Architecture of TAF11/TAF13/TBP complex suggests novel regulation properties of general transcription factor TFIID
Figures
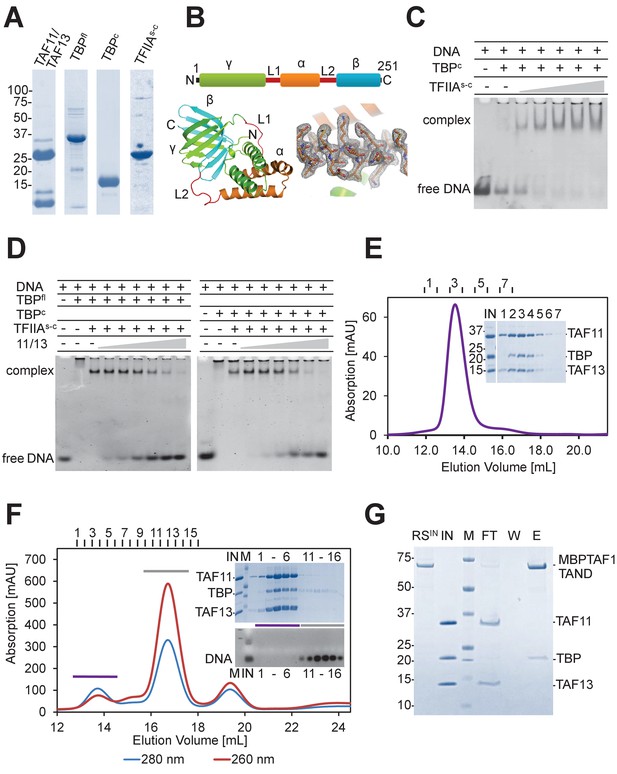
TAF11/TAF13 and TBP form ternary complex.
(A) Human TAF11/TAF13 complex, TBP full-length (TBPfl) and core (TBPc), and a single-chain version of TFIIA (TFIIAs-c) were purified to homogeneity as shown by SDS-PAGE. (B) TFIIA subunits α(ΑΑ2–59), β(ΑΑ302–376) and γ(ΑΑ2–110)(Bleichenbacher et al., 2003) were connected with linkers L1(-DGKNTANSANTNTV-) and L2(-SRAVDGELFDT-) as indicated (top). TFIIAs-c crystallized during purification (Figure 1—figure supplement 1). The 2.4 Å X-ray structure is shown in a cartoon representation with a section of electron density at 1σ(bottom). (C) TFIIAs-c was assayed by band-shift for activity. DNA, Adenovirus major late promoter (AdMLP) DNA. (D) Probing formation of putative pentameric TAF11/TAF13/TBPTFIIA/DNA complex by band-shift assay (Kraemer et al., 2001; Robinson et al., 2005). TAF11/TAF13 titration to the TFIIA/TBP/DNA complex results in DNA release. (E) SEC analysis reveals a stable TAF11/TAF13/TBP ternary complex. Elution fractions (1-7) were analyzed by SDS-PAGE (inset). IN, equimolar mixture of TAF11/TAF13 and TBP. No interactions were found between TAF11/TAF13/TBP complex and TFIIA (Figure 1—figure supplement 2). (F) TAF11/TAF13 competes with DNA for TBP binding, evidenced by SEC. Elution fractions (1-6, 11-16) were analyzed by SDS-PAGE and ethidium-bromide stained agarose gel (inset). IN, preformed TAF11/TAF13/TBP complex; DNA, AdMLP DNA; M, protein molecular weight marker and DNA ladder, respectively. (G) Immobilized human TAF1 N-terminal domain (TAF1-TAND) (Anandapadamanaban et al., 2013) efficiently depletes TBP from preformed TAF11/TAF13/TBP complex. RSIN, amylose resin with MBP-tagged TAF1-TAND bound; IN, preformed TAF11/TAF13/TBP; M: protein marker; FT, flow-through fraction; W, wash fraction; E, maltose elution fraction.
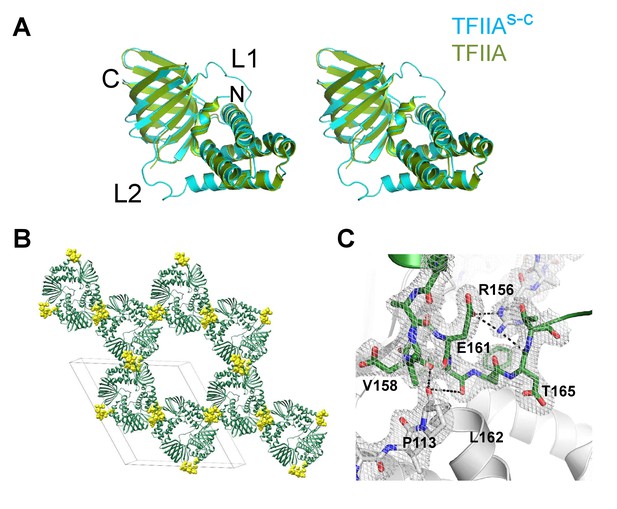
Crystal structure of TFIIAs-c at 2.4 Å resolution.
(A) Superimposition of human TFIIA from a TFIIA/TBP/DNA ternary complex (green, PDB ID: 1NVP) and uncomplexed TFIIAs-c in our crystal (blue) is shown in a cartoon representation in a stereo view. The structures are virtually identical (rms deviation 0.753 Å). C, N are C- and N-terminal end of single-chain; L1, L2 are Linker regions. (B) Crystal packing of TFIIAs-c, highlighting the critical role of engineered linkers (yellow spheres) in mediating the majority of crystal contacts. The unit cell is indicated (grey lines). (C) A close up highlights the interactions mediated by the linker region between symmetry related molecules in the crystal lattice. Symmetry related molecules are colored in grey or green, respectively. Hydrogen bonds (distance cut-off 3.7 Å) are shown as black dashed lines. Amino acid residue numbers are indicated. A section of the 2Fo-Fc electron density map contoured at 1σ is shown (grey mesh). Crystal statistics are provided in Table 1.
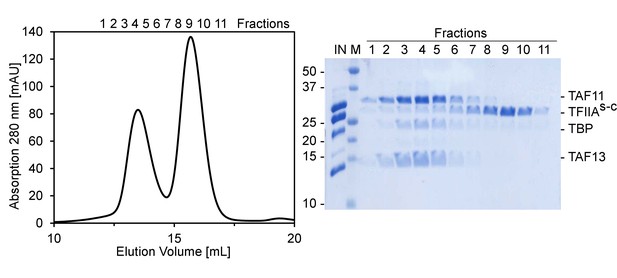
Size exclusion chromatography (SEC) analysis of TAF11/TAF13/TBP and TFIIAs-c.
