Mechanism of environmentally driven conformational changes that modulate H-NS DNA-bridging activity
Figures
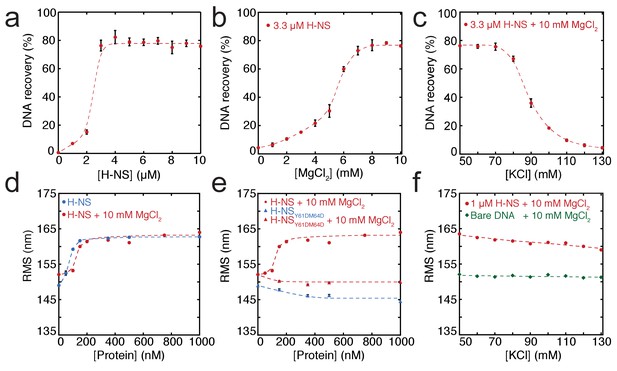
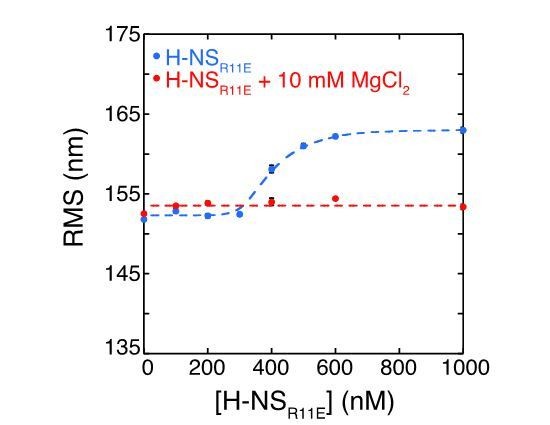
Modulation of H-NS function by ionic conditions.
(a) DNA-bridging efficiency as a function of H-NS concentration in the presence of 10 mM MgCl2. (b) DNA-bridging efficiency as a function of MgCl2 concentration. (c) DNA-bridging efficiency as a function of the KCl concentration. (d) Root Mean Square displacement (RMS) as a function of H-NS concentration, in the presence and absence of 10 mM MgCl2 (N > 70, per data point). (e) RMS of DNA as a function of H-NSY61DM64D in the presence and absence of MgCl2 (N > 70, for each point). (f) Extension of DNA as a function of the KCl concentration (N > 70, for each point). Error bars indicate standard deviation. Dashed lines are to guide the eye.
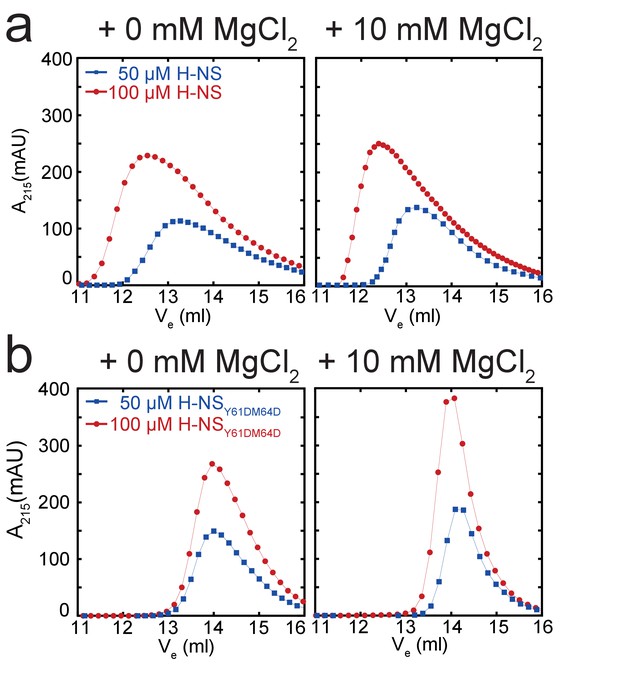
Multimeric state of H-NS measured using size exclusion chromatography.
(a) Elution pattern of 50 and 100 µM of H-NS as a function of Mg2+. (b) Elution pattern of 50 and 100 µM of H-NSY61DM64D as a function of Mg2+. Lines are to guide the eye.
