Salient experiences are represented by unique transcriptional signatures in the mouse brain
Figures
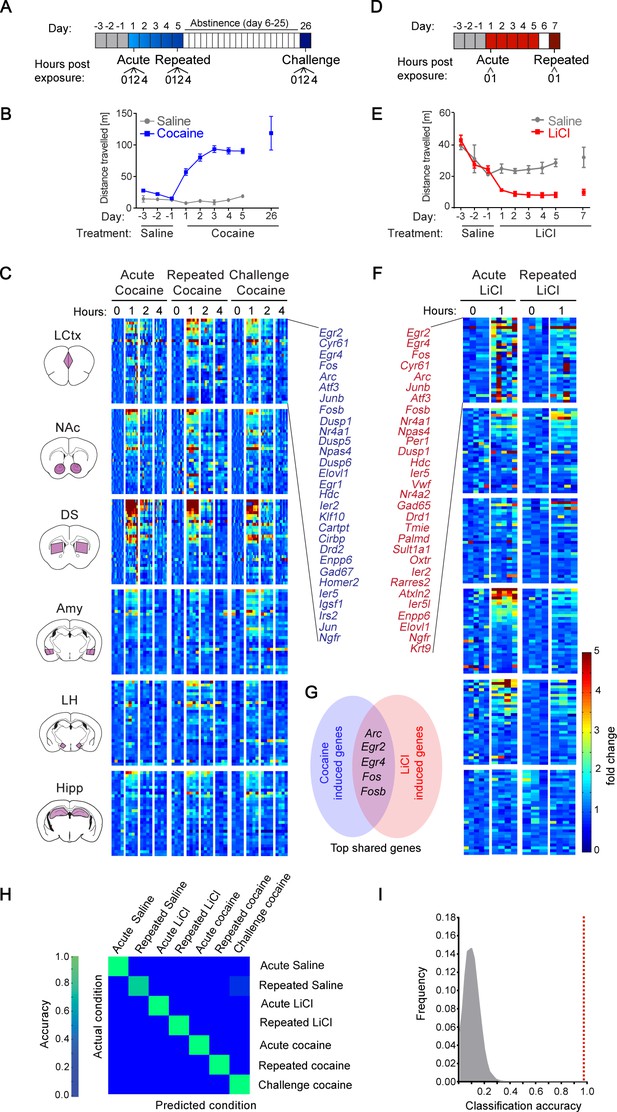
Transcriptional signatures representing recent experience.
(A) Schematic of experimental paradigm for cocaine sensitization. Mice were exposed to cocaine (i.p., 20 mg/kg) or saline, either acutely, repeatedly or re-exposed following abstinence (challenge), with transcriptional dynamics studied at 0, 1, 2 or 4 hr. (B) Locomotor activity of mice following acute, repeated or challenge cocaine experiences (compared to saline). Sample size: acute and repeated saline n = 4; acute cocaine n = 30; repeated cocaine n = 22; challenge cocaine n = 19 mice. Results indicate mean ± s.e.m. (C) Expression matrix of IEG induction dynamics following cocaine experiences. Individual animals are represented in columns sorted according to time points of cocaine experiences [sample numbers per time point - LCtx: limbic cortex (n = 5–11), NAc: nucleus accumbens (n = 5–12), DS: dorsal striatum (n = 5–12), Amy: amygdala (n = 3–4), LH: lateral hypothalamus (n = 2–4), Hipp: hippocampus (n = 2–4)]. Fold induction is graded from blue (low) to red (high). Genes represented were induced at least 2-fold over control in any one of the brain regions studied. Genes were sorted according to peak induction in the DS. (D) Schematic of experimental paradigm for LiCl exposure. Mice were exposed to LiCl (i.p.) or saline, either acutely (150 or 250 mg/kg) or repeatedly (150 mg/kg). (E) Locomotor activity of mice following acute or repeated LiCl exposure (as in panel C). Sample size: n = 4–5. (F) Expression matrix of IEG induction dynamics following LiCl experiences (n = 4–5). Genes were sorted according to peak induction in the Amy. (G) Venn diagram indicating the identity of the most robustly induced genes common to cocaine and LiCl experiences (most appearances and lowest variance). (H) Confusion matrix representing the classification accuracy of decoding the recent experience (acute, chronic and challenge cocaine, acute and repeated LiCl and acute and repeated saline) of individual mice based on expression of Arc, Egr2, Egr4, Fos and Fosb induction in the LCtx, NAc, DS, Amy and LH using a KNN classifier. Accuracy is scaled from blue to green, with bright green corresponding to 100% accuracy (n = 37 mice). Overall accuracy = 97.3%. (I) Results of a permutation test for verifying classification. A randomization test was performed, in which the classifier was run on 105 random permutations of the association of individual mice to the appropriate experience, and the frequency of classification accuracies is plotted in grey, while the red dotted line represents the classification accuracy obtained for non-randomized data (97.3%).
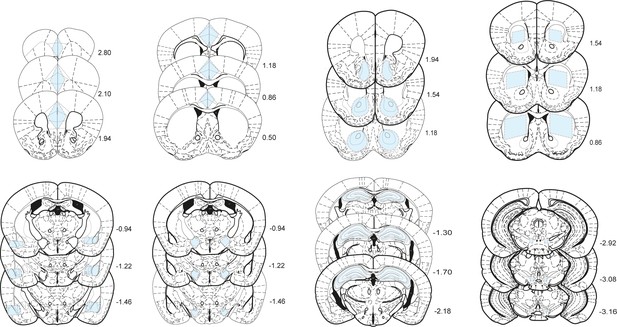
Boundaries of dissected structures.
