Cleavage activates dispatched for Sonic Hedgehog ligand release
Figures
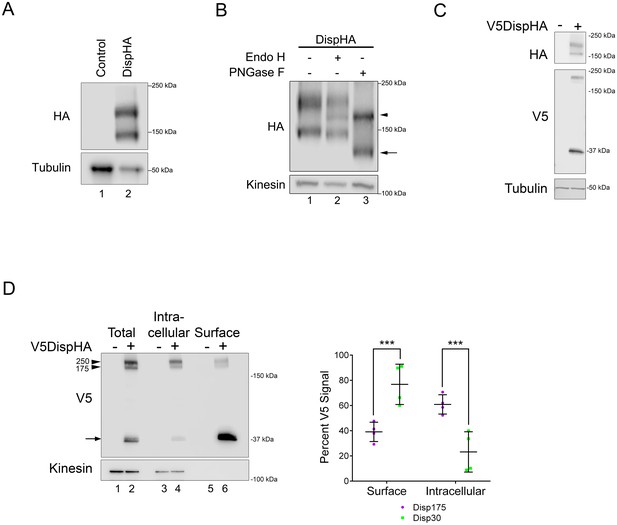
Disp is cleaved.
(A) Lysates prepared from vector control and DispHA-expressing NIH3T3 cells were analyzed by SDS-PAGE and western blot against the HA tag. (B) Lysates from DispHA-expressing NIH3T3 cells were treated with Endo H or PNGase F enzymes. The arrowhead marks deglycosylated Disp175 and the arrow marks deglycosylated Disp145. (C) V5DispHA was expressed in NIH3T3 cells and cell lysates analyzed by western blot. (D) Lysates were prepared from DispHA-expressing NIH3T3 cells treated with biotin-containing culture medium for 30 min at 4°C prior to lysis. Lysates were incubated with streptavidin beads and bound (surface) and unbound (intracellular) DispHA proteins were analyzed by western blot. Combined densitometry analysis of four independent labeling experiments is shown. Densitometry is presented as percentage of the sum total of Disp175 or Disp30 signals across the two fractions. Significance was determined using a paired Student’s t-test. For all statistical analyses *p≤0.05 and ***p≤0.005. Error bars indicate standard deviation (s.d.). For all western blots Kinesin or Tubulin serve as loading controls.
-
Figure 1—source data 1
Data for Figure 1D.
- https://doi.org/10.7554/eLife.31678.004
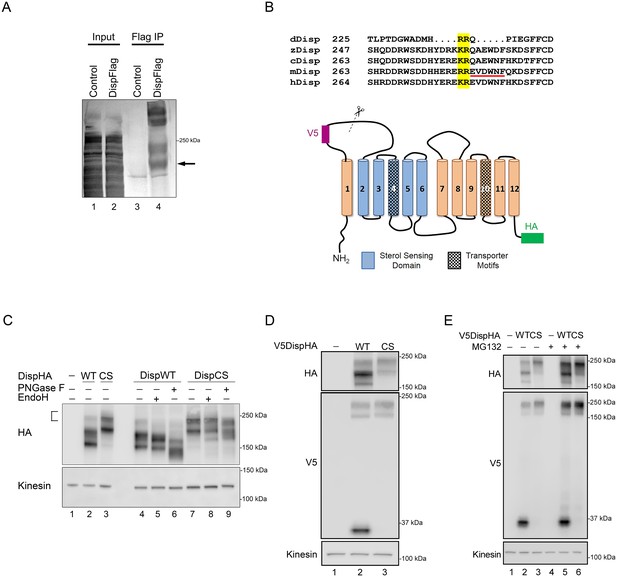
Disp is cleaved at a Furin consensus sequence in EC1.
