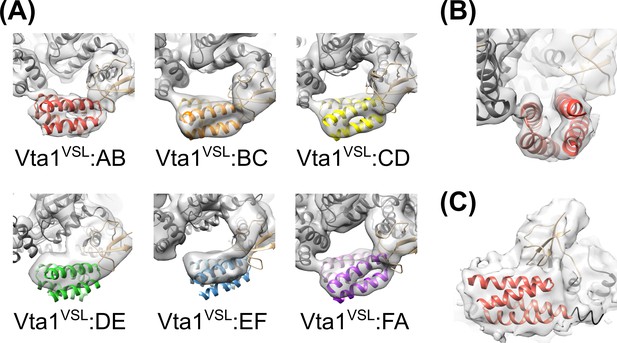
Rigid-body fitting of Vps4 β domain-Vta1VSL complexes into each density map.
(A) The A-B, B-C, C-D, and F-A Vta1VSL dimers show density for distinct helices. Density for D-E and E-F Vta1VSL dimers is poor and only S and β domain coordinates were used for rigid body fitting at these two interfaces. (B) Side view of the fitting at the A-B interface as in panel (A) showing distinct density for each of the four Vta1VSL helices. (C) Low threshold view of the A-B map reveals extensions of the Vta1VSL density that can accommodate the additional N-terminal residues included in our Vta1 construct (black ribbon).
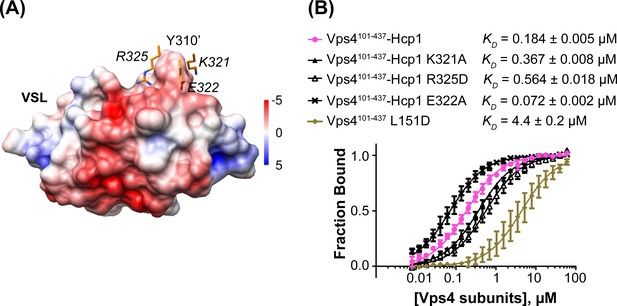
Mutations at the Vta1 interface with the second Vps4 subunit alter binding affinity.
(A) K321, E322 and R325 of the Vps4 small AAA ATPase domain (orange, labels in italic font) contact Y310’ of Vta1VSL (surface representation colored by electrostatic potential, kT/e) in the interaction shown in Figure 4C. (B) Binding of fluorescently labeled Vta1VSL to the Vps4101-437-Hcp1 hexamer (pink circles) is ~24x tighter than binding to a monomeric Vps4 construct, Vps4101-437 L151D (gold diamonds). Consistent with the Vta1 surface electrostatic potential, point mutations K321A and R325D weaken Vta1VSL binding 2-fold and 3-fold, respectively, while E322A strengthens binding 2-fold. Means and standard deviations are from at least three biological replicates.
-
Figure 5—source data 1
Binding of Vta1VSL to Vps4, related to Figure 5B.
- https://doi.org/10.7554/eLife.24487.021
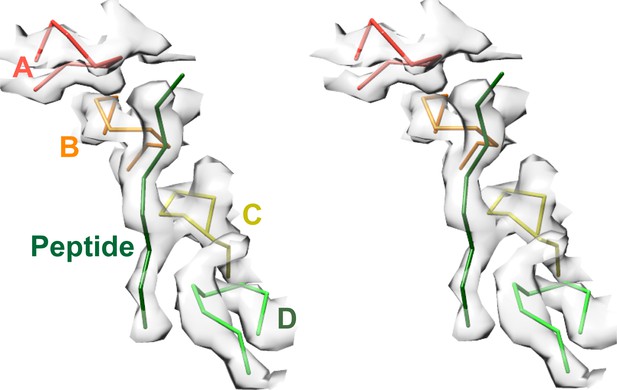
Pore loops of Vps4 form a spiral staircase to coordinate the substrate peptide.
Stereo view of the peptide and pore loop 1 (residues 203–210) of subunits A-D with density.
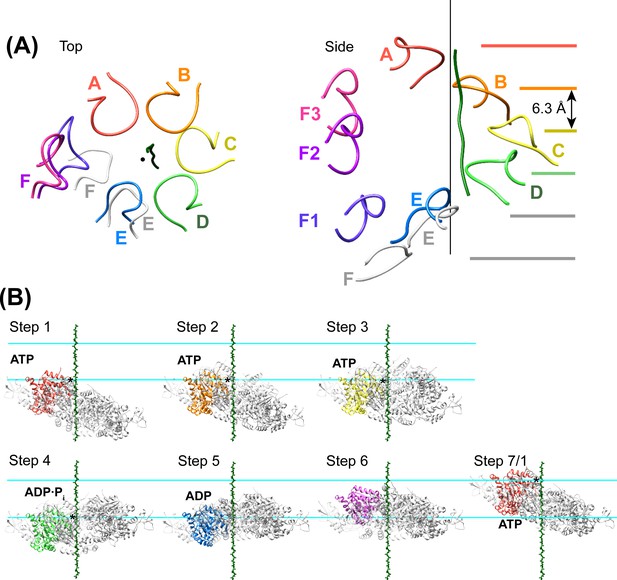
Peptide binding and mechanism of translocation.