Shaded regions in blue represent areas cut out from 400 µm slices for LCtx (limbic cortex), NAc (nucleus accumbens), DS (dorsal striatum), Amy (amygdala), LH (lateral hypothalamus), and Hipp (hippocampus) and 200 µm slices for VTA (ventral tegmental area), providing reproducible tissue samples for analysis of transcription. Distance from Bregma on the rostral-caudal axis is indicated in mm.
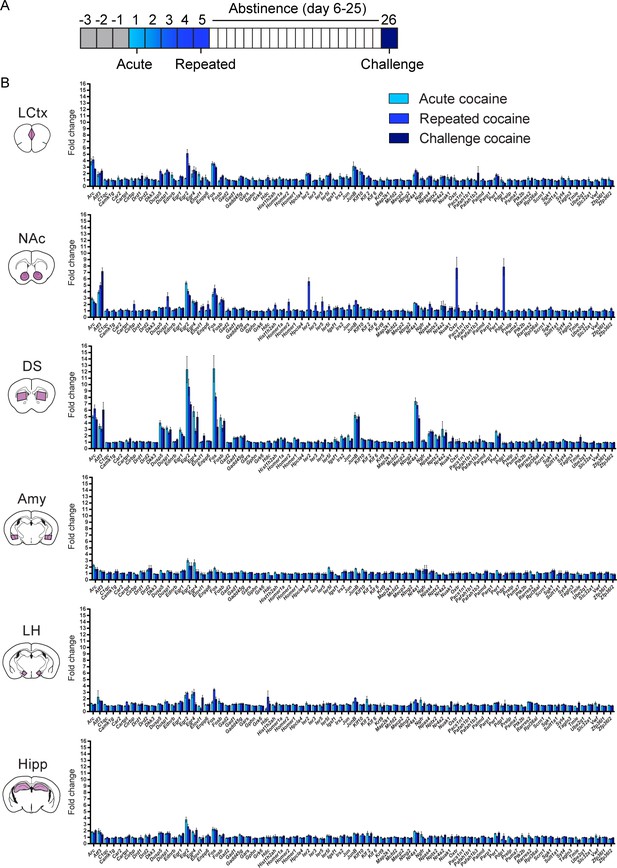
Acute, repeated and challenge cocaine experiences induce distinct transcriptional programs across brain structures.
(A) Schematic of the experimental paradigm. (B) Average transcriptional induction of 78 genes one hour following acute, repeated or challenge cocaine experiences in six brain nuclei [LCtx: limbic cortex (n = 6–10), NAc: nucleus accumbens (n = 6–10), DS: dorsal striatum (n = 6–10), Amy: amygdala (n = 4), LH: lateral hypothalamus (n = 4), Hipp: hippocampus (n = 4)]. Genes are sorted alphabetically. Results indicate mean ± s.e.m.
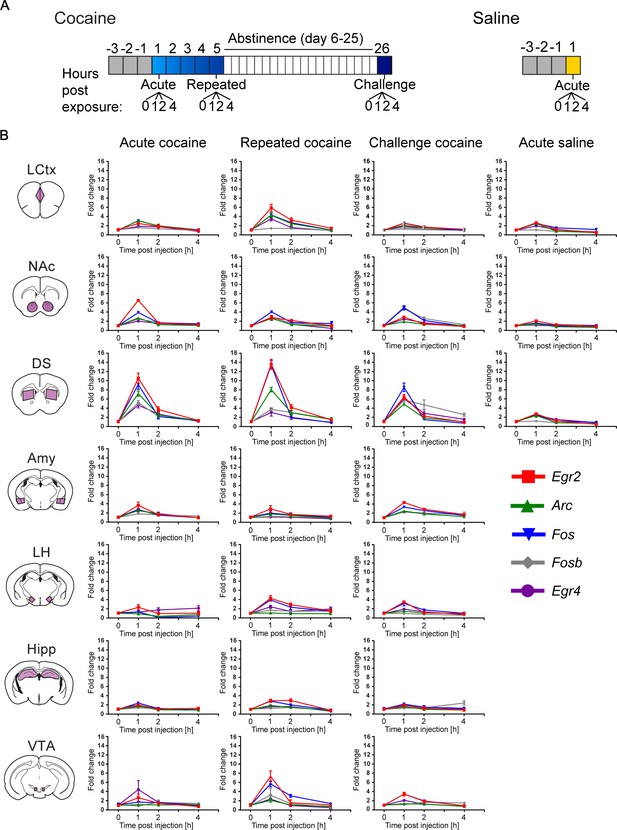
Time course of transcriptional induction of a minimal set of markers representing cocaine-induced transcriptional dynamics.
(A) Schematic of the experimental paradigms. (B) Arc, Egr2, Egr4, Fos and Fosb induction (0, 1, 2, 4 hr) following acute, repeated and challenge cocaine in seven brain nuclei (LCtx: limbic cortex, NAc: nucleus accumbens, DS: dorsal striatum, Amy: amygdala, LH: lateral hypothalamus, Hipp: hippocampus and VTA: ventral tegmental area), as well as acute saline (in the LCtx, NAc and DS). Subject numbers for each time point: n = 6–16 (LCtx, NAc and DS), n = 4–8 (Amy, LH and Hipp) and n = 2–4 (VTA) for acute, chronic and challenge cocaine. Acute saline included 6–8 animals in each time point for LCtx, NAc and DS. Results indicate mean ± s.e.m.