(A) Disp-Flag was expressed in HEK293T cells, immunopurified from cell lysates on Flag beads and the 145 kDa species (arrow) excised and analyzed by Edman degradation. (B) Alignment of EC1 sequence from Disp proteins of Drosophila, zebrafish, chick, mouse and human. Edman sequencing of murine Disp145 revealed EVDWNF (red line) to be the amino terminal sequence, suggesting that cleavage occurs adjacent to dibasic residues R279 and E280 (yellow box). A diagram of murine Disp shows V5 and HA epitope tag insertion sites, functional domains and the approximate location of the identified cleavage motif in predicted EC1 (scissors). (C) Cell lysates from NIH3T3 cells expressing wild type or cleavage site mutant (CS, R279A/E280A) were treated with deglycosylating enzymes. The bracket indicates the ~250 kDa fraction. (D) Lysates from NIH3T3 cells expressing wild type or cleavage site mutant V5DispHA proteins were examined for presence of the 30 kDa V5 fragment by western blot. (E) NIH3T3 cells expressing wild type or CS mutant Disp proteins were treated for ~6 hr with DMSO vehicle or MG132 proteasome inhibitor. Kinesin is the loading control.
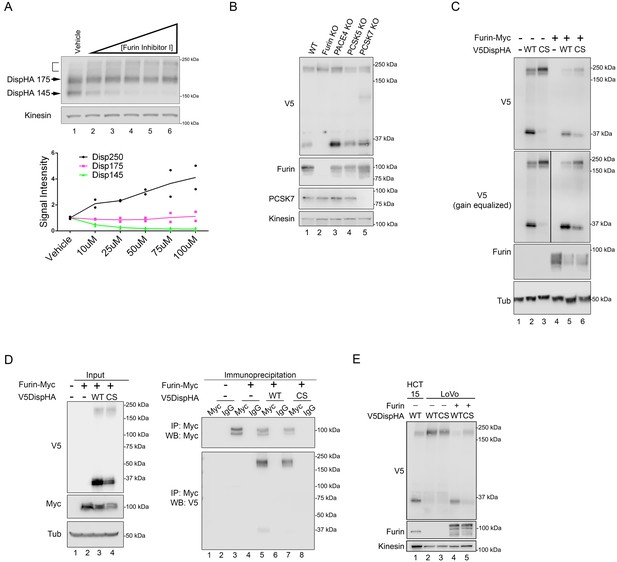
Disp is cleaved by Furin.
(A) NIH3T3 cells expressing DispHA were treated with increasing concentrations of Furin Inhibitor I (10, 25, 50, 75 and 100 μM). The bracket indicates the 250 kDa fraction. Graph shows Disp densitometry analysis normalized to Kinesin for two independent experiments. Normalized signal intensity for each DispHA species in treated conditions is shown relative to vehicle control intensity, which was set to 1. (B) CRISPR/Cas9 generated knockout lines for Furin, PACE4, PCSK5 and PCSK7 were transfected with V5DispHA-expression vector, and formation of the ~30 kDa V5 cleavage fragment was monitored by western blot of cell lysates from Clonal line 1. Furin and PCSK7 protein levels were examined by western blot. PACE4 and PCSK5 mutations were confirmed by deep sequencing as in Figure 3—figure supplement 1. (C) Lysates from cells co-expressing wild type or CS mutant V5DispHA and Furin-Myc proteins were examined for Disp cleavage by western blot for the V5 fragment. Co-expression of Furin-Myc reduced total Disp signal (top panel). Gain equalization of the V5 signal in Furin-Myc expressing cell lysates is shown for comparison. (D) Furin-Myc and V5DispHA were co-expressed in HEK293T cells and Furin-Myc was immunoprecipitated from lysates using anti-Myc. Input (left) and immunoprecipitates (right) are shown. (E) Wild type and CS mutant V5DispHA proteins were expressed in LoVo (lacking Furin) or HCT-15 (control) colon carcinoma cells and lysates were analyzed by western blot. Re-expression of Furin in LoVo cells rescued cleavage (lane 4 compared to 2). Kinesin and Tubulin are the loading controls for western blots.
-
Figure 3—source data 1
Data for Figure 3A.
