Serine ADP-ribosylation reversal by the hydrolase ARH3
Figures
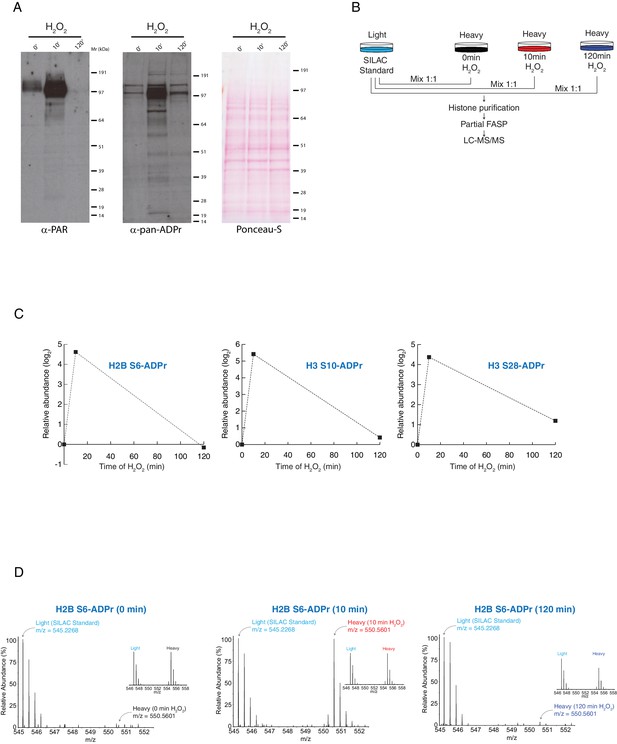
Histone serine ADP-ribose modification is reversible.
U2OS cells were treated with H2O2 and analysed at indicated time-points. (A) The samples were lysed and the proteins were separated by SDS-PAGE, analysed by western blot and probed for PAR (left) or pan-ADPr (middle). Ponceau-S staining was used as loading control (right). (B) Schematic representation of the SILAC-based strategy to quantify core histone Ser-ADPr marks after different time points of 2 mM H2O2–induced DNA damage. Light labeled (Lys0) cells treated for 10 min with 2 mM H2O2 were used as SILAC Standard. (C) The relative abundance of Ser-ADPr modification on histone proteins H2B (Ser6) and H3 (Ser10, Ser28) was calculated and plotted as a function against time of H2O2 treatment. (D) MS1s of a Ser-ADPr H2B peptide (H2B Ser6-ADPr) at different time points of 2 mM H2O2 treatment. The heavy peptide was derived from cells treated with H2O2 for the indicated time point, and the light peptide was derived from the SILAC Standard (10 min H2O2-treated cells). Each inset (right) shows a ∼1:1 ratio (heavy/light) of a non-ADP-ribosylated peptide from the same experiment.
-
Figure 1—source data 1
MaxQuant evidence table related to Figure 1—figure supplement 1.
ADPr identified peptide features from SILAC experiments performed to quantify ADPr after different time points of 2 mM H2O2–induced DNA damage (Figure 1—figure supplement 1).