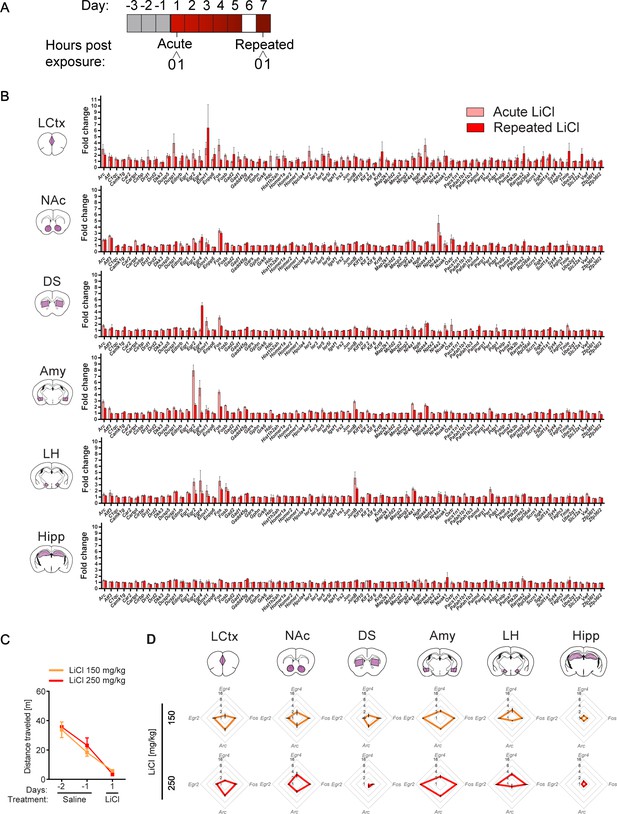
Transcriptional representation of LiCl experience.
(A) Schematic of experimental paradigm. (B) Comparison of average transcriptional induction of 78 genes following acute LiCl (n = 5) or repeated LiCl (n = 4) at 1 hr time point, in limbic cortex (LCtx), nucleus accumbens (NAc), dorsal striatum (DS), amygdala (Amy), lateral hypothalamus (LH), hippocampus (Hipp). Genes are sorted alphabetically. Results indicate mean ± s.e.m. (C) Locomotor activity of mice following acute i.p. exposure to 150 mg/kg (n = 4) or 250 mg/kg LiCl (n = 4). (D) Radar plots representing the transcriptional response 1 hr following 150 mg/kg and 250 mg/kg of LiCl in the LCtx, NAc, DS, Amy, LH and Hipp. Results indicate mean ± s.e.m.
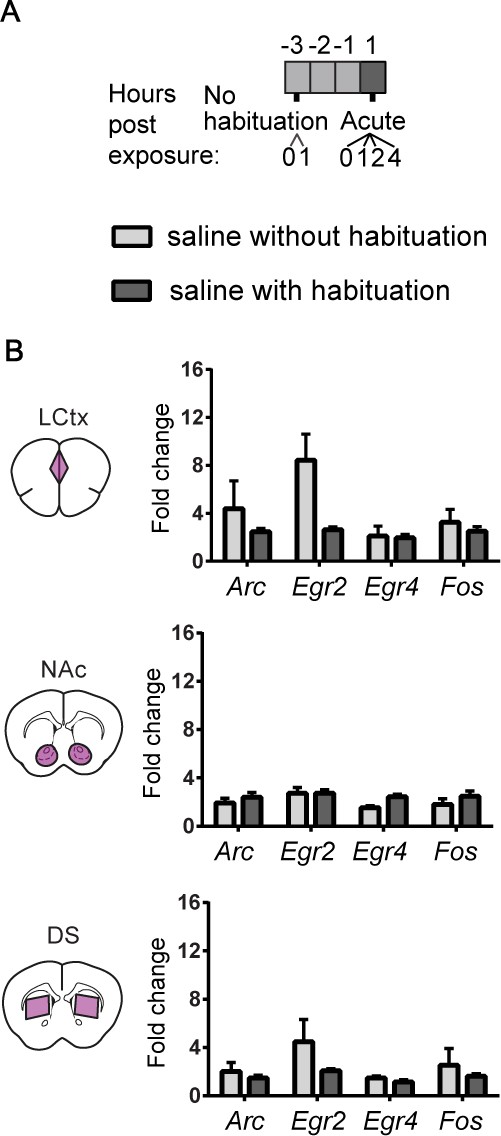
Characterization of repeated saline exposures illustrates the effect of habituation on induced transcription.
(A) Experimental paradigm for i.p. saline injections. (B) Bar graphs demonstrating the transcriptional induction of Arc, Egr2, Egr4 and Fos in the LCtx, NAc, and DS (limbic cortex, nucleus accumbens, and dorsal striatum, respectively) 1 hr following i.p. injection of saline with no prior habituation (n = 4), or following three days of habituation (n = 5). Results show mean ± s.e.m.
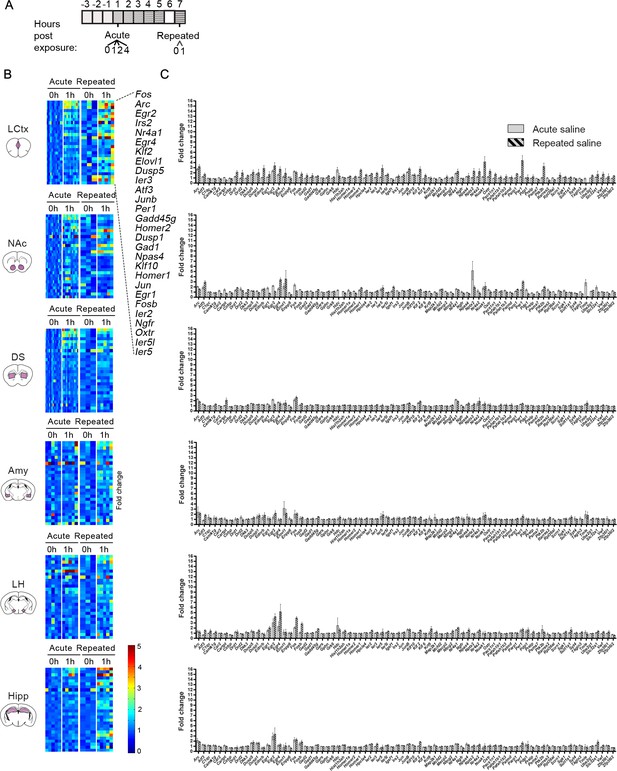
Characterization of the transcription induced by acute and repeated saline experiences.