Interaction analysis of TAF11/TAF13/TBP complex with TFIIAs-c by SEC. Preformed TAF11/TAF13/TBP complex and TFIIAs-c were combined and injected on a Superdex S200 10/300 column (left). Peak elution fractions were further analyzed by SDS-PAGE (right). TAF11/TAF13/TBP complex and TFIIAs-c eluted in separate peaks indicating a lack of interaction at the conditions studied. IN, Input sample; M, BioRad prestained marker (MW of marker bands are indicated). Elution fractions are numbered 1 to 11.
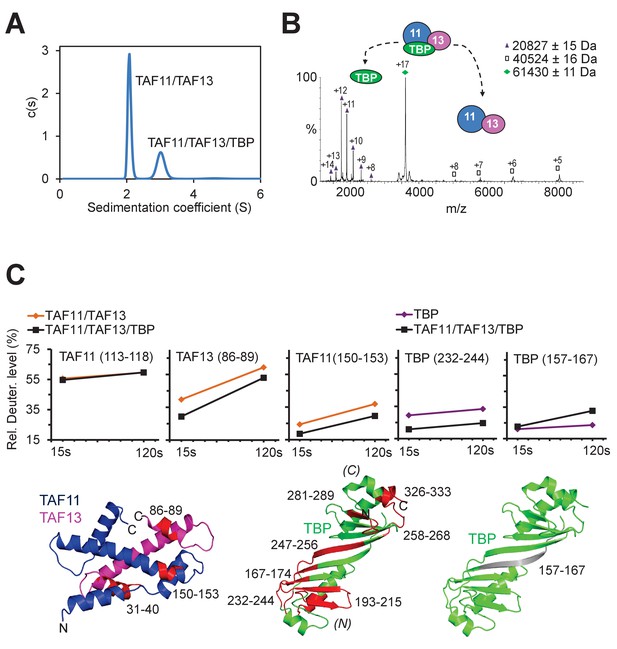
TAF11/TAF13/TBP interactions.
(A) Sedimentation velocity analytical ultracentrifugation (AUC) experiments reveal a major peak consistent with a 1:1:1 TAF11/TAF13/TBP ternary complex. The second, smaller peak corresponds to excess TAF11/TAF13. (B) Native mass spectrometry of the TAF11/TAF13/TBP complex confirms 1:1:1 stoichiometry. Collision induced dissociation (CID) results in TBP monomer and TAF11/TAF13. Experimental masses are provided (inset). Calculated masses: 20659 Da (TBP); 40691 Da (TAF11/TAF13); 61351 Da (TAF11/TAF13/TBP). Mass spectra of the TAF11/TAF13/TBP complex and TBP dimer are shown in Figure 2—figure supplement 1. (C) TAF11/TAF13/TBP and TAF11/TAF13 were analyzed by hydrogen-deuterium exchange/mass spectrometry (HDX-MS) (Rajabi et al., 2015). Changes in the deuteration level of selected peptides in TAF11, TAF13 or TBP are depicted in diagrams (top row). Peptides protected upon ternary complex formation are coloured in red in cartoon representations of TAF11/TAF13 and TBP (bottom row). One peptide in TBP (grey, far right) becomes more accessible, hinting at disassembly of a TBP dimer when TAF11/TAF13 binds (Figure 2—figure supplement 1). All peptides implicated in TAF11/TAF13-binding map to the concave DNA-binding surface of TBP. (N) and (C) indicate N- and C-terminal TBP stirrups, respectively.
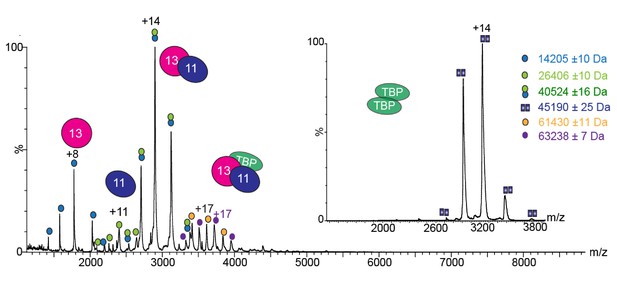
Native mass spectrometry (MS) analysis of TAF11/TAF13/TBP complex.
Mass spectrum of the TAF11/TAF13/TBP complex recorded from a buffer solution containing 150 mM ammonium acetate (pH7.5). Two charge state series are observed corresponding to complex with (circles filled in purple) and without (circles filled in orange) an oligo-histidine affinity purification tag, respectively. Calculated masses: 20659 Da (TBP); 22765 Da (TBP with His-tag); 45531 Da (TBP dimer); 40691 Da (TAF11/TAF13). The inset shows the native MS spectrum recorded for separately purified TBP alone, evidencing a TBP dimer.
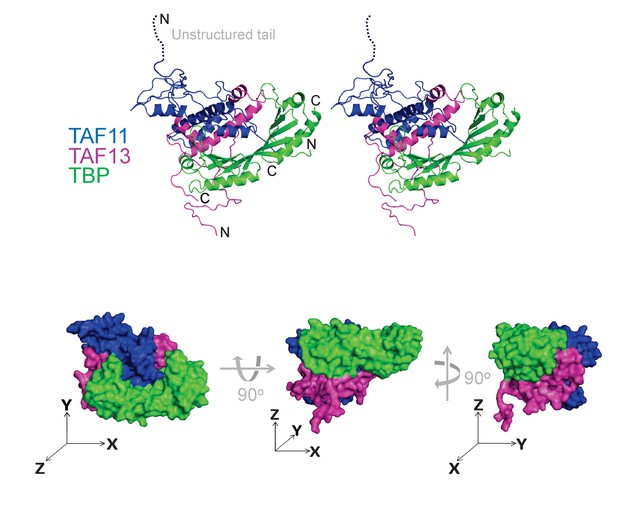
Architecture of TAF11/TAF13/TBP complex.