(A) The pore loop 1 residues of subunits A-D form a helix (axis, black line) that matches the symmetry of a canonical twisted β-strand, which rotates 60° and translates 6.3 Å every two residues. In white are the positions that subunits E and F would adopt if they continued this helix. The three positions seen for subunit F (Figure 2—figure supplements 7–8) appear to be snapshots along the return path from the end of the helix at subunit E to the start of the helix at subunit A. (B) Steps along the translocation cycle inferred from the cryo-EM structure. The peptide shown is modeled as a β-strand along the helix axis of subunits A-D. Vps4 maintains a constant interaction with the peptide through steps 1 to 4 before dissociating at step 5 and rebinding 12 residues further up the peptide at step 7, which is equivalent to step 1. Nucleotides suggested by density and coordination geometry are labeled. Pore loop 1 contacts with the substrate peptide in steps 1–4 are indicated with an asterisk. The two subunits closest to the view direction are included with 50% transparency. The two horizontal lines are separated by 37.8 Å (12 residues) and indicate points of substrate contact with pore loop 1 of the highlighted subunit.
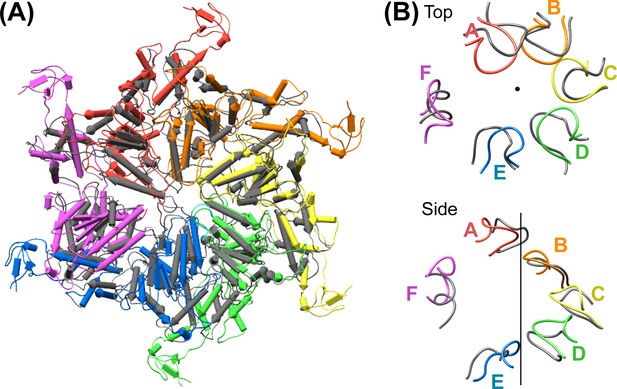
Comparison of the Vps4 hexamer with the ATPases of the 26S proteasome.
(A) Superposition of secondary structure elements reveals a similar overall structure between Vps4 and the ATPase subunits of the 26S proteasome (PDB 5GJQ, gray). (B) Top view and side view of the Vps4 pore loop 1 of Vps4 subunits (rainbow) and the proteasome (gray) shows close similarity about the helix axis (black).
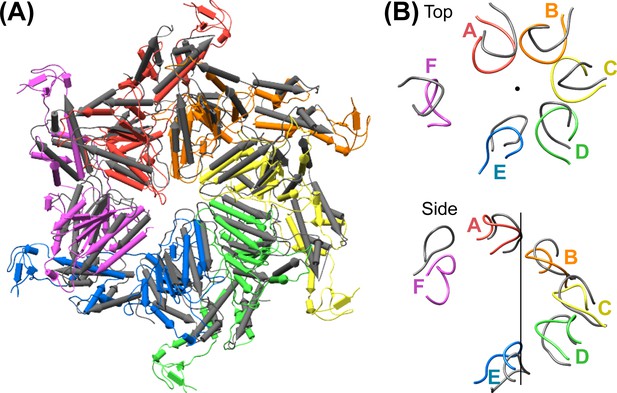
Comparison of the Vps4 hexamer with the NSF D1 ring.
(A) Superposition of secondary structure elements reveals a similar overall structure between Vps4 and the D1 ring of NSF (PDB 3J94, residues 215–489, gray). (B) Top view and side view of the Vps4 (rainbow) and NSF D1 (gray) subunit pore loop one elements shows close similarity about the helix axis (black).
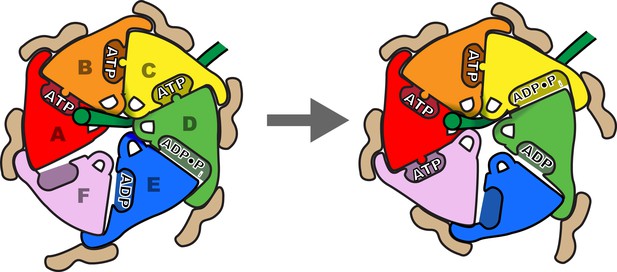
Schematic of one step in the translocation mechanism.