(A) Schematic of the experimental paradigm of i.p. injections of saline. (B) Expression matrix of the transcriptional induction of IEGs following acute or repeated saline experiences. Each column represents the transcriptional profile of an individual mouse. Sample numbers for each time point for acute saline: n = 10–12 in LCtx (limbic cortex); n = 12–13 in NAc (nucleus accumbens) and DS (dorsal striatum); n = 5 in Amy (amygdala), LH (lateral hypothalamus), Hipp (hippocampus). Sample numbers for each time point for repeated saline: n = 3–5 in LCtx, NAc, DS, Amy, LH and Hipp. Transcriptional induction is graded from blue (low) to red (high). Genes represented were induced on average at least 1.5-fold following either cocaine or LiCl experiences in any one of the studied brain nuclei. Genes are sorted according to peak induction in the LCtx. (C) Average transcriptional induction of 78 genes 1 hr following acute or repeated saline experience in the LCtx, NAc, DS, Amy, LH and Hipp. Genes are sorted alphabetically. Sample sizes: LCtx: n = 10 for acute saline and n = 5 for repeated saline; NAc: n = 12 for acute saline and n = 5 for repeated saline; DS: n = 12 for acute saline and n = 5 for repeated saline; Amy: n = 5 for acute saline and n = 5 for repeated saline; LH: n = 5 for acute saline and n = 5 for repeated saline; Hipp: n = 6 for acute saline and n = 5 for repeated saline. Results indicate mean ± s.e.m.
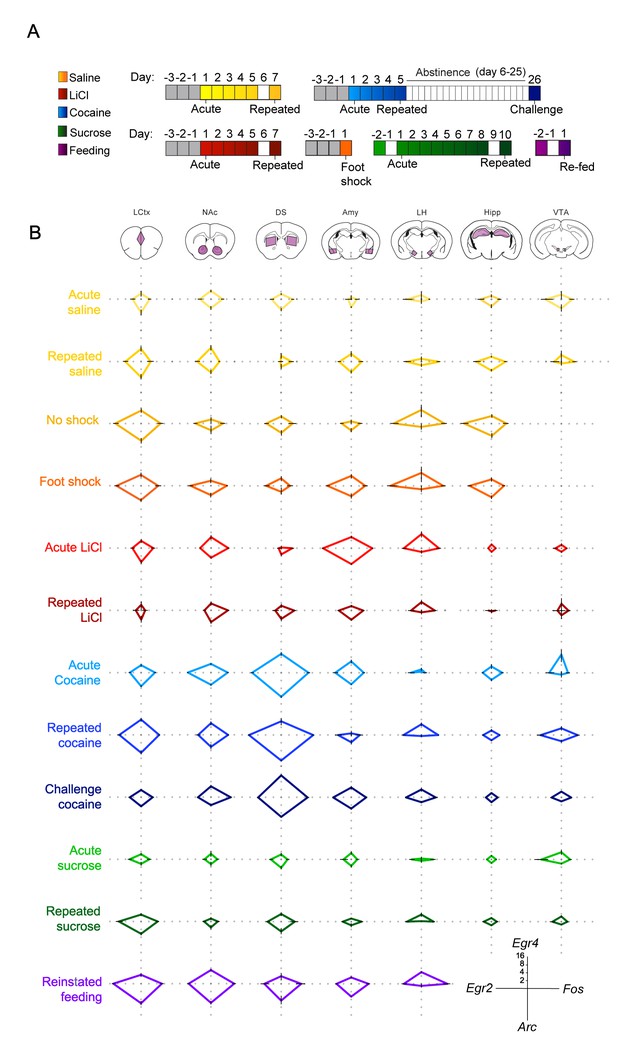
Salient experiences are represented by unique transcriptional signatures.
(A) Schematic of experimental paradigms. Experiences analyzed include saline (acute and repeated); foot shock (acute shock and no-shock controls exposed to the same environment); LiCl (acute and repeated); cocaine (acute, repeated and challenge following abstinence); sucrose (acute and repeated) and reinstatement of feeding (following 18 hr of deprivation). (B) Radar plots representing the transcriptional induction of Arc, Egr2, Egr4 and Fos across seven brain structures 1 hr after the different experiences [LCtx: limbic cortex (n = 4–14), NAc: nucleus accumbens (n = 4–14), DS: dorsal striatum (n = 4–14), Amy: amygdala (n = 4–9), LH: lateral hypothalamus (n = 3–9), Hipp: hippocampus (n = 4–9); VTA: ventral tegmental area (n = 2–8)]. Results are shown in log2 scale as mean ± s.e.m. of induction over baseline control.