TAF11/TAF13/TBP complex architecture was determined by using an integrative multi-parameter approach. We utilized the crystal structure of TBP (Nikolov et al., 1996) as well as the crystal structure of the TAF11/TAF13 dimer (Birck et al., 1998) combined with our native MS, SAXS, AUC and HDX-MS results as well as distance constraints from CLMS experiments (Figure 3—figure supplements 1–3). The structure of the TAF11/TAF13/TBP ternary complex is shown in a cartoon representation in stereo (top) and as a space filling model (devoid of unstructured regions) in three views (bottom). Three axes (x, y, z, drawn as arrows) illustrate the spatial relation between the views. TAF11 is colored in blue, TAF13 in magenta and TBP in green. This model satisfies >90% of the experimental constrains (Figure 3—figure supplement 4, Tables 2 and 4).
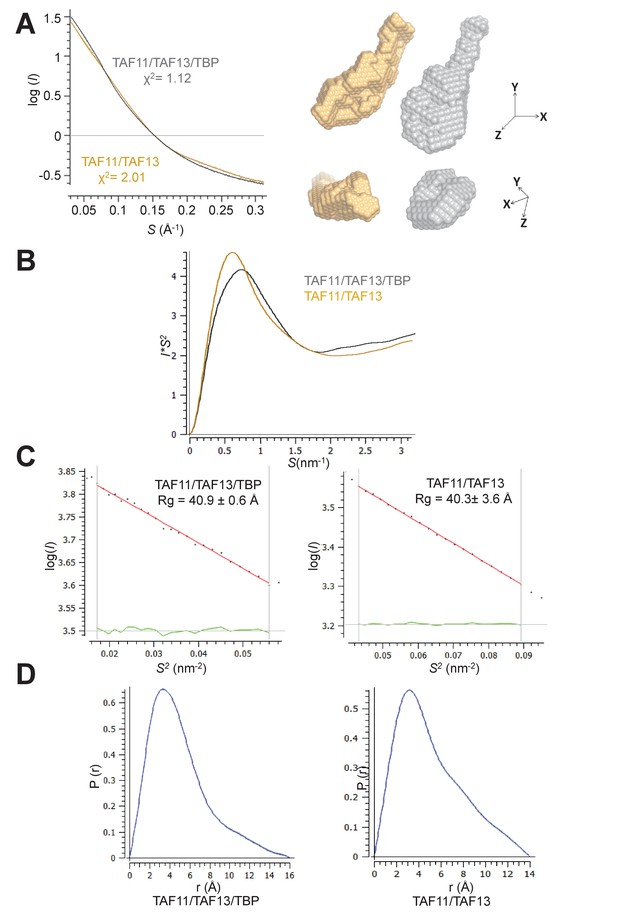
Small angle X-ray scattering (SAXS).
(A) Small-angle X-ray scattering (SAXS) data of TAF11/TAF13/TBP and TAF11/TAF13. Fit of experimental scattering (dots) and scattering calculated from ab initio models (solid lines) is provided in a log plot (left). SAXS envelopes of TAF11/TAF13 (yellow), TAF11/TAF13/TBP (grey) and an overlay are shown (right). (B) Kratky plots for TAF11/TAF13/TBP (light gray) and TAF11/TAF13 (light orange). (C) Guinier plots for TAF11/TAF13/TBP (left) and TAF11/TAF13 (right) are shown; Rg values are indicated. Residuals of the linear fit are represented as green lines. (D) Distance distribution analysis for TAF11/TAF13/TBP (left) and TAF11/TAF13 (right). SAXS statistics are provided in Table 3.
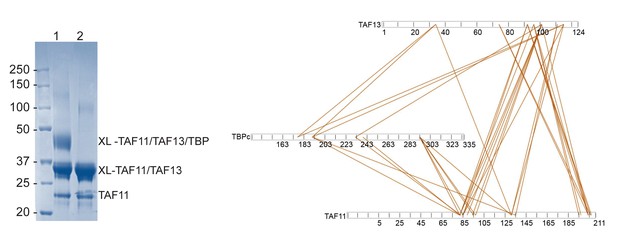
Cross-linking/mass spectrometry (CLMS).
TAF11/TAF13/TBP complex and TAF11/TAF13 complex were cross-linked separately using bissulfosuccinimidyl suberate (BS3) and then further purified by SEC using a Superdex 200 10/300 column. Cross-linked and purified TAF11/TAF13/TBP complex (lane 1) and TAF11/TAF13/TBP complex (lane 2) were analyzed by SDS-PAGE (left). The top band corresponds to cross-linked TAF11/TAF13/TBP, the middle band to TAF11/TAF13; the band on the bottom to uncross-linked TAF11. Cross-linked TAF11/TAF13 is present also in the cross-linked TAF11/TAF13/TBP sample. Bands were excised, reduced, alkylated, trypsin digested and desalted followed by mass spectrometric analysis. Cross-links were identified between TAF11 and TAF13, TAF11 and TBP as well as TAF13 and TBP. Cross-links between TAF11-TAF13, TAF11-TBP and TAF13-TBP are shown using 5% FDR cutoff data (right). The N-terminal region of TAF11 is lacking amino acids R and K, and consequently, no cross-links were observed with BS3. Observed cross-links are listed in Table 4.
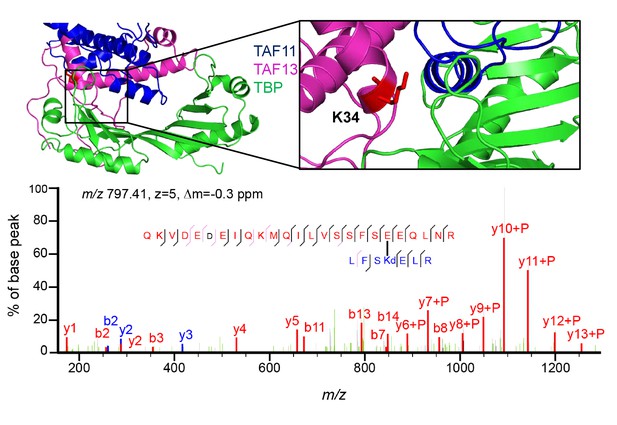
Site-specific cross-linking of TAF11 and TAF13 by Genetic Code Expansion (GCE).
