ER assembly of SNARE complexes mediating formation of partitioning membrane in Arabidopsis cytokinesis
Figures
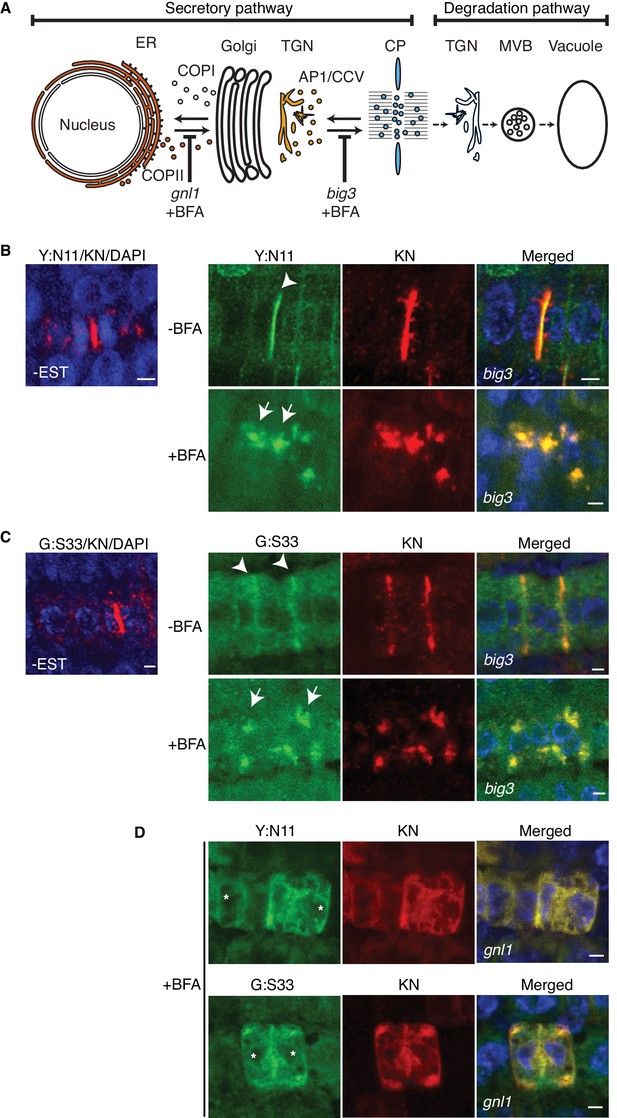
Site-specific inhibition of SNARE protein trafficking to the cell-division plane.
(A) Qa-SNARE KNOLLE trafficking route in cytokinesis (Reichardt et al., 2007). ER, endoplasmic reticulum; TGN, trans-Golgi network; CP, cell plate; MVB, multivesicular body; COPI, COPII, AP1/CCV, membrane vesicles with specific coat protein complexes; gnl1, big3, knockout mutations of ARF-GEFs rendering those trafficking steps sensitive to brefeldin A (BFA). (B–D) Subcellular localisation of estradiol-inducible YFP:NPSN11 (B, D; green) and GFP-SNAP33 (C, D; green), and KNOLLE (B–D; red) in big3 (B, C) and gnl1 GNL1BFA-sens. (D) mutant seedling roots treated with 50 µM BFA for 30 min, followed by 50 µM BFA + 20 µM estradiol for 210 min. Note that YFP:NPSN11 (Y:N11) or GFP:SNAP33 (G:S33) accumulates with KNOLLE (KN) at the BFA compartments in BFA-treated big3 mutant whereas YFP:NPSN11 or GFP:SNAP33 colocalises with KNOLLE at the ER in BFA-treated gnl1 GNL1BFA-sens. mutant. Note also no expression of YFP:NPSN11 (Y:N11, B) or GFP:SNAP33 (G:S33, C) without estradiol treatment. Nuclei of overlays (B–D) were counterstained with DAPI (blue). -BFA, mock treatment; +BFA, BFA treatment; -EST, no estradiol treatment. Arrowheads, cell plates; arrows, BFA compartments; asterisks, ER. Scale bar, 5 µm. The experiments were technically repeated three times.