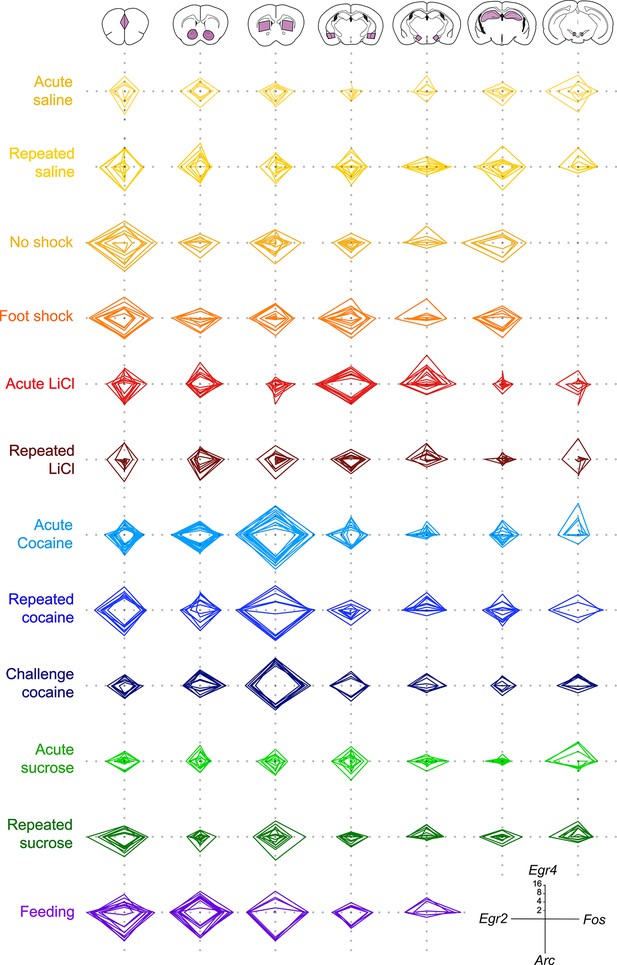
Low variability of the individual transcriptional representations of recent experience.
Radar plots representing the transcriptional induction of Arc, Egr2, Egr4 and Fos, in seven brain nuclei [LCtx: limbic cortex (n = 4–14), NAc: nucleus accumbens (n = 4–14), DS: dorsal striatum (n = 4–14), Amy: amygdala (n = 4–9), LH: lateral hypothalamus (n = 3–9), Hipp: hippocampus (n = 4–8), VTA: ventral tegmental area, (n = 2–8)] 1 hr following the defined experiences. Results indicate fold induction at 1 hr in log2 scale, of individual subjects over relevant control conditions.
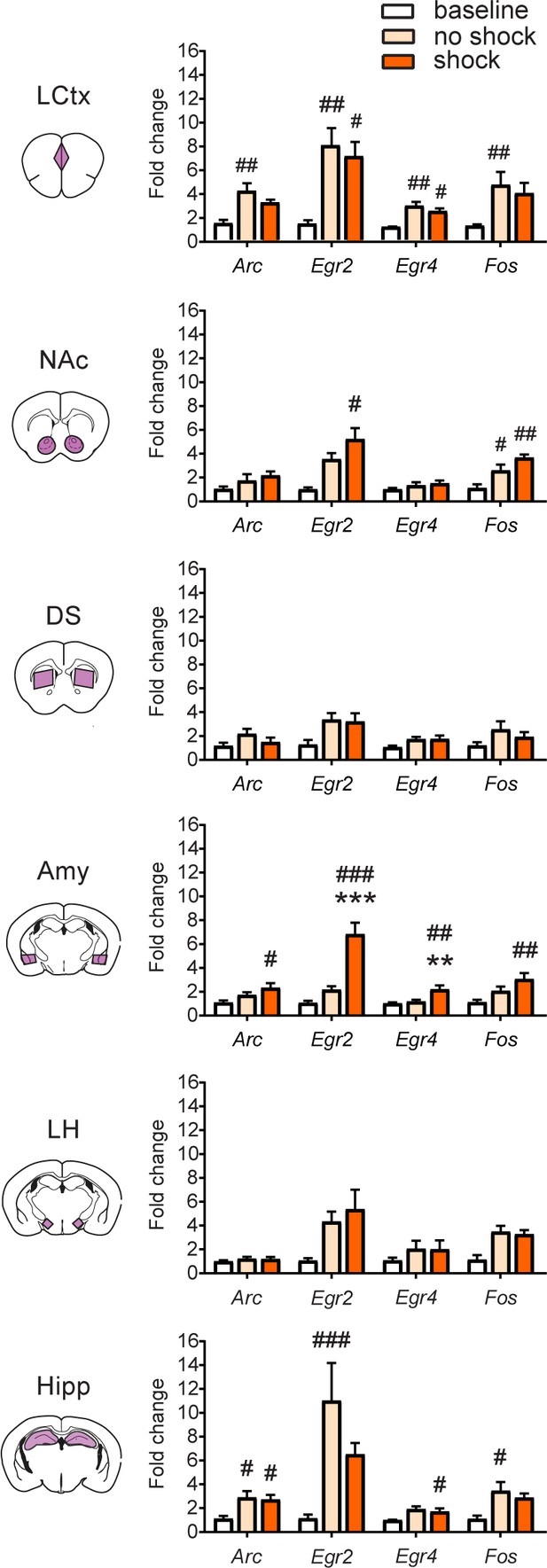
Transcriptional representation of negative valence in the amygdala.
Comparison of average transcriptional induction of Arc, Egr2, Egr4 and Fos in the LCtx, NAc, DS, Amy, LH and Hipp, 1 hr following acute exposure to the fear-conditioning chamber (no shock) or brief foot shock within the chamber (shock), to mice not exposed to chamber (baseline controls). Sample sizes across structures n = 3–8. Results indicate mean ± s.e.m. One-way ANOVA with Tukey post hoc comparison: **p<0.01, ***p<0.001 vs no shock control, #p<0.05, ##p<0.01, ###p<0.001 vs baseline control.
