Functional genomics of lipid metabolism in the oleaginous yeast Rhodosporidium toruloides
Figures
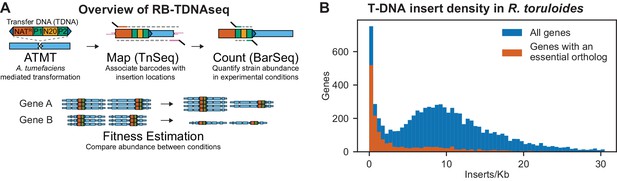
Overview of RB-TDNAseq and T-DNA insert density in R. toruloides coding regions.
(A) General strategy of RB-TDNAseq. A library of binary plasmids bearing an antibiotic resistance cassette (NATR) and a random 20 base-pair sequence ‘barcode’ (N20) flanked by specific priming sites (P1/P2) is introduced into a population of A. tumefaciens carrying a vir helper plasmid. A. tumefaciens efficiently transforms a T-DNA fragment into the target fungus (ATMT). NATR colonies are then combined to make a mutant pool. T-DNA-genome junctions are sequenced by TnSeq, thereby associating barcodes with the location of the insertion (Map). The mutant pool is then cultured under specific conditions and the relative abundance of mutant strains is measured by sequencing a short, specific, PCR on the barcodes (BarSeq) and counting the occurrence of each sequence (Count). Finally, for each gene, count data is combined across all barcodes mapping to insertions in that gene to obtain a robust measure of relative fitness for strains bearing mutations in that gene (Fitness Estimation). (B) Histogram of insert density in coding regions (start codon to stop codon) for all genes, and genes with orthologs reported to be essential in A. nidulans, C. neoformans, N. crassa, S. cerevisiae, or S. pombe. The following figure supplements are available for Figure 1.
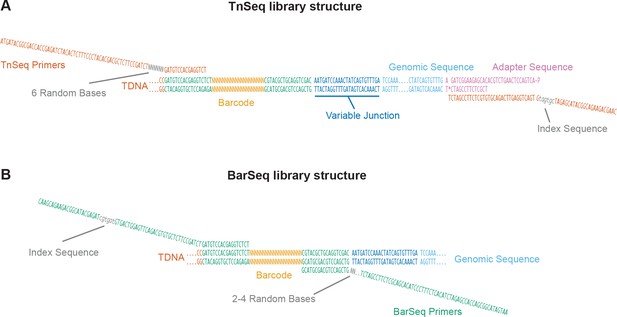
Schematic of TnSeq and BarSeq libraries generated using RB-TDNAseq.
(A) In the TnSeq protocol, genomic DNA is sheared into ~300 bp fragments, and Illumina TruSeq adapters are ligated on both ends. T-DNA junctions are then specifically enriched by PCR with a T-DNA-specific and an adapter-specific primer. (B) In the BarSeq protocol, genomic DNA is used as a template for a more robust and quantitative PCR on the barcoded region of the T-DNA insert. Phasing error caused by the identical T-DNA sequences flanking the random barcodes was reduced by adding sequence diversity at the beginning of each read, either by the introduction of a short random 6 bp sequence or a 2–4 bp random sequence for TnSeq and BarSeq, respectively.
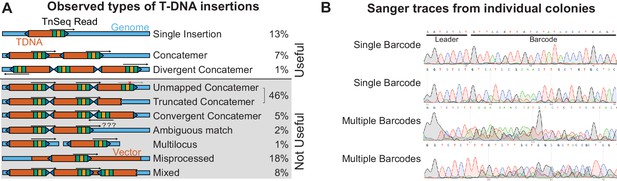
Complexities of T-DNA insertions.
(A) Inferred topology of T-DNA insertions from associations of barcodes and adjacent genomic or T-DNA sequence. Only three of the observed insertion types could be mapped using the TnSeq protocol. (B) Sanger sequencing of barcodes from single colonies isolated from the pool. Multiple overlapping peaks in the barcode region suggest multiple T-DNAs are present in a single strain. Note that these T-DNAs may be integrated at the same, or different loci. Inherent noise in barcode amplification and sequencing introduces significant ambiguity in this analysis. The inferred rate of multiple barcode insertion (29%) should be considered a maximum estimate.
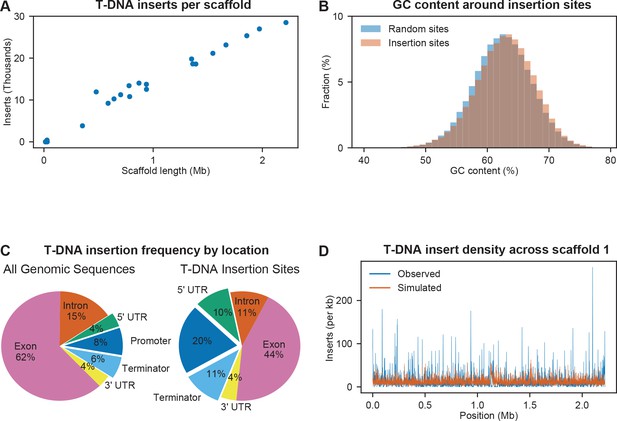
Observed biases in T-DNA insertion locations.
(A) Frequency of T-DNA insertion mapping was consistent across all 30 IFO 0880 scaffolds. (B) Histogram of GC content in 100 base pair regions flanking insertion sites and in random 100 base pair regions. (C) Proportion of the R. toruloides IFO 0880 genome in promoter regions, terminator regions, untranslated regions transcribed to mRNA, coding exons, and introns versus the proportion of T-DNA insertions mapped to those sequences. (D) Distribution of T-DNA insertion density across the length of scaffold 1. Total inserts were summed across a rolling 1,000 base pair window using the observed insertions and a simulated random mutant pool assuming biases for insertion in promoters, terminators and untranslated transcribed regions.
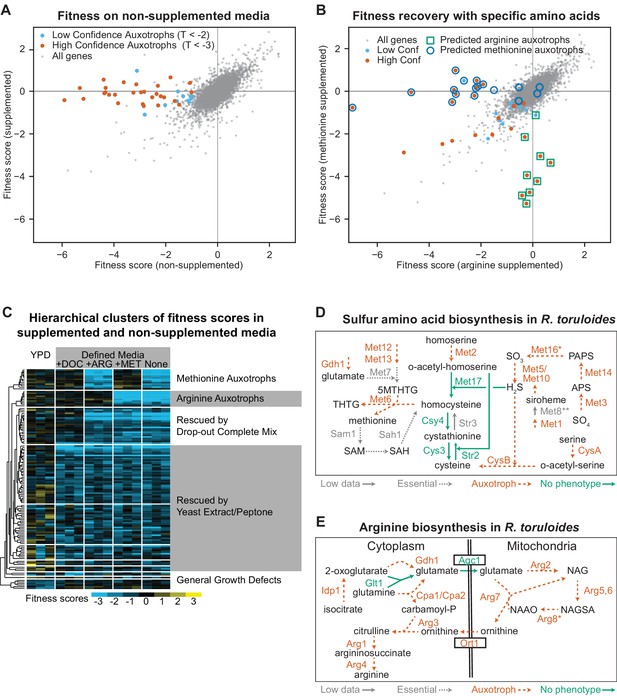
Confirmation of amino acid biosynthetic genes with high-throughput fitness experiments.
(A) Fitness scores for 6,558 genes in media with and without amino acid supplementation (drop-out complete mix). Gene fitness scores are log ratios of final versus starting abundance averaged over multiple barcoded insertions per gene across three biological replicates. Genes that had consistently different enrichment scores between treatments (∆F > 1, |T| statistic >3) are highlighted and represent genes for which mutant strains are auxotrophic for one or more amino acids, nucleotides, or vitamins present in the drop-out complete mixture. (B) Fitness scores in media supplemented with arginine or methionine. Highlighted genes are the same as highlighted in (A). Deletion strains for circled or boxed genes are auxotrophic for methionine or arginine, respectively, in S. cerevisiae or A. nidulans. See Supplementary file 2 for full fitness data. (C) Hierarchical clusters of fitness scores in supplemented and non-supplemented media. Fitness scores for each biological replicate versus its Time 0 replicate for genes with a consistent fitness defect (F < −1, T < −3) in one or more of the following conditions: Yeast extract/Peptone/Dextrose media (YPD) or defined media (DM, composed of yeast nitrogen base plus glucose) with or without the following supplements: (+DOC), arginine (+ARG), or methionine (+MET). (D) Sulfur amino acid biosynthesis in R. toruloides as inferred from fitness experiments. CysA/CysB are named according to their A. nidulans orthologs, all others by orthologs in S. cerevisiae. Auxotrophic mutants had F < −1 in non-supplemented media (DM) and T < −3 in DM versus supplemented media (DOC). Multiple insertions were mapped in STR3, suggesting non-essentiality, but strain abundance was too low to estimate fitness in BarSeq data. *MET16 had fitness scores that clustered with the other auxotrophic mutants, but TDM-DOC was −2.7. **Fitness scores for insertions in MET8 were not inconsistent with auxotrophy, but only two insertions were abundant enough to be tracked. 5MTHTG: 5-methyltetrahydropteroyltri-L-glutamate, THTG: tetrahydropteroyltri-L-glutamate, SAM: S-adenosyl-L-methionine, SAH: S-adenosyl-homocysteine, APS: adenylyl-sulfate, PAPS: 3'-phosphoadenylyl-sulfate. (E) Arginine biosynthesis in R. toruloides as inferred from fitness experiments. *ARG8 had fitness scores that clustered with the other auxotrophic mutants, but TDM-DOC was −2.9. NAG: N-acetylglutamate, NAGSA: N-acetylglutamate semialdehyde, NAAO: N-alpha-acetylornithine. The following figure supplements are available for Figure 2.
