A population of innate myelolymphoblastoid effector cell expanded by inactivation of mTOR complex 1 in mice
Figures
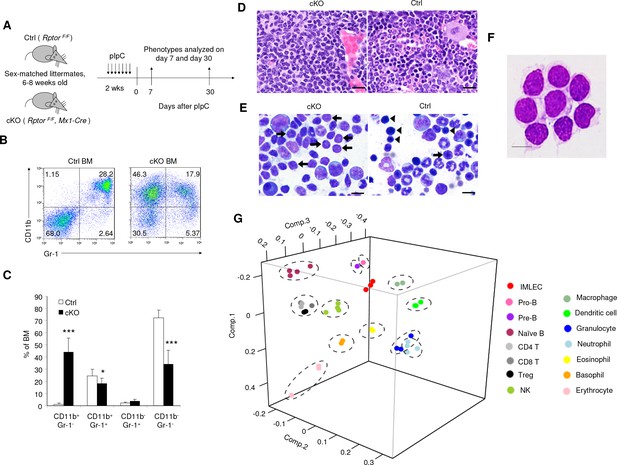
Targeted mutation of Rptor in hematopoiesis led to massive accumulation of IMLEC.
(A) Schematic of experimental design. Sex-matched 6–8 weeks old Ctrl (Rptor F/F) and cKO (Rptor F/F, Mx1-Cre) mice were treated with pIpC for seven times. The phenotypes were analyzed on day 7 or day 30 after the complement of pIpC treatment. (B) Representative flow cytometric analysis of myeloid cells from mice BM by CD11b and Gr-1 on day seven after pIpC treatment. Similar data were obtained on day 30. (C) Frequencies of various hematopoietic cell populations in BM based on CD11b and Gr-1 markers. n = 7 for Ctrl mice; n = 9 for cKO mice. Data are pooled from three independent experiments. (D) Expansion of lymphoblast population in cKO BM revealed by histology. Left, H&E staining showing prominence of cells with lymphoid morphology in the cKO BM; Right, normal BM histology from Ctrl mouse showing normal myeloid and erythroid lineages. Scale bar, 20 μm. (E) Expansion of lymphoblast revealed by cytology. Left, cytological preparation of BM smear from cKO mouse showing preponderance of lymphocytes (short arrows) and severe depletion of myeloid and erythroid lineages. There were some promyelocytes and myelocytes present (larger cells) but no mature neutrophils. Right, normal cytological preparation of BM from Ctrl mouse. There were numerous erythroid precursors with intensely basophilic, condensed chromatin (arrowheads) and a few lymphocytes (short arrow) with less condensed chromatin. Additionally, in Ctrl BM there were numerous mature neutrophils (ringed nucleus with constrictions) which were severely depleted in the cKO mouse. Scale bar, 10 μm. (F) Giemsa staining of FACS-sorted Rptor-deficient CD11b+Gr-1- BM cells. Scale bar, 10 μm. Similar morphology was observed in three independent experiments. (G) Principal component analysis (PCA) of gene expression in CD11b+ Gr-1- BM cells (IMLECs) and other hematopoietic cells. Numbers along axes indicate relative scaling of the principal variables. RNA-seq data from IMLECs obtained in our study were compared with those deposited in public database by others. Datasets are from known lymphoid (Pro-B, Pre-B, Naïve B, CD4 T, CD8 T, Treg and NK) or myeloid (macrophage, dendritic cell, granulocyte, neutrophil, eosinophil, basophil and erythrocyte) subsets.
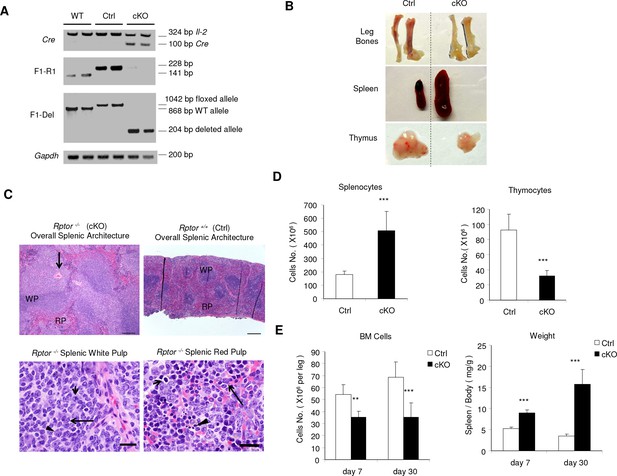
Conditional deletion of Rptor resulted in abnormal hematopoiesis.
