Stabilization and structural analysis of a membrane-associated hIAPP aggregation intermediate
Figures
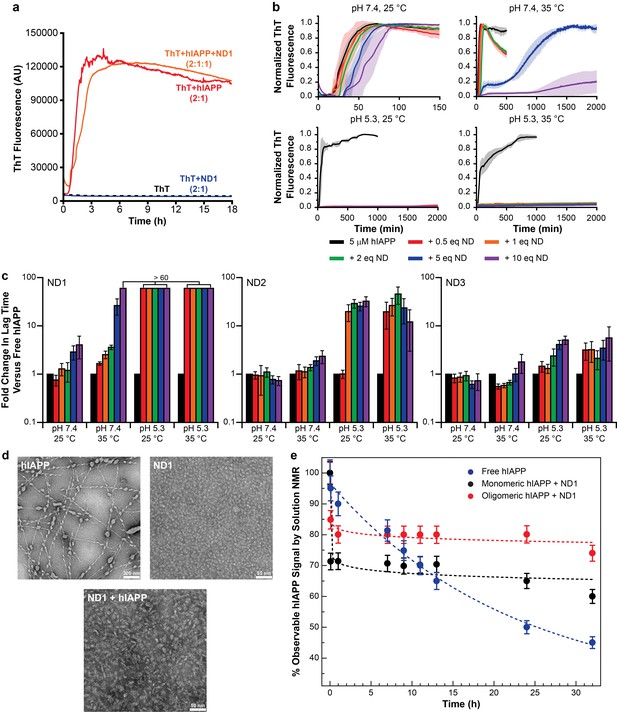
Nanodiscs modulate the kinetics of hIAPP aggregation.
(a) Thioflavin T (ThT) was determined to have no interactions with nanodiscs which could significantly alter the dye’s fluorescent properties (20 mM PO4 pH 7.4, 50 mM NaCl, 25°C). (b) ThT fluorescence was monitored as hIAPP (5 μM) was incubated with increasing concentration of ND1 under different conditions (either 20 mM PO4 pH 7.4 or 30 mM acetate pH 5.3, both with 50 mM NaCl at either 25 or 35°C). Solid curves represent the average of three independent trials while the shaded regions represent the standard deviations of those measurements. (c) Lag times were calculated for each individual kinetic trace for hIAPP incubated with ND1, ND2, and ND3 (Figure 2). The fold change in the lag time compared to untreated hIAPP are plotted with respect to both nanodisc concentration and sample conditions. (d) TEM was used to image samples of hIAPP (50 μM) fiber prepared in the absence of nanodisc, freshly prepared ND1 (50 μM), and hIAPP monomer (50 μM) incubated with ND1 (50 μM). All samples were prepared in 30 mM acetate pH 5.3 at 35°C. (e) The overall signal intensities measured from 2D 1H-15N HMQC spectra of hIAPP backbone amides in the absence or presence of ND1 were monitored over time. Peptide was prepared via both a monomeric and oligomeric methods (see Materials and methods for details) prior to treating with ND1.
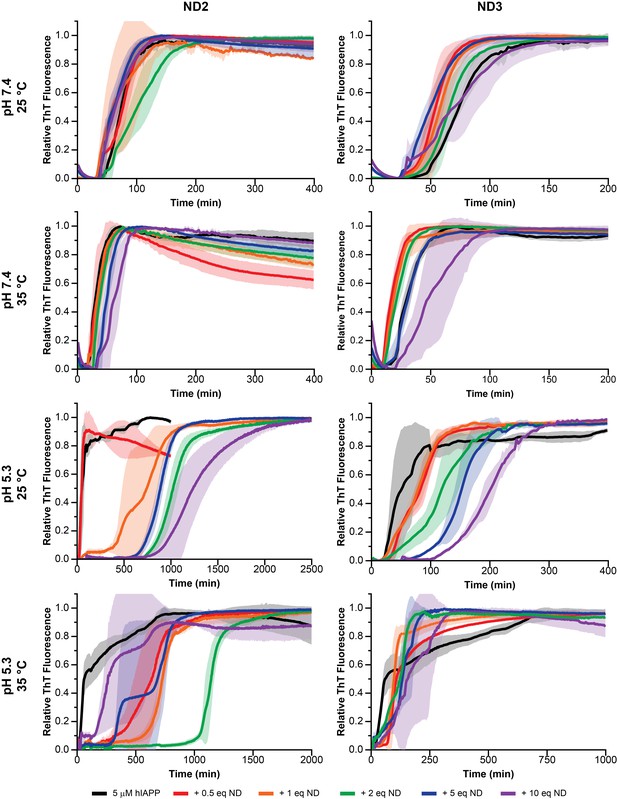
Nanodisc composition and environment dictates the extent of modulation on hIAPP aggregation.