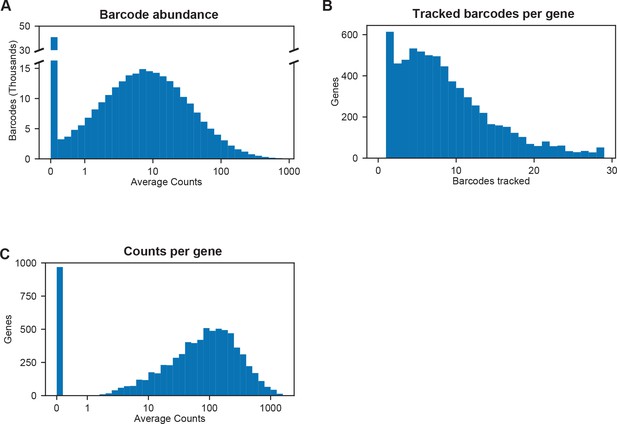
Barcode abundance in BarSeq experiments.
(A) Histogram of barcode abundance in a typical BarSeq experiment with 20 million reads per sample. (B) Histogram of tracked barcodes per gene in a typical BarSeq experiment. Median seven barcodes per gene, 68,021 total barcodes in 6,558 genes. See Supplementary file 1 for a full list of insert density by gene and orthologs reported as essential in model fungi. (C) Histogram of total reads per sample per gene in a typical experiment. This is the sum of counts for all the barcodes that were used in gene fitness estimation for each gene, averaged across all samples.
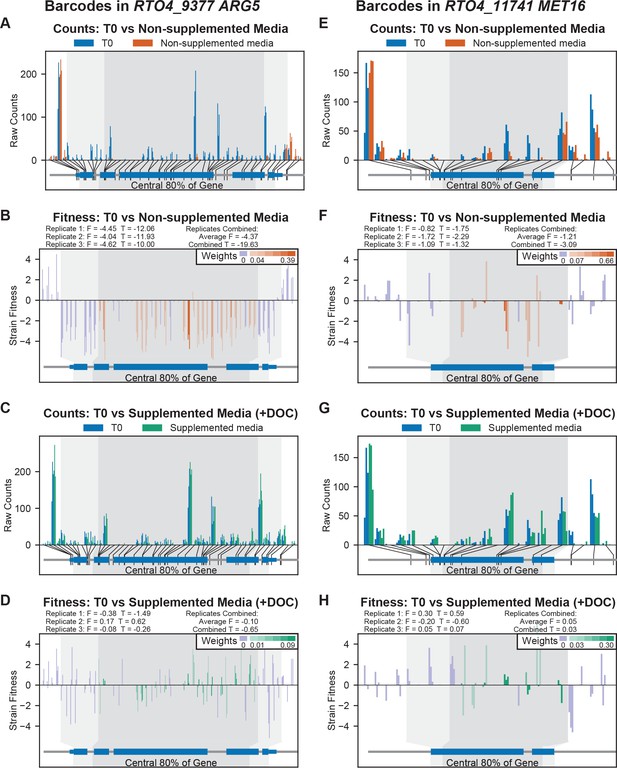
Contributions of individual strains to gene-level fitness scores.
(A) Raw counts for barcoded insertions inside and flanking the coding region for RTO4_9377 ARG5 in Time 0 samples and after growth in non-supplemented defined media (DM). The gene structure is shown below the plot with coding exons as large blue boxes and five prime/three prime untranslated regions as smaller blue boxes. The location of each insertion is noted with a black line between the gene model and the corresponding data on the bar chart. Counts from each biological replicate for each insertion are clustered together in the order they were mapped to the gene. The dark grey area of the plot indicates insertions in the central 80% of the coding region, and the flanking area in light grey indicates insertions in the first or last 10% of the coding region. (B) Fitness scores for individual insertions in ARG5, plotted in the same order as (A). The height of the bar indicates the fitness score derived from the log2 ratio of counts shown in (A). Shading of the bar indicates the weight assigned to each insertion in calculating F for the gene. F and T for each individual replicate and the average F and combined T for this condition are displayed above the plot. (C) Raw counts for insertions in ARG5 in Time 0 and after growth in arginine-supplemented media (+DOC: DM plus drop-out complete mix). (D) Fitness scores for ARG5 on arginine-supplemented media (+DOC). (E) Raw counts for insertions in RTO4_11741 MET16 on non-supplemented media. (F) Fitness scores for insertions in MET16 on non-supplemented media. (G) Raw counts for insertions in MET16 on methionine-supplemented media (+DOC). (H) Fitness scores for insertions in MET16 on methionine-supplemented media (+DOC).
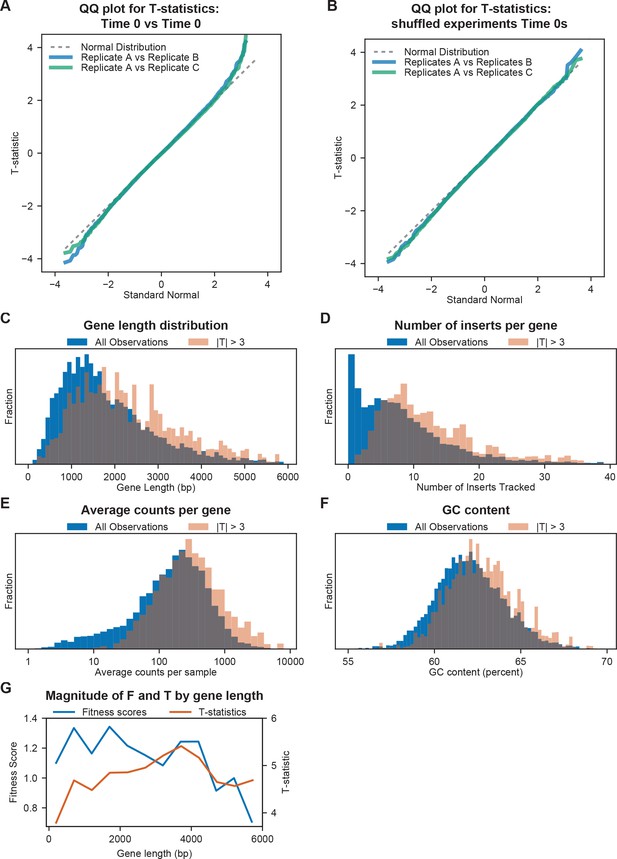
Properties of T-statistics.
(A) Distributions of T-statistics for mock comparisons between individual replicate Time 0 samples from our auxotrophy experiments, shown as a quantile-quantile (QQ) plot. A perfect fit to the standard normal distribution would be on the x = y line (dashed grey line). (B) Distributions of T-statistics for mock comparisons between shuffled sets of Time 0 samples from three experiments, shown as a quantile-quantile plot. (C) Histogram of gene lengths for all genes with sufficient data to compute fitness scores, and for observations with |T| > 3 versus Time 0 for combined T-statistics across biological triplicates. (D) Histogram of the number of inserts per gene with sufficient depth in BarSeq experiments to contribute to fitness estimation. All genes and observations with |T| > 3 versus Time 0 for combined T-statistics across biological triplicates. (E) Histogram of total counts per gene per sample for all genes and for observations with |T| > 3 versus Time 0 for combined T-statistics across biological triplicates. (F) Histogram of GC content for all genes and for observations with |T| > 3 versus Time 0 for combined T-statistics across biological triplicates. (G) Average fitness score and T-statistics for observations with |T| > 3 versus gene length. Fitness scores and T-statistics were binned by gene length in intervals of 500 bp and averaged across each bin.
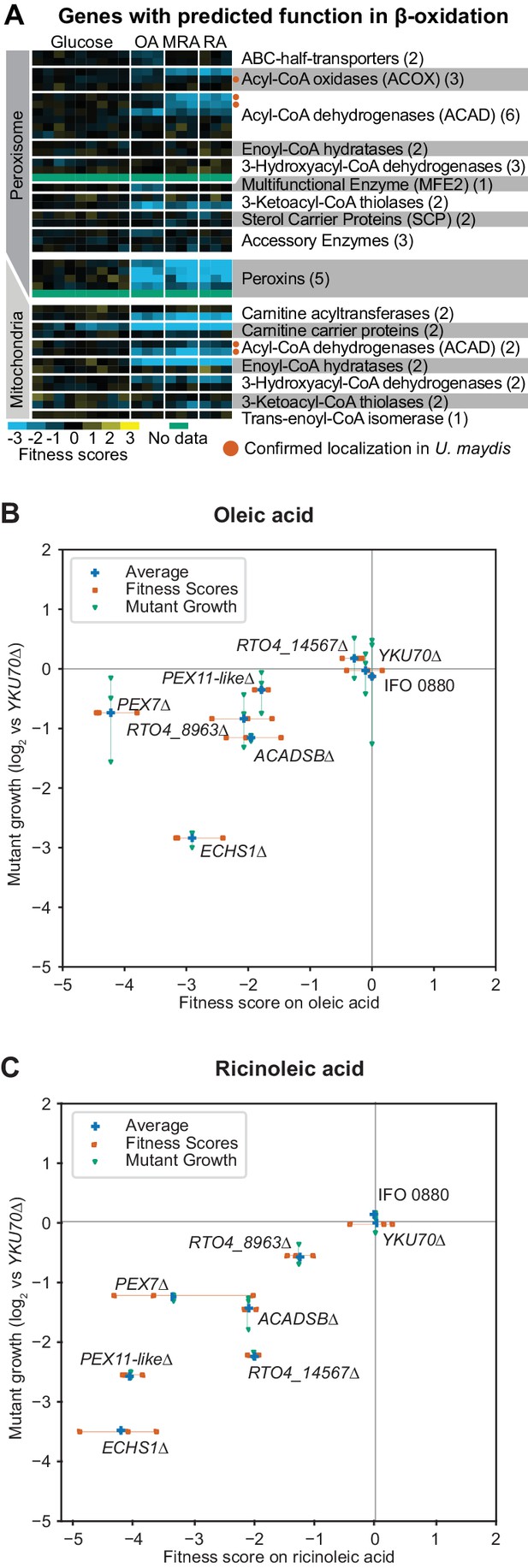
Genes with fitness defects on fatty acids.
