A reverse transcriptase ribozyme
Figures
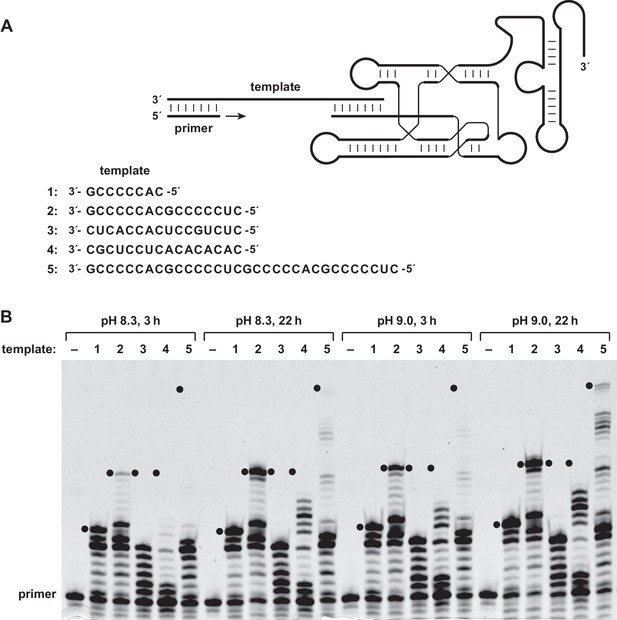
Reverse transcriptase activity of the 24–3 ribozyme.
(A) Secondary structure of the complex formed by the ribozyme, template, and primer (nucleotide sequences are listed in supplementary file 1). The template consists of four regions: primer binding site, sequence to be copied, A3 or A5 spacer, and ribozyme-pairing domain (listed 3´→5´). The ribozyme was tested for its ability to copy five different template sequences (1–5). For sequences of other regions of the template, see supplementary file 1. (B) Extension of a deoxynucleotide-terminated RNA primer on an RNA template. Reaction conditions: 100 nM ribozyme, 125 nM template, 125 nM primer, 2 mM each dNTP, 200 mM MgCl2, pH 8.3 or 9.0, 20°C, 3 or 22 hr. Black dots indicate the expected position of full-length products.
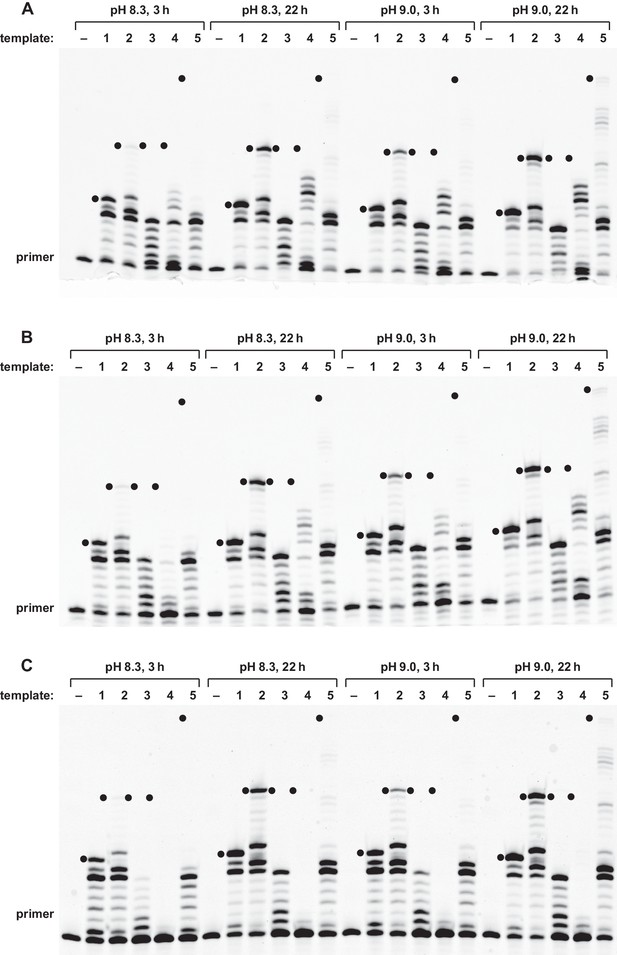
Reverse transcriptase activity of the 24–3 ribozyme.
Extension of (A) all-RNA primer, (B) deoxynucleotide-terminated RNA primer, or (C) all-DNA primer on an RNA template. Reaction conditions: 100 nM ribozyme, 125 nM template 1, 125 nM primer, 2 mM each dNTP, 200 mM MgCl2, pH 8.3 or 9.0, 20°C, 3 or 22 hr. Black dots indicate the expected position of full-length products.