- https://doi.org/10.7554/eLife.28533.004
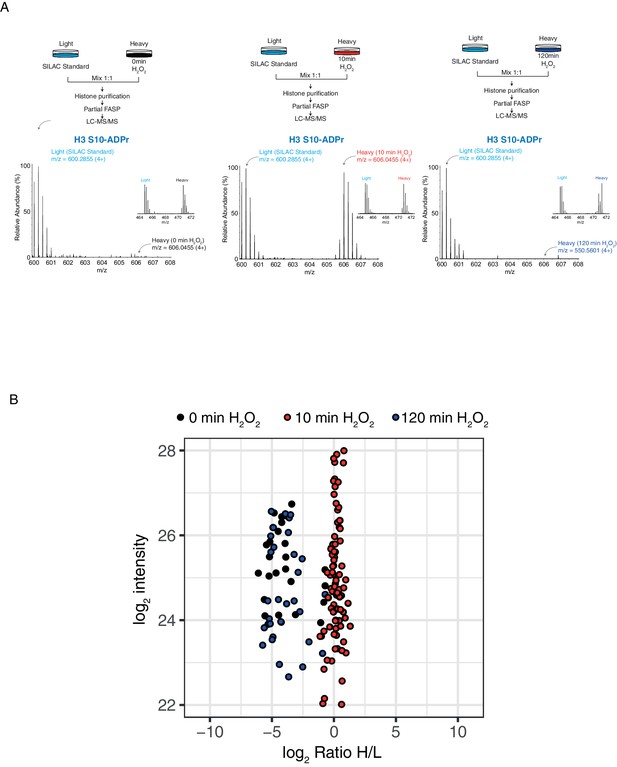
Histone serine ADP-ribose modification is reversible.
(A) Schematic representation of the SILAC-based strategy to quantify core histone Ser-ADPr marks after different time points of 2 mM H2O2–induced DNA damage. Light labeled (Lys0) cells treated for 10 min with H2O2 were used as SILAC Standard (top panels). MS1s of a Ser-ADPr H3 peptide (H3-Ser10-ADPr) at different time points of 2 mM H2O2 treatment (bottom panels). The heavy peptide was derived from cells treated with H2O2 for the indicated time point, and the light peptide was derived from the SILAC Standard (10 min H2O2-treated cells). Each inset (right) shows a ∼1:1 ratio (heavy/light) of a non-ADP-ribosylated peptide from the same experiment. (B) Log2 of summed ADPr peptide intensities were plotted against log2 of ADPr Heavy/Light SILAC ratios for each time point. ADPr peptides detected after 0 min of 2 mM H2O2 are colored in black. ADPr peptides detected after 10 min 2 mM H2O2 are colored in red. ADPr peptides detected after 120 min 2 mM H2O2 are colored in blue.
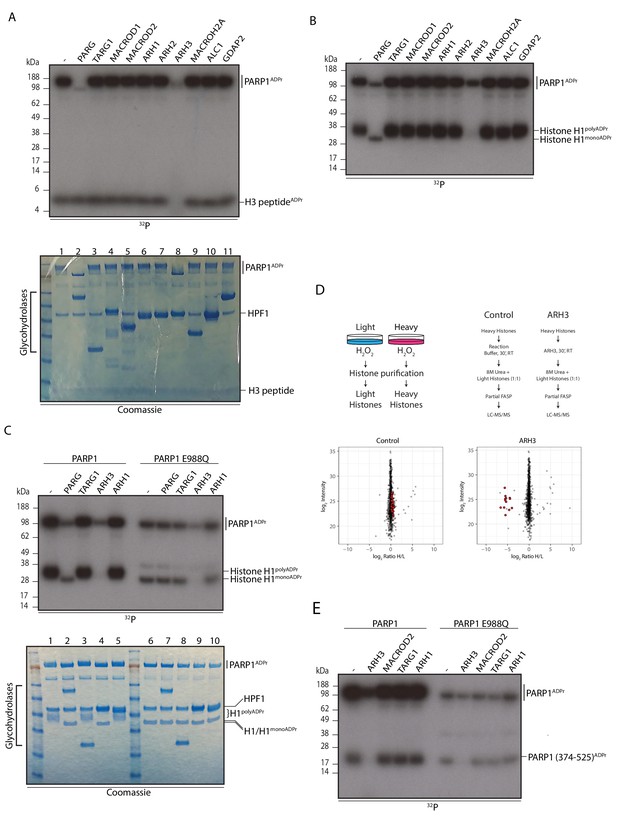
ARH3 hydrolyses Ser-ADPr.