(A) Heatmap of fitness scores for R. toruloides genes with predicted roles in beta-oxidation of fatty acids. Enzyme classes and predicted locations were inferred from homologous proteins in Ustilago maydis as reported by Camões et al. (Camões et al., 2015). See Supplementary file 2 for full fitness data. (B) Log2 optical density ratio for single deletion mutants versus the YKU70∆ control strain at mid-log phase on 1% oleic acid as carbon source are plotted against the fitness scores for each gene from BarSeq experiments on 1% oleic acid. (C) Log2 optical density ratio for single deletion mutants versus the YKU70∆ control strain at mid-log phase on 1% ricinoleic acid as carbon source are plotted against the fitness scores for mutants in each gene from BarSeq experiments on 1% ricinoleic acid. See Supplementary file 2 for a statistical summary for all strains shown in (B) and (C), including P values and effect sizes. The following figure supplements are available for Figure 3.
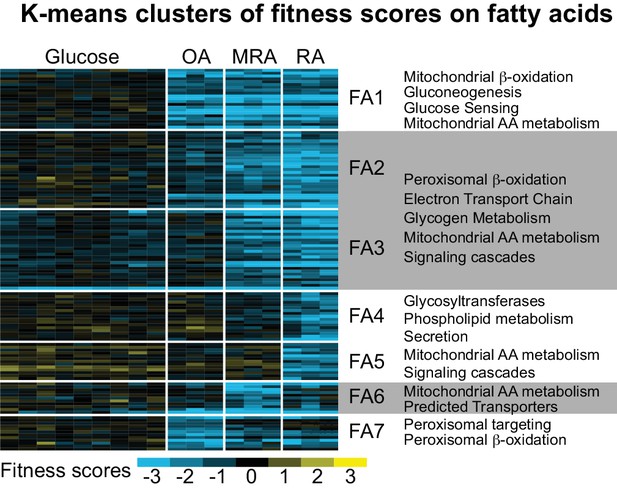
K-means clusters of fitness scores for 129 genes for which mutants have specific fitness defects on fatty acids.
Fitness scores for individual biological replicates were clustered in this analysis (six replicates on glucose, three for each fatty acid). OA: oleic acid, RA: ricinoleic acid, MRA: methyl ricinoleic acid. Seven clusters were identified based on carbon utilization patterns; FA1 - fitness defects on all fatty acids, FA2 and FA3 - fitness defects on MRA and RA, FA4 and FA5 – fitness defects on RA only, FA6 – fitness defects on MRA only, and FA7 – fitness defects on OA only. Major categories of predicted gene functions are summarized for the clusters. See Supplementary file 2 and 3 for full fitness data and gene ontology enrichments.
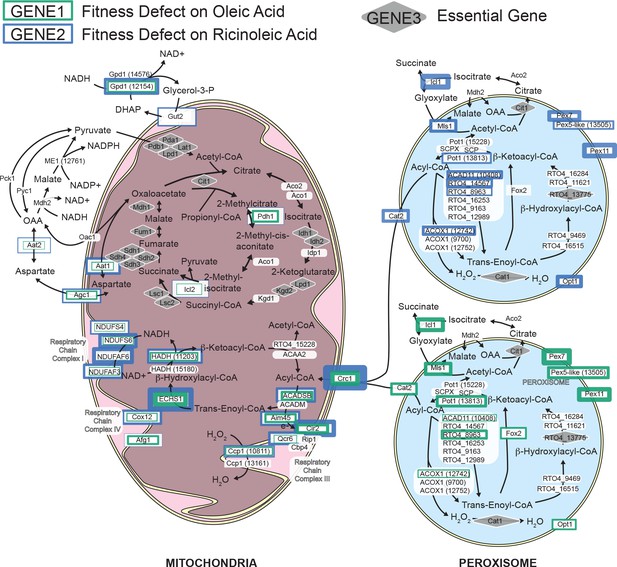
Model for beta-oxidation of fatty acids in R. toruloides.
Fitness scores for genes with predicted roles in mitochondrial and peroxisomal beta-oxidation are represented by the width of green or blue borders around each protein, with wider borders corresponding to lower fitness scores. Green and blue borders represent fitness on oleic and ricinoleic acid, respectively. Fitness scores on fatty acids were consistently most severe for a few genes predicted to mediate steps in mitochondrial beta-oxidation with little to no predicted redundancy in the R. toruloides genome. Fitness scores were more variable between different fatty acids for peroxisomal enzymes, for which more paralogs are present.
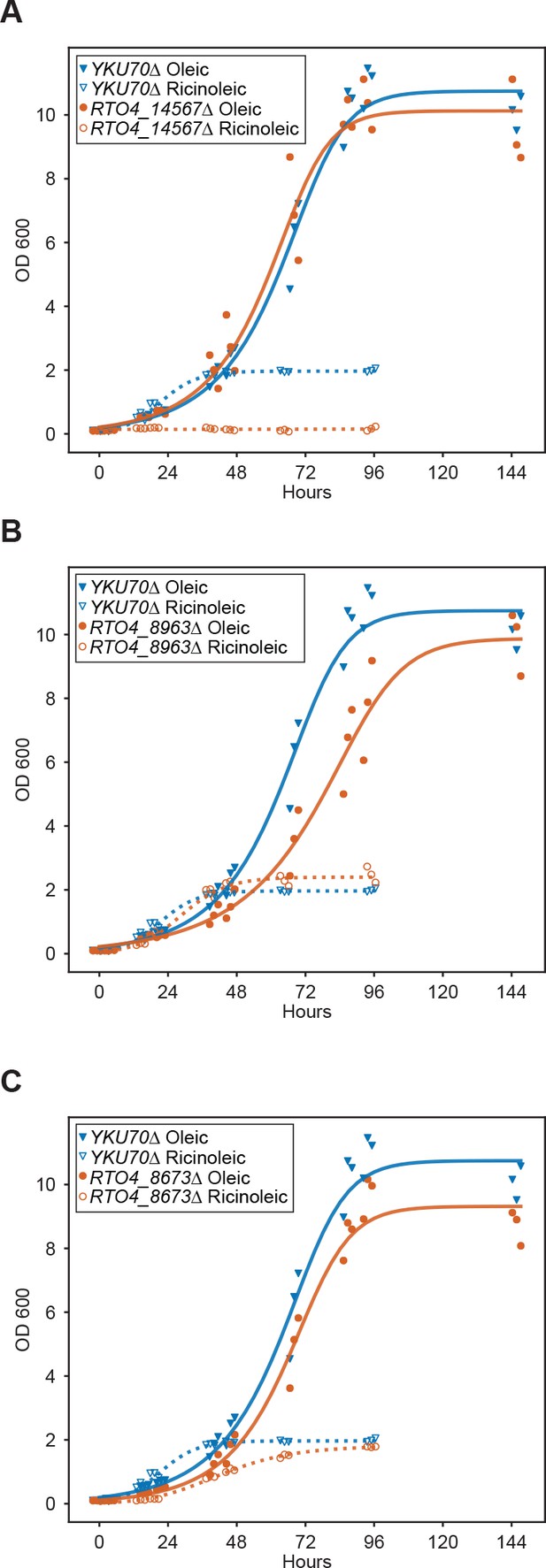
Extended growth curves for deletion mutants on fatty acids.
Growth curves for deletion mutants of (A) RTO4_14567 (similar to H. sapiens ACAD11), (B) RTO4_8963 (similar to H. sapiens ACAD11), and (C) RTO4_8673 (similar to PEX11) on 1% oleic acid and 1% ricinoleic acid as the sole carbon source. See Supplementary file 2 for a statistical summary for all strains including P values and effect sizes.