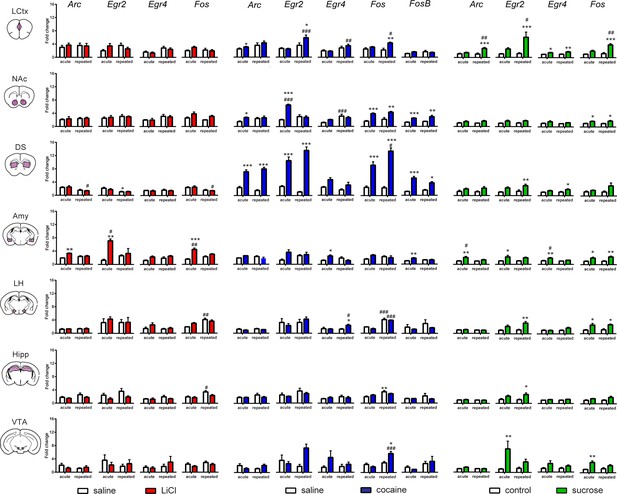
Transcriptional representation of habituation and reinforcement.
Comparison of the transcriptional profiles induced by acute and repeated aversive (LiCl) and rewarding (cocaine and sucrose) experiences reveal distinctions and commonalities in the encoding of an anticipated experience of defined valence. Results indicate mean ± s.e.m Sample numbers for each time point- LCtx: limbic cortex (n = 6–14), NAc: nucleus accumbens (n = 6–14), DS: dorsal striatum (n = 6–14), Amy: amygdala (n = 4–9), LH: lateral hypothalamus (n = 3–9), Hipp: hippocampus (n = 4–9); VTA: ventral tegmental area (n = 2–8)]. Two-way ANOVA followed by Tukey post hoc comparison: *p<0.05, **p<0.01, ***p<0.001 vs control (saline or water), #p<0.05, #p<0.01, ###p<0.001 vs acute/repeated treatment.
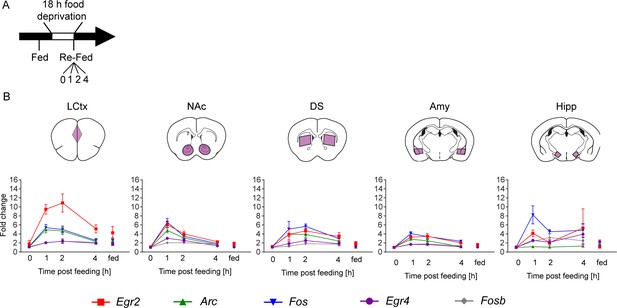
Reinstatement of feeding is represented by robust transcriptional dynamics.
(A) Schematic of experimental paradigm for reinstatement of feeding. Mice were continuously fed or food deprived for 18 hr, followed by analysis of transcription at 0, 1, 2, 4 hr after reinstatement of feeding. (B) Time course (at 0, 1, 2, 4 hr following reinstatement of feeding, in comparison to continuously fed mice, ‘fed’) of transcriptional induction of Arc, Egr2, Egr4, Fos and Fosb in five brain nuclei. Data for each group was normalized to the 0 hr baseline controls. Sample numbers for each time point: LCtx, limbic cortex n = 10–11; NAc, nucleus accumbens n = 10–11; DS, dorsal striatum n = 7–9; Amy, amygdala n = 3–4; LH, lateral hypothalamus n = 2–4. Results indicate mean ± s.e.m.
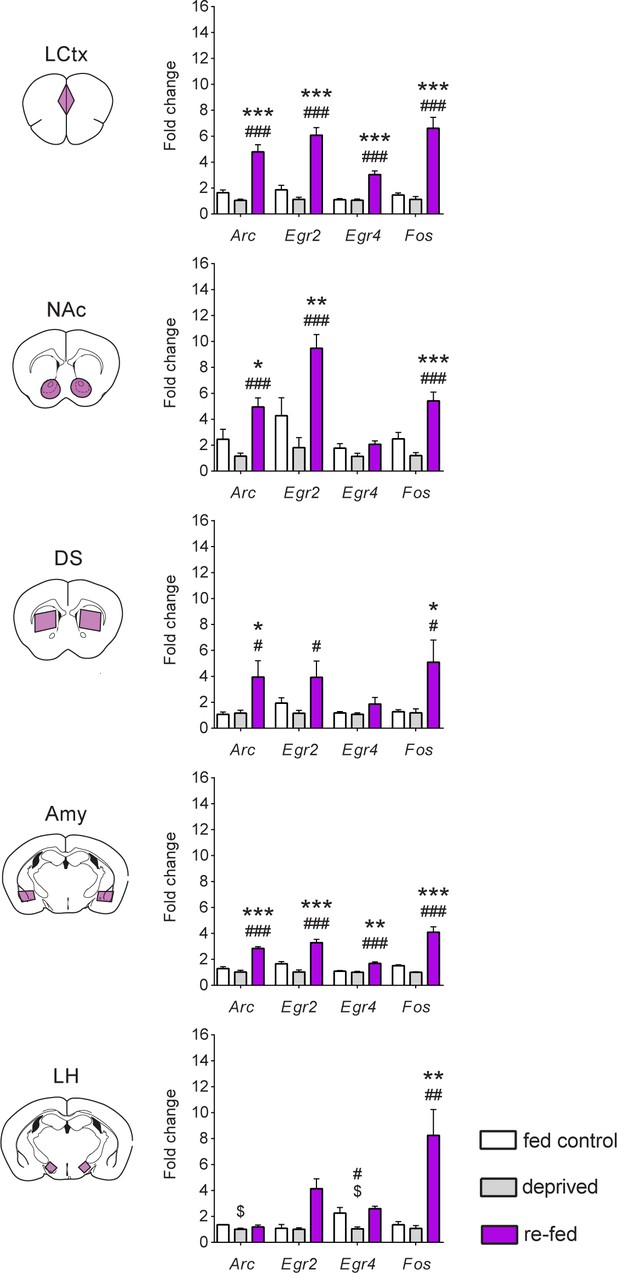
Reinstatement of feeding is represented by robust transcriptional dynamics.