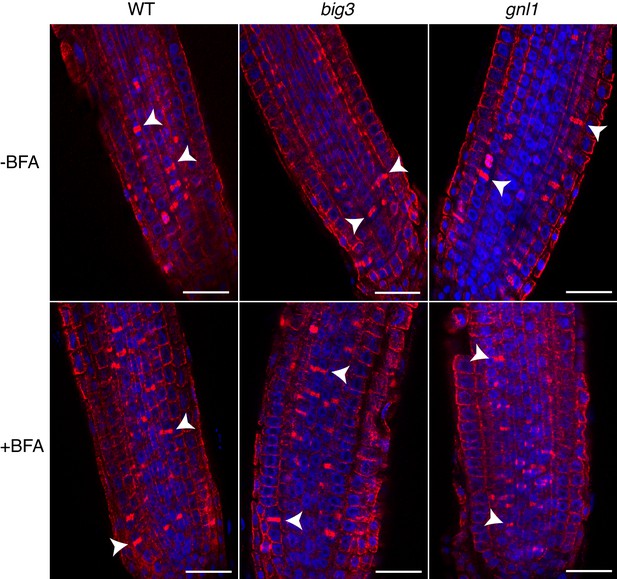
Cytokinetic cells in big3 and gnl1 GNL1BFA-sens. mutant seedling roots.
Mutant seedling roots were treated with 50 µM BFA for 30 min followed by 50 µM BFA + 20 µM estradiol for 210 min. Note that the population of cytokinetic cells was not altered by BFA treatment in mutant seedling roots, as evidenced by the anti-α-tubulin-labeled phragmoplast (red). See supplementary file 1 for quantification. Nuclei were counterstained with DAPI (blue). WT, wild-type; -BFA, mock treatment; +BFA, BFA treatment. Arrowhead, cell-plate with phragmoplast. Scale bar, 25 µm. The experiments were technically repeated twice.
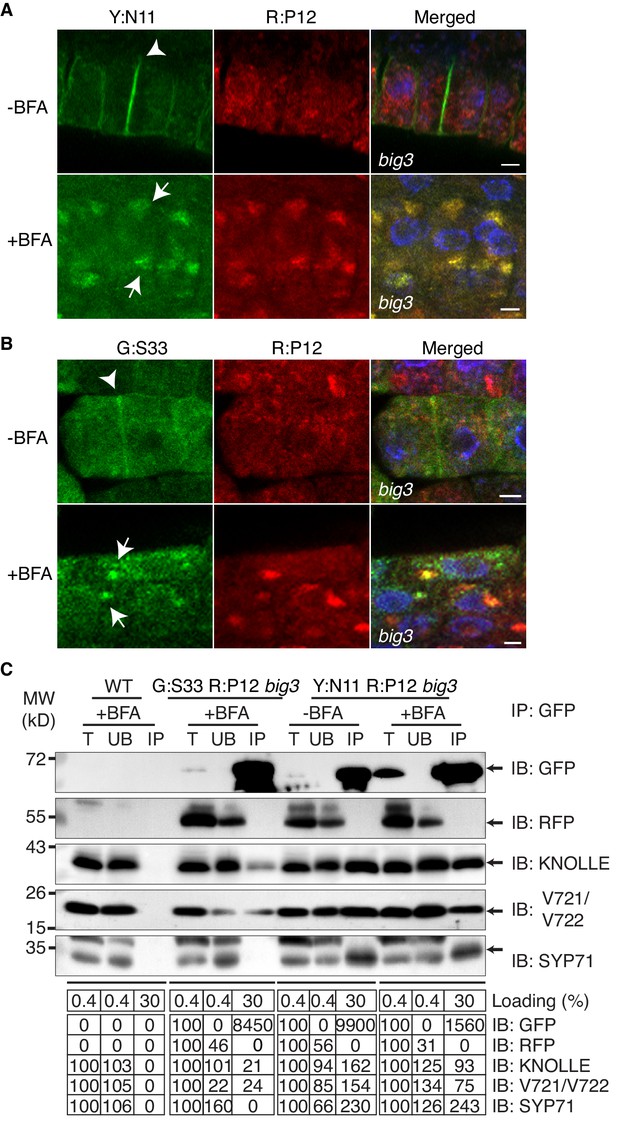
Subcellular localisation (A, B) and co-immunoprecipitation analysis (C) of pKNOLLE::mRFP:PEP12 (aka SYP21) (red) in big3 mutant seedling root cells expressing estradiol-inducible YFP:NPSN11 (A, C) and GFP:SNAP33 (B, C).