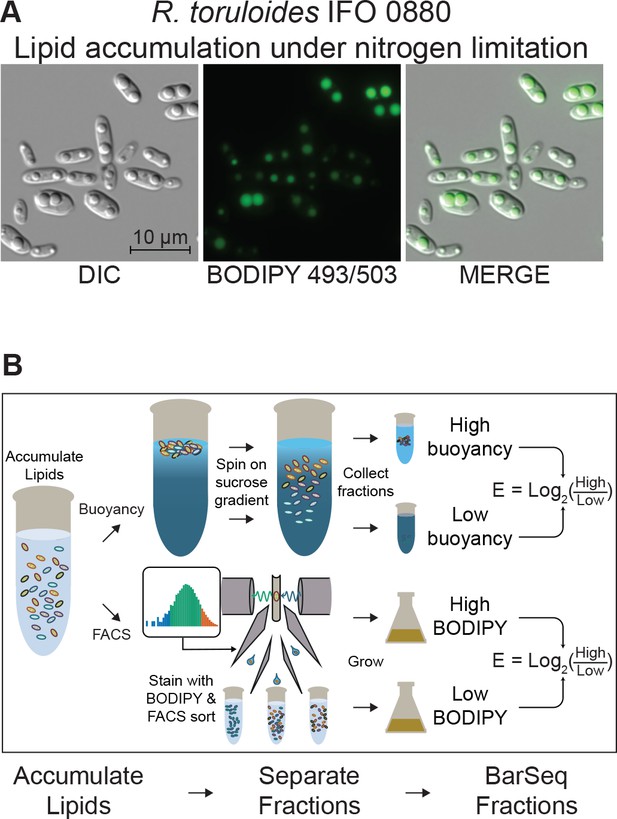
Detecting mutants with altered lipid accumulation.
(A) Lipid accumulation in R. toruloides under nitrogen limitation. DIC microscopy of R. toruloides grown in low nitrogen media for 40 hr and stained with BODIPY 493/503 to label lipid droplets. (B) Two strategies to enrich populations for high or low TAG content cells. (Top) Buoyant density separation on sucrose gradients. Lipid accumulated cells are loaded onto a linear sucrose gradient and centrifuged. Cells settle at their neutral buoyancy, with the size of the low-density lipid droplet as the main driver of buoyancy differences. The gradient is then split into several fractions, and fractions representing the most and least buoyant 5–10% of the population, as well as a no-separation control are subjected to DNA extraction and strain quantification with BarSeq. For each gene an enrichment score is calculated as the log2 ratio of mutant abundance in the high buoyancy versus low buoyancy fractions. (Bottom) FACS sorting on BODIPY signal. Cells cultured in lipid accumulation conditions (limited nitrogen) are stained with BODIPY 493/503, then sorted in a FACS system. The 10% of the population with the highest and lowest BODIPY signal are sorted into enriched populations, as well as non-gated control. These small populations (10 million cells each) are then cultured for additional biomass and subjected to DNA extraction and strain quantification with BarSeq. For each gene, a FACS enrichment score is calculated as the log2 ratio of mutant abundance in the high BODIPY versus low BODIPY fractions. The following figure supplements are available for Figure 4.
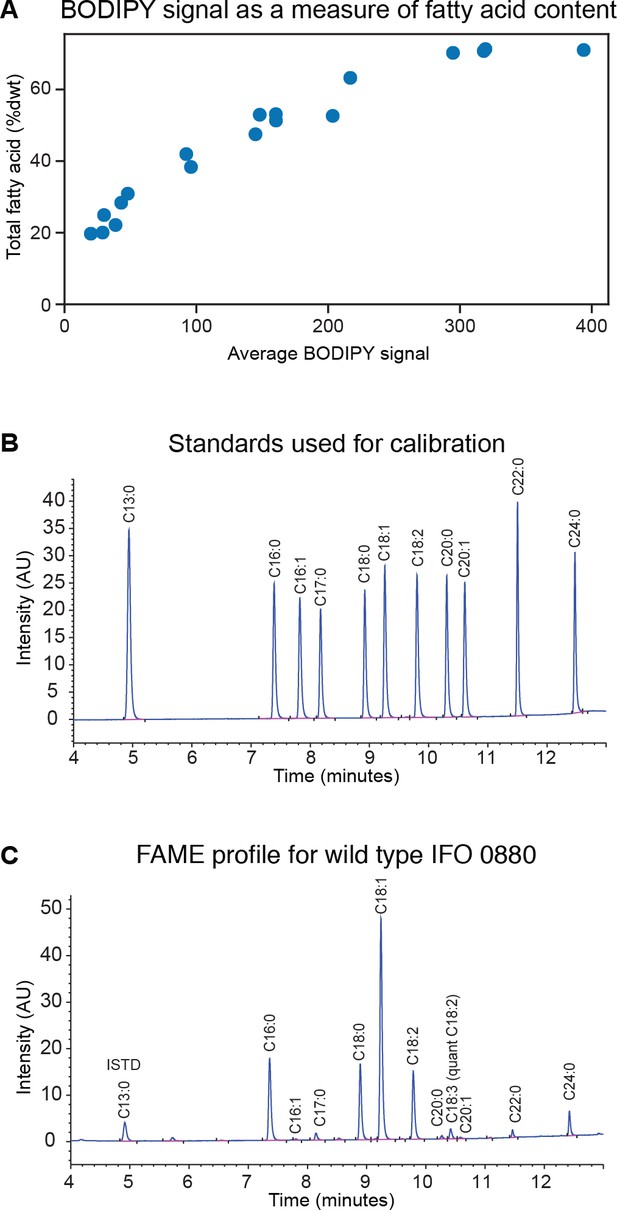
Measuring lipid accumulation under nitrogen limitation.
(A) Total fatty acid methyl ester (FAME) content in R. toruloides cultures, quantified using gas chromatography and flame ionization detection (GC-FID), correlates with average cellular BODIPY signal determined by flow cytometry. (B) Standards used for quantification of FAME content. Peak area/concentration ratios for ten commercially available fatty acid standards were used to quantify FAME peaks from experimental samples. (C) Example FAME profile for IFO 0880. Peak area/concentration ratios for C18:2 were used to quantify C18:3.
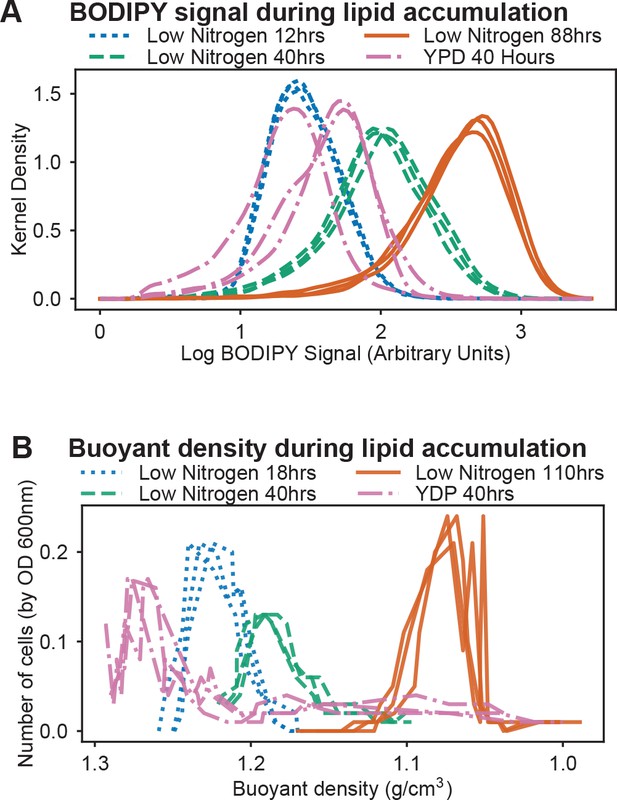
Lipid accumulation and buoyancy changes under nitrogen limitation.
(A) Time course of lipid accumulation (measured by BODIPY intensity) in nitrogen limited media (C/N 120; 12, 40, and 88 hr). Rich media control shown for comparison (YPD at 40 hr). Kernel Density plots for three biological replicates are shown for each growth condition. (B) Time course of buoyant density on sucrose gradients in nitrogen limited media (C/N 120; 18, 40, and 110 hr). Rich media control shown for comparison (YPD at 40 hr). Relative cell numbers were measured by OD 600 nm. Density was measured directly by weight of a 100 µl sample.
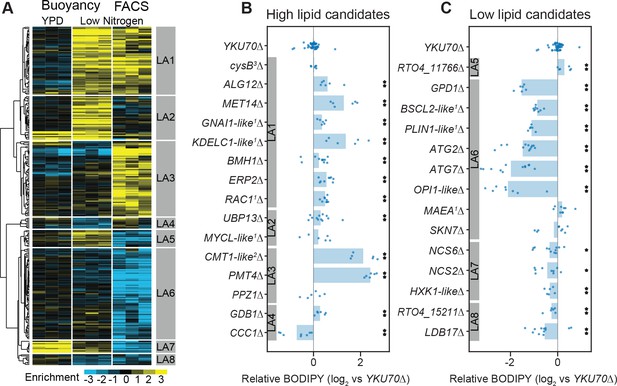
RB-TDNAseq on enriched populations identifies genes affecting lipid accumulation.
(A) Hierarchical clusters of enrichment scores for 271 genes with consistent enrichment (|E| > 1, |T| > 3) in high/low fractions separated by buoyant density or FACS sorting of BODIPY stained cells after lipid accumulation on low nitrogen media. Enrichment scores for individual biological replicates (three per condition) were clustered in this analysis. Eight major clusters were identified (LA1-LA8). See Supplementary file 2 for full enrichment data. (B and C) Relative BODIPY signal for deletion mutants. Points are the average BODIPY/cell for 10,000 cells from independent biological replicate cultures normalized to three control YKU70∆ cultures processed on the same day. Three biological replicates were processed for each strain in any given experiment and each strain was included in at least two experiments processed on different days (N ≥ 6). A statistical summary for all strains including N, P values, and effect sizes is included in Supplementary file 2. **p<0.01, *p<0.05 by homoscedastic T-test versus YKU70∆. 1Human homolog, 2C. neoformans homolog, 3A. nidulans homolog. The following figure supplements are available for Figure 5.
