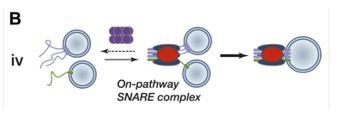
Sec17 (α-SNAP) and an SM-tethering complex regulate the outcome of SNARE zippering in vitro and in vivo
Figures
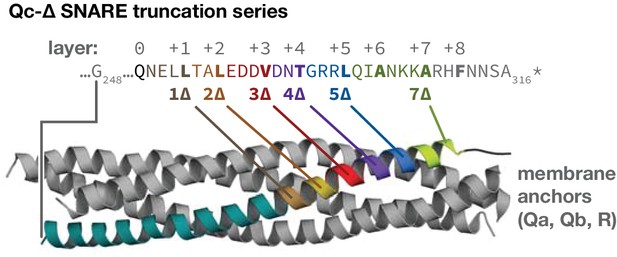
The set of Qc-∆ truncation mutants, mapped onto the structure of a quaternary SNARE bundle.
The C-terminus of each Qc-∆ is indicated with boldface type, except for Qc1∆, which contains one additional aminoacyl residue that remains after cleavage of the C-terminal affinity tag used for purification. The transmembrane anchors of the Qa, Qb, and R-SNAREs are not depicted.
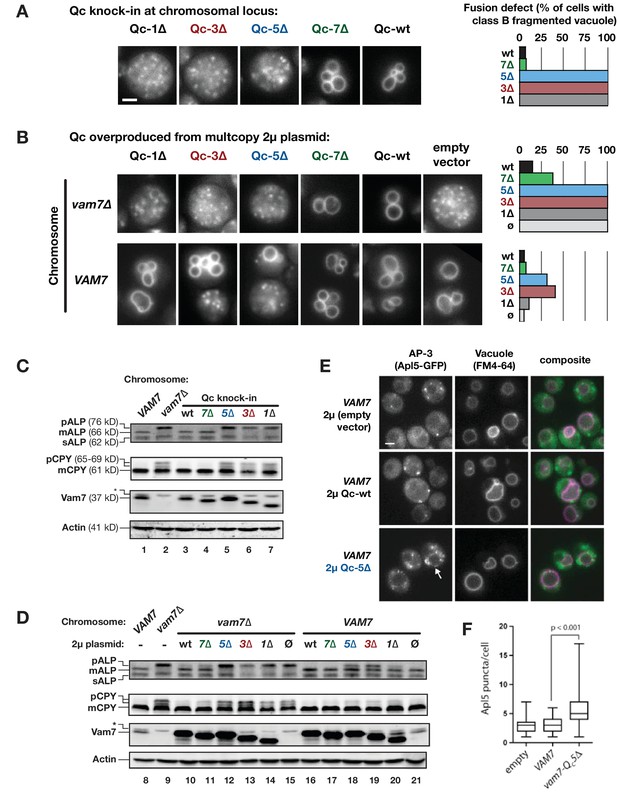
Characterization of Qc-∆ zippering mutants in vivo.
(A and B) Vacuoles were labeled by pulse-chase loading with the styryl dye FM4-64, and observed by wide-field epifluorescence microscopy. Defects in vacuole morphology are quantified in the graphs to the right. 106–411 cells of each genotype were scored in at least two independent experiments. (C and D) Cargo trafficking defects of Qc-∆ mutants. Cell lysates were prepared and separated by SDS-PAGE, then analyzed by immunoblot using polyclonal antibodies against Vam7 (Qc), or monoclonal antibodies against ALP or CPY. Polyclonal anti-actin was used for the loading controls. The slower-than-expected migration of Qc-5∆ was also observed with recombinant Qc-5∆ purified from E. coli cells. A non-specific band in the Vam seven blots was present in lysates from all strains including vam7∆ null mutants and is indicated by (*). (E and F) Overproduction of Qc-5∆ causes dominant partial accumulation of AP-3 vesicles. In some cells, the vacuole is not fragmented, and AP-3 vesicles (Apl5-GFP punctae) accumulate at the vacuole limiting membrane, as shown in panel E. In panel F, AP-3 vesicles are quantified (Mann-Witney U test; n = 100 cell profiles per strain in two independent experiments). Scale bars (A,B,E) indicate 2 µm.