Transgenic seedlings were treated with 50 µM BFA for 30 min followed by 50 µM BFA + 20 µM estradiol for 210 min. Protein extracts were subjected to immunoprecipitation with anti-GFP beads, protein blots were probed with the antisera indicated on the right (IB): GFP, anti-GFP; RFP, anti-RFP; KN, anti-KNOLLE; V721/V722, anti-VAMP721/722; SYP71, anti-SYP71. G:S33, GFP:SNAP33; R:P12, mRFP:PEP12; Y:N11, YFP:NPSN11. kDa, protein size (left); MW, molecular weight; -BFA, mock treatment; +BFA, BFA treatment; T, total extract; UB, unbound; IP, immunoprecipitate. Loading (%), relative loading volume to total volume; relative signal intensity (input signal = 100% for UB and IP). Nuclei of merged images (A, B) were counterstained with DAPI (blue). Arrowheads, cell plates; arrows, BFA compartments. Scale bar, 5 µm. The experiments were technically repeated three times.
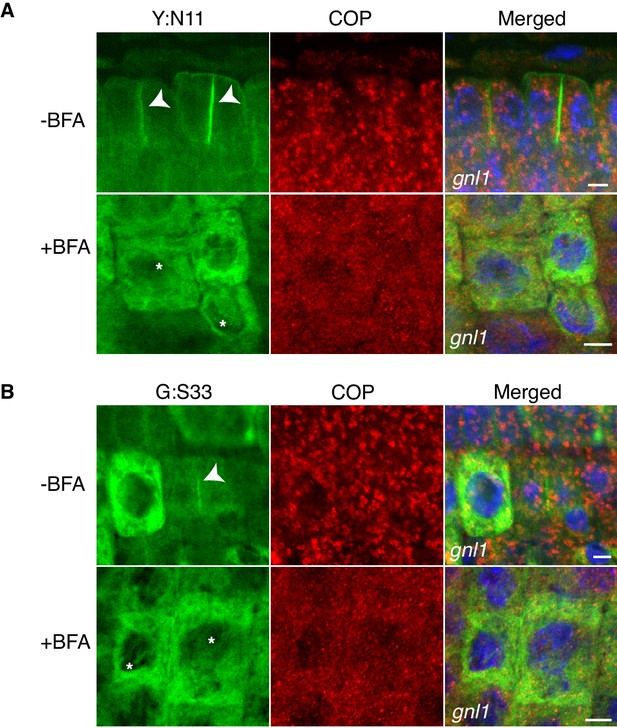
Site-specific inhibition of SNARE protein trafficking to the cell-division plane and loss of COPI from Golgi membrane in gnl1 GNL1BFA-sens. seedlings.
Mutant seedling roots were treated with 50 µM BFA for 30 min, followed by 50 µM BFA + 20 µM estradiol for 210 min. Note cytosolic localisation of γCOP in BFA-treated cells whereas YFP:NPSN11 (Y:N11, A) and GFP:SNAP33 (G:S33, B) in the same condition were trapped in the ER. Nuclei of merged images were counterstained with DAPI (blue). -BFA, mock treatment; +BFA, BFA treatment. Arrowheads, cell plates; asterisks, ER. Scale bar, 5 µm. The experiments were technically repeated three times.
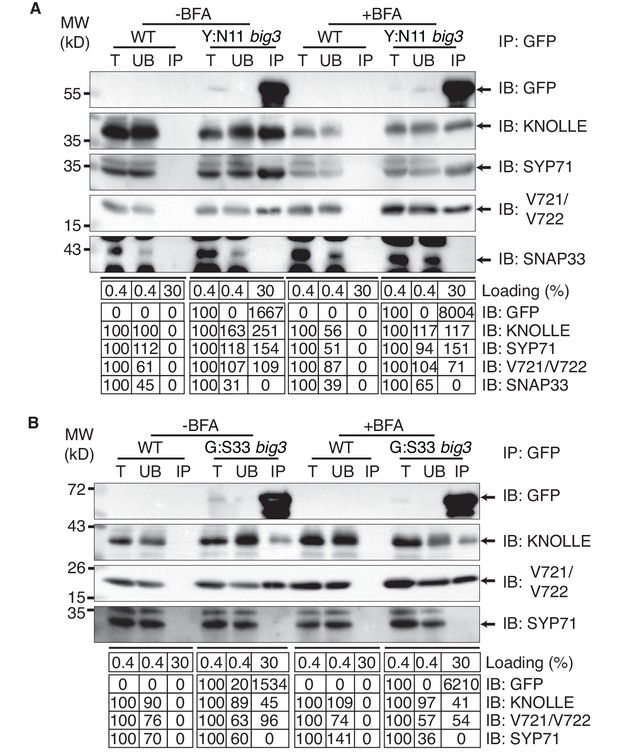
Interaction analysis of cytokinetic SNAREs with traffic blocked at the TGN.