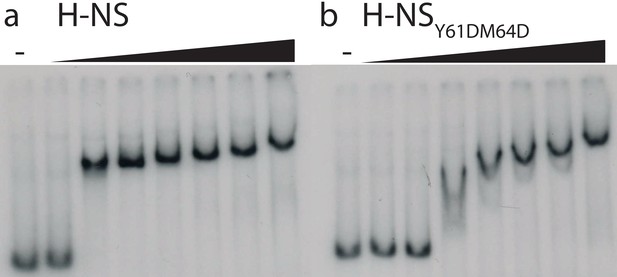
Electrophoretic Mobility shift assay.
(a) H-NS and (b) H-NSY61DM64D. The proteins were added to 32P-labeled 685 bp DNA substrate at 0.4, 0.8, 1.2, 1.6, 12.4, 3.3, 6.6 µM concentrations. H-NS and H-NSY61DM64D exhibit similar DNA-binding affinity. The difference in mobility shift pattern is attributed to a difference in cooperativity of DNA binding between the two H-NS variants.
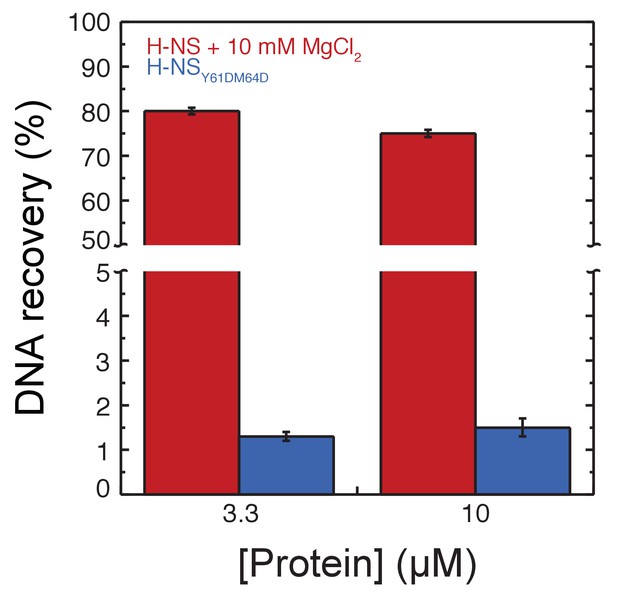
DNA recovery of H-NS and.
H-NSY61DM64D in the DNA-bridging assay.
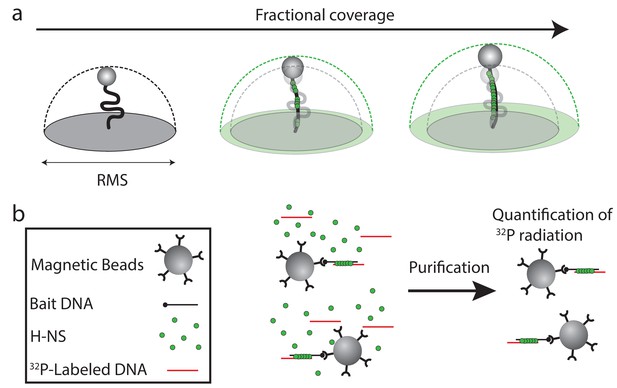
Schematic depiction of techniques used in this study.
(a) Schematic depiction of TPM and the effect of increased protein binding to DNA on the Root Mean Squared (RMS) motion of the bead. (b) Schematic depiction of the DNA-bridging assay. Streptavidin-coated paramagnetic beads are coupled to a bait DNA, 5’ labeled with biotin. The beads bound with DNA are then incubated in the presence of 32P-radiolabeled DNA strand and H-NS. Next, using a magnetic rack, the beads are pulled down and the amount of recovered prey DNA is quantified.