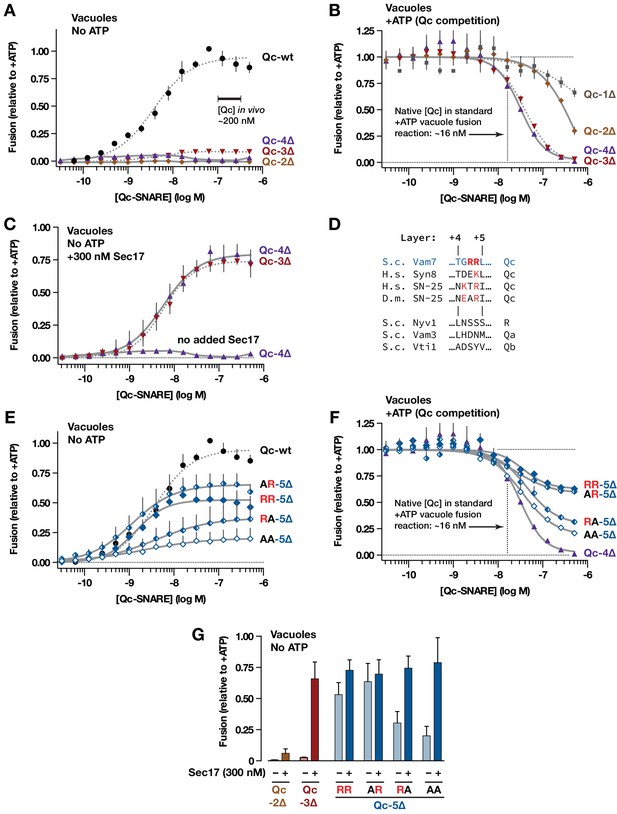
Interplay of SNARE zippering and Sec17 in cell-free assays of homotypic vacuole fusion.
The content-mixing assay and reaction schemes are diagrammed in Figure 3—figure supplement 1. Curves in the dose-response and dose-inhibition experiments are nonlinear fits of the Hill equation. Dashed lines denote data re-plotted from Schwartz and Merz (2009) to facilitate comparison. For all panels, points and bars denote the mean (+or ± s.e.m.) of ≥3 independent experiments. (A) Recombinant Qc-2∆, Qc-3∆, and Qc-4∆ proteins are nonfusogenic in no-ATP ‘bypass’ gain-of-function assays. In these assays no ATP, Sec17, or Sec18 are added to the vacuoles (as shown in Figure 3—figure supplement 1, reaction ii). The approximate concentration of endogenous cytoplasmic Vam7 in vivo is indicated (Thorngren et al., 2004). (B) Qc-3∆ and Qc-4∆ are efficient competitive inhibitors of native Qc-wt. In these ATP-containing reactions, endogenous Sec17 and Sec18 are active and fusion is driven by native Qc-wt (Vam7) liberated from cis-SNARE complexes on isolated vacuoles, as diagrammed in Figure 3—figure supplement 1, reaction i. The approximate concentration of native Vam7 in a standard +ATP vacuole fusion reaction is indicated (~16 nM; Thorngren et al., 2004). (C) Added Sec17 restores fusion activity to Qc-3∆ and −4∆ in no-ATP ‘bypass’ assays. The reactions were set up as in panel A, except that the reactions were supplemented with 300 nM Sec17. (D) Sequence alignment of SNAREs in the layer +4 to+5 region. S.c., Saccharomyces cerevisiae; H.s., Homo sapiens; D.m., Drosophila melanogaster. A conserved arginyl (R) residue is indicated in red. (E) Ability of Qc-5∆ variants to promote fusion in no-ATP ‘bypass’ assays. The reactions were set up as in panel A. (F) Competitive inhibition of fusion by Qc-5∆ variants. The reactions were set up as in panel B. (G) Sec17 rescue in gain of function assays with Qc-2∆, Qc-3∆, and Qc-5∆ variants. The no-ATP ‘bypass’ reactions were set up as in panel A, except that the Qc-∆ proteins were always used at 100 nM, and a subset of the reactions were supplemented with 300 nM Sec17, as indicated.
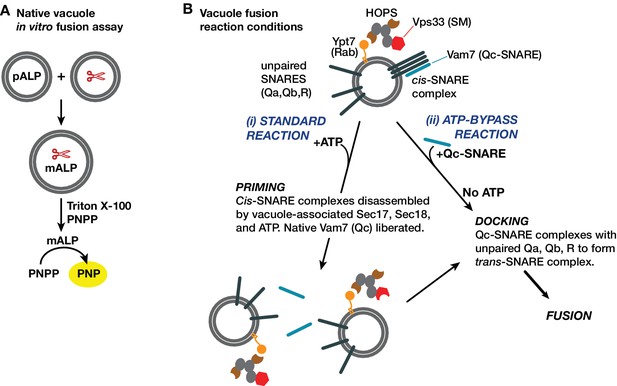
Yeast vacuole fusion assay and reaction schemes used in this study.