(A) Deletion of Rptor in BM cells. PCR were performed to check the deletion in BM from mice 2 weeks after pIpC treatment (for Ctrl and cKO mice, no treatment for WT mice). (B) Representative pictures of leg bones (tibiae and femurs), spleen, and thymus harvested from mice on day 30 post pIpC treatment. (C) Histology findings in the cKO spleen by H&E staining. Up left panel: a spleen histological section showing expanded white pulp areas (WP) and compressed intervening red pulp (RP). The white pulp contains an increased population of lightly staining cells that sometimes is situated in the marginal zones and follicular centers (B cell areas) and sometimes infiltrates the periarteriolar sheaths (T cell area, long arrow). Up right panel: normal splenic architecture from control mouse. Scale bar, 200 μm. Down left panel: higher magnification of expanded cell population within the splenic white pulp. Morphologically these cells resemble germinal center lymphocytes. There are both smaller cells with cleaved nuclei (long arrow) resembling centrocytes and larger cells with 1–2 prominent nucleoli resembling centroblasts (short arrow). There are small numbers of plasmacytoid cells (arrowhead). Down right panel: histological section of splenic red pulp Red pulp of the spleen showing foci of extramedullary hematopoiesis with decreased erythroid (short arrow) and myeloid (long arrow) progenitors. The erythroid lineage contains occasional foci of pyknotic or karyorrhectic (fragmented) nuclei consistent with cell death (asterisk). Scale bar, 20 μm. (D) The cellularities of splenocytes (left) and thymocytes (right) from mice at day 30 post pIpC treatment. n = 6 for Ctrl mice; n = 5 for cKO mice. (E) BM cellularities (left) and spleen/body weight ratios (right) for mice on day 7 and day 30 after pIpC treatment. n = 3 ~ 8 for Ctrl mice; n = 5 ~ 6 for cKO mice.
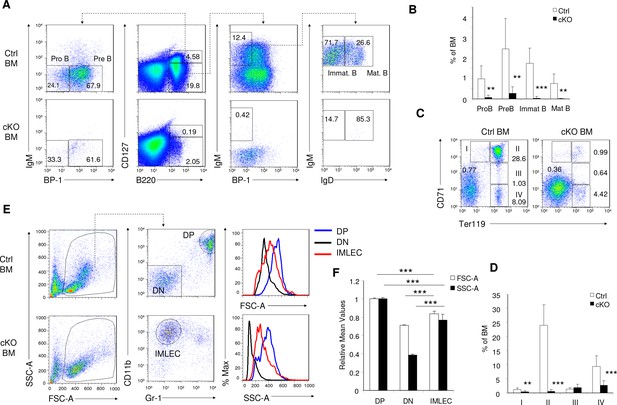
Raptor deletion led to impaired developments of B lymphoid, erythroid and myeloid compartments in BM.
(A) Representative FACS profiles for Pro B (CD127+B220+BP-1-IgM-IgD- cells), Pre B (CD127+B220+BP-1+IgM-IgD- cells), Immature B (CD127-B220+BP-1-IgM+IgD- cells), Mature B (CD127-B220+BP-1-IgM+IgD+ cells) in mice BM on day seven after pIpC treatment. (B) Frequencies of BM B cells subsets as in (A). n = 5 for Ctrl mice and n = 6 for cKO mice. (C) Representative FACS analysis of erythroid populations in BM by Ter119 and CD71. Roman numerals and numbers indicate the identity and percentages of the developmentally defined subpopulations: I, proerythroblasts; II, basophilic erythroblasts; III, polychromatophilic erythroblasts; IV, orthochromatophilic erythroblasts. (D) Frequencies of erythroblast subsets in mice BM. n = 5 for Ctrl mice and n = 6 for cKO mice. (E, F) Size and granularity of BM CD11b+ Gr-1- IMLECs from Rptor cKO mice. (E) Flow cytometric assays for size and granularity of Ctrl and cKO BM cells by FSC and SSC, respectively. Erythroid cells with smaller size and granularity were excluded in the initial gating. DP (double positive) represents CD11b+ Gr-1+ granulocytes, and DN (double negative) represents (CD11b- Gr-1-) cells. (F) Relative size (FSC-A) and granularities (SSC-A) of DN, DP and IMLEC. The mean values in Ctrl mice are artificially defined as 100%. n = 5 for Ctrl mice; n = 3 for cKO mice.