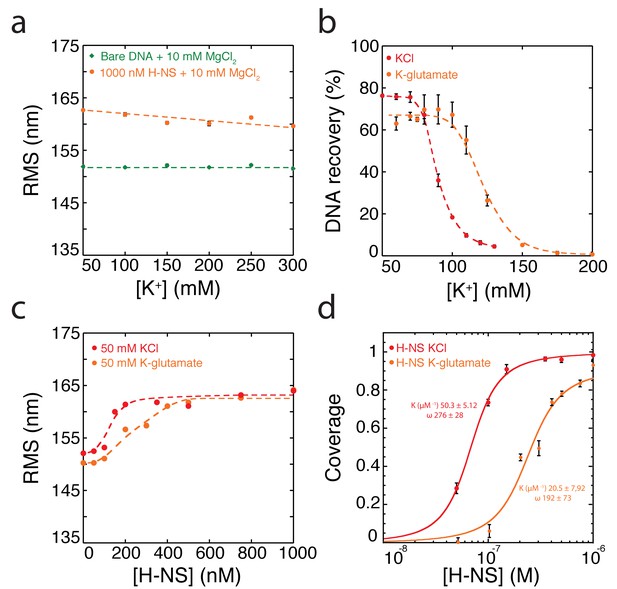
Modulation of H-NS by alternative anions.
(a) Extension of DNA as a function of the K-glutamate concentration (N > 70, for each point). (b) DNA-bridging efficiency as a function of the K-glutamate concentration. Error bars indicate standard deviation. (c) RMS as a function of H-NS concentration, in the presence of KCl and K-glutamate (N > 70, for each data point) (d) McGhee-von Hippel analysis of H-NS DNA-binding curves based on TPM data. Binding curves of H-NS in the presence of KCl and K-glutamate. Fit variables for all datasets, the binding affinity (K), and cooperativity (ω). Dashed lines are to guide the eye and solid lines are fit curves.
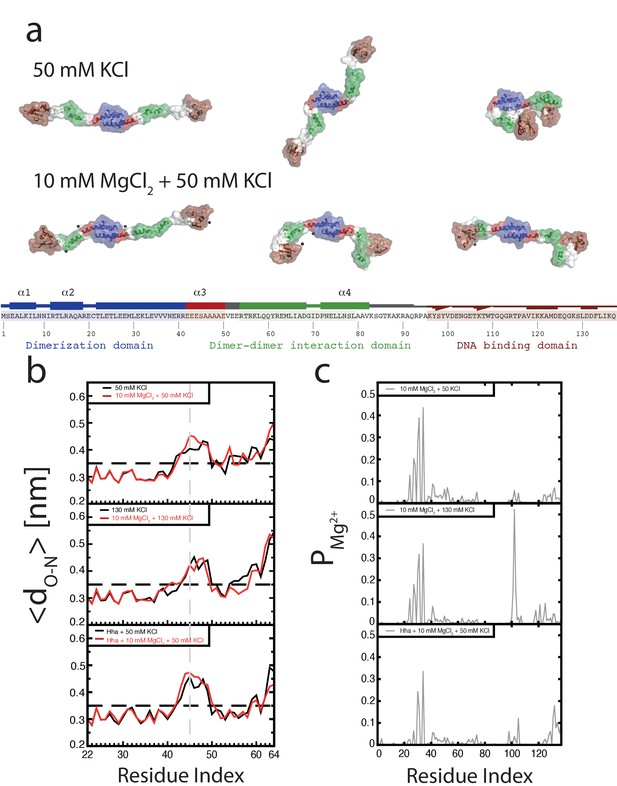
Conformation of the H-NS dimer as a function of osmolarity.
(a) Snapshots depicting representative conformations of H-NS in the simulations with 50 mM KCl (top) and 10 mM MgCl2 +50 mM KCl (bottom) (For the full movies depicting these effects see Figure 2—videos 1 and 2, for other examples of the ‘closed’ conformations of H-NS see Figure 2—figure supplement 1). The buckle region is highlighted in red and the domains are colored blue, green and brown for the dimerization domain, the dimer-dimer interface and the DNA-binding domain, respectively. The conserved motif involved in DNA binding is highlighted as ball-stick models. Mg2+ ions are shown as dark gray orbs. The protein is shown in ribbon representation with a transparent surface. The atomic radii in the protein were set to 3 A to smooth the surface. (b) Location of buckle in helix α3. The average distance dO-N between donor and acceptor in the helical hydrogen bond in helix α3 is plotted as a function of the residue index of the acceptor. The dashed black line in the graphs indicates the distance threshold for forming a hydrogen bond. Time traces of these distances are given in Figure 2—figure supplement 3. (c) Location of Mg2+ on H-NS. The probability of finding Mg2+ ions within 0.6 nm of an H-NS residue, PMg2+, is plotted as function of the residue index for the three systems containing Mg2+.