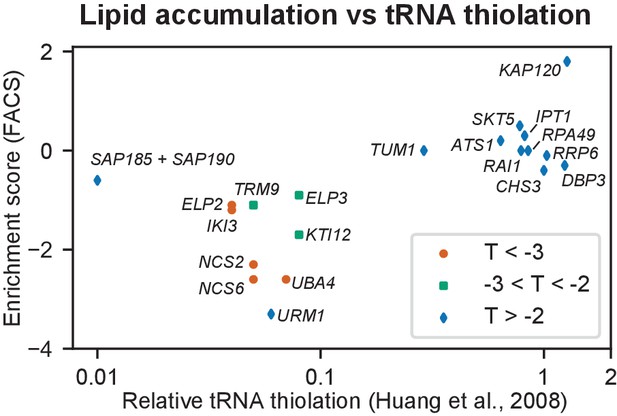
tRNA thiolation in S. cerevisiae versus lipid accumulation in R. toruloides.
Relative levels of tRNA thiolation for S. cerevisiae mutants as reported by Huang et al. (Huang et al., 2008) versus enrichment scores for orthologous R. toruloides genes in the FACS separation experiment after lipid accumulation. Low lipid content (i.e. negative enrichment scores) for R. toruloides mutants corresponds to lower levels of tRNA thiolation in S. cerevisiae mutants.
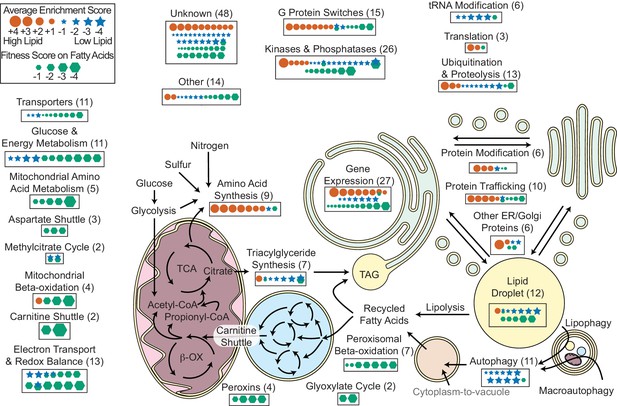
Overview of R. toruloides lipid metabolism.
Key metabolic pathways and cellular functions mediating lipid metabolism as identified from fitness scores on fatty acid and enrichment scores from lipid accumulation screens. Fitness and/or enrichment scores for individual genes are depicted graphically by relative size of hexagonal, circular or star icons respectively. Only fitness scores for genes with consistent growth defects on at least one fatty acid (see Supplementary file 2) and enrichment scores from high confidence clusters (see Figure 5 and Supplementary file 2) are shown. Enrichment scores were averaged between buoyancy and FACS experiments, except for genes with confounding enrichment scores in rich media conditions, for which only FACS data were averaged. Positive scores (orange circles) represent genes for which mutants have increased lipid accumulation. Negative fitness scores (blue stars) represent genes for which mutants have decreased lipid accumulation. Genes detected in proteomics of R. toruloides lipid droplets by Zhu et al. (RAC1, GUT2, PLIN1, EGH1, RIP1, MGL2, AAT1, CIR2, MLS1, and RTO4_8963) or found in lipid droplets of many organisms (DGA1 and BSCL2) (see Supplementary file 5) are depicted under ‘Lipid Droplet’ and also their molecular functions, e.g. ‘G Protein Switches’ for RAC1. The following figure supplements are available for Figure 6.
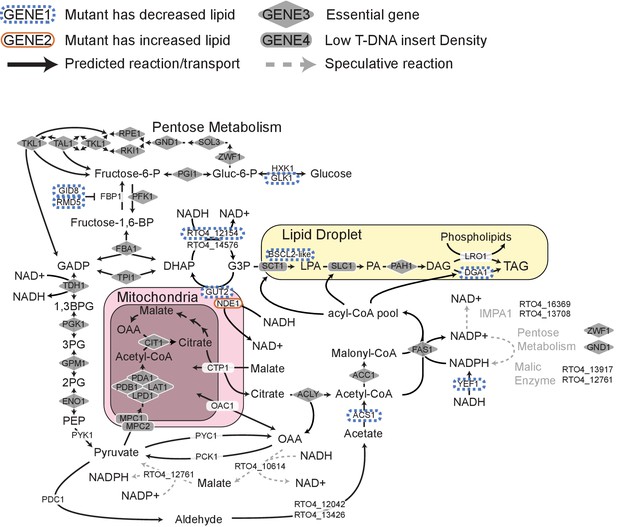
Genes directly effecting TAG biosynthesis in R. toruloides.
Model pathway illustrating genes involved in glycolysis, triacylglyceride (TAG) synthesis, and cytosolic NAD+/NADH balance during TAG synthesis. Genes for which mutants had altered lipid accumulation (enrichment scores in clusters LA1, LA6, LA7, or LA8) are highlighted in orange or blue. Genes with low rates of T-DNA insertion (essential genes and genes for which mutants have a strong growth defect) are highlighted in grey. The primary source of NADPH in R. toruloides remains unclear (see Appendix 1 for detail). Speculative pathways mediating NADPH production are indicated with dashed grey arrows. DAG: diacylglycerol, PA: phosphatidic acid, LPA: lysophosphatidic acid, G3P: glycerol-3-phosphate, DHAP: dihydroxy-acetone-phosphate, GADP: glycerate 3-phosphate, 1,3BPG: 1,3-bisphosphoglycerate, 3 PG: 3-phosphoglycerate, 2 PG: 2-phosphoglycerate, PEP: phosphoenolpyruvate, OAA: oxaloacetate.
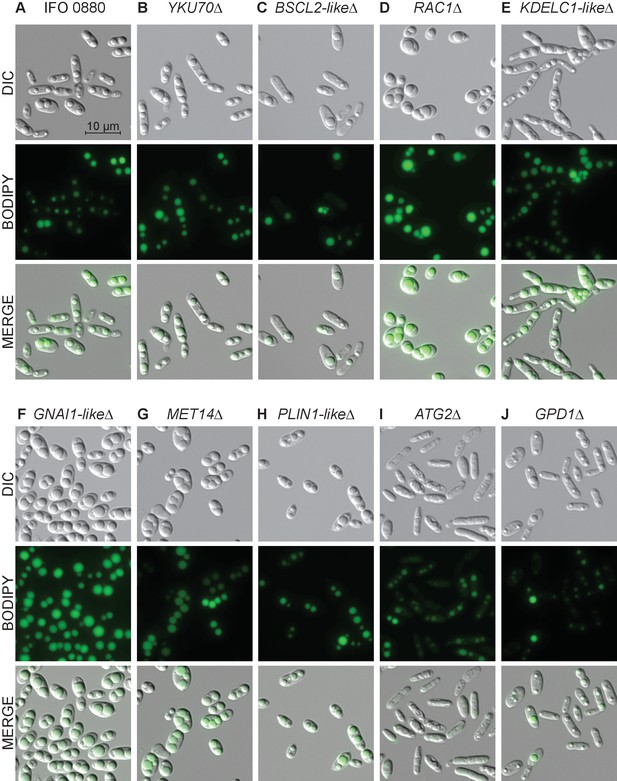
Light and fluorescence microscopy images of selected lipid accumulation mutants.
DIC microscopy on eight deletion mutants for lipid accumulation genes. All deletion mutants (C–J) were constructed in a YKU70∆ background to enable homologous recombination at the targeted locus. Cells were grown 40 hr in low nitrogen lipid accumulation media. DIC, BODIPY 493/503 fluorescence, and composite images are shown for ten strains. (A) R. toruloides IFO 0880 (WT). (B) RTO4_11920∆ ortholog of YKU70. (C) RTO4_11043∆ similar to H. sapiens BSCL2. (D) RTO4_14088∆ ortholog of H. sapiens RAC1. (E) RTO4_10371∆ similar to H. sapiens KDELC1. (F) RTO4_16215∆ similar to H. sapiens GNAI1. (G) RTO4_8709∆ ortholog of MET14. (H) RTO4_16381∆ similar to H. sapiens PLIN1. (I) RTO4_13598∆ ortholog of ATG2. (J) RTO4_12154∆ ortholog of GPD1. The following figure supplements are available for Figure 7.