Comparison of transcriptional induction of Arc, Egr2, Egr4 and Fos, at 0 (deprived) and 1 hr following reinstatement of feeding (re-fed), and continuously fed mice (fed control) across five brain nuclei. Sample numbers for each time point: LCtx, limbic cortex n = 10–11; NAc, nucleus accumbens n = 10–11; DS, dorsal striatum n = 7–9; Amy, amygdala n = 3–4; LH, lateral hypothalamus n = 2–4. Results indicate mean ± s.e.m. One-way ANOVA followed by Tukey post hoc comparison: *p<0.05, **p<0.01, ***p<0.001 vs fed control; #p<0.05, ##p<0.01, ###p<0.001 vs deprived; $p<0.05 vs fed control.
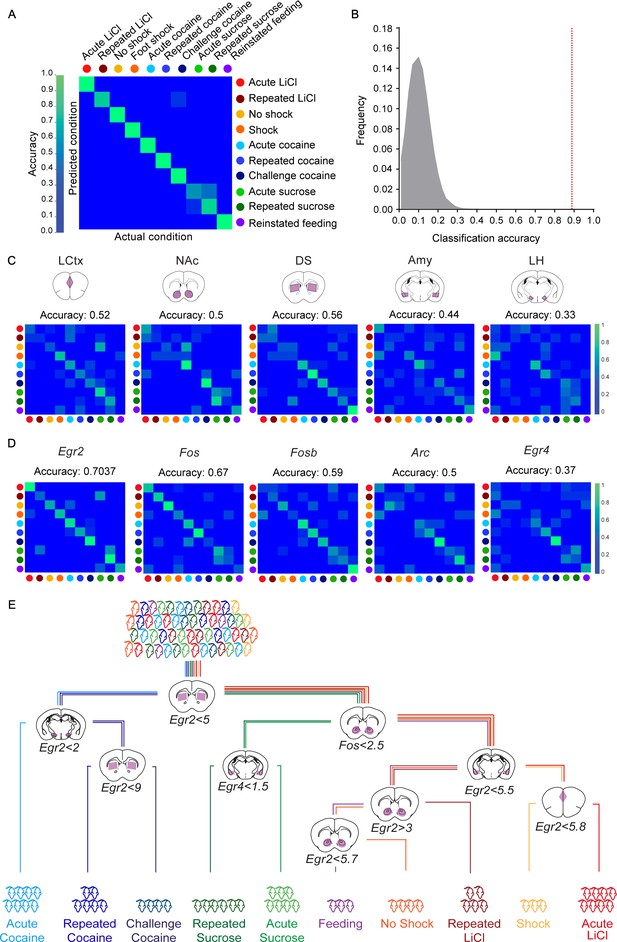
Decoding the recent experience of individual mice from minimal transcriptional signatures.
(A) Confusion matrix representing the classification accuracy (90.7%) of decoding the recent experience of individual mice based on 25 features. Efficiency is scaled from blue to green, with bright green corresponding to 100% efficiency (n = 54 mice). (B) Verification of classification validity. A randomization test was performed, in which the classifier was run on 105 random permutations of the association of individual mice to the appropriate experience, and the frequency of classification accuracies is plotted in grey, while the red dotted line represents the classification accuracy obtained for non-randomized data (90.7%). (C, D) Confusion matrices representing the classification accuracy of decoding utilizing transcriptional measurements from individual brain structures (five genes in one structure, (C) or individual genes (single genes across five structures, (D). Dots represent the identity of the experience, color-coded according to A. X and Y axes denote the actual and predicted conditions. (E) A decision tree enabling the classification of mice according to experience by minimal gene expression (one of many possible trees which can equivalently segregate the data). Mice are classified based on features that enable maximal segregation at each internal node. The thresholds define the allocation of mice to the left branch of the tree at each bifurcation. Mice are color-coded according to experience.
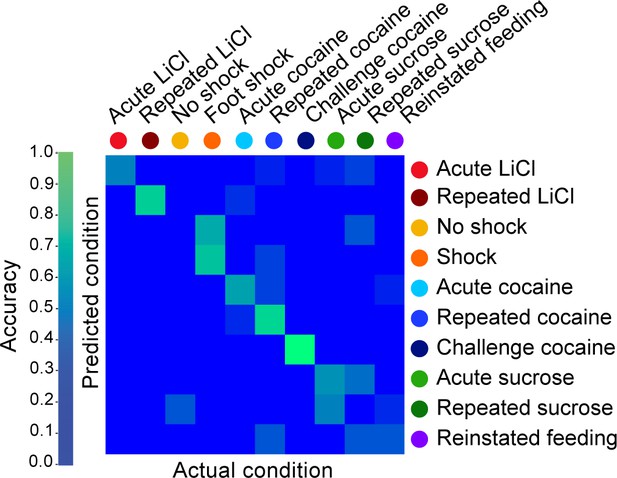
Decoding the recent experience of individual mice from averaged gene expression across different brain regions.