(A) Content mixing assay system. One set of vacuoles is isolated from a yeast strain containing the inactive vacuolar hydrolase proALP. The second set of vacuoles is isolated from a strain containing the maturases PrA and PrB (denoted by scissors). Content mixing causes proteolysis of proALP to mature mALP. The amount of mALP is assayed by lysis of the vacuole and addition of the colorimetric substrate para-nitrophenolphosphate, which is hydrolyzed to a yellow product by mALP. The formation of 4-nitrophenolate (PNP) is measured by spectrophotometry. (B) Fusion assay reaction configurations used in this study. The isolated vacuole bears active Ypt7-GTP, the SM-tether complex HOPS, and small amounts of both Sec17 and Sec18. SNAREs are present in two forms: unpaired SNAREs, and cis-SNARE complexes. In purified vacuole preparations, the Qc Vam7 is present only in the cis-SNARE complexes (Boeddinghaus et al., 2002). Thus, there are two ways to drive fusion of isolated vacuoles. In the standard ‘+ATP’ reaction (i), ATP and Sec18 liberate SNAREs including native Vam7 from cis-complexes, leading to docking and fusion (Boeddinghaus et al., 2002; Mayer et al., 1996). In the ‘ATP bypass’ reaction (ii), purified Vam7 (or Qc-∆) is supplied in the absence of ATP, allowing docking and fusion to proceed.
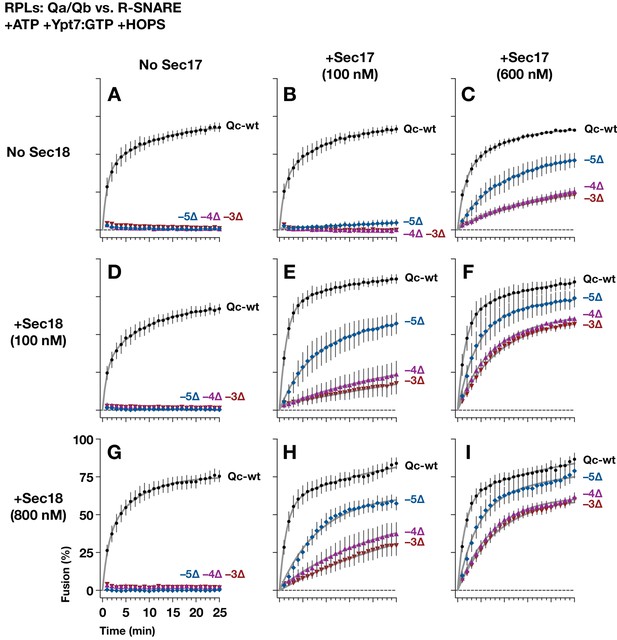
Sec17 allows partially-zipped SNARE complexes to drive fusion with or without Sec18.
The chemically defined RPL fusion system is diagrammed in Figure 4—figure supplement 1. RPLs bearing Ypt7-GTP (Rab), and either the Qa- and Qb-SNARES, or the R-SNARE, were incubated with HOPS (100 nM) and the indicated Qc-SNAREs (250 nM). Reactions were performed in the absence or presence of Sec17 and Sec18, as indicated. 1 mM ATP and HOPS were present under all conditions. The Rab Ypt7 was present on both vesicle populations and was loaded with GTP (Materials and methods). On the vertical axes, 100% fusion indicates complete association of the FRET probes encapsulated within the two vesicle populations, as determined in control reactions. Each data point shows the mean content mixing signal ± s.e.m. for three independent experiments. The lines show nonlinear best-fits of a second-order kinetic model.
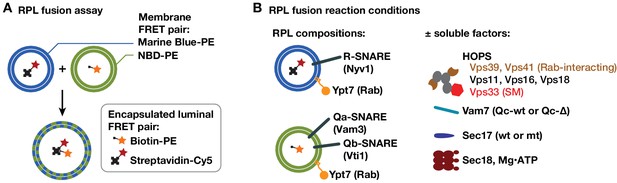
RPL fusion assay system.
(A) The RPL fusion assay simultaneously monitors lipid and content mixing using orthogonal FRET pairs (Zucchi and Zick, 2011). Lipid mixing is monitored by FRET between phosphatidylethanolamine (PE) lipid derivitized with either NBD or Marine Blue. Content mixing is monitored through the association of encapsulated biotin and avidin conjugates (to phycoerythrin and Cy5 fluorophores, respectively). Although both lipid and content mixing signals were collected in the experiments shown, only content mixing signals are presented because the two signals were highly correlated, and because content mixing is the reaction's biologically relevant endpoint. (B) The RPLs used in this study contained a lipid mixture approximating that of the yeast vacuole membrane (Zick et al., 2014), native purified Ypt7-GTP, and either Qa and Qb, or R-SNAREs. The asymmetric SNARE topology allows docking and fusion without any prior requirement for Sec18-mediated cis-SNARE complex disassembly.