Wild-type (WT) and big3 mutant seedlings carrying estradiol-inducible YFP:NPSN11 (A) or GFP:SNAP33 (B) transgenes were treated with 50 µM BFA for 30 min followed by 50 µM BFA + 20 µM estradiol for 210 min (see Figure 1B–C). Protein extracts were subjected to immunoprecipitation with anti-GFP beads, protein blots were probed with the antisera indicated on the right (IB): GFP, anti-GFP; KN, anti-KNOLLE; V721/V722, anti-VAMP721/722; SYP71, anti-SYP71; SNAP33, anti-SNAP33; kDa, protein size (left); MW, molecular weight; -BFA, mock treatment; +BFA, BFA treatment; T, total extract; UB, unbound; IP, immunoprecipitate. Loading (%), relative loading volume to total volume; relative signal intensity (input signal = 100% for UB and IP). The experiments were technically repeated more than six times.
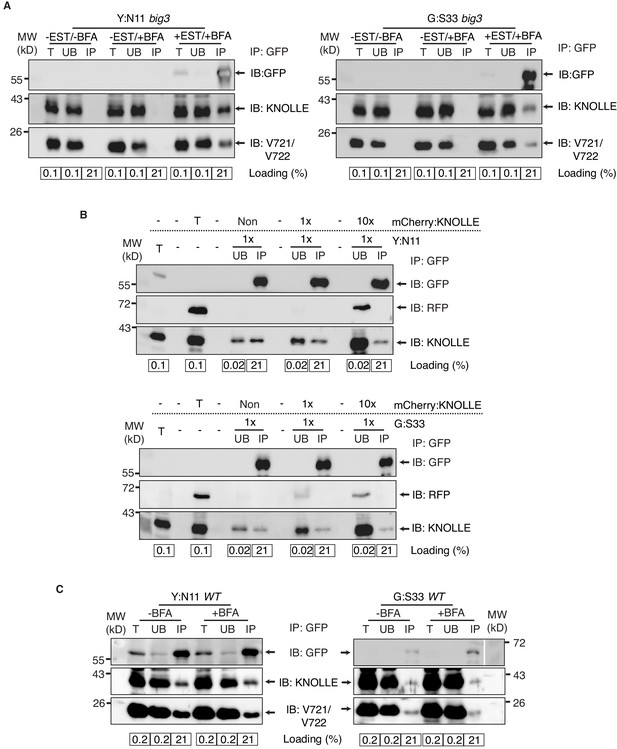
Control experiments for co-immunoprecipitation analysis of cytokinetic SNAREs.
(A) No EST-induction of expression of KNOLLE SNARE partners. big3 mutant seedlings carrying YFP:NPSN11 (left) or GFP:SNAP33 (right) transgenes were treated or not treated with 50 µM BFA for 240 min. Seedlings treated with 20 µM estradiol and 50 µM BFA for 210 min were used as positive control. Protein extracts were subjected to immunoprecipitation with anti-GFP beads, protein blots were probed with the antisera indicated on the right (IB). (B) In vitro mixing of experimental extract with different amounts of extract from tagged KNOLLE seedlings. Cleared protein extracts of YFP:N11 or GFP:SNAP33 and the cleared protein extracts of mCherry:KNOLLE were mixed and subjected to immunoprecipitation with anti-GFP beads. Protein blots were probed with the antisera indicated on the right (IB). Note that the addition of mCherry:KNOLLE protein does not alter the amount of endogenous KNOLLE co-immunoprecipitated with YFP:N11 or GFP:SNAP33. Non, no addition of mCherry:KNOLLE protein lysate; 1x, equal amounts; 10x, 10x excess. (C) Seedlings of wild-type (WT) background carrying YFP:NPSN11 or GFP:SNAP33 transgenes were treated with 50 µM BFA + 20 µM estradiol for 210 min. Protein extracts were subjected to immunoprecipitation with anti-GFP beads, protein blots were probed with the antisera indicated on the right (IB). GFP, anti-GFP; RFP, anti-RFP; KN, anti-KNOLLE; V721/V722, anti-VAMP721/722; kDa, protein size (left); MW, molecular weight; -EST, no estradiol; -BFA, mock treatment; +BFA, BFA treatment; T, total extract; UB, unbound; IP, immunoprecipitate; Loading (%), relative loading volume to total volume; relative signal intensity (input signal = 100% for UB and IP).