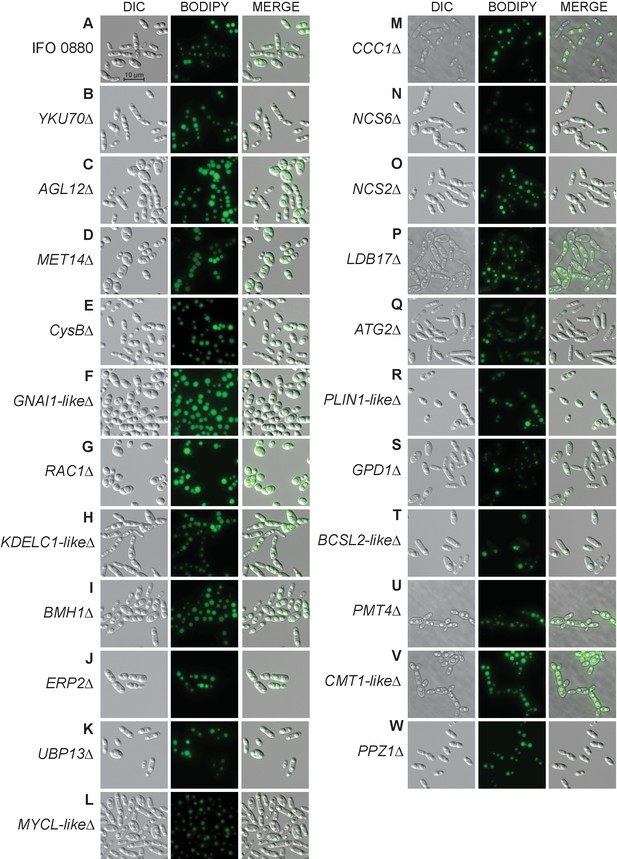
Additional light and fluorescence microscopy images.
DIC microscopy on 21 deletion mutants for lipid accumulation genes. All deletion mutants (C–W) were constructed in a YKU70∆ background to enable homologous recombination at the targeted locus. Cells were grown 40 hr in low nitrogen lipid accumulation media. DIC, BODIPY 493/503 fluorescence, and composite images are shown for 23 strains. (A) R. toruloides IFO 0880 (WT). (B) RTO4_11920∆ ortholog of YKU70. (C) RTO4_11272∆ ortholog of ALG12. (D) RTO4_8709∆ ortholog of MET14. (E) RTO4_12031∆ ortholog of A. nidulans CysB. (F) RTO4_16215∆ similar to H. sapiens GNAI1. (G) RTO4_14088∆ ortholog of H. sapiens RAC1. (H) RTO4_10371∆ similar to H. sapiens KDELC1. (I) RTO4_16644∆ ortholog of BMH1. (J) RTO4_16731∆ ortholog of ERP2. (K) RTO4_9026∆ ortholog of UBP13. (L) RTO4_15890∆ similar to H. sapiens MYCL. (M) RTO4_8506∆ ortholog of CCC1. (N) RTO4_12817∆ ortholog of NCS6. (O) RTO4_10764∆ ortholog of NCS2. (P) RTO4_9970∆ ortholog of LDB17. (Q) RTO4_13598∆ ortholog of ATG2. (R) RTO4_16381∆ similar to H. sapiens PLIN1. (S) RTO4_12154∆ ortholog of GPD1. (T) RTO4_11043∆ similar to H. sapiens BSCL2. (U) RTO4_12121∆ ortholog of PMT4. (V) RTO4_10302∆ similar to C. neoformans CMT1. (W) RTO4_11380∆ ortholog of PPZ1.
Tables
Predicted gene function: Mutants with increased lipid accumulation.
Predicted functions for genes for which mutants were high-confidence candidates for increased lipid accumulation (enrichment scores clustered in LA1, Figure 5).
| Gene ID | Short name | Annotation from | Description | Enrichment | ||
|---|---|---|---|---|---|---|
| BD | FACS | |||||
| G Protein Switches | ||||||
| * | RTO4_15883 | RAS1 | S. cerevisiae | GTPase | 2.0 | 2.3 |
| RTO4_14088 | RAC1 | H. sapiens | GTPase | 2.0 | 0.9 | |
| * | RTO4_16215 | GNAI1-like | H. sapiens | GTPase | 1.6 | 1.0 |
| RTO4_11402 | gapA | A. nidulans | GTPase-activating protein | 0.6 | 1.4 | |
| RTO4_13336 | RIC8A | H. sapiens | Guanine nucleotide exchange factor | 1.3 | 1.4 | |
| RTO4_16170 | sif-like | D. melanogaster | Guanine nucleotide exchange factor | 1.5 | 0.9 | |
| RTO4_16644 | BMH1 | S. cerevisiae | 14-3-3 protein | 1.3 | 2.2 | |
| RTO4_16068 | BMH1 | S. cerevisiae | 14-3-3 protein | 0.7 | 1.2 | |
| Kinases and Phosphatases | ||||||
| RTO4_13246 | CNA1 | S. cerevisiae | Phosphatase (Calcineurin catalytic subunit) | 0.8 | 1.2 | |
| RTO4_11675 | CNB1 | S. cerevisiae | Phosphatase (Calcineurin regulatory subunit) | 1.1 | 1.2 | |
| RTO4_11667 | PTC1 | S. cerevisiae | Phosphatase | 0.9 | 1.2 | |
| RTO4_10638 | CLA4 | S. cerevisiae | Kinase | 3.4 | 4.5 | |
| * | RTO4_16605 | TPK1 | S. cerevisiae | Kinase | 1.1 | 0.5 |
| Gene Expresssion | ||||||
| RTO4_10333 | SET1 | S. cerevisiae | Chromatin modifying | 3.0 | 1.1 | |
| RTO4_10279 | BRE2 | S. cerevisiae | Chromatin modifying | 2.5 | 1.0 | |
| RTO4_12689 | SPP1 | S. cerevisiae | Chromatin modifying | 2.0 | 1.3 | |
| RTO4_15412 | RCO1 | S. cerevisiae | Chromatin modifying | 3.5 | 1.6 | |
| RTO4_10209 | MIT1-like | S. cerevisiae | Transcripition factor | 1.4 | 0.3 | |
| RTO4_14550 | CYC8 | S. cerevisiae | Transcription factor | 3.7 | 3.8 | |
| RTO4_10274 | SKN7-like | S. cerevisiae | Transcription factor | 2.2 | 1.5 | |
| RTO4_13346 | CBC2 | S. cerevisiae | RNA splicing factor | 1.6 | 1.2 | |
| Protein Modification | ||||||
| RTO4_11272 | ALG12 | S. cerevisiae | Alpha-1,6-mannosyltransferase | 3.5 | 1.7 | |
| RTO4_14881 | CAP10-like | C. neoformans | Xylosyltransferase | 1.5 | 2.0 | |
| RTO4_16598 | LARGE1 | H. sapiens | N-acetylglucosaminyltransferase-like protein | 1.8 | 1.3 | |
| Protein Trafficking | ||||||
| RTO4_12145 | ERP1 | S. cerevisiae | COPII cargo adapter protein (p24 family) | 2.4 | 2.7 | |
| RTO4_16731 | ERP2 | S. cerevisiae | COPII cargo adapter protein (p24 family) | 1.7 | 2.0 | |
| RTO4_12521 | EMP24 | S. cerevisiae | COPII cargo adapter protein (p24 family) | 1.9 | 2.4 | |
| RTO4_14054 | BST1 | S. cerevisiae | GPI inositol-deacylase | 1.5 | 0.2 | |
| * | RTO4_15883 | RAS1 | S. cerevisiae | GTPase | 2.0 | 2.3 |
| Other ER/Golgi Proteins | ||||||
| RTO4_10371 | KDELC1-like | H. sapiens | Endoplasmic reticulum protein EP58 | 3.1 | 6.0 | |
| RTO4_15763 | SH3 Domain-containing ER Protein | 1.0 | 1.5 | |||
| Amino Acid Biosynthesis | ||||||
| RTO4_11050 | MET1 | S. cerevisiae | Uroporphyrinogen III transmethylase | 3.8 | 2.0 | |
| RTO4_8744 | MET5 | S. cerevisiae | Sulfite reductase | 4.4 | 2.1 | |
| § | RTO4_10374 | MET10 | S. cerevisiae | Sulfite reductase | 2.5 | 1.3 |
| RTO4_8709 | MET14 | S. cerevisiae | Adenylylsulfate kinase | 4.1 | 1.1 | |
| RTO4_11741 | MET16 | S. cerevisiae | Phosphoadenosine phosphosulfate reductase | 1.7 | 1.1 | |
| RTO4_12031 | cysB | A. nidulans | Cysteine synthase A | 3.3 | 2.1 | |
| * | RTO4_16196 | ARG1 | S. cerevisiae | Argininosuccinate synthase | 1.3 | 1.8 |
| Translation | ||||||
| RTO4_12273 | MRN1 | S. cerevisiae | RNA-binding protein | 2.5 | 1.6 | |
| RTO4_8595 | EIF4E2 | H. sapiens | Translation initiation factor | 2.0 | 0.5 | |
| Ubiquitination and Proteolysis | ||||||
| RTO4_11150 | Mub1-like | S. cerevisiae | Ubiquitin ligase complex member | 3.8 | 2.0 | |
| RTO4_15576 | CDC4 | S. cerevisiae | Ubiquitin ligase complex member | 1.7 | 1.8 | |
| Triacylglyceride Synthesis | ||||||
| † | RTO4_8972 | NDE1 | S. cerevisiae | NADH dehydrogenase | 1.6 | 1.9 |
| Lipid Droplet Associated | ||||||
| RTO4_14088 | RAC1 | H. sapiens | GTPase | 2.0 | 0.9 | |
| Mitochondrial Beta-oxidation | ||||||
| RTO4_16284 | HSD17B10 | H. sapiens | 3-hydroxyacyl-CoA dehydrogenase | 1.6 | 0.5 | |
| Other | ||||||
| RTO4_12175 | mesA | A. nidulans | Myosin binding protein | 1.3 | 1.8 | |
| RTO4_8401 | SHE4 | S. cerevisiae | Transmembrane protein involved in cell polarity | 1.0 | 1.3 | |
| Unknown Function | ||||||
| RTO4_16524 | Protein of unknown function | 3.1 | 1.9 | |||
| RTO4_11613 | Protein of unknown function | 2.5 | 1.7 | |||
| RTO4_12505 | Protein of unknown function | 2.1 | 2.1 | |||
| RTO4_13512 | Protein of unknown function | 1.5 | 1.9 | |||
| RTO4_10805 | Protein of unknown function | 1.2 | 1.8 | |||
| RTO4_15251 | Protein of unknown function | 1.6 | 1.3 | |||
| RTO4_15358 | Protein of unknown function | 2.0 | 0.5 | |||
| RTO4_13513 | Protein of unknown function | 1.3 | 1.2 | |||
| RTO4_12461 | Protein of unknown function | 1.5 | 0.8 | |||
| RTO4_13351 | Protein of unknown function | 1.2 | 1.0 | |||
-
Cellular processes grouped as in Figure 6. BD: Enrichment score from buoyant density separation. FACS: Enrichment score from fluorescence activated cell sorting.