By changing the concentration of negatively charged DMPG lipids in the nanodisc (25% in ND2% and 50% in ND3), the pH of the surrounding buffer (7.4 with 20 mM PO4 or 5.3 with 30 mM acetate) and the solution temperature, a wide range of kinetic behaviors can be observed for hIAPP.
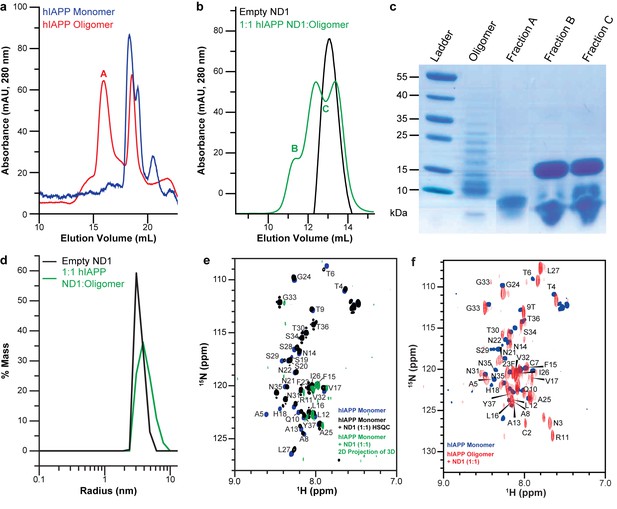
Peptide preparation impacts the stabilization of a folded hIAPP species by ND1.
(a) Freshly dissolved hIAPP (blue) and the oligomer preparation of hIAPP (red) indicate two distinct populations of the peptide. (b) When the oligomeric hIAPP was incubated with ND1, a larger peptide-ND1 complex was stabilized. (c) Gel electrophoresis highlights changes in the oligomer population before and after incubation with ND1 and purification by SEC. (d) DLS confirms the findings of SEC; treatment of ND1 with oligomeric hIAPP promotes a larger, stabilized, species. (e) When ND1 is added to monomeric hIAPP (black), there is minimal spectral perturbation in the 2D 15N/1H HMQC spectrum, suggesting minimal change in the structure. Additionally, when HNCA triple-resonance NMR experiments were performed on the same sample and the spectrum was compressed into the N-H dimensions, a dramatic reduction in signal intensity and disappearance of peaks were observed (green), further suggesting a lack of structural changes in the peptide. (f) Compression of the HNCA spectrum into N-H dimensions yields a full 2D spectrum with an increased dispersion, indicative of a more folded state, which can be completely assigned, facilitating further structural analysis.
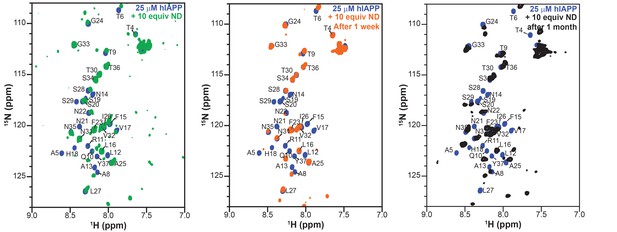
Stability and lifetime of NMR visible hIAPP-ND complexes.
1H/15N HMQC spectra were observable with only modest changes over the course of one month under quiescent conditions at room temperature, suggesting that the sample was amenable for very long (spanning to many days to a week) experiments that were needed for resonance assignment and structural characterization reported in this study.
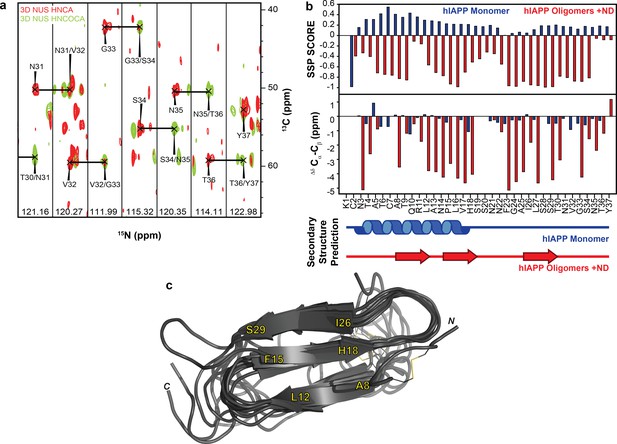
NMR characterization of hIAPP-ND1 interaction.
(a) Triple-resonance (HNCA and HNCOCA) spectra of hIAPP oligomers in the presence of ND1 were utilized for resonance assignment and chemical shift determination (all strips can be found in Figure 6). (b) Secondary structure prediction performed using both Secondary Structure Propensity from Julie Forman-Kay’s Laboratory and the ∆δ 13Cα-Cβ secondary chemical shifts suggest a structure consisting of three β-strands (Marsh et al., 2006). (c) The 10 lowest energy structures were produced by CS-ROSETTA. The average Cα-RMSD of lowest energy structure for residues 6–34 is 1.946 ± 0.521 and for all residues is 3.534 ± 0.489.