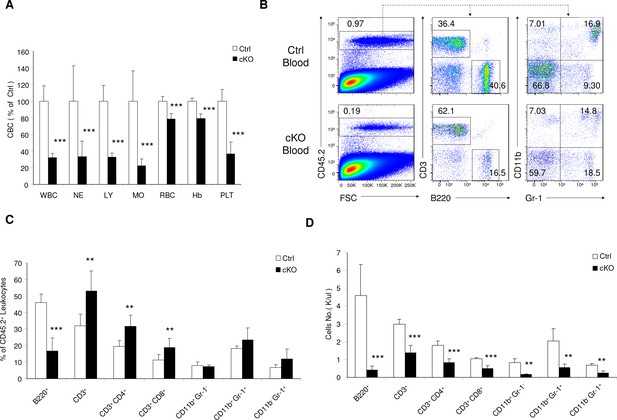
Rptor cKO mice are pancytopenic.
(A) Complete blood cell counts (CBC) data for Ctrl and cKO mice on day 30 after pIpC treatment. WBC, white blood cells; NE, neutrophils; LY, lymphocytes; MO, monocytes; RBC, red blood cells; Hb, hemoglobin; PLT, platelets. Means of the Ctrl mice in each experiment are artificially defined as 100%. (B) Representative FACS profiles showing the percentages of different leukocyte populations in peripheral blood from Ctrl and cKO mice on day 30 after pIpC treatment. (C, D) Reduction of leukocyte populations in the peripheral blood of cKO mice. Frequencies among CD45.2+ leukocytes (C) and absolute numbers (D) of various cell types in blood on day 30 after pIpC treatment are shown. n = 7 for Ctrl mice; n = 6 for cKO mice. Data represent one of three independent experiments with similar results.
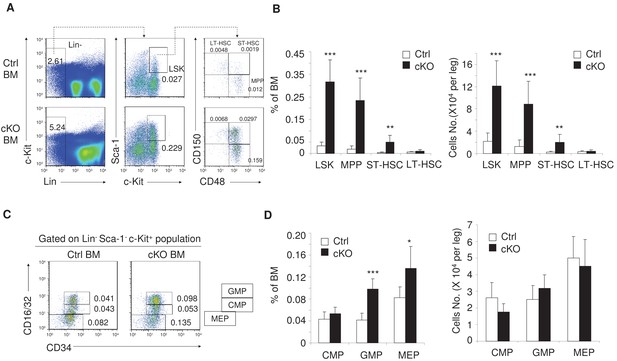
Rptor deletion increased hematopoietic stem and progenitor cells in BM.
(A) Representative FACS profiles for Lin- Sca-1+ c-Kit+ cells (LSK, identified as CD3- B220- Ter119- CD11b- Gr-1- Sca-1+ c-Kit+ cells), multipotent progenitor cells (MPP, identified as Lin- Sca-1+ c-Kit+ CD150- CD48+ cells), short-term HSC (ST-HSC, identified as Lin- Sca-1+ c-Kit+ CD150+ CD48+ cells) and long-term HSC (LT-HSC, identified as Lin- Sca-1+ c-Kit+ CD150+ CD48- cells) in the BM from Raptor Ctrl and cKO mice on day seven after pIpC treatment. Numbers indicate the percentages of gated populations in total BM cells. (B) Frequencies (left) and absolute numbers (right) of stem and progenitor cells as in (A) are shown. n = 6 for Ctrl mice; n = 7 for cKO mice. (C) Representative FACS profiles for common myeloid progenitor (CMP, identified as Lin- Sca-1- c-Kit+ CD34MediumCD16/32Medium cell), granulocyte/macrophage progenitor (GMP, identified as Lin- Sca-1- c-Kit+ CD34+ CD16/32 + cell) and megakaryocyte/erythroid progenitor (MEP, identified as Lin- Sca-1- c-Kit+ CD34- CD16/32cell) in BM. (D) Frequencies (left) and absolute numbers (right) of myeloid progenitors as in (C) are shown. n = 5 for Ctrl mice; n = 5 for cKO mice.