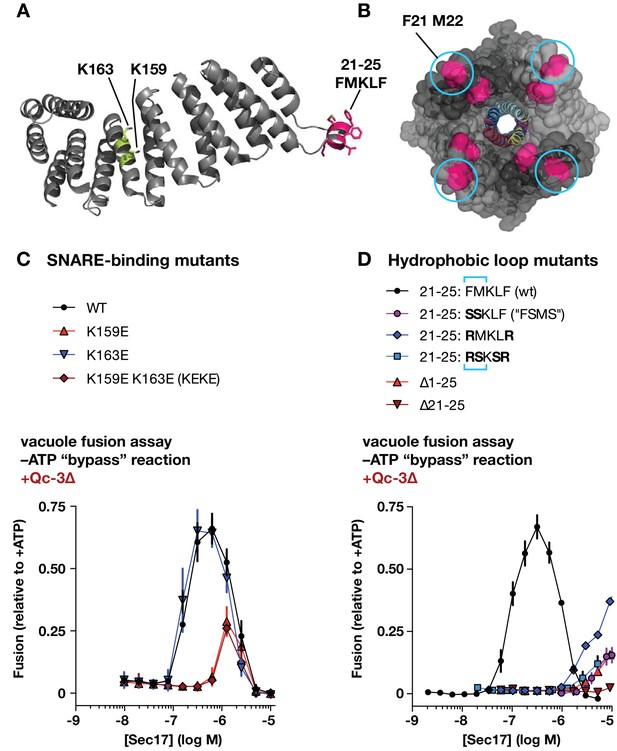
Effects of Sec17 mutations on stimulation of fusion.
(A) Locations of Sec17 mutations. The diagram is a rendering of PDB 1QQE (Rice and Brunger, 1999). The N-terminal hydrophobic loop is shaded magenta. Two highly conserved lysine residues are shaded green. (B) Position of Sec17 hydrophobic loop relative to SNARE core complex. Rendering shows a SNARE complex with four bound α-SNAP molecules, from the perspective of the membrane-proxiimal SNARE domain C-termini. Hydrophobic loop residues are colored magenta. The two residues homologous to those mutated in Sec17-FSMS are circled. The rendering is based on PDB 3J96 (Zhao et al., 2015). (C and D) Ability of Sec17 mutants to rescue Qc-3∆ trans-SNARE complexes in vitro. Vacuole fusion reactions were assembled in the gain-of-function –ATP (‘bypass’) configuration (Figure 3—figure supplement 1, reaction ii), with 75 nM Qc-3∆ and the indicated concentrations of Sec17 or its mutants. Fusion is normalized relative to the signals from standard ATP-driven reactions without added Sec17. Each point denotes the mean ± s.e.m. of three independent experiments.
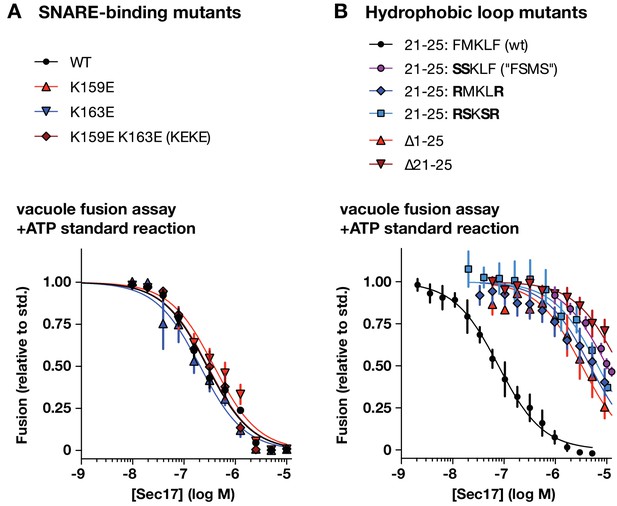
Effects of Sec17 mutations on inhibition of fusion.
(A and B) Ability of Sec17 mutants to inhibit standard ATP-driven vacuole fusion reactions. Vacuole fusion reactions were assembled in standard configuration with 1 mM ATP (Figure 3—figure supplement 1, reaction i), and the indicated concentrations of Sec17 or its mutants. Fusion is normalized relative to the signals from standard ATP-driven reactions without added Sec17. Each point denotes the mean ±s.e.m. of three independent experiments.
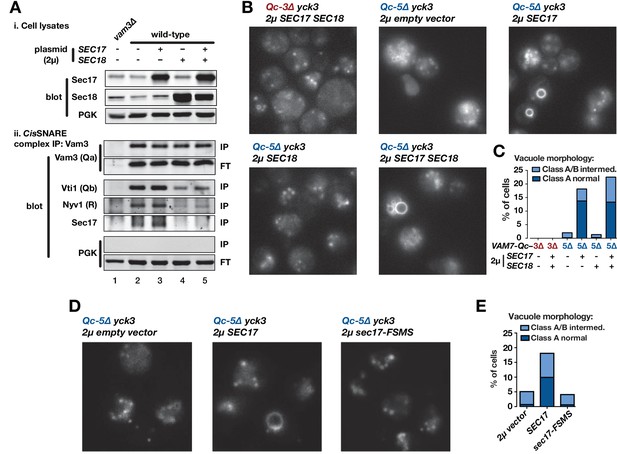
Sec17 overproduction partially rescues in vivo activity of Qc-5∆.