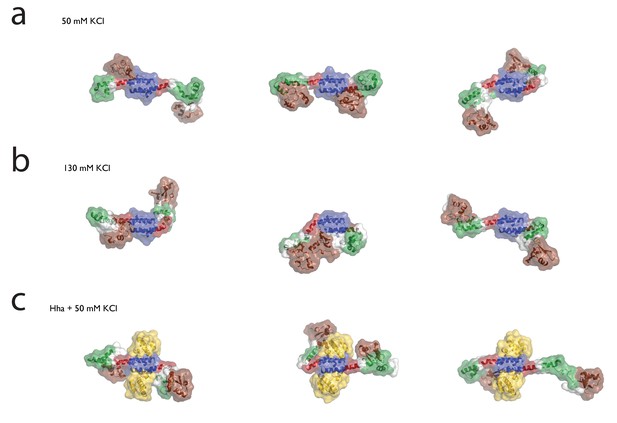
Examples of ‘closed’ H-NS conformations in the presence of (a) 50 mM KCl, (b) 130 mM KCl, or (c) Hha.
https://doi.org/10.7554/eLife.27369.010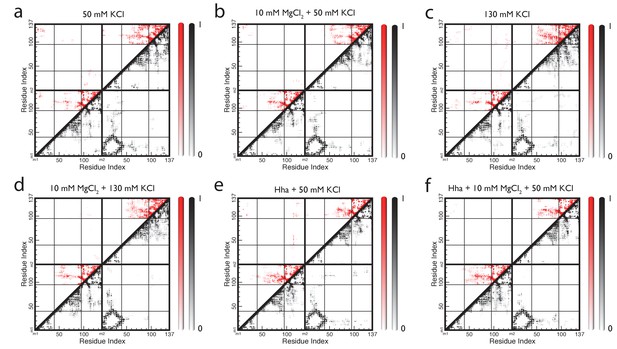
Contact maps of full-length H-NS dimers simulations in different conditions.
(a) 50 mM KCl, (b) 10 mM MgCl2 +50 mM KCl, (c) 130 mM KCl, (d) 10 mM MgCl2 +130 mM KCl, (e) Hha +50 mM KCl, (f) Hha +10 mM MgCl2 +50 mM KCl, Residues are considered in contact if the smallest distance between two atoms is 0.6 nm or less. The maps show the probability of finding a contact between residues, averaged over the last 40 ns of 8 molecular dynamics runs per system. White represents a probability of zero, darker colors indicates higher probability. The color scale runs from 0 to 1. Contacts with the DNA-binding domain are highlighted in red.
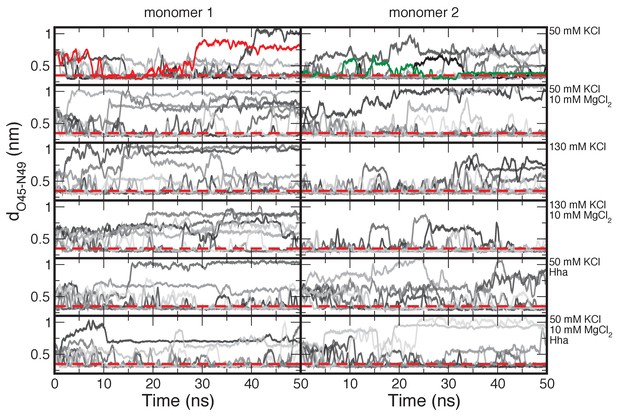
Time traces of the O-H distance between residues 45 and 49.
The time traces are shown in different shades of gray to indicate the different runs. The red dashed line indicates the distance threshold for forming a hydrogen bond. The red and green lines are examples of the formation of irreversible and reversible buckles respectively. The time traces are shown as a running average of 50 ps to reduce the noise from thermal fluctuations.