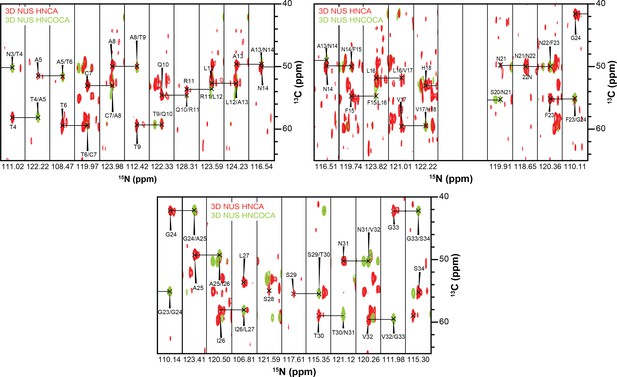
3D strips used for resonance assignment.
The majority of resonances from the 37 residues of hIAPP were resolvable in both HNCA and HNCOCA experiments that were performed on a 1:1 ratio of oligomeric-hIAPP:ND1. Chemical shift values were measured based on these resonance assignments for structural calculation reported in this study.
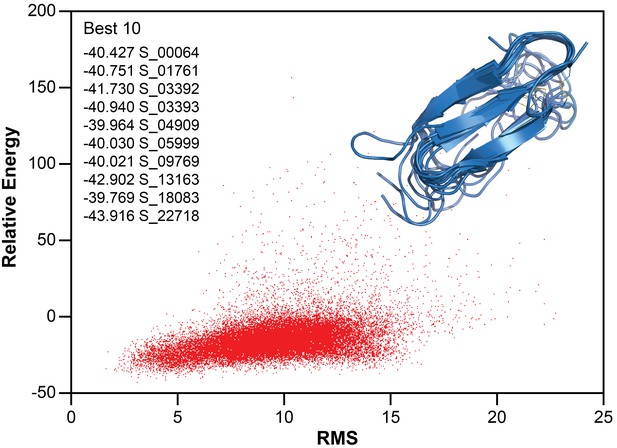
Structure calculation of membrane-associated hIAPP.
The relative energy plot of the CS-Rosetta calculation, including an overlay of 10 lowest energy structures.
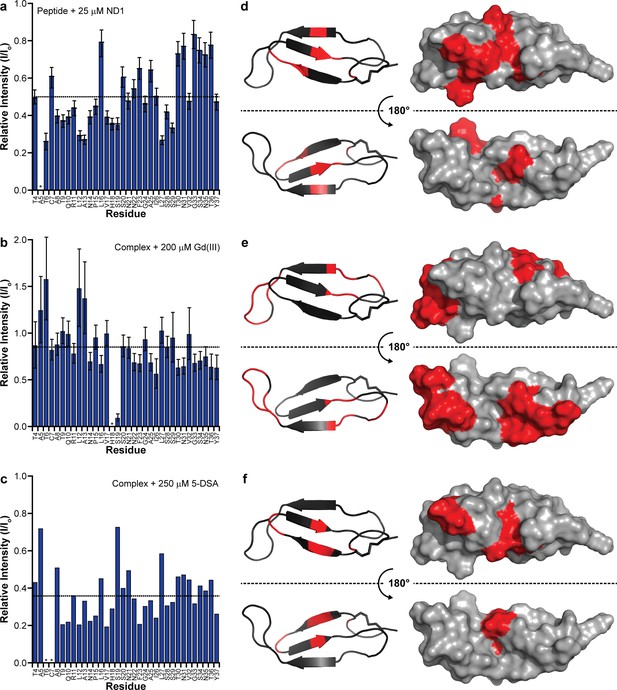
Identifying membrane-associated hIAPP interfaces.
(a–c) Line broadening and signal reduction obtained from 1H-15N HMQC spectra were used to identify the residues interacting directly with ND1 and compared to the average signal reduction for the sample (dashed line). (d–f) Highlighted in red are residues whose signal intensities were reduced more than the average and are mapped onto the structure. The addition of 1 equiv. ND1 (25 μM) to hIAPP identifies the residues directly interacting with the nanodisc surface (a,d) while the titration of Gd(III) (200 μM) into a solution of premixed hIAPP (50 μM) and ND1 (50 μM) selectively reduces the signal intensity of solvent accessible residues that are not interacting with ND1(b,e). Titrating 5-DSA (250 μM) into an identical sample containing a 1:1 ratio of hIAPP:ND1 selectively quenches the residues residing near the surface of ND1 (c,f).