The UV-activatable amino acid DiAzKs (Diazirine-Lysine) was introduced at position K34 in TAF13 by genetic code expansion (MultiBacTAG) (Koehler et al., 2016). Thus, labeled TAF11/TAF13 complex and TBP were reconstituted into TAF11/TAF13/TBP complex, which was then exposed to UV light. A cartoon representation of TAF/TAF13/TBP is shown (left) with a zoom-in on the site of ncAA introduction (right). K34 of TAF13 was chosen as it appears to be located at/near the interfaces between TAF13, TAF11 and TBP in our model of the ternary complex. Cross-linked sample then was separated by SDS-PAGE, excised from gel, reduced, alkylated, trypsin digested and desalted followed by mass spectrometry analysis. Specific cross-linking patterns were obtained upon TAF11/TAF13 complex formation with TBP. Example of mass spectrum for a cross-linked peptide is shown below.
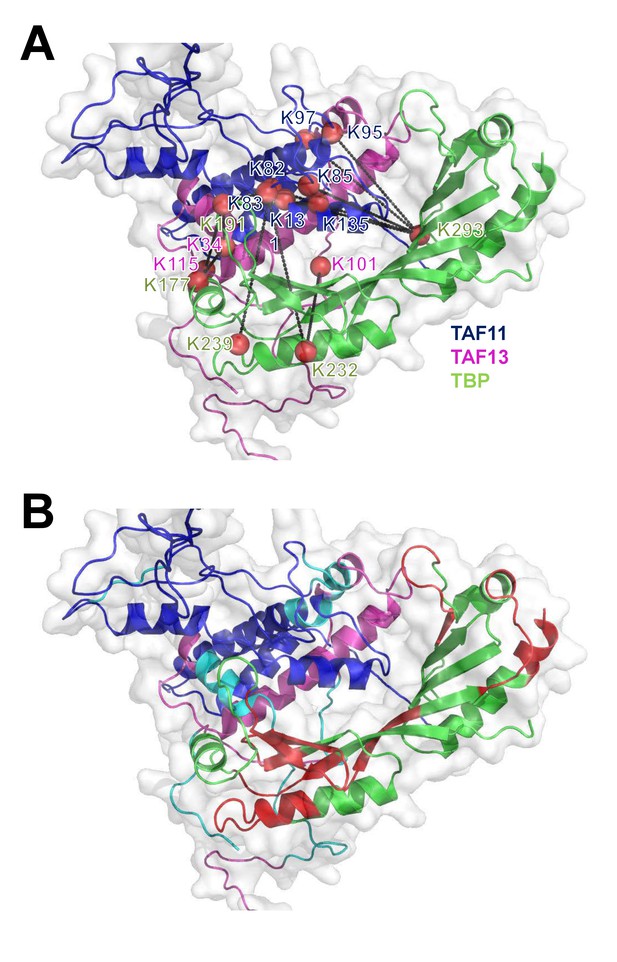
Mapping CLMS and HDX-MS data on TAF11/TAF13/TBP complex.
(A) Cross-links identified by BS3 CLMS mapped on the TAF11/TAF13/TBP ternary complex. The current model satisfies >90% of all observed cross-links. Cross-linking sites are depicted as red balls. (B) Regions on TBP exhibiting changes in deuteration levels upon complex formation with TAF11/TAF13 are highlighted in red.
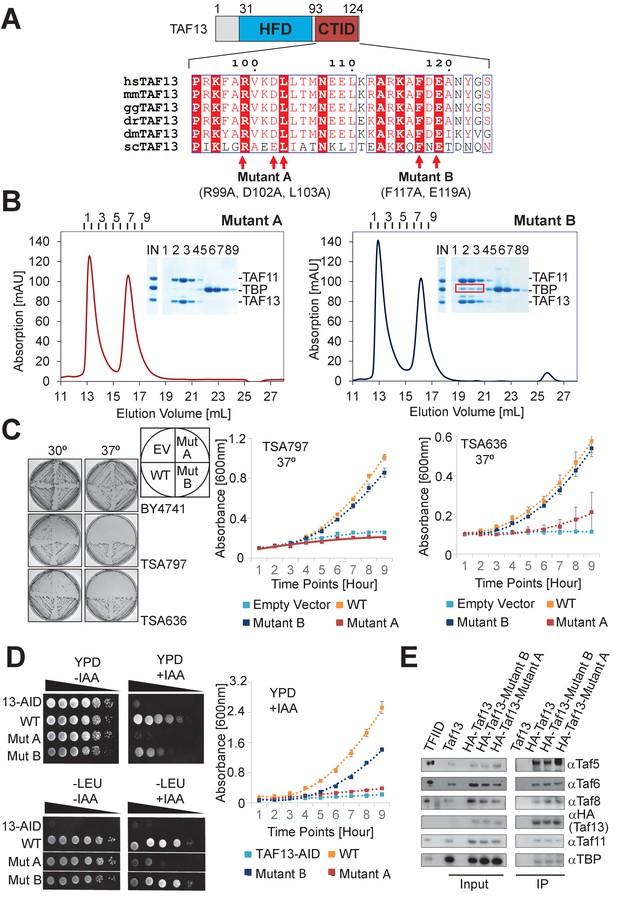
Highly conserved C-terminal TBP-interaction domain (CTID) in TAF13 required for survival.
(A) Sequence alignments reveal a highly conserved C-terminal TBP interaction domain (CTID) in TAF13 comprising virtually identical signature residues in TAF13 from yeast to humans. Residues that were mutated in the CTID of TAF13 are indicated by arrows, giving rise to two mutant TAF13 proteins (Mutant A, B). The locations of the mutated residues in TAF11/TAF13/TBP are illustrated in Figure 4—figure supplement 1A. (B) SEC analysis demonstrates complete abolition of the TBP binding by TAF11/TAF13 in case of Mutant A. In case of Mutant B, residual interaction with TBP is observed (marked by red box). Elution fractions (1-8) were analyzed by SDS-PAGE (inset). IN, equimolar mixture of TAF11/TAF13 and TBP. (C) Cell growth experiments in yeast containing temperature-sensitive (ts) Taf13 on solid media plates at permissive (30○C) and non-permissive (37○C) temperatures are shown on the left. EV, empty vector; WT, wild-type Taf13; MutA, MutB, Taf13 mutants A and B; TSA797, TSA636, yeast strains harboring distinct temperature-sensitive Taf13 mutants (Shen et al., 2003; Lemaire and Collart, 2000). Corresponding absorbance plots displaying growth curves of temperature-sensitive strains in liquid media at the non-permissive temperature (37°C) are provided on the right. Polynomial fits are shown as dotted lines. Standard errors of mean (SEM) are shown as bars. The corresponding growth curves for strain BY4741 used as a control, are shown in Figure 4—figure supplement 1B. (D) Cell growth experiments in yeast containing Taf13 fused to an auxin-inducible degron tag (AID) are shown in spot assays on solid media plates (YPD, -LEU) on the left, in presence or absence of indole-3-acetic acid (IAA) which activates Taf13-AID depletion. 13-AID, Taf13 degron-tag fusion (Warfield et al., 2017); YPD, yeast total media; -LEU, synthetic drop-out media. Corresponding absorbance plots displaying growth curves in presence of IAA are shown on the right. Absorbance plots in absence of IAA are provided in Figure 4—figure supplement 1C. (E) Western blots from co-immunoprecipitations (co-IPs) from yeast are shown of HA-tagged wild-type and mutant Taf13 proteins, probed with specific antibodies against Taf5, Taf6, Taf8, Taf11, TBP and the HA tag on Taf13. Purified yeast holo-TFIID (marked as TFIID) and extract from yeast transformed with untagged wild-type Taf13 (marked Taf13) were used as controls. All TFIID subunits probed are equally present in all HA co-IPs.