Protein abundance under nitrogen limitation: * increased; † increased 10-fold or more; ‡ decreased; § decreased 10-fold or more (Zhu et al., 2012).
Predicted gene function: Mutants with decreased lipid accumulation.
Predicted functions for genes for which mutants were high-confidence candidates for decreased lipid accumulation (enrichment scores clustered in LA6 - LA8, Figure 5).
| Gene ID | Short name | Annotation from | Description | Cluster | Enrichment | ||
|---|---|---|---|---|---|---|---|
| BD | FACS | ||||||
| tRNA thiolation | |||||||
| RTO4_10764 | NCS2 | S. cerevisiae | tRNA 2-thiolation protein | LA7 | 0.5 | −2.3 | |
| RTO4_12817 | NCS6 | S. cerevisiae | tRNA 2-thiolation protein | LA7 | 0.7 | −2.6 | |
| RTO4_14918 | ELP2 | S. cerevisiae | Elongator complex protein | LA7 | 0.7 | −1.2 | |
| RTO4_14716 | IKI3 | S. cerevisiae | Elongator complex protein | LA7 | 0.4 | −1.1 | |
| RTO4_11341 | UBA4 | S. cerevisiae | Adenylyltransferase and sulfurtransferase | LA7 | 0.6 | −2.6 | |
| G Protein Switches | |||||||
| † | RTO4_15198 | Rab6 | H. sapiens | GTPase | LA6 | −1.3 | −1.6 |
| RTO4_14622 | RGP1 | H. sapiens | Guanine nucleotide exchange factor | LA6 | −1.4 | −1.5 | |
| Kinases and Phosphatases | |||||||
| RTO4_10698 | VHS1 | S. cerevisiae | Kinase | LA6 | 0.8 | −3.7 | |
| RTO4_16375 | HRK1 | S. cerevisiae | Kinase | LA6 | 0.4 | −2.2 | |
| * | RTO4_11453 | GLC7 | S. cerevisiae | Kinase | LA8 | −1.2 | −0.9 |
| RTO4_16810 | KIN1 | S. cerevisiae | Kinase | LA6 | 0.1 | −1.1 | |
| RTO4_10025 | SAT4 | S. cerevisiae | Kinase | LA7 | 1.6 | −3.6 | |
| RTO4_13327 | ATG1 | S. cerevisiae | Kinase | LA6 | 0.1 | −2.5 | |
| RTO4_14907 | SCH9 | S. cerevisiae | Kinase | LA6 | −0.6 | −2.0 | |
| RTO4_14906 | kinase-like | S. cerevisiae | Kinase | LA6 | −0.3 | −1.8 | |
| RTO4_13290 | YAK1 | S. cerevisiae | Kinase | LA8 | −1.1 | −0.9 | |
| RTO4_11732 | PPH3 | S. cerevisiae | Phosphatase 4 catalytic subunit | LA6 | 0.9 | −3.6 | |
| RTO4_12586 | PSY2 | S. cerevisiae | Phosphatase 4 regulatory subunit | LA6 | 0.2 | −1.2 | |
| RTO4_16463 | PTC7-like | S. cerevisiae | Phosphatase | LA6 | 0.1 | −2.0 | |
| Autophagy | |||||||
| RTO4_13327 | ATG1 | S. cerevisiae | Kinase | LA6 | 0.1 | −2.5 | |
| RTO4_13598 | ATG2 | S. cerevisiae | Membrane protein | LA6 | −0.6 | −3.4 | |
| RTO4_12968 | ATG3 | S. cerevisiae | Ubiquitin-like-conjugating enzyme | LA6 | −0.8 | −4.5 | |
| RTO4_13496 | ATG4 | S. cerevisiae | Cysteine protease | LA6 | −0.1 | −2.3 | |
| RTO4_11901 | ATG7 | S. cerevisiae | Ubiquitin-like modifier-activating enzyme | LA6 | −0.8 | −4.2 | |
| RTO4_13543 | ATG8 | S. cerevisiae | Ubiquitin-like protein | LA6 | −1.0 | −4.2 | |
| RTO4_11326 | ATG9 | S. cerevisiae | Membrane protein | LA6 | 0.0 | −1.3 | |
| RTO4_9008 | ATG14 | S. cerevisiae | Autophagy-specific subunit of PtdIns3P-kinase complex | LA6 | 0.0 | −5.0 | |
| RTO4_16723 | ATG18 | S. cerevisiae | Phosphoinositide binding protein | LA6 | −0.9 | −5.8 | |
| Ubiquitination and Proteolysis | |||||||
| † | RTO4_16672 | PRB1 | S. cerevisiae | Vacuolar proteinase | LA6 | −0.2 | −1.7 |
| RTO4_15345 | SIS1 | S. cerevisiae | Protein chaperone | LA6 | −0.4 | −1.2 | |
| RTO4_10423 | RMD5 | S. cerevisiae | GID complex E3 ubiquitin ligase | LA6 | −0.4 | −2.0 | |
| RTO4_11737 | GID8 | H. sapiens | GID complex member | LA6 | −0.1 | −1.5 | |
| RTO4_9816 | LONRF1 | H. sapiens | E3 ubiquitin ligase | LA6 | −0.5 | −4.5 | |
| RTO4_15320 | USP48 | H. sapiens | Ubiquitin carboxyl-terminal hydrolase | LA6 | 0.0 | −1.2 | |
| RTO4_9600 | COPS3 | H. sapiens | COP9 signalosome complex subunit | LA1 | 1.4 | 0.6 | |
| RTO4_11569 | GPS1 | H. sapiens | COP9 signalosome complex subunit | LA6 | 0.7 | −2.1 | |
| Triacylglyceride Synthesis | |||||||
| † | RTO4_12154 | GPD1 | S. cerevisiae | Glycerol-3-phosphate dehydrogenase | LA6 | −1.7 | −4.0 |
| RTO4_11043 | BCSL2-like | H. sapiens | Seipin | LA6 | −0.8 | −2.9 | |
| RTO4_16460 | DGA1 | H. sapiens | Diacylglycerol acyltransferase | LA6 | −0.7 | −4.0 | |
| RTO4_14597 | ACS1 | S. cerevisiae | Acetyl-CoA synthetase | LA8 | −1.7 | −1.0 | |
| RTO4_10182 | YEF1 | S. cerevisiae | NAD+/NADH kinase | LA6 | −0.1 | −1.6 | |
| ‡ | RTO4_11039 | GUT2 | S. cerevisiae | Glycerol-3-phosphate dehydrogenase | LA6 | −0.2 | −1.1 |
| Lipid Droplet Associated | |||||||
| RTO4_16381 | PLIN1-like | S. cerevisiae | Perilipin | LA6 | −1.7 | −4.3 | |
| ‡ | RTO4_11039 | GUT2 | S. cerevisiae | Glycerol-3-phosphate dehydrogenase | LA6 | −0.2 | −1.1 |
| RTO4_15372 | EGH1 | S. cerevisiae | Steryl-beta-glucosidase | LA6 | 0.7 | −2.5 | |
| RTO4_13614 | RIP1 | S. cerevisiae | Mitochondrial complex III iron-sulfur protein | LA6 | −0.5 | −2.8 | |
| RTO4_11043 | BCSL2-like | H. sapiens | Seipin | LA6 | −0.8 | −2.9 | |
| RTO4_16460 | DGA1 | H. sapiens | Diacylglycerol acyltransferase | LA6 | −0.7 | −4.0 | |
| Protein Modification | |||||||
| RTO4_12670 | B3GALT1-like | H. sapiens | Beta-1,3-Galactosyltransferase | LA6 | −0.9 | −3.1 | |
| Protein Trafficking | |||||||
| † | RTO4_15198 | Rab6 | H. sapiens | GTPase | LA6 | −1.3 | −1.6 |
| Other ER/Golgi Proteins | |||||||
| RTO4_8838 | DNAJC4 | H. sapiens | DnaJ family chaperone | LA6 | −0.8 | −1.3 | |
| RTO4_13971 | DNAJC3 | H. sapiens | DnaJ family chaperone | LA6 | −1.1 | −2.2 | |
| Gene Expression | |||||||
| RTO4_11333 | KLF18-like | H. sapiens | Transcription factor | LA6 | −0.2 | −1.1 | |
| RTO4_15641 | SKN7 | S. cerevisiae | Transcription factor | LA6 | 0.9 | −2.9 | |
| RTO4_14676 | LHX5-like | H. sapiens | Transcription factor | LA6 | −0.2 | −2.8 | |
| RTO4_11891 | HAP2 | S. cerevisiae | Transcription factor | LA6 | −0.8 | −2.4 | |
| RTO4_12420 | OPI1-like | S. cerevisiae | Transcription factor | LA6 | 0.0 | −3.7 | |
| RTO4_14100 | HAPX | C. neoformans | Transcription factor | LA8 | −1.2 | −1.7 | |
| RTO4_13255 | SGF73 | S. cerevisiae | SAGA-associated factor | LA6 | 0.4 | −1.5 | |
| Methylcitrate Cycle | |||||||
| RTO4_14162 | ICL2 | S. cerevisiae | 2-methylisocitrate lyase | LA6 | −0.3 | −1.8 | |
| RTO4_12642 | PDH1 | S. cerevisiae | 2-methylcitrate dehydratase | LA6 | −0.1 | −1.7 | |
| Electron Transport and Redox Balancing | |||||||
| RTO4_11165 | CBP4 | S. cerevisiae | Mitochondrial complex III assembly factor | LA6 | −0.4 | −2.5 | |
| RTO4_13614 | RIP1 | S. cerevisiae | Mitochondrial complex III iron-sulfur protein | LA6 | −0.5 | −2.8 | |
| RTO4_13902 | AFG1 | S. cerevisiae | Mitochondrial complex IV assembly factor | LA6 | −0.3 | −1.3 | |