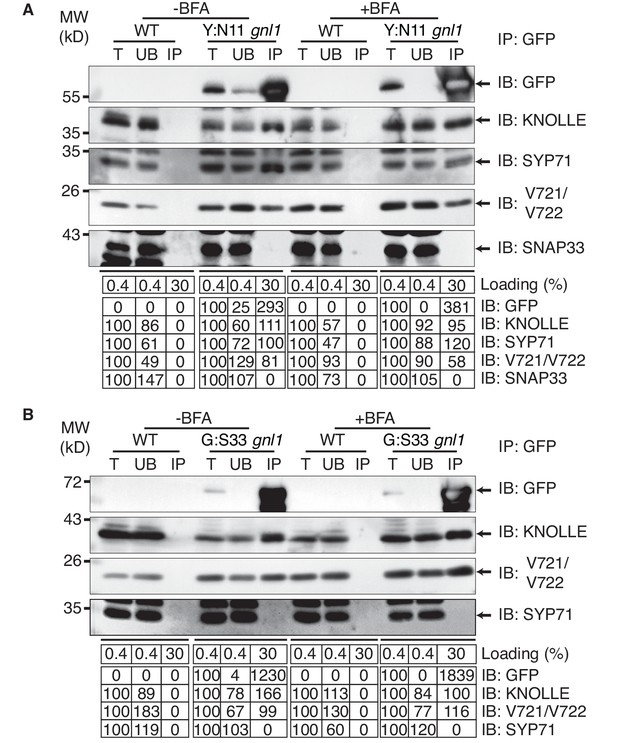
Interaction analysis of cytokinetic SNAREs with traffic blocked at the ER.
Wild-type (WT) and gnl1 mutant seedlings complemented with GNL1BFA-sens. encoding a BFA-sensitive variant of GNL1 and carrying estradiol-inducible YFP:NPSN11 (A) or GFP:SNAP33 (B) transgenes were treated with 50 µM BFA for 30 min followed by 50 µM BFA + 20 µM estradiol for 210 min (see Figure 1D). Protein extracts were subjected to immunoprecipitation with anti-GFP beads, protein blots were probed with the antisera indicated on the right (IB): GFP, anti-GFP; KN, anti-KNOLLE; V721/V722, anti-VAMP721/722; SYP71, anti-SYP71; SNAP33, anti-SNAP33; kDa, protein size (left); MW, molecular weight; -BFA, mock treatment; +BFA, BFA treatment; T, total extract; UB, unbound; IP, immunoprecipitate. Loading (%), relative loading volume to total volume; relative signal intensity (input signal = 100% for UB and IP). The experiments were technically repeated more than six times.
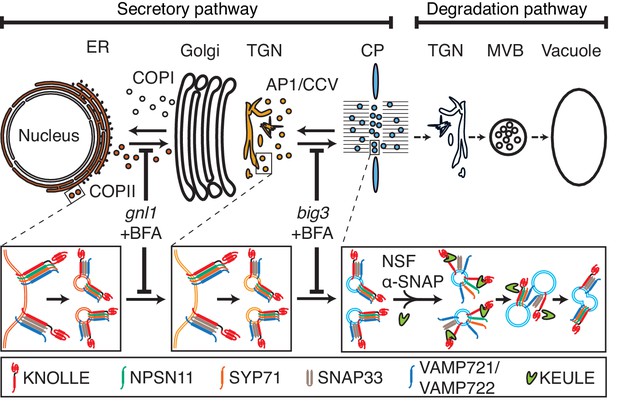
Trafficking of cis-SNARE complexes during cytokinesis (model).
Two different types of cytokinetic cis-SNARE complexes are assembled on the ER, recruited into COPII vesicles and passed on to the Golgi stack/TGN. At the TGN, they are incorporated into AP1/CCV vesicles for delivery to the division plane. Following their disassembly by NSF ATPase, monomeric Qa-SNARE KNOLLE is assisted by SM protein KEULE in the formation of trans-SNARE complexes mediating fusion of adjacent vesicles during cell-plate formation and expansion (Park et al., 2012).
Additional files
-
Supplementary file 1
(a) Frequency of cytokinetic cells in mutant seedling roots.
(b) Primers used in this study
- https://doi.org/10.7554/eLife.25327.010