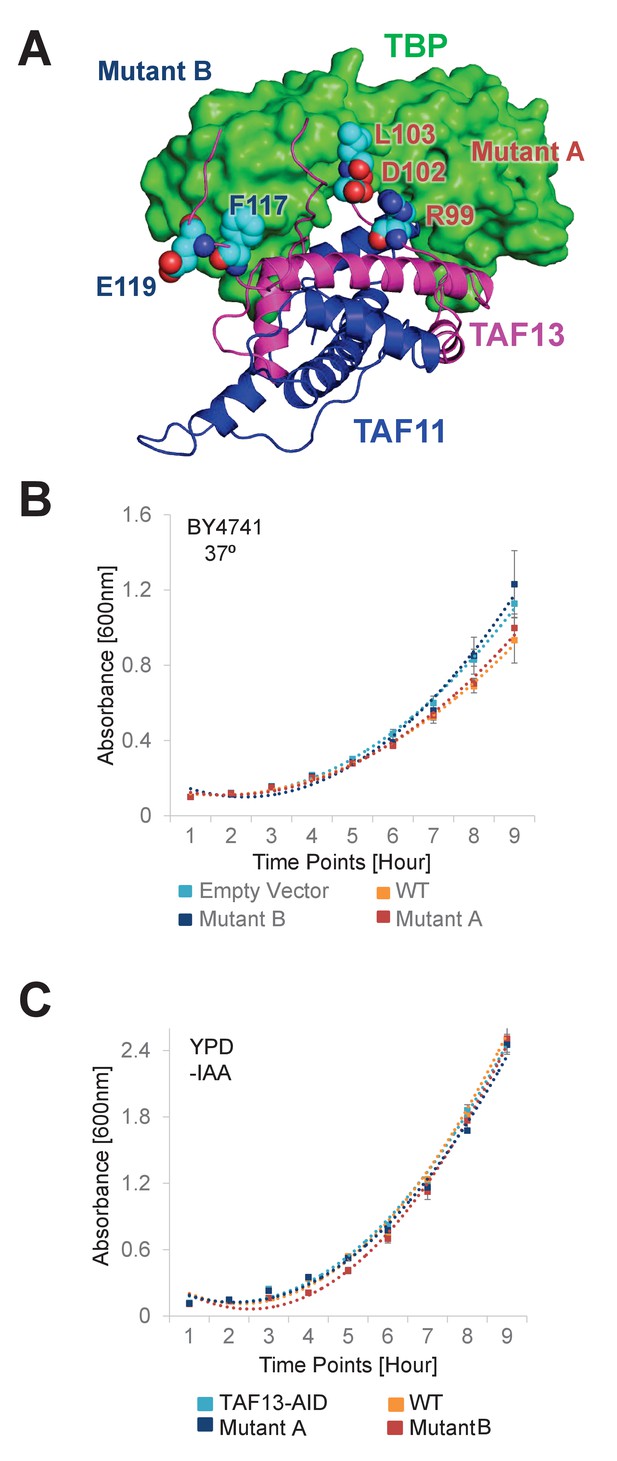
TAF11 CTID mutant studies (A).
Location of amino acid residues mutated in TAF11 Mutant A and Mutant B, respectively, mapped on the TAF11/TAF13/TBP architecture. Mutated amino acids are drawn in van-der-Waals representation. TAF11 and TAF13 are shown as ribbons colored in blue and magenta, respectively. TBP is shown in surface representation colored in green. Amino acids mutated in Mutant A are at the center of the TAF11/TAF13 interaction surface with TBP, while amino acids mutated in Mutant B are more in the periphery of the interaction. (B) Growth curves for the wild-type strain, BY4741, are shown. BY4741 was transformed with empty vector (EV), vector containing the gene for wild-type TAF13 (WT) and the two mutants (A, B), respectively, and grown at 37°C. Polynomial fits for each growth curve are shown as dotted lines. Standard errors of mean (SEM) calculated from three independent experiments are shown as bars. (C) Growth curves for the Taf13 degron strain in absence of IAA are shown. Yeast containing the Taf13-degron fusion was transformed with empty vector (EV), vector containing the gene for wild-type TAF13 (WT) and the two mutants (A, B), respectively, and grown at 37°C. Polynomial fits for each growth curve are shown as dotted lines. Standard errors of mean (SEM) calculated from three independent experiments are shown as bars.
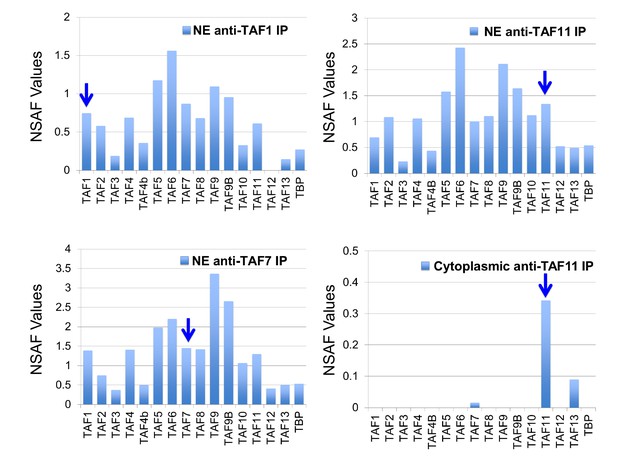
Human TAF/TFIID co-immunoprecipitations.