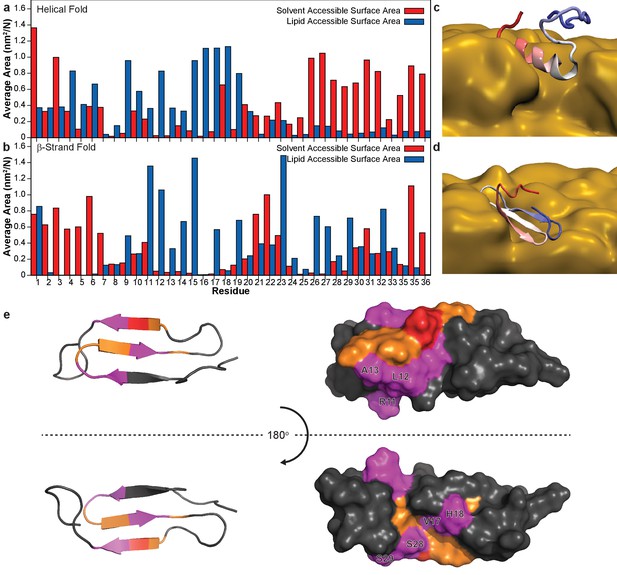
Simulation of membrane-hIAPP interactions.
Molecular dynamic simulations with the Martini force field were performed to evaluate the preferential interactions of hIAPP with ND1 (Abraham et al., 2015; de Jong et al., 2013; Marrink et al., 2007). The average surface area of both a partially folded monomer (Rodriguez Camargo et al., 2017) (a) and the β-strand model (b) plotted for each residue. The preference for the solvent or lipid accessibility was mapped onto both simulated structures in the presence of the membrane (c,d). (e) When the hIAPP residues observed to interact with ND1 by NMR (red) are compared to those observed by MD simulation (orange), significant overlap is observed (magenta).
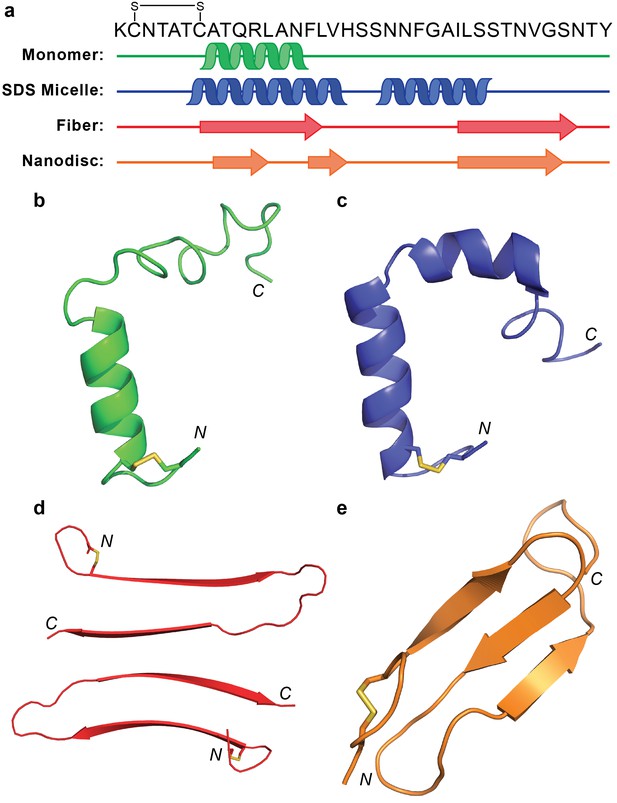
Comparison of hIAPP structures.
(a) Known structures and models of hIAPP suggest partial folding of the monomeric subunit, although the folding varies with the sample preparation and environment. (b) Monomeric hIAPP prepared at pH 5.3 demonstrates a partial helical fold spanning C7-F15 (PDB: 5MGQ) (Rodriguez Camargo et al., 2017). (c) Monomeric hIAPP stabilized by SDS micelles adopts a similar N-terminal helix and a second helical region near the C-terminus (PDB: 2L86) (Nanga et al., 2011). (d) The striated ribbon morphology of hIAPP fibers shows twoβ-hairpins interacting through their C-terminal β-strands (Luca et al., 2007). (e) The folded hIAPP monomer interacting with the surface of ND1 possesses three antiparallel β-strands.
Tables
Nanodisc identity and composition.
All nanodiscs were formed at a protein (MSP):lipid ratio of 1:50 and purified by size exclusion chromatography prior to use.
| Nanodisc | Lipid composition |
|---|---|
| ND1 | 90% DMPC/10% DMPG |
| ND2 | 75% DMPC/25% DMPG |
| ND3 | 50% DMPC/50% DMPG |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.31226.013