(A) Analysis of cis-SNARE complex abundance in lysates of cells overproducing Sec17 and Sec18. The top panel shows immunoblots of cell lysates. In the bottom panel, anti-Vam3 (Qa-SNARE) was immunoprecipitated from detergent lysates from the indicated strains under non-denaturing conditions. The precipitated material was separated by SDS-PAGE and analyzed by immunoblot, as indicated. IP, immunoprecipitate; FT, flow-through. PGK, phosphoglycerate kinase (control). Additional experimental details are provided in the Materials and methods. (B) Vacuoles in the indicated cell lines were labeled by pulse-chase with FM4-64 dye and observed by epifluorescence. (C) Quantification of phenotypes in B. Bars show mean scores from three independent experiments (n = 88–354 cells per genotype per experiment). (D) Vacuoles in the indicated cell lines were labeled by pulse-chase with FM4-64 dye and observed by epifluorescence. (E) Quantification of phenotypes in D. Bars show mean scores as in C.
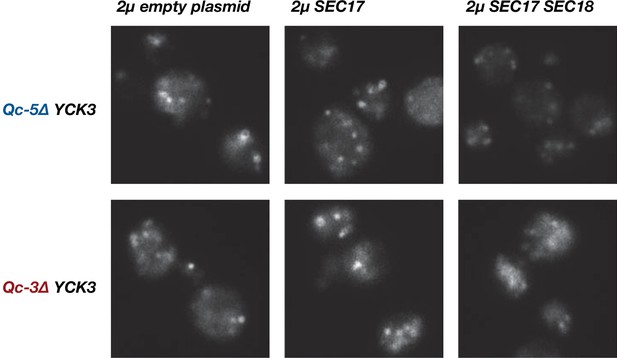
Little or no rescue of fragmented vacuole morphology was observed with Qc-3∆ or Qc-5∆ chromosomal integrants when Sec17, or Sec17 and Sec18, were overproduced in a YCK3 genetic background.
Vacuoles were labeled by pulse-chase with FM4-64 dye and observed by epifluorescence. Representative fields are shown.
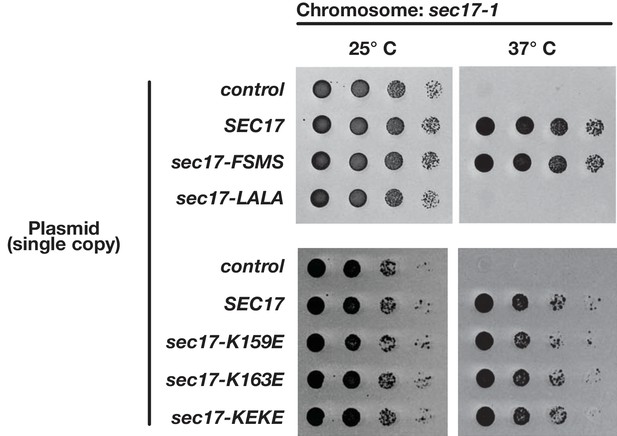
Ability of SEC17 mutants to support growth in sec17-1 mutant cells at nonpermissive temperature.
Liquid cultures were adjusted to equivalent cell densities, serial dilutions were spotted onto synthetic complete agar plates lacking uracil to select for plasmid retention. The plates were incubated at permissive (25°C) or nonpermissive (37°C) temperature for growth of sec17-1 cells, and photographed.
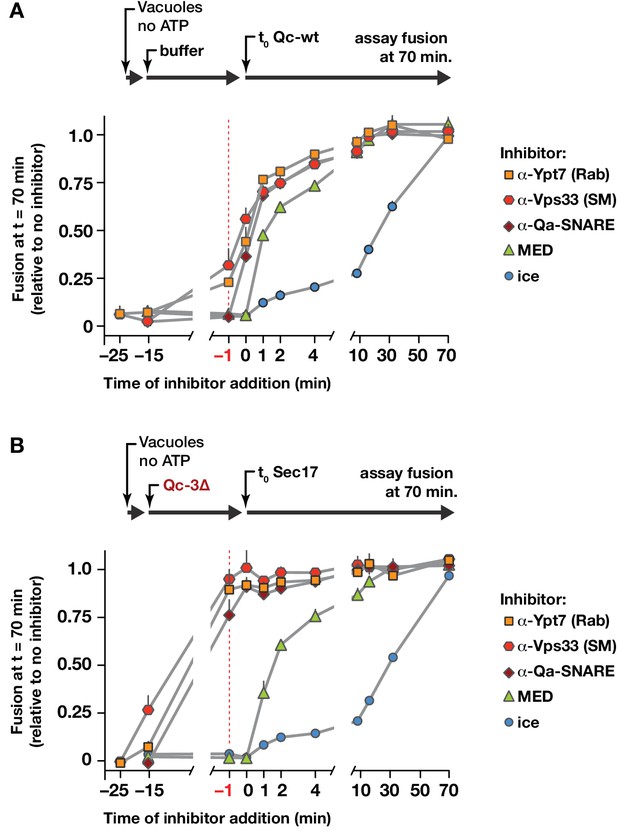
Staging of Sec17 rescue in cell-free assays of vacuole fusion.