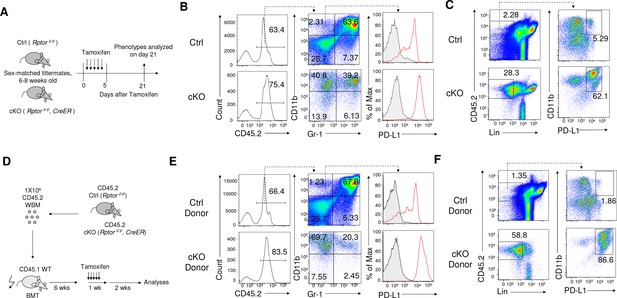
Tamoxifen induced conditional deletion of Rptor in hematopoiesis also led to massive accumulation of IMLECs.
(A) Schematic of experimental design. Sex-matched 6–8 weeks old Ctrl (Rptor F/F) and cKO (Rptor F/F, CreER) mice were treated with Tamoxifen for five consecutive days. The phenotypes were analyzed 3 weeks after the first treatment. (B) Representative flow cytometric analysis of myeloid cells from Ctrl and cKO mice BM by CD11b, Gr-1 and PD-L1. (C) Rptor deletion caused expansion of Lin- (B220- CD3- Ter119- NK1.1- Gr-1- F4/80- CD115-) CD11b+ PD-L1+ IMLECs in BM. (D) Schematic of experimental design. CD45.2+ Ctrl BM and cKO BM were transplanted into lethal dose irradiated CD45.1+ recipient mice. Tamoxifen treatment of recipient mice was started 6 weeks after fully reconstitution of donor-derived cells. The phenotypes were analyzed 3 weeks after the first treatment. (E, F) Representative flow cytometrical profiles of donor-derived CD45.2+ Myeloid cells (E) and IMLECs (F) in BM of recipient mice are shown as in (B) and (C), respectively.
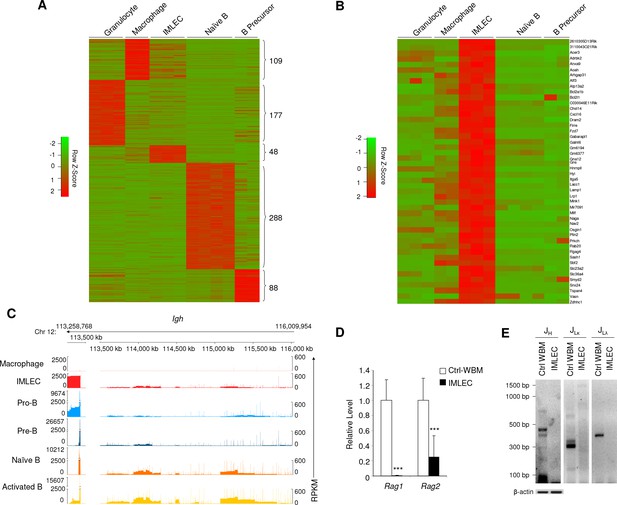
Unique gene signature of Rptor-/- CD11b+ Gr-1- BM IMLECs.
(A) Heat map representing the relative expression levels for indicated population-specific genes. Genes up-regulated for at least 4-fold with FDR-adjusted p-value<0.01 were considered population-specific. Numbers on right indicate amounts of population-specific genes. (B) Heat map representing expression levels of 48 CD11b+Gr-1- BM IMLEC-specific genes among indicated populations including macrophage, B precursor, naive B and granulocyte. (C) Genome browser display of transcript structure and gene expression quantity for immunoglobulin heavy chain (IgH) complex gene region from indicated populations. For each track, the normalized numbers of aligned reads count or Reads Per Kilobase of transcript per Million mapped RNA (RPKM) are shown in y-axis, while the gene positions for the sterile transcripts are shown in x-axis. Numbers on top indicate the start and end loci on bases along chromosome 12 for specified gene region. Similar profiles of cKO IMLEC were observed in two other samples analyzed. (D) mRNA levels of Rag1 and Rag2 in cKO CD11b+ Gr-1- BM IMLECs in comparison to Ctrl whole BM cells. n = 6 for Ctrl-WBM; n = 8 for IMLEC. Data are pooled from three independent experiments. (E) IMLECs did not display immunoglobulin gene rearrangement. FACS-sorted CD11b+ Gr-1- cells (IMLECs) from cKO BM were dissected for Ig gene rearrangement patterns of heavy chain (JH), light chains Igk (JLκ) and Igl (JLλ) by PCR. WBM cells from Raptor Ctrl mice were used as positive control. These data have been repeated twice.