- https://doi.org/10.7554/eLife.31678.008

(A) Genotype of PC family knockout clones. Red arrows indicate the Cas9 cleavage site. sgRNA sequence (5’ to 3’) is shown for each targeted gene along with corresponding deletions (black dashes) and insertions (red letters) identified by deep sequencing of knockout clonal lines. Insertion/deletion sizes are indicated in parenthesis and resulting premature truncation amino acid is shown in red. (B) CRISPR/Cas9 generated knockout lines for Furin, PACE4, PCSK5 and PCSK7 were transfected with V5DispHA-expression vector, and formation of the ~30 kDa V5 cleavage fragment was monitored by western blot of cell lysates from Clonal line 2. Furin and PCSK7 protein levels were examined by western blot. Kinesin is loading control. PACE4 and PCSK5 were assessed by deep sequencing as in A.
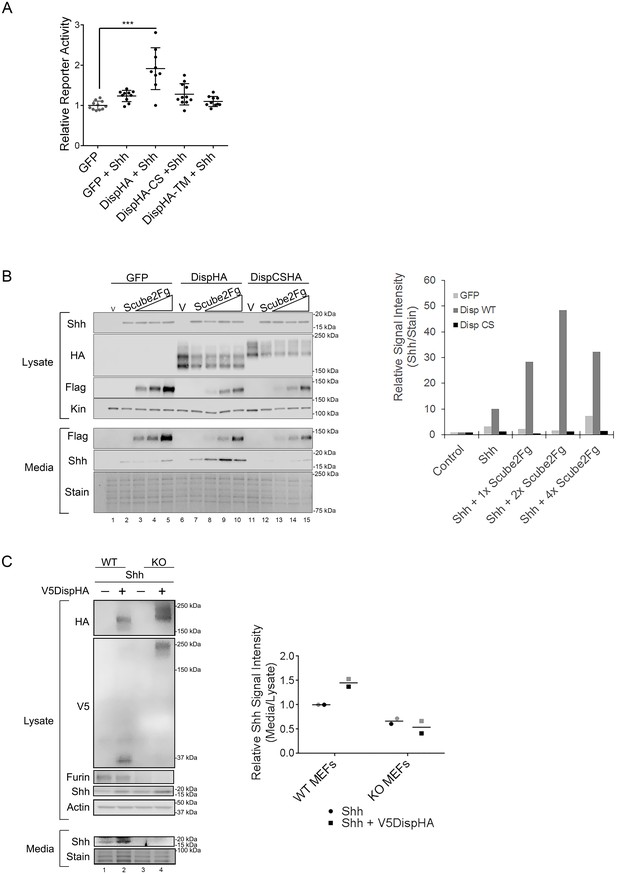
Disp cleavage is required for Shh release.
(A) Disp-/- cells stably transfected with Shh or empty vector control were transiently transfected with GFP or the indicated DispHA expression vectors, and then co-cultured with LightII reporter cells. Reporter activity normalized to tk-renilla and relative to GFP control (set to 1) is shown. Assays were performed four times in duplicate or triplicate and all data points pooled. Error bars represent s.d. Significance was determined by one-way ANOVA. p***≤0.005. (B) Disp-/- mouse embryonic fibroblasts were stably transfected with empty vector control (V) or vector encoding Shh. Wild type or CS mutant Disp proteins were transiently expressed in Disp-/- cells alone or with Scube2, and lysates and media were examined by SDS-PAGE and western blot. Equal protein amounts (25 µg) from TCA precipitates of conditioned media were analyzed. Kinesin serves as loading control for lysate. Coomassie stain of membrane is shown as loading control for conditioned media (bottom). The graph (right) represents densitometry analysis of Shh media signal intensity normalized to media coomassie stain. The experiment was repeated three times. A representative experiment is shown. (C) Furin-/- cells were transiently transfected with Shh alone or with V5DispHA expression vector, and tested for Shh release as in (B). The experiment was repeated twice for Furin-/- Clone 2 and once for Clone 1, and a representative blot for Furin-/- Clone 2 is shown. The graph (right) represents densitometry analysis of Shh media signal normalized to media coomassie stain, and plotted relative to lysate Shh signal normalized to Actin. Analysis of the representative Clone 2 western is black. Densitometry analysis of the corresponding Clone 1 release assay is shown in gray.
-
Figure 4—source data 1
Data for Figure 4A.
- https://doi.org/10.7554/eLife.31678.010
-
Figure 4—source data 2
Data for Figure 4B.