(A) Glycohydrolysis of mono-ADP-ribosylated H3 histone peptide (aa 1-21). ADP-ribosylated H3 peptide was obtained by incubation with PARP1 and HPF1 using radioactively labelled NAD+ as an ADPr donor. After stopping the modification reaction with PARP inhibitor, the indicated glycohydrolases and related proteins were added to the substrate as shown on the figure. Lower panel – Coomassie stained gel of the proteins used in the reaction that also serves as a loading control. (B) Removal of poly-ADP-ribosylation from the recombinant histone H1 substrate by various glycohydrolases. Reaction performed as described in A. (C) Comparison of the hydrolase reactions on the poly- and mono-ADP-ribosylated histone H1 substrates. Lower panel – Coomassie stained gel of the proteins used in the reaction that also serves as a loading control. (D) Schematic representation of the SILAC-based strategy to quantify ADPr removal upon in vitro incubation of purified Ser-ADPr histones with or without recombinant ARH3 (top panels). Log2 of summed peptide intensities were plotted against log2 Heavy/Light SILAC ratios for each condition (bottom panels). ADPr peptides are colored in red. (E) Autoradiogram analysis of Ser mono-ADPr hydrolysis with a non-histone protein substrate. The ADP-ribosylated automodification domain of PARP1 (aa 374-525) was used in this assay and the reactions were supplemented with the indicated hydrolases. In panel panels A, B, C and E, the reaction products were separated by SDS-PAGE and analysed by autoradiography. The signals relating to the specific ADP-ribosylated protein are indicated in each panel.
-
Figure 2—source data 1
MaxQuant evidence table related to Figure 2D.
ADPr identified peptide features from SILAC experiments performed to quantify ADPr removal upon in vitro incubation of purified Ser-ADPr histones with or without recombinant ARH3 (Figure 2D).
- https://doi.org/10.7554/eLife.28533.007
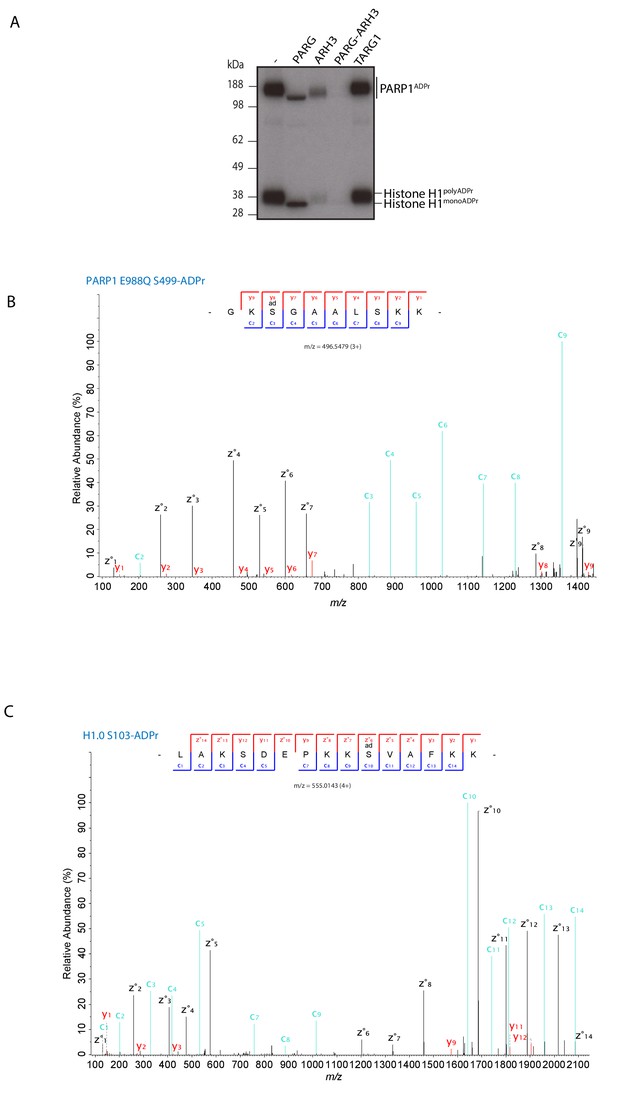
PARP1 E988Q generates mono Ser-ADPr on itself and on substrates.