Orbitrap mass spectroscopic analyses of proteins co-immunoprecipitated from nuclear (NE) or cytoplasmic HeLa cell extracts using mouse monoclonal antibodies against the indicated TAFs. The stoichiometry of the TAFs and TBP in the purified TFIID complexes was calculated by determining normalized spectral abundance factors (NSAFs) (Sardiu et al., 2008; Zybailov et al., 2007). Each column is the average of three independent MS runs. Blue arrows indicate the bait in each immunoprecipitation.
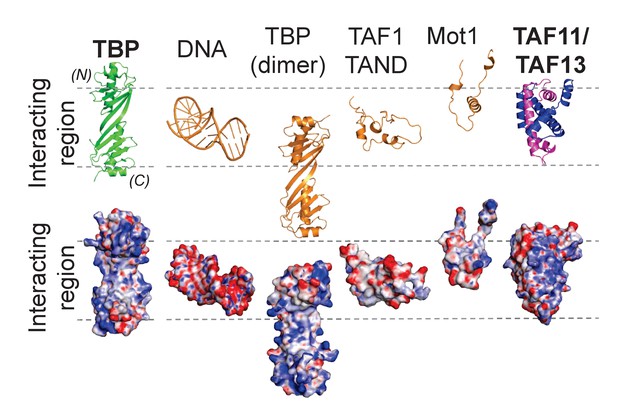
Distinct modes of TBP binding involving the concave DNA-binding surface.
The interaction interfaces of TBP binders are shown in a cartoon representation (top). Interactors shown are TATA-box DNA and protein interactors including the TAF11/TAF13 dimer. The binding modes are further illustrated using space-filling models depicting the corresponding electrostatic surface potentials (bottom). The interacting region representing the concave DNA-binding surface of TBP is delimited by dashed lines. Structures shown are TBP on one hand, as well as TATA-DNA (PDB ID 1CDW), a second copy of TBP from the crystal structure of unliganded TBP (PDB ID 1TBP), TAF1-TAND (PDB ID 1TBA), Mot1 (PDB ID 3OC3) and TAF11/TAF13, respectively, on the other. (N) and (C) indicate N- and C-terminal TBP. The ‘TATA-box mimicry’ by TAF1-TAND in shape and charge distribution is evident. TAF11/TAF13 engage the entire concave DNA-binding surface of TBP including the stir-ups.
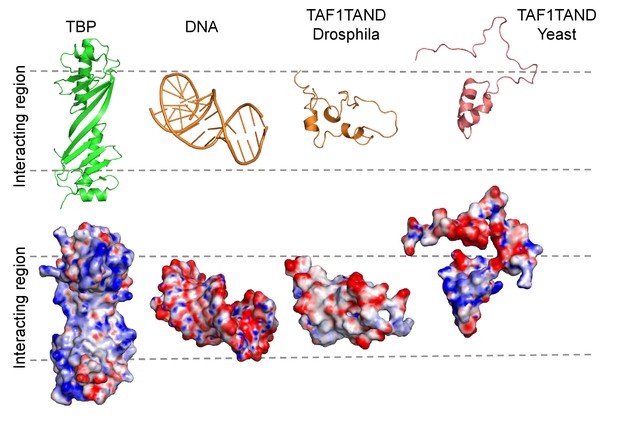
TAF1-TAND TATA-box mimicry in Drosophila and Yeast.
Cartoon representation (top row) and electrostatic surface potential (bottom row) of TBP (left, PDB ID: 1TBP), TATA-DNA (PDB ID: 1CDW), and TAF1-TAND from Drosophila (PDB ID: 1TBA) and Yeast (PDB ID: 4B0A). The DNA-binding surface of TBP is shown (left), as well as the corresponding complementary surfaces of the interactors (right). Red indicates regions with negative charge, blue indicates regions with positive charge. The shape and surface charge similarity between Drosophila TAF1-TAND and TATA-DNA is evident (‘TATA-box mimicry’) but is less pronounced in TAF1-TAND from yeast. Yeast TAF1-TAND binding is more extensive involving also the convex surface of TBP (Table 5).
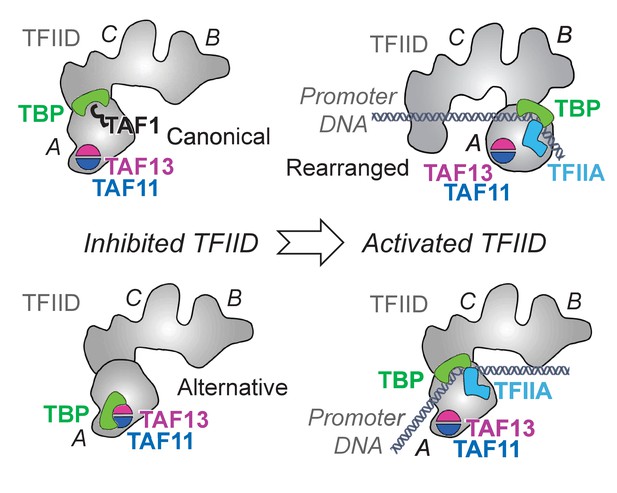
Novel TFIID regulatory state comprising TAF11/TAF13/TBP.
. TFIID is thought to exist in an inhibited ‘canonical’ state with TAF1-TAND bound to TBP’s DNA binding surface (bottom left). Activated states of TFIID (right) bind promoter DNA stabilized by TFIIA (Cianfrocco et al., 2013; Louder et al., 2016; Papai et al., 2010). Our results suggest a novel, alternative TFIID inhibited state comprising TAF11/TAF13/TBP (top left). TFIID is shown in a cartoon representation based on previous EM studies (Cianfrocco et al., 2013; Louder et al., 2016; Papai et al., 2010). TFIID lobes A, B and C are indicated. TAF11/TAF13 are placed in lobe A as suggested by immune-labeling analysis (Leurent et al., 2002). Promoter DNA is colored in grey.