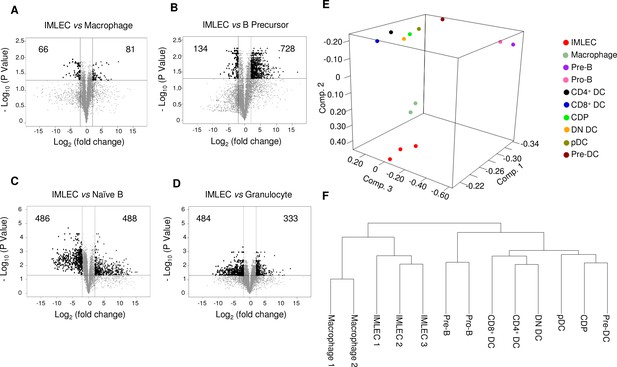
In silico pair wise comparisons between IMLECs and other closely related leukocytes based on our RNA-seq data and the publically available RNA-seq data.
(A–D) Based on the space proximity in the PCA data, macrophage (A), B precursor (B), naive B (C) and granulocyte (D) were selected for one-to-one comparison. Numbers in corners indicate amounts of genes significantly (FDR-adjusted p-value<0.05) up-regulated (right) or down-regulated (left) by at least 4-fold. (E, F) IMLECs in Raptor cKO BM are distinct from all DC subsets tested. (E) Principal component analysis (PCA) of gene expression in Raptor cKO CD11b+ Gr-1- BM cell (IMLEC), macrophage, B precursors and various ex vivo DC subsets. Numbers along axes indicate relative scaling of the principal variables. (F) Clustering of IMLEC, macrophage, B precursors with other DC subsets based on their gene expression profiles. Hierarchical clustering with complete linkage was carried out using the R software. The public RNA-seq datasets for DC subsets used are as followings: CDP (common DC precursor, GSM1531794); pDC (GSM1531795); preDC (GSM1531796); DN DC (CD4 and CD8 double negative DC, GSM1531797); CD4+ DC (GSM1531798) and CD8+ DC (GSM1531799).
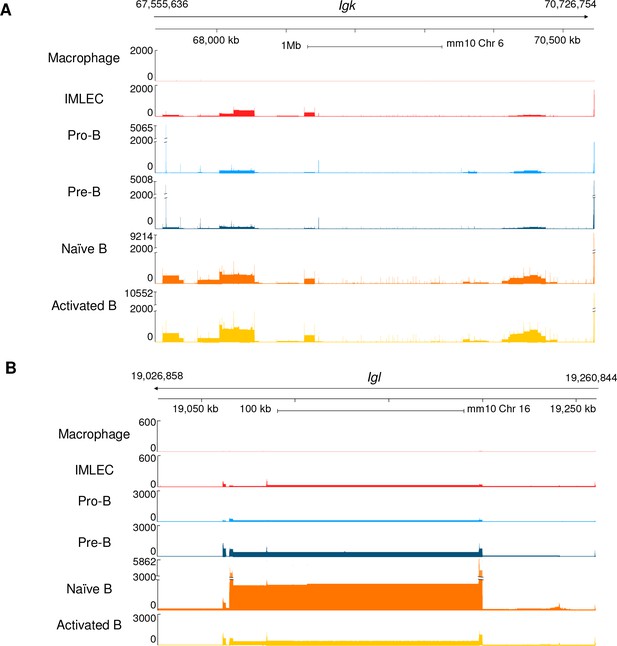
cKO IMLECs have a similar expression patterns as that of B-lymphoid subsets in the sterile transcripts from Igk and Igl loci.
(A, B) Genome browser display of transcript expression of Igk locus in chromosome 6 (A) and Igl locus in chromosome 16 (B). For each track, the y-axis is the normalized number of aligned reads counts of gene region, whereas the x-axis depicts physical distance in base-pairs (bp). Numbers on top indicate the start and end positions of specified gene regions.