(A) Removal of poly-ADP-ribosylation from the recombinant histone H1 substrate by the glycohydrolases PARG, ARH3, PARG and ARH3 together, and TARG1 as indicated. ADP-ribosylated histone H1 was obtained by incubation with PARP1 and HPF1 using radioactively labelled NAD+ as an ADPr donor. After stopping the modification reaction with PARP inhibitor, the indicated glycohydrolases were added to the substrate as shown on the figure. (B) High-resolution ETD fragmentation spectrum of a PARP1 E988Q peptide modified by ADP-ribose on Ser499. (C) High-resolution ETD fragmentation spectrum of a H1.0 peptide modified by ADP-ribose on Ser103.
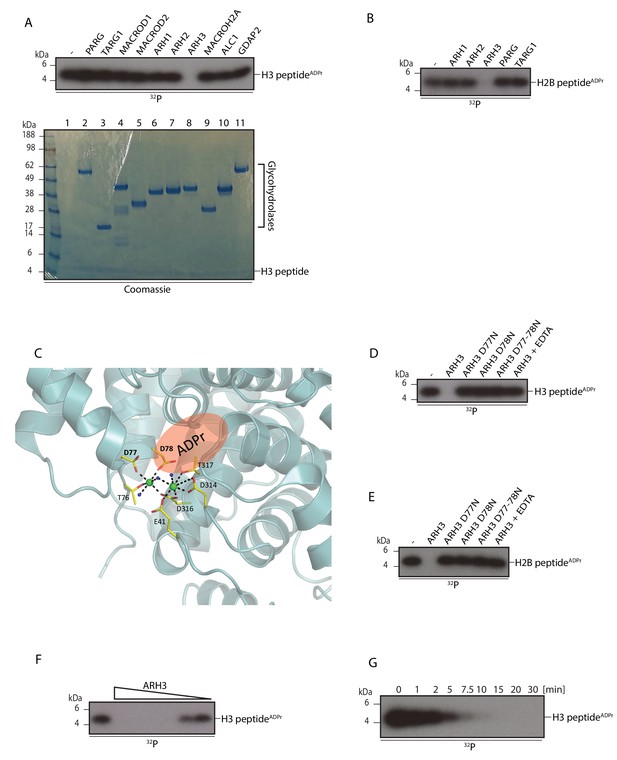
Characterisation of the serine ADP-ribosylhydrolase activities of ARH3.
Glycohydrolase assay using the purified serine ADP-ribosylated histone peptides as substrates. Reactions were performed and analysed as described in Figure 2A. (A) Activity of different hydrolases against ADP-ribosylated H3 peptide (aa 1-21). (B) The same as in (A) except purified histone H2B peptide was used. (C) ARH3 structure (PDB ID: 2FOZ) showing the amino acid residues in the catalytic site coordinating Mg ions (green). The putative ADPr binding site is highlighted in red. (D) Mutating ARH3 catalytic residues or addition of EDTA to the reaction abolishes ARH3 activity. (E) Reaction performed as (D) except histone H2B peptide was used. (F) ARH3 activity is enzyme concentration dependent. ARH3 enzyme concentrations were in the range from 3 μM to 5 nM. (G) ARH3 activity is time dependent. 0.2 µM ARH3 was used in this assay.
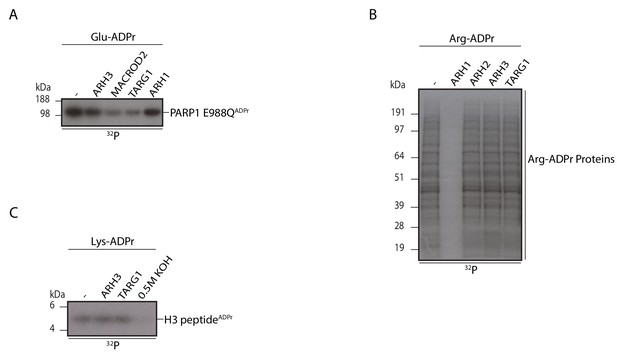
ARH3 specifically cleaves Ser mono-ADPr, but not Glu, Arg or Lys mono-ADPr.