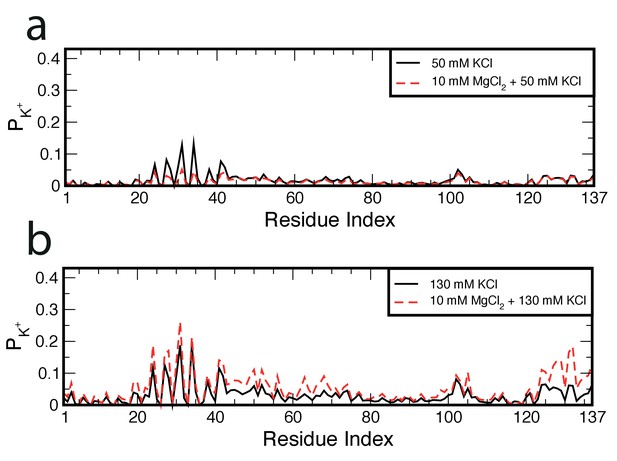
Location of K+on hr-NS.
The probability of finding K+ ions within 0.6 nm of an H-NS residue, PK+, is plotted as function of the residue index for (a) 50 mM KCl and (b) 130 mM KCl in the presence and absence of MgCl2.
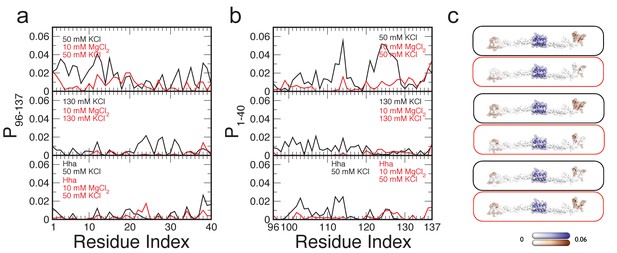
Contact maps of H-NS dimers in different conditions, focused on the interactions between the dimerization domain and the DNA-binding domain.
(a) The probability of finding the DNA-binding domain within 0.6 nm of residues in the dimerization domain, P96-137 (b) The probability of finding the dimerization domain within 0.6 nm of residues in the DNA-binding domain, P1-40. Residues are considered in contact if the smallest distance between two atoms is 0.6 nm or less. (c) P96-137 and P1-40 are highlighted on a three-dimensional structural model of H-NS. The protein dimer is shown in ribbon representation and transparent surface representation, with white indicating no interactions, blue indicating values for P96-137 and brown indicating P96-137. The color scale runs from 0 to 0.06.
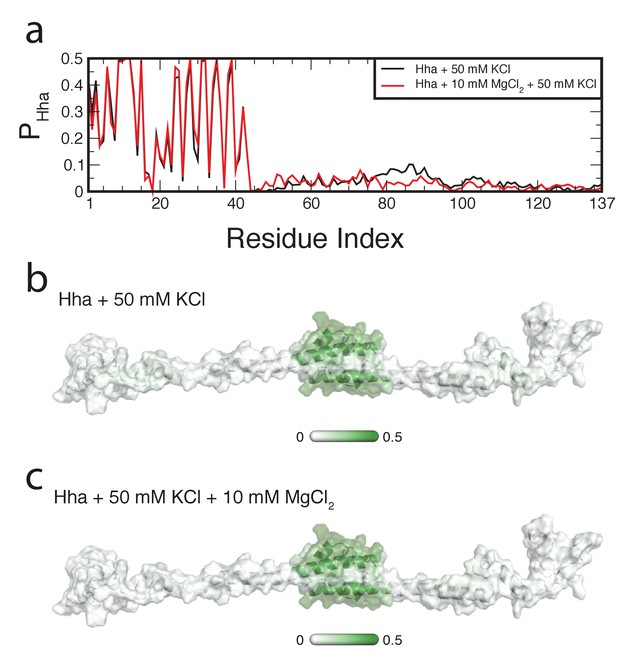
Location of Hha on H-NS.
(a) The probability of finding Hha within 0.6 nm of an H-NS residue, PHha, is plotted as a function of the residue index. The PHha is indicated on a surface representation of an H-NS dimer in open conformation in the (b) absence and (c) presence of MgCl2, ranging from 0 (white) to 0.5 (dark green).