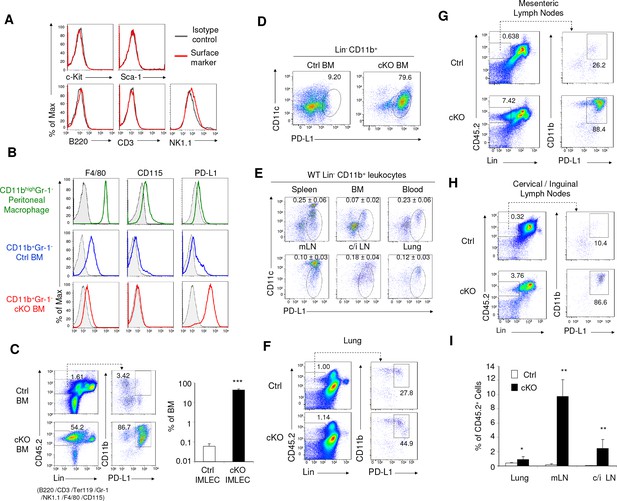
Identification of IMLEC by cell surface markers in both Raptor cKO and WT mice.
(A) IMLECs do not express surface markers for conventional lymphocytes (CD3, B220 and NK1.1) and stem and progenitor cells (c-Kit and Sca-1). CD11b+ Gr-1- IMLECs from Raptor cKO BM were tested for their expression of lymphocytes and progenitor cell markers. These data have been repeated three times. (B) IMLECs do not express markers for monocytes and macrophages but surprisingly express high levels of PD-L1. Filled gray areas indicate distributions of fluorescence from stainings by control antibodies. One representative result of at least five experiments is shown. (C) Rptor deletion causes expansion of Lin- (B220- CD3- Ter119- NK1.1- Gr-1- F4/80- CD115-) CD11b+ PD-L1+ IMLECs in BM. Representative flow staining profiles for BM IMLECs (left) and their abundance summary data (right) are presented. n = 5 for Ctrl BM; n = 5 for cKO BM. (D) Lin-CD11b+ PD-L1+ BM IMLECs are CD11c-/low. One representative result of two experiments is shown. (E) The Lin- CD11b+ PD-L1+ CD11c-/low IMLECs are found in various lymphoid and non-lymphoid organs from WT mice. mLN, mesenteric lymph nodes; c/i LN, cervical and inguinal lymph nodes. The numbers (Mean ±SD) are summarized abundances of IMLECs among mononuclear cells (MNCs) from 3 WT mice. (F–H) Rptor deletion caused a broad accumulation of IMLECs in lung (F), mesenteric lymph nodes (G) and peripheral (cervical and inguinal) lymph nodes (H). Results shown are representative of three mice in each group. (I) Summary data showing increased IMLECs in the lymphoid and non-lymphoid organs. n = 3 for both groups.
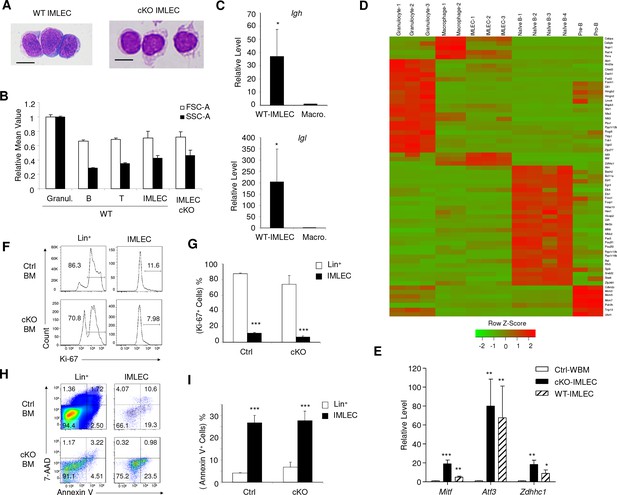
Comparison of IMLECs identified in both WT and Raptor-deficient BM.
(A) Giemsa staining of Lin- CD11b+ PD-L1+ IMLECs from WT BM and Rptor-cKO BM. One representative result of 3 independent experiments is shown. Scale bar, 10 μm. (B) Size and granularity of IMLECs from Raptor Ctrl and cKo BM were evaluated by FSC and SSC, respectively. n = 5 for each group. (C) IMLEC from WT BM expresses sterile transcripts from the Igh and Igl loci. Ig transcript levels in macrophage are artificially defined as 1. (D) Heat map representing the relative expression levels for indicated population specific transcription factor genes. Genes up-regulated for at least 4-fold with FDR-adjusted p-value<0.01 were considered population-specific. (E) IMLECs from both WT and Raptor-cKO BM exhibit high levels of mRNA transcripts for the specified transcription factors (Mitf, Atf3 and Zdhhc1). n = 5 for each group. (F, G) Ctrl and cKO IMLECs are non-cycling cells in contrast to Lin+ BM cells. (H, I) IMLECs are prone to apoptosis based on their binding to Annexin V. Results in (C), (G) and (I) represent one of 2 independent experiments with each involving three mice per group.