Reactions were analysed as in Figure 2A. (A) Analysis of ARH3 activity on a glutamate-ADP-ribosylated protein. The mono-ADP-ribosylated recombinant PARP1 E988Q protein was treated with the indicated glycohydrolases. (B) Analysis of ARH3 activity on arginine-ADP-ribosylated proteins. Cellular extracts from K562 cells were ADP-ribosylated by recombinant ARTC2.2 protein. The reactions were then supplemented with the indicated glycohydrolases. (C) Analysis of ARH3 activity on a lysine-ADP-ribosylated peptide. A chemically modified histone H3 peptide with mono-ADPr on lysine residues was incubated with the indicated glycohydrolases or 0.5 M KOH for 30 min.
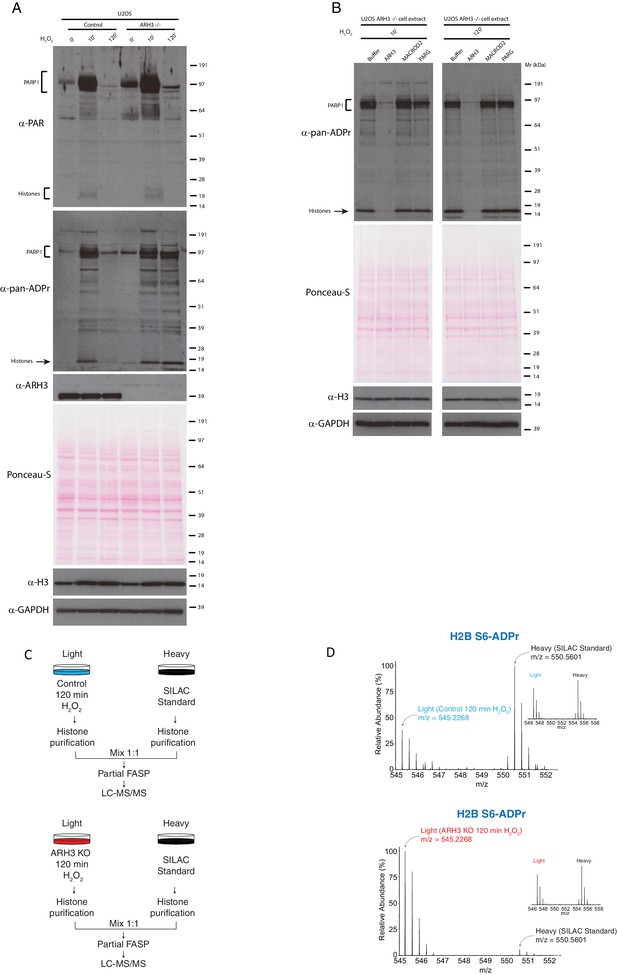
ARH3 is necessary for removal of Ser-ADPr in cells.