(A) Master vacuole fusion reactions were assembled under no-ATP ‘bypass’ conditions (Figure 3—figure supplement 1), and pre-incubated for 25 min at 27°C. Fusion was then initiated (t0) by adding Qc-wt to 20 nM final concentration. At the indicated time points, an aliquot was withdrawn from the master reaction and added to a tube containing the indicated inhibitor (or placed on ice), and incubated for the duration of the experiment. At t = 70 min, each reaction aliquot was assayed for content mixing. (B) Fusion reactions assembled under no-ATP ‘bypass’ conditions were pre-incubated for 10 min at 27°C. At t = −15 min, 75 nM Qc-3∆ was added and the reactions were incubated for an additional 15 min. Fusion was initiated by adding Sec17 (300 nM). At the indicated time points, an aliquot was withdrawn from the master reaction and added to a tube containing the indicated inhibitor (or placed on ice), and incubated for the duration of the experiment. At t = 70 min, each reaction aliquot was assayed for content mixing. For both panels, each point indicates mean + s.e.m. for two to six independent experiments.
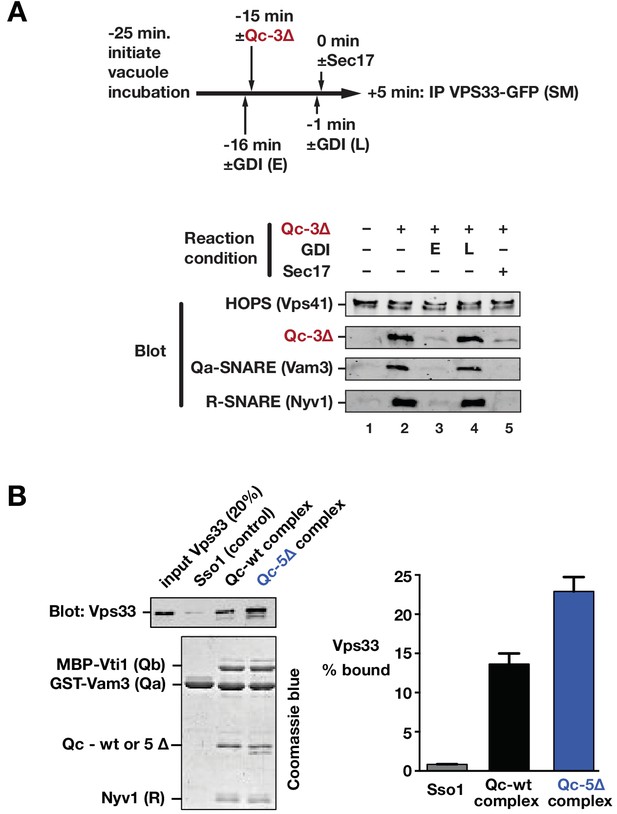
Sec17 and SNARE zippering regulate SNARE interactions with HOPS and Vps33.
(A) Analysis of HOPS-SNARE interactions during a Qc-3∆ block and Sec17 rescue. No-ATP ‘bypass’ reactions were set up similarly to the fusion reactions in Figure 8B, except that only vacuoles isolated from protease-deficient cells expressing Vps33-GFP were used. The vacuoles were pre-incubated for 10 min at 27°C. Qc-3∆ (75 nM) was then added to initiate the assembly of partially zipped trans-complexes. The reactions were incubated for an additional 10 min, and fusion was triggered by adding Sec17 (300 nM). The Rab inhibitor GDI (2.4 µM) was added at early (E) or late (L) time points (lanes 3 and 4). 5 min after Sec17 addition, the vacuoles were sedimented and dissolved in nonionic detergent, and Vps33-GFP was immunoprecipitated with anti-GFP antibodies. The precipitates were separated by SDS-PAGE and analyzed by immunoblot. (B) Interactions of purified Vps33 monomer with vacuolar SNARE complexes. Complexes containing either Qc-wt or Qc-5∆, or the Golgi SNARE Sso1 (control) were assembled on glutathione-agarose resin, and used in pulldowns with soluble, purified recombinant Vps33 as described previously (Lobingier et al., 2014). After washing, the resin-associated material was separated by SDS-PAGE and analyzed either by immunoblotting for Vps33-GFP or by Coomassie blue staining. The band intensities for Vps33 were quantified in three independent experiments and are plotted as mean ± s.e.m.
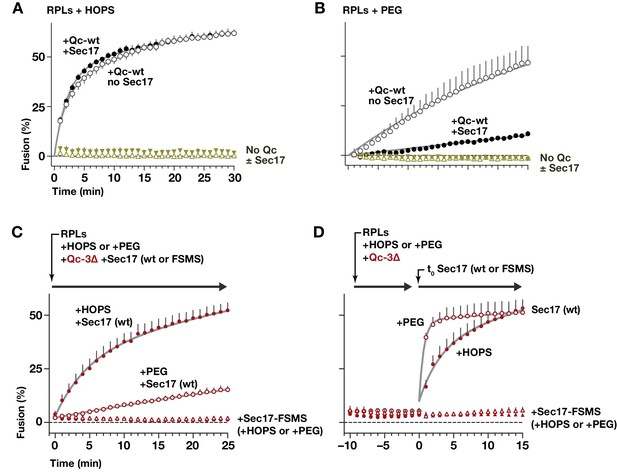
HOPS-SM gates Sec17 function.