Confusion matrix representing the classification accuracy (55.6%) of decoding the recent experience of individual mice based on averaged transcriptional induction of Arc, Egr2, Egr4, Fos and Fosb) across five structures (LCtx, limbic cortex; NAc, nucleus accumbens; DS, dorsal striatum; Amy, amygdala; LH, lateral hypothalamus and Hipp, hippocampus). Classification was performed using k-Nearest Neighbor Algorithm with a leave one out approach. Accuracy is scaled from blue to green, with bright green corresponding to 100% accuracy (n = 54 mice).
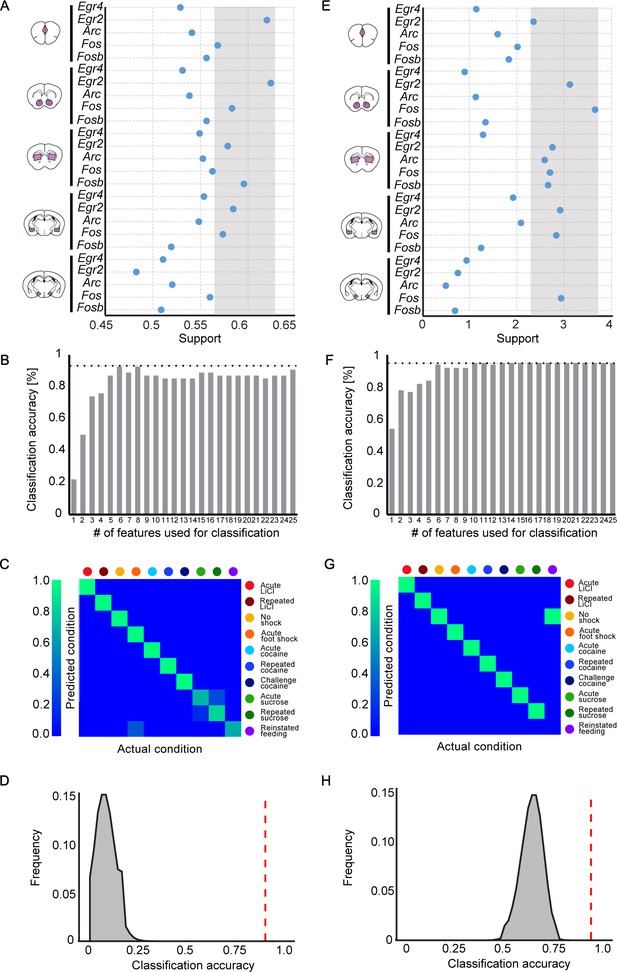
Feature selection to identify the features contributing most significantly to decoding.
(A, E) Selection of features (expression of IEG in a brain structure) with the highest potential to contribute to the classification of recent experiences (‘support’). Support was defined by running 106 classifiers with varying composition of features, and calculating the mean accuracy for each feature over all the iterations to which it contributed, using the Random k-Nearest Neighbors algorithm (RKNN; A) or by calculating the decrease in Gini gain for each feature, using the Random Forest algorithm (RF; E). The grey zone defines the features with the highest support values, selected for classification. (B, F) Accuracy of classification obtained with an increase in the number of features. For each # (subset) of features, the features ranking highest in support were chosen and a KNN (B) or RF (F) classifier was evaluated using only those features. (C, G) Confusion matrix representing the classification accuracy of decoding the recent experience of individual mice based on the most informative features for KNN (Accuracy-93.6%; C) or most informative features for RF (Accuracy: 94.4%; G). Efficiency is scaled from blue to green, with bright green corresponding to 100% efficiency (n = 54 mice). (D, H) Verification of classification validity. A randomization test was performed, in which the classifier was run on 106 random permutations of the association of individual mice to the appropriate experience, and the frequency of classification accuracies is plotted in grey, while the red dotted line represents the classification accuracy obtained for non-randomized data.
Additional files
-
Source Code 1
Classfication Tests_averageGenes
- https://doi.org/10.7554/eLife.31220.019
-
Source Code 2
Classfication Tests_averageRegions
- https://doi.org/10.7554/eLife.31220.020
-
Source Code 3
Classfication Tests_pcaOverRegions
- https://doi.org/10.7554/eLife.31220.021
-
Source Code 4
ClassificationAndConfusionMatrices_confMat
- https://doi.org/10.7554/eLife.31220.022
-
Source Code 5
Classification And Confusion Matrices_knnEvaluate
- https://doi.org/10.7554/eLife.31220.023
-
Source Code 6
Feature Selection RKNN_find Features
- https://doi.org/10.7554/eLife.31220.024
-
Source Code 7
Feature Selection RKNN_find Interesting Genes
- https://doi.org/10.7554/eLife.31220.025
-
Source Code 8
Features Selection Random Forest_DecisionTree
- https://doi.org/10.7554/eLife.31220.026
-
Source Code 9
Linear Projections_RegionsScatter
- https://doi.org/10.7554/eLife.31220.027
-
Source Code 10
Linear Projections
- https://doi.org/10.7554/eLife.31220.028
-
Source Code 11
Randomization_permTest
- https://doi.org/10.7554/eLife.31220.029
-
Supplementary file 1
Primer sequences and efficiency calculations.
- https://doi.org/10.7554/eLife.31220.030
-
Supplementary file 2
Raw data.
- https://doi.org/10.7554/eLife.31220.031
-
Supplementary file 3
Statistics.
- https://doi.org/10.7554/eLife.31220.032
-
Supplementary file 4
Animal numbers
- https://doi.org/10.7554/eLife.31220.033
-
Transparent reporting form
- https://doi.org/10.7554/eLife.31220.034