Tables
X-ray data collection and refinement statistics.
https://doi.org/10.7554/eLife.30395.005| TFIIAs-c | |
|---|---|
| Data collection | |
| Space group | P65 |
| Cell dimensions | |
| a, b, c (Å) | 123.3, 123.3, 34.8 |
| a, b, g (°) | 90, 90, 120 |
| Resolution (Å) | 53.4–2.4 |
| Last resolution bin (Å) | 2.52–2.38 |
| Rmeasure (%) | 12.9 (64.8) |
| I / σI | 11.5 (2.72) |
| Completeness (%) | 99.8 (99.9) |
| Multiplicity | 6.7 (6.8) |
| Refinement | |
| Resolution (Å) | 40.36–2.38 (2.44–2.38) |
| No. reflections | |
| Work set | 11859 |
| Free set | 601 |
| Rwork | 0.18 (0.27) |
| Rfree | 0.24 (0.36) |
| No. of atoms | |
| Protein | 1689 |
| Water | 50 |
| r.m.s deviations | |
| Bond lengths (Å) | 0.0223 |
| Bond angles (°) | 2.088 |
-
*Values in parentheses are for highest resolution shell.
Peptide deuteration level changes upon complex formation in HDX-MS
https://doi.org/10.7554/eLife.30395.008| TAF11 | |||
|---|---|---|---|
| Residues numbers | Sequence | Δ%D (15 s)† | Δ%D (120 s)† |
| 93–96 | EKKQ | −8.465 | −7.309 |
| 105–109 | KMQIL | −7.014 | |
| 150–153 | VVIA | −7.295 | |
| TAF13 | |||
| Residues numbers | Sequence | Δ%D (15 s)† | Δ%D (120 s)† |
| 14–31 | NEEIGGGAEGGQGKRKRL | −7.507 | −7.192 |
| 32–35 | FSKE | −8.794 | |
| 36–40 | LRCMM | −7.0 | |
| 86–88 | IVF | −8.598 | |
| 86–89 | IVFL | −10.366 | |
| 97–104 | FARVKDLL | −7.223 | |
| 116–124 | AFDEANYGS | 10.89 | 8.721 |
| TBP | |||
| Residues numbers | Sequence | Δ%D (15 s)† | Δ%D (120 s)† |
| 157–167* | IVPQLQNIVST | 9.038 | |
| 167–174 | TVNLGCKL | −9.675 | −10.171 |
| 193–197 | FAAVI | −14.068 | −12.267 |
| 197–208 | IMRIREPRTTAL | −7.785 | |
| 199–208 | RIREPRTTAL | −9.661 | |
| 209–215 | IFSSGKM | −8.655 | −8.75 |
| 232–244 | KYARVVQKLGFPA | −21.337 | −18.382 |
| 233–242 | YARVVQKLGF | −7.128 | |
| 247–252 | LDFKIQ | −7.271 | |
| 250–256 | KIQNMVG | −8.913 | |
| 259–266 | DVKFPIRL | −11.932 | −10.109 |
| 259–268 | DVKFPIRLEG | −13.79 | −11.479 |
| 260–266 | VKFPIRL | −10.393 | |
| 260–268 | VKFPIRLEG | −16.066 | −13.002 |
| 281–287 | PELFPGL | −7.811 | |
| 285–289 | PGLIY | −8.061 | −7.339 |
| 326–335 | PILKGFRKTT | −7.077 | −7.902 |
-
*TBP peptide with increasing deuteration level upon complex formation.
†Peptides exhibiting changes in deuteration level ≥7% are shown.
Data collection and refinement statistics SAXS.
https://doi.org/10.7554/eLife.30395.009| TAF11/TAF13/TBP | TAF11/TAF13 | |
|---|---|---|
| Data collection parameters | ||
| Beamline | ESRF-BM29 | ESRF-BM29 |
| Beam size at sample | ~700 µm x 700 µm | ~700 µm x 700 µm |
| Wavelength (Å) | 0.931 | 0.931 |
| S range (Å−1) | 0.003–0.497 | 0.003–0.497 |
| Concentration range (mg ml-1) | 0.3–7.11 | 0.53–7.48 |
| Temperature (°C) | 4 | 4 |
| Beamline | ESRF-BM29 | ESRF-BM29 |
| Beam size at sample | ~700 µm x 700 µm | ~700 µm x 700 µm |
| Wavelength (Å) | 0.931 | 0.931 |
| Structural parameters* | ||
| I(0) (arbitrary units) [from P(r)] | 49.63 | 43.65 |
| Rg (Å) [from P(r)] | 41 | 41.2 |
| I(0) (arbitrary units) (from Guinier) | 50.21 ± 0.33 | 44.15 ± 0.08 |
| Rg (Å) (from Guinier) | 40.9 ± 0.6 | 40.3 ± 3.6 |
| Dmax (Å) | 160 | 140 |
| Porod volume estimate (Å3) | 120110 | 89850 |
| Molecular mass Mr [from porod volume] | 70.65 kDa | 52.91 kDa |
| I(0) (arbitrary units) [from P(r)] | 49.63 | 43.65 |
| Rg (Å) [from P(r)] | 41 | 41.2 |
| I(0) (arbitrary units) (from Guinier) | 50.21 ± 0.33 | 44.15 ± 0.08 |
| Rg (Å) (from Guinier) | 40.9 ± 0.6 | 40.3 ± 3.6 |
| Dmax (Å) | 160 | 140 |
| Porod volume estimate (Å3) | 120110 | 89850 |
| Molecular mass Mr [from porod volume] | 70.65 kDa | 52.91 kDa |
-
*Reported for experimental merged data.
Cross-links observed by BS3 CLMS.