(A) Control and ARH3 KO U2OS (ARH3-/-) cells were treated with 2 mM H2O2 for the indicated time points. After treatment, cells were lysed and proteins were separated by SDS-PAGE, analysed by western blot and probed for PAR, pan-ADPr, ARH3, H3, and GAPDH antibodies. Additionally, Ponceau-S staining was used as loading control (B) Cell extracts obtained from ARH3 KO cells treated with 2 mM H2O2 for 10 and 120 min were incubated with buffer or ADP-ribosylhydrolases ARH3, MACROD2 and PARG. Samples were separated by SDS-PAGE, analysed by Western blot and probed for pan-ADPr, H3, GAPDH, and Ponceau-S staining were used as loading control. (C) Schematic representation of the SILAC-based strategy to quantify core histone Ser-ADPr marks from Control U2OS cells (top panel) or ARH3 KO U2OS cells (bottom panel) after H2O2 treatment for 120 min. Heavy labeled (Lys8) Control U2OS cells treated for 10 min with H2O2 were used as SILAC Standard. (D) MS1s of a Ser-ADPr H2B peptide (H2B Ser6-ADPr) from Control U2OS cells (top panel) or ARH3 KO U2OS cells (bottom panel) after H2O2-treatment for 120 min. The light peptide was derived from Control (top panel) or ARH3 KO (bottom panel) cells treated with 2 mM H2O2 for 120 min, and the heavy peptide was derived from the SILAC Standard (10 min H2O2 treated cells). Each inset shows a ∼1:1 ratio (heavy/light) of a non-ADP-ribosylated peptide from the same experiment. In the ARH3 KO U2OS cells, the chosen H2B mark (H2B S6-ADPr) resulted ~40 times more abundant than in the Control U2OS cells.
-
Figure 5—source data 1
MaxQuant ADPr sites table related to Figure 5C–D.
List of identified Ser-ADPr sites from SILAC experiments performed to quantify core histone Ser-ADPr marks from Control and ARH3 KO U2OS cells (Figure 5C–D).
- https://doi.org/10.7554/eLife.28533.012
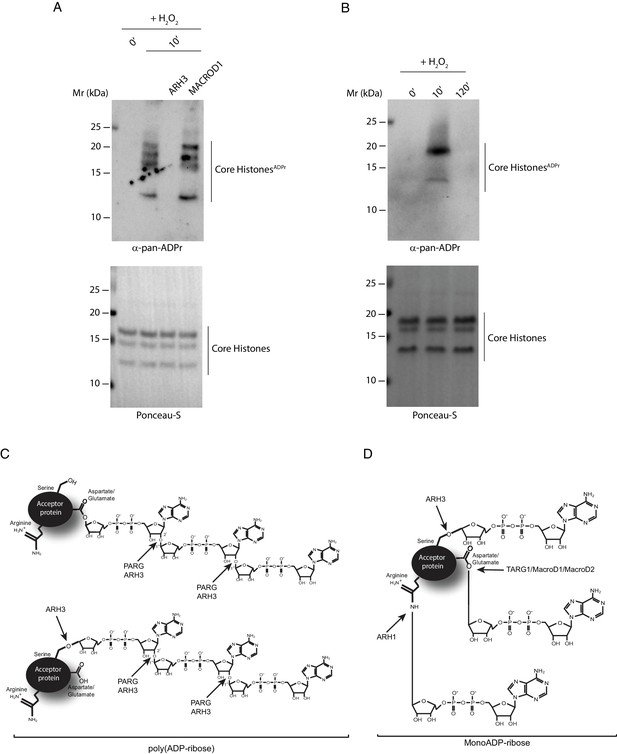
ARH3 as a tool for recognizing histone Ser-ADPr.
(A) Core histones were purified from U2OS cells treated with H2O2 for the indicated time points. Recombinant ARH3 or MACROD1 or reaction buffer was added to histones purified from 10 min H2O2-treated cells. After treatment, proteins were separated by SDS-PAGE, analysed by western blot and probed for pan-ADPr. Ponceau-S staining was used as loading control. (B) Purified core histones from U2OS cells treated with H2O2 for the indicated time points were separated by SDS-PAGE, analysed by Western blot and probed for pan-ADPr. Ponceau-S staining was used as loading control. (C and D) A schematic representation of the specificity of the ADP-ribosylhydrolases PARG, ARH3, TARG1, MACROD1, MACROD2 and ARH1 for PARylated (C) and MARylated (D) proteins.
Additional files
-
Source code 1
R-script used to analyse the MaxQuant data.
- https://doi.org/10.7554/eLife.28533.014