- https://doi.org/10.7554/eLife.31678.011
-
Figure 4—source data 3
Data for Figure 4C.
- https://doi.org/10.7554/eLife.31678.012
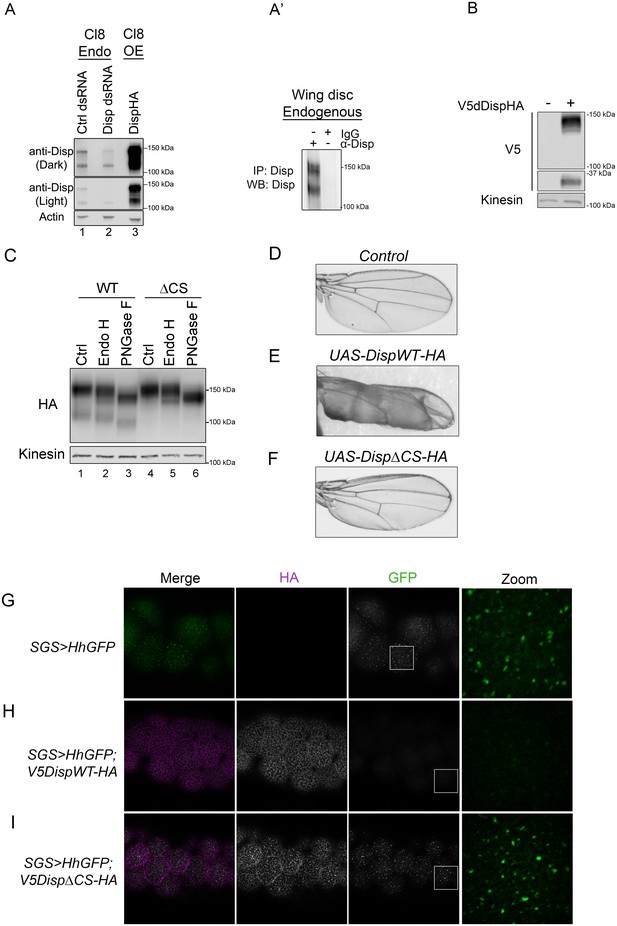
Cleavage is required for Disp activity in vivo.
(A) Lysates from Drosophila Cl8 cells treated with control or disp dsRNA or transfected with pAc-dispHA were analyzed by western blot using anti-dDisp. Actin is the loading control. (A’) Endogenous dDisp150 and dDisp110 were specifically immunoprecipitated with anti-dDisp from wing imaginal disc lysate. (B) V5dDispHA was expressed in Cl8 cells and lysates were analyzed by western blot to confirm generation of the V5 fragment. Kinesin is the loading control. (C) Lysates were prepared from Cl8 cells expressing Δ206–238 (ΔCS) dDispHA protein and analyzed by western blot. Lysates were treated with Endo H or PNGase F. Kinesin is the loading control. (D–F) Wild type or ΔCS dDispHA proteins were expressed dorsally in wing imaginal discs using apterous-GAL4. Representative male wings are shown. (G–I) WT or ΔCS V5dDispHA proteins (magenta) were expressed with HhGFP (green) in salivary glands using SGS-GAL4. Maximum intensity projections of basolateral and basal optical sections of salivary glands are shown. Square in the GFP images indicates zoom area.