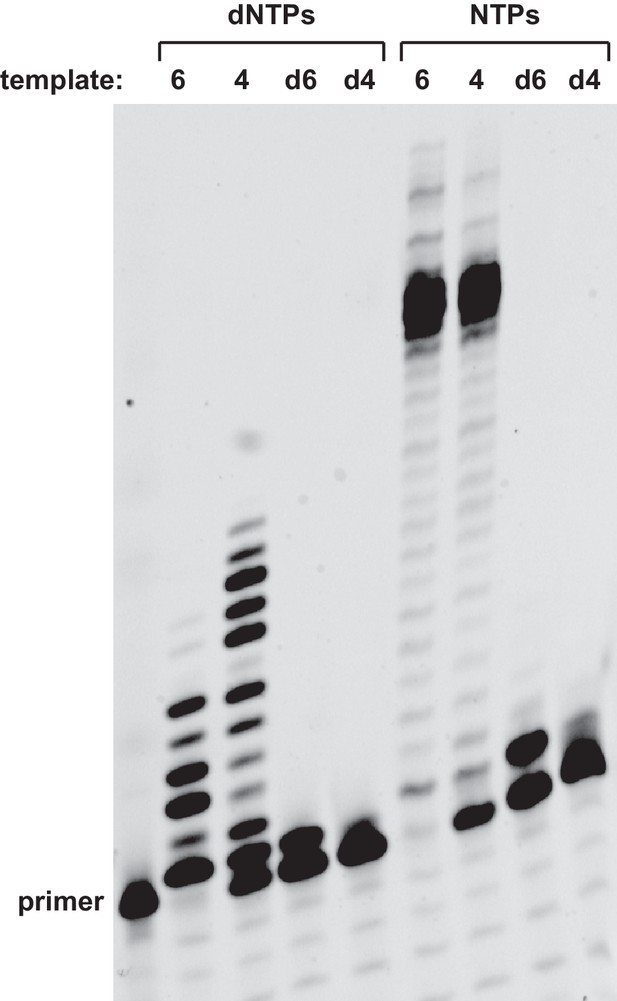
Lack of DNA-dependent polymerase activity of the 24–3 ribozyme.
Extension of an all-RNA primer on either an RNA or DNA template, employing either dNTPs or NTPs. The sequences of the RNA templates (6 and 4) and corresponding DNA templates (d6 and d4) are listed in supplementary file 1. Reaction conditions: 100 nM ribozyme, 125 nM template, 125 nM primer, 2 mM each dNTP or NTP, 200 mM MgCl2, pH 8.3, 20°C, 21 hr.
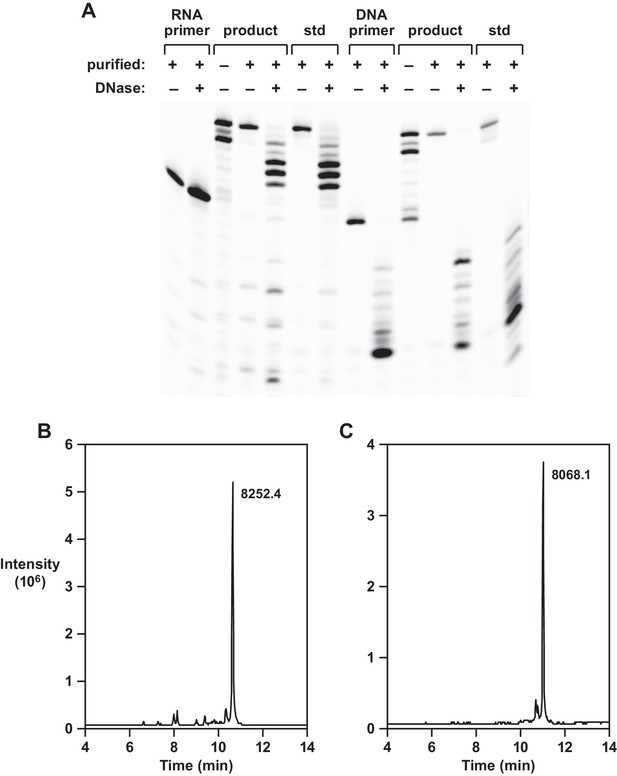
Analysis of reverse transcription products.
(A) Partial DNase I digestion of full-length products obtained using template 1, in comparison to authentic materials, with either an RNA primer (left 7 lanes) or DNA primer (right 7 lanes). For the RNA-primed reaction, only the extended portion is cleaved; for the DNA-primed reaction, both the primer and extended portion are cleaved. (B,C) LC/MS analysis of purified full-length products obtained using template 1 and either an RNA or a DNA primer, respectively.
-
Figure 2—source data 1
LC/MS analysis of full-length products.