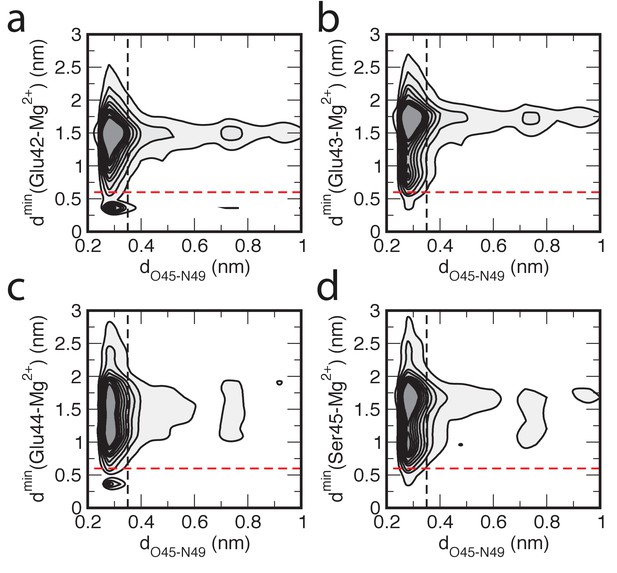
Correlation between hydrogen bond distance and proximity of Mg2+to the buckle in helix α3.
The correlation between the hydrogen bond distance dO45-N49 and the minimum distance between Mg2+ ions and a residue dmin is shown as a contour plot. The darker colors indicate higher probabilities of finding that particular configuration. Contours are drawn for every 0.2%. A distance larger than 0.35 nm indicates a broken hydrogen bond (black dashed line). A Mg2+ ion is close if this distance is less than 0.6 nm (red dashed line). The panels show the contour plots for four residues: (a) Glu42, (b) Glu43, (c) Glu44 and (d) Ser45.
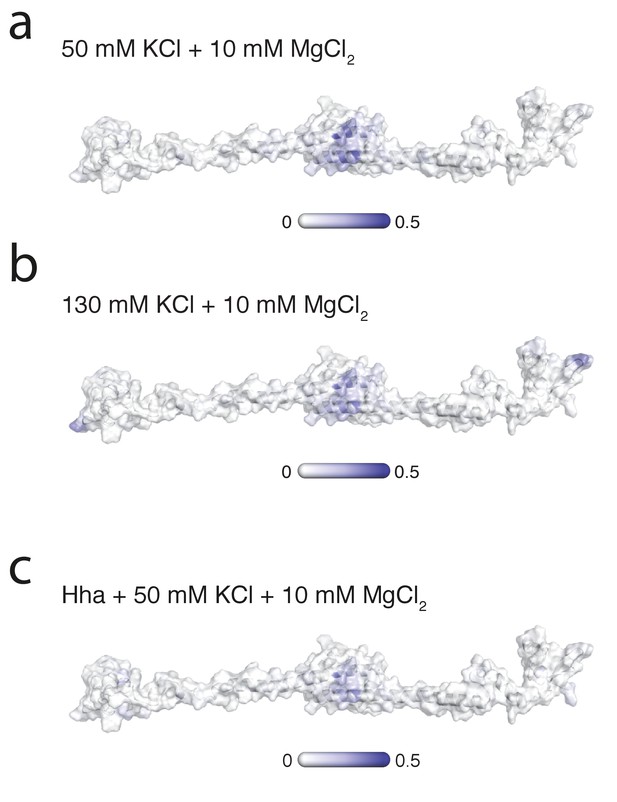
Mg2+localization on H-NS.
The probability of finding Mg2+ within 0.6 nm of H-NS in the presence of (a) 50 mM KCl, (b) 130 mM KCl, or (c) Hha. This probability is indicated on a surface representation of an H-NS dimer in open conformation, ranging from 0 (white) to 0.5 (dark blue).