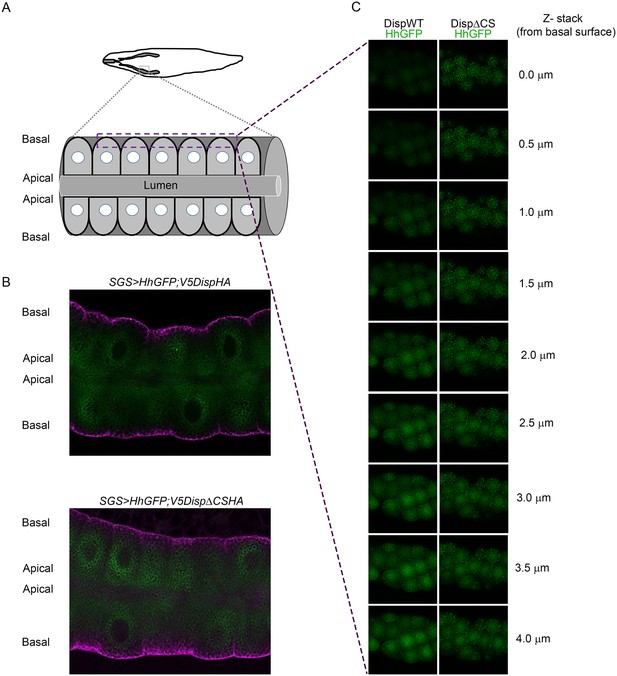
(A) The top graphic shows the location of salivary glands in a third instar larvae. The lower graphic shows organization of epithelial cells within the salivary gland. (B) Sagittal sections of the salivary glands expressing wild type (top) or CS mutant (bottom) V5DispHA proteins (magenta) and Hh-GFP (green) are shown. (C) Optical slices from z-stack images are shown. Z-slices were imaged from the basal surface (top) into the salivary gland. The dashed square in A shows the approximate location where images were acquired.
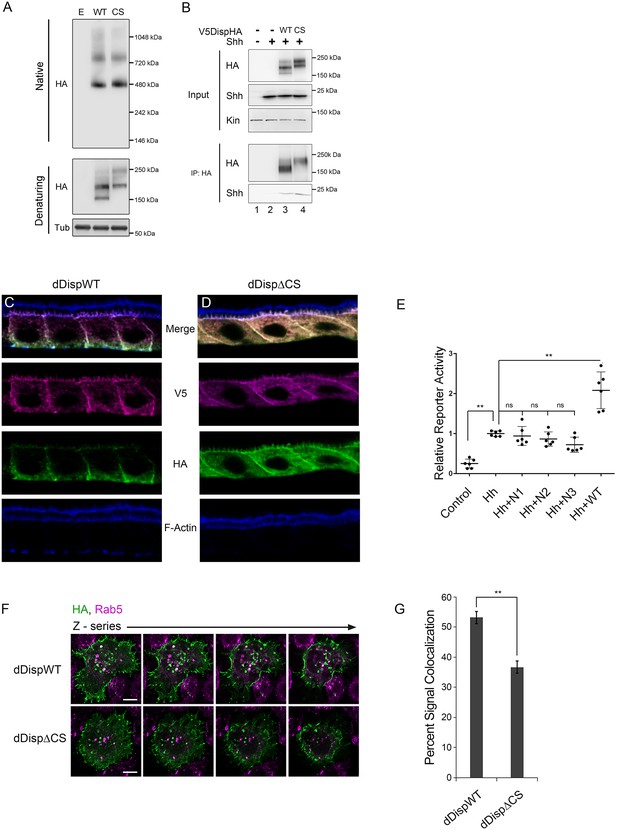
Processing impacts Disp membrane localization.
(A) Lysates from NIH3T3 cells expressing wild type or CS mutant Disp proteins were analyzed by native (upper) and denaturing (lower) gel electrophoresis and western blot. The predicted molecular weight of the Disp trimer is ~480 kDa. Tubulin is the loading control. (B) DispHA proteins were immunoprecipitated with anti-HA antibody from lysates of Disp-/- cells expressing Shh alone or with wild type or CS DispHA. Wild type and CS mutant DispHA proteins co-immunoprecipitate Shh (bottom). (C–D) Wild type and ΔCS V5dDispHA proteins were expressed in ovarian follicle cells using the C204-GAL4. F-actin marks apical membrane in follicle cells (blue). The dDisp amino-terminal region is indicated by V5 (magenta) and carboxyl domain by HA (green). (E) V5dDispHA or increasing amounts of the amino-terminal V5 fragment were expressed with Hh in ligand producing cells, and then co-cultured with Cl8 cells transfected with Hh-responsive ptcΔ136-luciferase reporter and actin-renilla control. Luciferase reporter activity in ligand receiving cells was measured, and is shown normalized to renilla and relative to the Hh response in reporter cells co-cultured with empty vector control transfected cells. The experiment was repeated three times in duplicate and all data pooled. Error bars indicate s. d. Significance was determined by a one-way ANOVA. (F–G) S2 cells expressing wild type or ΔCS V5dDispHA proteins were analyzed for colocalization (white) between HA (green) and endogenous Rab5 signals (magenta) by confocal microscopy. Serial z-sections are shown. Scale bar is 5 µm. Percent colocalization between HA and Rab5 signals was determine by Imaris image analysis software (H). Error bars indicate s. e. For all panels, **p≤0.01; ns, not significant.