https://doi.org/10.7554/eLife.30395.010| TAF11-TBP | |||
|---|---|---|---|
| TAF11 residue | TBP residue | No. of matches | Highest score |
| K97 | K293 | 5 | 9.416 |
| K85 | K293 | 2 | 8.818 |
| K82 | K239 | 2 | 8.314 |
| K135 | K191 | 1 | 8.085 |
| K82 | K293 | 2 | 7.926 |
| K82 | K191 | 1 | 7.684 |
| K131 | K293 | 1 | 6.928 |
| K83 | K232 | 1 | 6.124 |
| K95 | K293 | 1 | 4.145 |
| (9 unique links) | |||
| TAF13-TBP | |||
| TAF13 residue | TBP residue | No. of matches | Highest score |
| K34 | K177 | 1 | 10.578 |
| K34 | K191 | 1 | 7.856 |
| K101 | K232 | 1 | 7.584 |
| K115 | K191 | 1 | 7.519 |
| K115 | K177 | 1 | 5.742 |
| (5 unique links) | |||
| TAF11-TAF13 | |||
| TAF11 residue | TAF13 residue | No. of matches | Highest score |
| K195 | K96 | 2 | 12.795 |
| K85 | K101 | 2 | 12.014 |
| K204 | K96 | 3 | 11.053 |
| K89 | K101 | 5 | 10.98 |
| K204 | K101 | 6 | 10.909 |
| K131 | K34 | 3 | 9.627 |
| K82 | K92 | 2 | 9.321 |
| K197 | K96 | 2 | 9.308 |
| K131 | K111 | 10 | 9.226 |
| K135 | K111 | 2 | 8.371 |
| K131 | K115 | 1 | 8.326 |
| K207 | K92 | 1 | 8.292 |
| K82 | K96 | 1 | 7.981 |
| K85 | K115 | 1 | 7.943 |
| K94 | K101 | 2 | 7.934 |
| K97 | K101 | 1 | 7.867 |
| K204 | K92 | 4 | 6.872 |
| K85 | K92 | 1 | 6.817 |
| K82 | K101 | 1 | 6.437 |
| K206 | K92 | 1 | 5.77 |
| K204 | S74 | 1 | 3.42 |
| K206 | S74 | 1 | 3.42 |
| (22 unique links) | |||
Interaction surfaces in TBP complexes
https://doi.org/10.7554/eLife.30395.021| Interactor | Interface (Å2) |
|---|---|
| TBP dimer | 3010.2 |
| DNA | 4020.1 |
| TAF1-TAND (D. melanogaster) | 3287.8 |
| TAF1-TAND (Yeast) | 7483.1* |
| Mot1 (E. cuniculi) | 4300.0 |
| TAF11/TAF13 | 3305.2 |
-
Calculated with PyMol v1.8.2.0 (www.pymol.org).
*Includes TAND1 and TAND2.
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| gene (Homo Sapiens TAF11) | TAF11 | This paper | TAF11_uniprot:Q15544 | Synthesized by Genescript and cloned with his-tag as described in methods. |
| gene (Homo Sapiens TAF13) | Taf13 (WT; Mutant A; Mutant B) | This paper | TAF13_uniprot:Q15543 | Synthesized by Genescript and cloned with hisTag as described in methods. |
| gene (Homo Sapiens TBP) | TBP;TBPc | This paper | TBP_uniprot:P20226 | Synthesized by Genescript and cloned with hisTag as described in methods. |
| gene (Homo Sapiens TAF1) | TAF1; TAF1_TAND | This paper | TAF1_uniprot:P21675 | Synthesized by Genescript and cloned with MBP tag as described in methods. |
| gene (Homo Sapie TFIIAαβ) | TFIIA; TFIIAαβ | This paper | TFIIAαβ_unirprot: P52655 | Synthesized by Genescript and cloned with his tag as described in methods. |
| gene (Homo Sapiens TFIIAγ) | TFIIA; TFIIAγ | This paper | TFIIAγ_uniprot: P52657 | Synthesized by Genescript and cloned with his tag as described in methods. |
| gene (Saccharomyces cerevisiae Taf13) | Taf13 (WT; Mutant A; Mutant B) | This paper | TAF13_uniprot:P11747 | Synthesized by Genescript and cloned with hisTag as described in methods. |
| strain, strain background is BY4741 | TSA797 | EuroSCARF SRD GmbH, Germany | Y41183 | Tested for growth at permissive and non-permissive temperatures exhibiting expected phenotype. |
| strain, strain background is BY4741 | TSA636 | EuroSCARF SRD GmbH | Y41075 | Tested for growth at permissive and non-permissive temperatures exhibiting expected phenotype. |
| strain, strain background | Taf13-AID auxin- inducible degron strain | Warfield et al., 2017 | Prof. Steven Hahn Lab (http://research.fhcrc.org/hahn/en.html) | |
| cell line (Spodoptera frugiperda 21) | Sf21 | Invitrogen, Carlsbad, CA, USA | ||
| cell line (Henrietta Lachs) | HeLa | Mycoplasma-free HeLa cells obtained from Betty Heller, IGBMC cell line resource www.igbmc.fr | The cell line used was Hela WS, also called HeLa S3 or HeLa CCL-2.2 (RRID:CVCL_0058). The STR Profile report (SOH32553) stated that the "submitted sample (STRA0021) is an exact match to ATCC cell line HeLa CCL- 2.2. The mycoplasma contamination of this cell line is regularly tested using the VenorTMGeM Mycoplasma Detection Kit (from Sigma Aldrich,Catalog Number MP0025). The used cells were mycoplasma free. | |
| antibody | Rabbit anti-HA antibody | Sigma Aldrich | H6908 | |
| antibody | Rabbit anti-Taf5 | From Prof Tony Weil, Vanderbuilt University | ||
| antibody | Rabbit anti-Taf6 | From Prof Tony Weil, Vanderbuilt University | Dilution 1:1000 | |
| antibody | Rabbit anti-Taf8 | From Prof Tony Weil, Vanderbuilt University_ | Dilution 1:2000 | |
| antibody | Rabbit anti-Taf11 | From Prof Tony Weil, Vanderbuilt University | Dilution 1:1000 | |
| antibody | Rabbit anti-TBP | From Prof Tony Weil, Vanderbuilt University_ | Dilution 1:2000 | |
| antibody | Goat anti-rabbit HRP | Thermo Fisher Scientific Inc. | 31463 | Dilution 1:10000 |
| recombinant DNA reagent | MultiBac system | Berger et al. (2004) | ||
| recombinant DNA reagent | MutiBacTAG system | Koehler et al. (2016) | ||
| chemical compound, drug | DiAzKs | Koehler et al. (2016) | ||
| software, algorithm | XDS | Kabsch, 2010 | ||
| software, algorithm | PHASER | McCoy (2007) | ||
| software, algorithm | CCP4 suite | Winn et al. (2011) | ||
| software, algorithm | ATSAS | Petoukhov et al. (2012) | ||
| software, algorithm | MSCovert | Kessner et al. (2008) | ||
| software, algorithm | Xi software | ERI Edinburgh | ||
| software, algorithm | HADDOCK | de Vries et al. (2010) | ||
| software, algorithm | MassLynx | Waters | ||
| software, algorithm | Mass Hunter | Agilent Technologies Inc. | ||
| software, algorithm | HD Examiner | Sierra Analytics Inc. | ||
| DNA for EMSA | AdMLP TATA-DNA | This paper | Synthesized by BioSpring GmbH |
Additional files
-
Source code 1
Crystal structure coordinates of human TFIIAs-c
- https://doi.org/10.7554/eLife.30395.023
-
Transparent reporting form
- https://doi.org/10.7554/eLife.30395.024