(A and B) RPLs bearing Ypt7:GTP and either the Qa and Qb SNAREs, or the R SNARE, were incubated with Sec17 (0 or 600 nM) and Qc-wt (80 and 400 nM in panel A and B, respectively). In panel A, HOPS was present (100 nM). In panel B, the HOPS requirement was bypassed by adding 2% PEG. In control reactions, Qc-wt or Sec17 were omitted. No Sec18 was present under any condition. (C) RPLs bearing Ypt7:GTP and either the Qa and Qb SNAREs, or the R SNARE, were incubated with Qc-3∆ (80 nM), Sec17 or Sec17 FSMS (600 nM), and HOPS (50 nM) or PEG (2%). All soluble components were added together to the vesicles at t = 0. (D) Reactions were set up as in panel C, except that Sec17 was omitted from the initial reaction mix. After a 10 min pre-incubation at 27°C, Sec17 (wt or the FSMS loop mutant) was added at t = 0. The final reagent concentrations were the same as in panel C. For panels A-D, points denote mean plus or minus s.e.m. of 2 (A,B) or 6 (B,C) independent experiments. Lines show nonlinear best-fits of a second-order kinetic model.
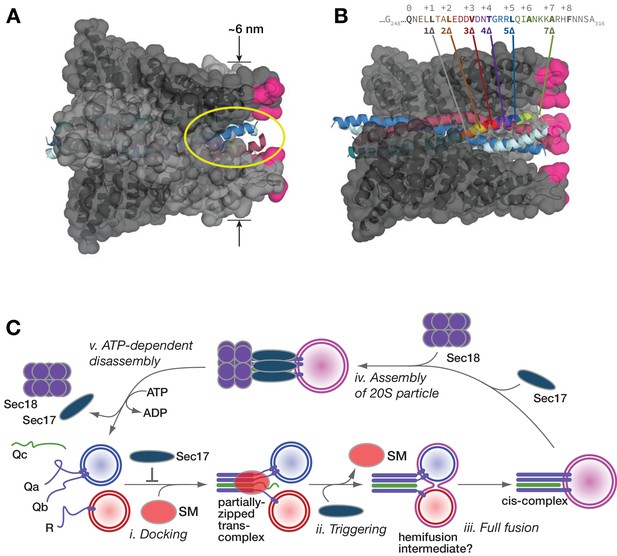
Working model.
(A and B) Sec17/α-SNAP on quaternary SNARE complex (renderings based on PDB 3J96; (Zhao et al., 2015). In B, note the tapered overall shape of the SNARE-Sec17 complex, which forms a roughly triangular structure that could fit between two docked membranes bearing partially zipped SNARE complexes. The hydrophobic loop is magenta. The yellow ellipse shows one of four portals between adjacent Sec17 molecules, through which unstructured SNARE juxta-membrane regions could pass when SNAREs are complexed in the trans configuration. In C, one of four Sec17 molecules is omitted to reveal the Qc packing layers. The portal openings begin at approximately layer +6. Sec17 partially contacts the SNAREs to layer +7 or +8. (C) Interplay of Sec17 and SM-tethering complex on the forward fusion pathway. The early inhibitory function of Sec17 is likely to involve sequestration of SNAREs or SNARE complexes prior to the onset of trans-SNARE interactions. This early inhibition is suppressed by the SM (in our experiments, HOPS-Vps33). The late, stimulatory function of Sec17 can occur in either the absence or presence of HOPS/Vps33. When HOPS-Vps33 is present, Vps33 (SM) ejection from the SNARE complex is a prerequisite for completion of C-terminal SNARE zippering and fusion.