-
Figure 6—source data 1
Data for Figure 6E.
- https://doi.org/10.7554/eLife.31678.016
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (Drosophila melanogaster) | Oregon-R | Bloomington Stock Center, Bloomington, IN | ||
| Genetic reagent (D. melanogaster) | apterous-Gal4 | Bloomington Stock Center | ||
| Genetic reagent (D. melanogaster) | SGS-Gal4 | Bloomington Stock Center | ||
| Genetic reagent (D. melanogaster) | UAS-HhGFP | Hartman et al. (2013), 201(5) 741–57. | ||
| Genetic reagent (D. melanogaster) | UAS-dispWT | This study | ||
| Genetic reagent (D. melanogaster) | UAS-dispΔCS | This study | ||
| Cell line (Mus musculus) | NIH3T3 | ATCC, Manassas, VA | CRL-1658 | |
| Cell line (Homo sapiens) | HEK293T | ATCC | CRL-11268 | |
| Cell line (M. musculus) | Dispatched KO MEFs | Ma et al. (2002) 111(1): 63–75 | ||
| Cell line (H. sapiens) | LoVo | ATCC | CCL-229 | |
| Cell line (H. sapiens) | HCT-15 | ATCC | CCL-225 | |
| Cell line (M. musculus) | Light II | ATCC | JHU-68 | |
| Cell line (D. melanogaster) | Cl8 | DGRC, Bloomington, IN | stock # 151 | |
| Cell line (D. melanogaster) | S2 | Thermo Fisher, Waltham, MA | R69007 | |
| Cell line (M. musculus) | Furin-/- | This paper | CRISPR/Cas9 | C57BL/6 MEF cells |
| Cell line (M. musculus) | Pcsk5-/- | This paper | CRISPR/Cas9 | C57BL/6 MEF cells |
| Cell line (M. musculus) | Pace4-/- | This paper | CRISPR/Cas9 | C57BL/6 MEF cells |
| Cell line (M. musculus) | Pcsk7-/- | This paper | CRISPR/Cas9 | C57BL/6 MEF cells |
| Antibody | anti-Kif5b | Abcam, Cambridge, MA | ab167429 | 1:5000 (WB) |
| Antibody | anti-tubulin | Cell Signaling, Danvers, MA | 3873 | 1:10000 (WB) |
| Antibody | anti-Myc Affinity gel | Sigma, St Louis, MO | E6654 | 25 ul slurry for IP |
| Antibody | anti-HA affinity gel | Sigma | E6779 | 25 ul slurry for IP |
| Antibody | anti-Shh | Santa Cruz Biotechnolgy, Dallas, TX | sc-9024 | 1:2000 (WB) |
| Antibody | anti-Myc | Roche, Basal, Switzerland | 11667149001 | 1:1000 (WB) |
| Antibody | anti-Disp | This Study | 1:1000 | |
| Antibody | anti-mouse HRP | Jackson Immuno, West Grove, PA | 715-035-151 | 1:10000 |
| Antibody | anti-Rabbit HRP | Jackson Immuno | 711-035-152 | 1:10000 |
| Antibody | anti-Rat HRP | Jackson Immuno | 112-035-175 | 1:10000 |
| Antibody | AlexaFluor 488 | Life technologies, Carlsbad, CA | A11029 (Mouse) A11034 (Rabbit) A11006(Rat) | 1:1000 |
| Antibody | AlexaFluor 555 | Life technologies | A21424 (Mouse) A21429 (Rabbit) A21434(Rat) | 1:1000 |
| Antibody | AlexaFluor 633 | Life technologies | A21236 (Mouse) A21245 (Rabbit) A21247(Rat) | 1:1000 |
| Antibody | anti-Mouse IR800 | LiCor, Lincoln, NE | 92632212 | 1:10000 |
| Antibody | anti-Rabbit IR800 | LiCor | 92632213 | 1:10000 |
| Antibody | anti-Rat IR800 | LiCor | 92632219 | 1:10000 |
| Antibody | ||||
| Transfected construct (M. musculus) | Dispatched | This paper | RIKEN | http://dna.brc.riken.jp/ |
| Transfected construct (M. musculus) | pCDNA3 Dispatched WT HA | This paper | pCDNA3 from Invitrogen | |