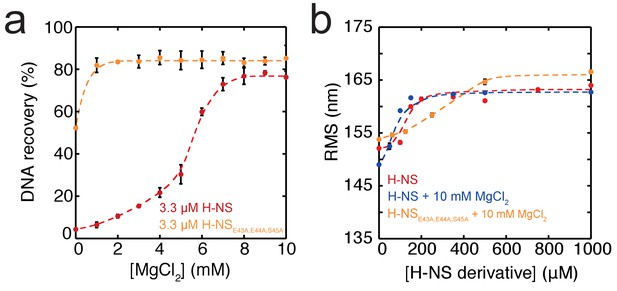
Function of the H-NS derivative, H-NSE43A,E44A,S45A.
(a) DNA-bridging efficiency of H-NSE43A,E44A,S45A as a function of the Mg2+ concentration. (b) Root Mean Square displacement (RMS) of DNA in the presence of H-NSE43A,E44A,S45A.
Conformational flexibility of H-NS.
The protein is rendered in cartoon representation with a transparent surface. The color code indicates the domain organization of H-NS: blue - site 1 (residue 1–40), red - the buckle region in helix 3 (residue 42–49), green - site2 (residue 55–83) and brown - the DNA-binding domain (residue 96–137). The DNA-binding site on H-NS is shown in sticks. This movie contains snapshots at every 100 ps from run5 of the MD simulations of H-NS at 50 mM KCl (see Figure 2—figure supplement 3 for time traces of the helical hydrogen bond distance between residue 45 and 49). The snapshots are rendered with PyMol.
The effect of magnesium on the conformational flexibility of H-NS.
The protein is rendered in cartoon representation with a transparent surface. The color code indicates the domain organization of H-NS: blue - site 1 (residue 1–40), red - the buckle region in helix 3 (residue 42–49), green - site2 (residue 55–83) and brown - the DNA-binding domain (residue 96–137). The DNA-binding site on H-NS is shown in sticks. This movie contains snapshots at every 100 ps from run7 of the MD simulations of H-NS at 50 mM KCl +10 mM MgCl2 (see Figure 2—figure supplement 3 for time traces of the helical hydrogen bond distance between residue 45 and 49 and the minimum distance between residue 45 and the magnesium ions). The snapshots are rendered with PyMol.
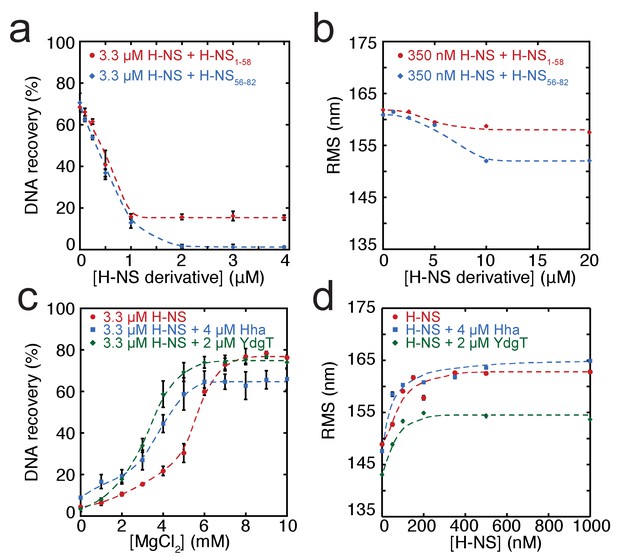
Modulation of H-NS function by protein cofactors.
(a) DNA bridging efficiency as a function of inhibiting peptides targeting either the dimerization domain (H-NS1-58) and multimerization domain (H-NS56-82). (b) Root Mean Square displacement (RMS) as a function of inhibiting peptides targeting either the dimerization (H-NS1-58) and multimerization (H-NS56-82) (N > 70, for each point). (c) DNA bridging efficiency as a function of Mg2+ concentration in the presence and absence of 4 µM Hha or 2 µM YdgT. (d) RMS of DNA in the presence of H-NS, H-NS-Hha, and H-NS-YdgT. Dashed lines are to guide the eye (N > 60, for each point).