| ‡ | RTO4_10010 | NDUFS4 | H. sapiens | Mitochondrial complex I accessory factor | LA8 | −1.3 | −0.1 |
| RTO4_13925 | NDUFAF3 | H. sapiens | Mitochondrial complex I assembly factor | LA8 | −1.0 | −1.6 | |
| Amino Acid Biosynthesis | |||||||
| † | RTO4_12302 | CPA2 | S. cerevisiae | Large subunit of carbamoyl phosphate synthetase | LA6 | −0.4 | −2.4 |
| Glucose and Energy Metabolism | |||||||
| RTO4_10423 | RMD5 | S. cerevisiae | GID complex E3 ubiquitin ligase | LA6 | −0.4 | −2.0 | |
| RTO4_11737 | GID8 | H. sapiens | GID complex member | LA6 | −0.1 | −1.5 | |
| RTO4_12034 | TPS2 | S. cerevisiae | Trehalose 6-phosphate synthase | LA6 | 0.0 | −3.8 | |
| * | RTO4_10264 | GLK1 | S. cerevisiae | Hexokinase | LA7 | 2.1 | −2.0 |
| Transporters | |||||||
| † | RTO4_12909 | OAT1 | C. neoformans | Nucleobase transporter | LA6 | −0.2 | −1.1 |
| RTO4_11397 | COT1 | S. cerevisiae | Vacuolar zinc transporter | LA6 | −0.2 | −1.1 | |
| RTO4_11924 | SNF3 | S. cerevisiae | Plasma membrane low glucose sensor | LA6 | 0.0 | −2.8 | |
| Other | |||||||
| RTO4_12512 | cry | N. crassa | Blue-light photoreceptor cryptochrome | LA7 | 0.6 | −1.6 | |
| RTO4_14974 | Steroidogenesis/phosphatidylcholine transfer domain | LA6 | −0.3 | −1.2 | |||
| RTO4_15889 | MAEA | H. sapiens | EMP macrophage erythroblast attacher | LA6 | −0.1 | −1.7 | |
| RTO4_16287 | CDD1 | S. cerevisiae | Cytidine deaminase | LA6 | 0.3 | −2.3 | |
| RTO4_15247 | WDR26 | H. sapiens | WD repeat protein | LA6 | −0.9 | −1.3 | |
| RTO4_8764 | MGS1 | S. cerevisiae | DNA-dependent ATPase and ssDNA annealing protein | LA6 | 0.2 | −1.2 | |
| Unknown | |||||||
| RTO4_10431 | Protein of unknown function | LA6 | 0.7 | −1.6 | |||
| RTO4_8973 | Protein of unknown function | LA8 | −0.2 | −1.1 | |||
| RTO4_13195 | Protein of unknown function | LA6 | −0.2 | −1.1 | |||
| RTO4_10367 | Protein of unknown function | LA6 | −0.1 | −1.3 | |||
| RTO4_10102 | Protein of unknown function | LA6 | −0.3 | −1.2 | |||
| RTO4_14926 | Protein of unknown function | LA6 | 0.2 | −1.7 | |||
| RTO4_12045 | Protein of unknown function | LA6 | 0.0 | −1.5 | |||
| RTO4_13600 | Protein of unknown function | LA6 | −0.3 | −1.3 | |||
| RTO4_10976 | Protein of unknown function | LA6 | −0.2 | −1.5 | |||
| RTO4_9970 | LDB17 | S. cerevisiae | Protein of unknown function | LA8 | −1.3 | −0.5 | |
| RTO4_13435 | Protein of unknown function | LA7 | 0.2 | −2.0 | |||
| RTO4_9692 | Protein of unknown function | LA6 | −0.5 | −1.4 | |||
| RTO4_15521 | Protein of unknown function | LA6 | 0.2 | −2.2 | |||
| RTO4_8769 | Protein of unknown function | LA6 | −0.5 | −1.6 | |||
| RTO4_8770 | Protein of unknown function | LA6 | −0.5 | −1.9 | |||
| RTO4_11259 | Protein of unknown function | LA7 | 0.7 | −3.3 | |||
| RTO4_9490 | Protein of unknown function | LA6 | −0.6 | −2.4 | |||
| RTO4_15520 | Protein of unknown function | LA6 | −0.5 | −2.5 | |||
| RTO4_8771 | Protein of unknown function | LA6 | −0.6 | −2.5 | |||
| RTO4_13452 | Protein of unknown function | LA6 | −1.3 | −4.0 | |||
| RTO4_15211 | Protein of unknown function | LA8 | −1.1 | −1.5 | |||
-
Cellular processes grouped as in Figure 6. BD: Enrichment score from buoyant density separation. FACS: Enrichment score from fluorescence activated cell sorting.
Protein abundance under nitrogen limitation: * increased; † increased 10-fold or more; ‡ decreased; § decreased 10-fold or more (Zhu et al., 2012).
Sucrose density gradients used in this study
https://doi.org/10.7554/eLife.32110.026| Media | Time | Sucrose range (Density)* | Average density ±StDev | High buoyancy fractions (Density) | Median buoyancy fractions (Density) | Low buoyancy fractions (Density) |
|---|---|---|---|---|---|---|
| Low Nitrogen | 40 hr | 50–20% (1.22–1.10) | 1.177 ±0.003 | 17–20 (<1.11) | 6–7 (1.18–1.19) | 1–2 (>1.21) |
| YPD | 40 hr | 80–50% (1.29–1.16) | 1.234 ±0.012 | 19–22† (<1.14) | 4–8† (1.24–1.27) | 1 (>1.28) |
-
All density measurements in g/mL
*Highest and lowest specific density measured in any collected fraction in the linear portion of the gradient.
-
†Some biological replicates differ in exact fractions collected. Fractions were collected within this range such that the high buoyancy fraction constituted the most buoyant 5–10% of the population, the median buoyancy fraction constituted the median 30–50% of the population and the low buoyancy fraction constituted the least buoyant 5–10% of the population.
Additional files
-
Supplementary file 1
R. toruloides Genome Summary.
Overview of the revised genome assembly of R. toruloides IFO 0880 with summary information on gene models, predicted functions, orthologous genes, manually curated annotations for genes with lipid utilization and accumulation phenotypes observed in this study, and likely essential genes identified by recalcitrance to T-DNA insertion.
- https://doi.org/10.7554/eLife.32110.027
-
Supplementary file 2
BarSeq and Mutant Data.
Fitness and enrichment scores from BarSeq data in all conditions/fractions tested in this study; fitness/enrichment scores used in clustering analysis for Figures 2, 3 and 5; raw data for growth of validation deletion mutant strains on fatty acids; and summary statistics for all validation experiments with deletion mutants, including flow cytometry on lipid accumulation mutants.
- https://doi.org/10.7554/eLife.32110.028
-
Supplementary file 3
GO Enrichments.
GO term enrichments for genes clustered in Figure 3—figure supplement 1 and Figure 5A.
- https://doi.org/10.7554/eLife.32110.029
-
Supplementary file 4
Strains, Primers, and Libraries.
High throughput sequencing libraries used in this study, strains constructed in this study, and primers used for strain construction on RB-TDNAseq analysis.
- https://doi.org/10.7554/eLife.32110.030
-
Supplementary file 5
Functional Lipid Surveys.
Summary of genes identified in 35 systems-level studies on lipid accumulation, lipid catabolism, and lipid droplet biology in fungi and other eukaryotes with functional genomics screens and proteomics.
- https://doi.org/10.7554/eLife.32110.031
-
Supplementary file 6
FAME versus BODIPY.
Raw data for Figure 4—figure supplement 1 comparing total lipid from gas chromatography of fatty acid methyl esters to average BODIPY intensity in samples of wild type and mutant strains of R. toruloides.
- https://doi.org/10.7554/eLife.32110.032
-
Transparent reporting form
- https://doi.org/10.7554/eLife.32110.033