| Transfected construct (M. musculus) | pCDNA3 V5 Dispatched WT HA | This paper | V5 introduced following A106 | |
| Transfected construct (M. musculus) | pCDNA3 V5 Dispatched CS HA | This paper | Mutate R279, E280 to alanine | |
| Transfected construct (M. musculus) | MSCV Hygro Shh-FL | This paper | MSCV Hygro from Clontech | |
| Transfected construct (M. musculus) | pCDNA3-V5 DispTM4/TM10HA | This paper | ||
| Transfected construct(M. musculus) | pFLC-I-Scube2 | SourceBioscience, Nottingham, UK | Clone E030016G24 | |
| Transfected construct(M. musculus) | pCDNA3-Scube2 Flag | This paper | ||
| Transfected construct (H. sapiens) | pCMV6 - huFurin Myc DDK | This paper | Origene, RC204279 | |
| Transfected construct (M. musculus) | pCDNA3 - Shh-FL | This paper | Gift from P. Beachy Lab | |
| Transfected construct (Aequorea victoria) | pCDNA3 - GFP | This paper | ||
| Transfected construct (D. melanogaster) | pFLC-I-disp cDNA | This paper | DGRC | Supported by NIH grant 2P40OD010949 |
| Transfected construct (D. melanogaster) | pAc-dispHA | This paper | Cloned into pAc5.1 Vector with HA tag | |
| Transfected construct (D. melanogaster) | pAc-V5dispHA | This paper | V5 introduced following V108 | |
| Transfected construct (D. melanogaster) | pAc-disp Δ206–238 HA | This paper | ||
| Transfected construct (D. melanogaster) | pUAS-aatB-V5dispWTHA | This paper | ||
| Transfected construct (D. melanogaster) | pUAS-aatB-V5dispΔ206–238 HA | This paper | ||
| Transfected construct (D. melanogaster) | pUAS-HhGFP | This paper | ||
| Transfected construct (Photinus pyralis) | ptcΔ 136-luciferase | This paper | ||
| Transfected construct (Renilla reniformis) | pAc-renilla | This paper | ||
| Transfected construct (D. melanogaster) | pAC-hh | This paper | ||
| Software, algorithm | Adobe Photoshop CS4 | Adobe, San Jose, CA | for making figures | |
| Software, algorithm | LAS X | Leica, Wetzlar, Germany | image analysis | |
| Software, algorithm | Prism | GraphPad, La Jolla, CA | for statistical analysys and graphs | |
| Software, algorithm | Huygens Professional software | for decovolution images | ||
| Commercial assay or kit | Quickchange II XL Kit | Agilent, Santa Clara, CA | 200522 | |
| Commercial assay or kit | Lipofectamine 2000 | ThermoFisher Scientific | 11668027 | |
| Commercial assay or kit | Lipofectamine 3000 | ThermoFisher Scientific | L3000008 | |
| Commercial assay or kit | FuGene HD | ThermoFisher Scientific | PRE2311 | |
| Commercial assay or kit | ECL Prime Western Blotting Detection Reagent | Fisher Scientific, Hampton, NH | RPN2232 | |
| Commercial assay or kit | Dual Luciferase Reporter Assay Kit | Promega, Madison, WI | PRE1960 | |
| Chemical compound, drug | Furin I Inhibitor | Enzo Life Sciences, Farmingdale, NY | ALX-260–022 M005 | |
| Chemical compound, drug | MG-132 | EMD Chemicals Inc., St. Louis, MO | 474790 |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.31678.017