Tables
Strains and plasmids used in this study
https://doi.org/10.7554/eLife.27396.017| Strain/plasmid | Description/genotype | Reference/source |
|---|---|---|
| S. cerevisiae strains | ||
| DKY6281 | MATα pho8∆::TRP1 leu2-3,112 ura3-52 his3-200 trp1-901 lys2-801 suc2-9 | |
| BJ3505 | MATα pep4∆::HIS3 prb1-∆1.6R ura3-52 his3-200 trp1-∆101 lys2-801 can1 gal2 | (Jones, 2002) |
| BY4741 | MATa his3∆one leu2∆0 met15∆0 ura3∆0 | ATCC |
| BY4741 vam7∆::KAN | Invitrogen | |
| AMY1018 | BY4741 VAM7::NAT (Qc-wt knock-in; control wt strain) | This study |
| AMY1022 | BY4741 vam7(1-289)::NAT (Qc-1∆ knock-in) | This study |
| AMY1021 | BY4741 vam7(1-295)::NAT (Qc-3∆ knock-in) | This study |
| AMY1020 | BY4741 vam7(1-302)::NAT (Qc-5∆ knock-in) | This study |
| AMY1019 | BY4741 vam7(1-309)::NAT (Qc-7∆ knock-in) | This study |
| DNY588 | BY4741 VAM7::NAT yck3∆::KAN | This study |
| DNY591 | BY4741 vam7(1-289)::NatR yck3∆::KAN | This study |
| DNY589 | BY4741 vam7(1-295)::NatR yck3∆::KAN | This study |
| DNY590 | BY4741 vam7(1-302)::NatR yck3∆::KAN | This study |
| GOY23 | MATα pep4∆::LEU2 prb1∆::LEU2 leu2-3,112 ura3-52 his3-200 trp1-901 lys2-801 suc2-9 | (Odorizzi et al., 1998) |
| AMY87 | BJ3505 vam3∆ | Merz Lab collection |
| RSY269 | sec17-1 | (Novick et al., 1981) |
| Yeast plasmids | ||
| pDN516 | ApR 2µ URA3 | (Nickerson et al., 2012) |
| pDN313 | pDN516::SEC18 | (Lobingier et al., 2014) |
| pDN314 | pDN516::SEC17 | (Lobingier et al., 2014) |
| pDN315 | pDN516::SEC17 SEC18 | (Lobingier et al., 2014) |
| pDN365 | pDN516::sec17-(F21S, M22S — ‘FSMS’) | This study |
| pDN366 | pDN516::sec17-(L291A, L292A —‘LALA’) | This study |
| pRS416 | ApR CEN URA3 | (Sikorski and Hieter, 1989) |
| pRS426 | ApR 2µ URA3 | (Sikorski and Hieter, 1989) |
| pMS120 | VAM7 (pRS426; Qc-wt overproduction) | This study |
| pMS121 | vam7(1-289) (pRS426; Qc-1∆ overproduction) | This study |
| pMS122 | vam7(1-295) (pRS426; Qc-3∆ overproduction) | This study |
| pMS123 | vam7(1-302) (pRS426; Qc-5∆ overproduction) | This study |
| pMS124 | vam7(1-309) (pRS426; Qc-7∆ overproduction) | This study |
| pMS125 | VAM7 (pAG25) | This study |
| AMP1492 | pRS416::SEC17 | This study |
| AMP1774 | pRS416::sec17-K159E | |
| AMP1774 | pRS416::sec17-K163E | This study |
| AMP1774 | pRS416::sec17-K159E,K163E (‘KEKE’) | This study |
| E. coli plasmids | ||
| AMP356 | pET41::His6-TEV-VAM7-intein-CBD | (Schwartz and Merz, 2009) |
| AMP359 | pET41::His6-TEV-VAM7(1–289, 1∆)-intein-CBD | (Schwartz and Merz, 2009) |
| AMP1809 | pET41::His6-TEV-VAM7(1–291, 2∆)-intein-CBD | This study |
| AMP358 | pET41::His6-TEV-VAM7(1–295, 3∆)-intein-CBD | (Schwartz and Merz, 2009) |
| AMP1810 | pET41::His6-TEV-VAM7(1–299, 4∆)-intein-CBD | This study |
| AMP357 | pET41::His6-TEV-VAM7(1–302, 5∆)-intein-CBD | (Schwartz and Merz, 2009) |
| AMP1806 | pET41::His6-TEV-VAM7(1–302, AR-5∆)-intein-CBD | This study |
| AMP1807 | pET41::His6-TEV-VAM7(1–302, RA-5∆)-intein-CBD | This study |
| AMP1808 | pET41::His6-TEV-VAM7(1–302, AA-5∆)-intein-CBD | This study |
| AMP360 | pET41::His6-TEV-VAM7(1–309, 7∆)-intein-CBD | (Schwartz and Merz, 2009) |
| AMP1547 | pTYB12::intein-CBD-SEC17 | (Schwartz and Merz, 2009) |
| AMP1547 | pTYB12::intein-CBD-SEC17 (F21R, M22S, L24S, F25R) | This study |
| AMP1548 | pTYB12::intein-CBD-SEC17 (F21R, F25R) | This study |
| AMP1549 | pTYB12::intein-CBD-SEC17 (∆21–25) | This study |
| AMP1550 | pTYB12::intein-CBD-SEC17 (F21S, M22S — ‘FSMS’) | This study |
| AMP1551 | pTYB12::intein-CBD-SEC17 (∆1–26) | This study |
| AMP1777 | pTYB12::intein-CBD-SEC17 (K159E) | This study |
| AMP1778 | pTYB12::intein-CBD-SEC17 (K163E) | This study |
| AMP1779 | pTYB12::intein-CBD-SEC17 (K159E, K163E — ‘KEKE’) | This study |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.27396.018