Left, Subunits A-D form a helical surface of pore loop 1 residues that binds substrate in a β conformation along or close to the helix axis. The helix is stabilized by Vta1VSL binding to adjacent subunits and by ATP binding at subunit interfaces. Right, next step in the cycle where subunit F has bound ATP and assembled on the growing end of the Vps4 helix, ATP has been hydrolysed at the C-D interface, and the nucleotide-binding site of subunit E has been opened to allow ADP·Pi release and rebinding of ATP.
Videos
Top view of the proposed translocation mechanism.
Vps4 reaction cycle modeled by linear interpolation between the six different states represented in the cryo-EM structure. The ESCRT-III substrate is modeled as a β-strand lying along the axis of the helix defined by Vps4 subunits A-D. Nucleotides are shown in pink (ATP) and gray (ADP).
Side view of the proposed translocation mechanism.
As for Video 1.
Model of Vps4 assembly and ESCRT-III disassembly.
Vps4 (purple) is recruited to the ESCRT-III lattice (green) through binding of its N-terminal MIT domain to MIT Interacting Motifs (MIMs) (Kieffer et al., 2008; Obita et al., 2007; Stuchell-Brereton et al., 2007), which are sequences at the ends of the long, flexible C-terminal tails of ESCRT-III subunits. The avidity effect of the ESCRT-III polymer promotes Vps4 hexamerization around flexible ESCRT-III sequences. The hexamer is further stabilized by the dimeric Vta1 protein (tan), which also concentrates at the ESCRT-III polymer through its N-terminal t-MIT domain (not shown) (Guo and Xu, 2015; Skalicky et al., 2012; Vild et al., 2015). The Vps4 hexamer hydrolyzes ATP and translocates the substrate through the central pore, thereby destabilizing the ESCRT-III structure and removing the subunit from the lattice. We speculate that Vps4 and Vta1 remain bound to the ESCRT-III lattice via their MIT domains such that they are in position to process additional ESCRT-III subunits until the polymer is disassembled. This animation was created using Autodesk Maya (Autodesk, Inc.) from protein structural models exported from UCSF Chimera (Pettersen et al., 2004).
Tables
Reconstruction, refinement, and model statistics of Vps4.
| Vps4101-437-Hcp1, whole particle | Hcp1-subtracted Vps4 | |
|---|---|---|
| Reconstruction | ||
| Particle images | 58,155 | 39,417 |
| Resolution (unmasked, Å) | 6.7 | 5.7 |
| Resolution (masked, Å) | 5.2 | 4.3 |
| Map sharpening B-factor (Å2) | - | −208 |
| EM Databank Accession Number | EMD-8551 | EMD-8550 |
| Refinement and validation of Vps4 subunits A-E | ||
| Resolution used for refinement (Å) | - | 4.3 |
| Number of atoms | - | 10604 |
| R.M.S deviation | ||
| Bond length (Å) | - | 0.01 |
| Bond angles (°) | - | 0.92 |
| Ramachandran | ||
| Favored (%) | - | 91.13 |
| Allowed (%) | - | 8.87 |
| Outlier (%) | - | 0.00 |
| Molprobity score / percentile (%) | - | 1.94/100th |
| Clashscore / percentile (%) | - | 7.75/97th |
| PDB | - | 5UIE |
Reconstruction statistics of Vps4-Vta1 classes.
| Vps4 Vta1VSL (A-B) | Vps4 Vta1VSL (B-C) | Vps4 Vta1VSL (C-D) | Vps4 Vta1VSL (D-E) | Vps4 Vta1VSL (E-F) | Vps4 Vta1VSL (F-A) | |
|---|---|---|---|---|---|---|
| Particle images | 26,243 | 13,066 | 10,700 | 11,684 | 14,274 | 26,964 |
| Resolution (unmasked, Å) | 6.9 | 7.8 | 7.8 | 7.8 | 7.5 | 6.9 |
| Resolution (masked, Å) | 5.4 | 6.7 | 7.2 | 6.9 | 6.5 | 5.3 |
| EMDB ID | EMD-8552 | EMD-8553 | EMD-8554 | EMD-8555 | EMD-8556 | EMD-8557 |
Reconstruction statistics of Vps4-Subunit F classes.
| Vps4 F1 | Vps4 F2 | Vps4 F3 | |
|---|---|---|---|
| Particle images | 6908 | 6794 | 7093 |
| Resolution (unmasked, Å) | 7.8 | 8.1 | 7.8 |
| Resolution (masked, Å) | 6.9 | 7.2 | 6.9 |
| EMDB ID | EMD-8572 | EMD-8571 | EMD-8570 |
Additional files
-
Supplementary file 1
Proteins and expression vectors.
- https://doi.org/10.7554/eLife.24487.032