- https://doi.org/10.7554/eLife.31153.008
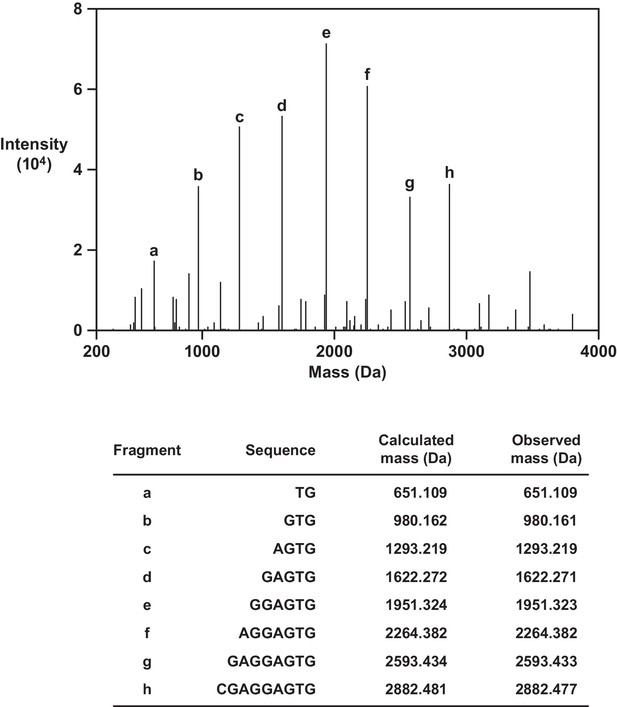
High-resolution MS/MS analysis of reverse transcription product.
Extension of an all-RNA primer on template four yielded a product with a calculated mass of 8874.432 and observed mass of 8874.441. The parent ion was fragmented at internucleotide linkages within the DNA portion of the molecule to generate secondary ions that were analyzed by tandem MS. Successive fragments a–h correspond to 3´-terminal subsequences within the DNA portion of the molecule.
-
Figure 2—figure supplement 1—source data 1
High-resolution MS/MS analysis of reverse transcription product (related to Figure 2—figure supplement 1).
- https://doi.org/10.7554/eLife.31153.009
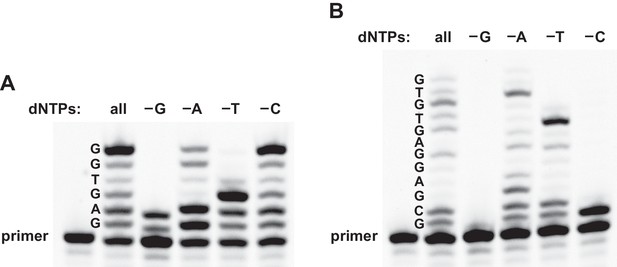
Reverse transcription in reaction mixtures lacking one of the four dNTPs.
(A) For the sequence 5´-GAGTGG-3´ (template 3), expected-length material was obtained in the presence of either all four dNTPs or a mixture lacking dCTP, expected-length material of altered mobility was obtained in a mixture lacking dATP, and only partial-length material was obtained in a mixture lacking either dGTP or TTP. (B) For the sequence 5´-GCGAGGAGTGTG-3´ (template 4), expected-length material was obtained in the presence of all four dNTPs, expected-length material of altered mobility was obtained in a mixture lacking dATP, and only partial-length material was obtained in a mixture lacking dGTP, TTP, or dCTP. Reaction conditions: 100 nM ribozyme, 125 nM template, 125 nM primer, 2 mM dNTPs, 200 mM MgCl2, pH 8.3, 20°C, 22 hr.
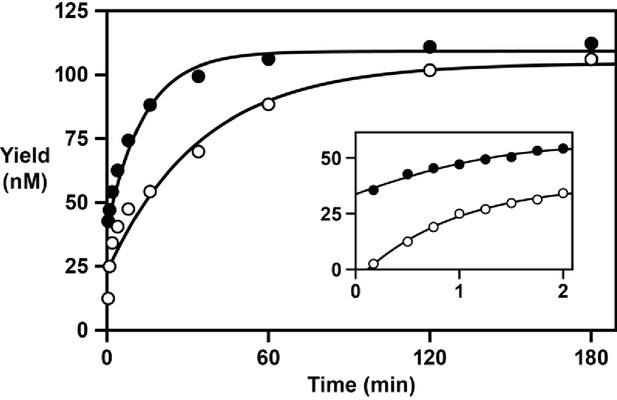
RNA-dependent RNA and DNA polymerase activity of the 24–3 ribozyme.
Time course of the reaction using either NTPs (filled circles) or dNTPs (open circles), measuring the rate of single-nucleotide addition to a deoxynucleotide-terminated primer on template 1. The data were fit to a double exponential rise to maximum (r = 0.996 for RNA polymerization; r = 0.983 for DNA polymerization). Inset depicts the data over the first 2 min of the reaction. Reaction conditions: 100 nM ribozyme, 125 nM template, 125 nM primer, 2 mM NTPs or dNTPs, 200 mM MgCl2, pH 8.3, 20°C.
-
Figure 4—source data 1
RNA-dependent RNA and DNA polymerase activity.
- https://doi.org/10.7554/eLife.31153.012
Additional files
-
Supplementary file 1
Sequences of RNA and DNA molecules used in this study.
- https://doi.org/10.7554/eLife.31153.013
-
Transparent reporting form
- https://doi.org/10.7554/eLife.31153.014