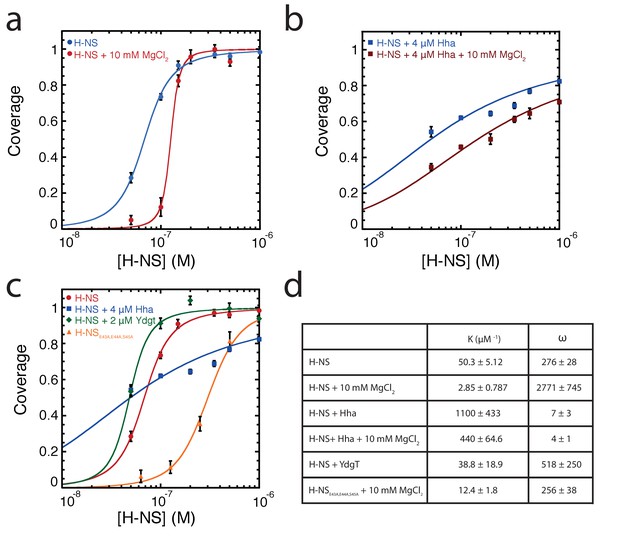
McGhee-von Hippel analysis of H-NS DNA binding curves based on TPM data.
Binding curves of (a) H-NS, (b) H-NS + Hha TPM data as a function of Mg2+, and (c) H-NS, H-NS + Hha, H-NS + YdgT, and H-NSE43A,E44A,S45A (d) Fit variables for all datasets, the binding affinity (K), and cooperativity (ω). Fit curves are shown as solid lines.
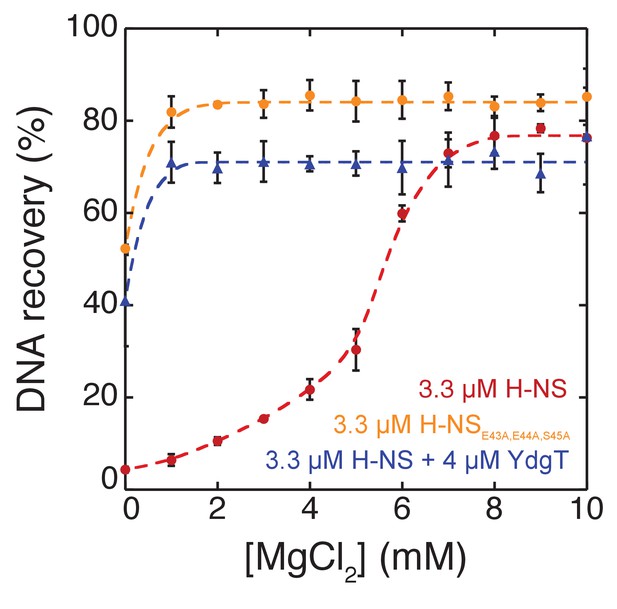
DNA-bridging efficiency of H-NS (red), H-NS + 4 µM YdgT (blue) and H-NSE43A,E44A,S45A (orange) as a function of MgCl2 concentration.
https://doi.org/10.7554/eLife.27369.023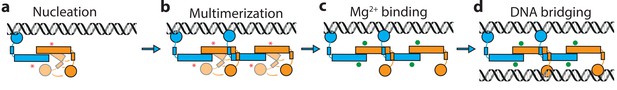
Model of H-NS complex assembly.
(a) H-NS nucleates at preferred DNA sequences in the genome. (b) H-NS laterally multimerizes laterally along the DNA in the ‘closed’ conformation. (c) In the presence of Mg2+ or other H-NS modulators such as Hha, H-NS switches to the ‘open’, bridging capable conformation. (d) H-NS forms DNA bridges in trans. The red asterisk indicates the buckle location. Mg2+ ions are shown as green orbs.
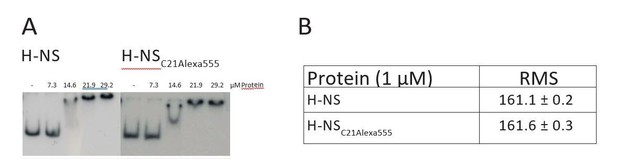
DNA binding and activity of H-NS and H-NSC21Alexa555.
(A) Electrophoretic mobility shift assay using a 32P labeled (AT-rich) curved DNA substrate as described in Dame et al., Bioch., 2001. (B) The root mean squared (RMS) displacement of a DNA tether bound by H-NS and H-NSC21Alexa555 as measured by TPM.
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.27369.025