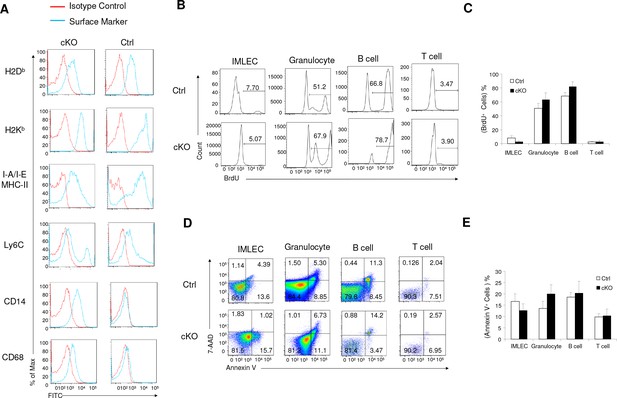
Comparisons of surface markers, viability and cellular proliferation among IMLECs and other defined lineages.
(A) IMLECs from Raptor Ctrl and cKO BM displayed comparable expression levels of MHC-I, MHC-II and other surface markers (Ly6C, CD14 and CD68). Data shown are representative of 3 independent experiments. (B, C) IMLECs from Raptor Ctrl and cKO BM exhibited much slower proliferation than other BM lineages. Representative flow profile (B) and summary data (C) are from BrdU incorporation assays. (D, E) IMLECs from Raptor Ctrl and cKO BM were more prone to apoptosis than other BM lineages. Representative flow profile (D) and summary data (E) are from Annexin V and 7-AAD staining. B cells, B220+ BM cells; T cells, CD3+ BM cells; Granulocytes, CD11b+Gr-1+ BM cells. n = 5 for Ctrl mice; n = 5 for cKO mice.
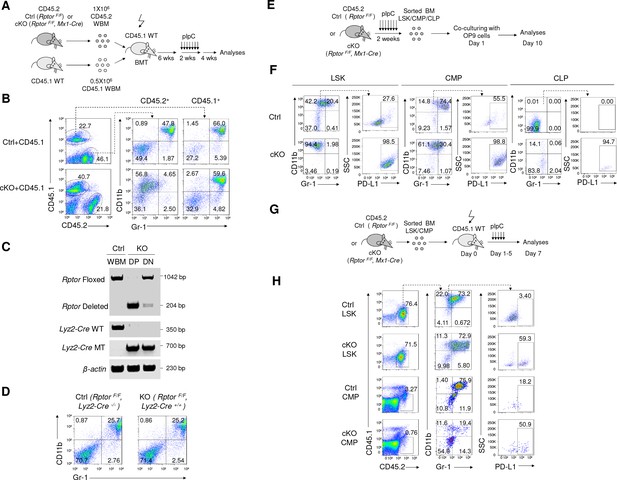
Raptor represses differentiation of IMLEC from CMP.
(A, B) IMLEC accumulation in Rptor-/- BM is cell autonomous. (A) Diagram depicting generation of mixed BM chimera, induction of Rptor deletion and analyses for IMLEC. (B) In mixed BM chimera mice, IMLEC was accumulated only in the Raptor-deficient hematopoietic cells. Mice were sacrificed and analyzed at 4 weeks post pIpC treatment. CD45.1+ and CD45.2+ donor-derived BM cells were gated to show the myeloid subsets based on CD11b and Gr-1. Similar data were obtained in three independent experiments, each involving at least three mice per group. (C, D) BM IMLECs are not converted from CD11b+ Gr-1- granulocytes. (C) Genotyping and deletion efficacy. CD11b+Gr-1+ cells (DP) and CD11b-Gr-1- cells (DN) were sorted from BM of KO (RptorF/F, Lyz2-Cre+/+) mice. Floxed and deleted Rptor alleles, as well as Lyz2-Cre wild type (WT) and mutated (MT) alleles were confirmed by PCR. (D) Efficient deletion of Rptor in the CD11b+Gr-1+ compartment failed to cause IMLEC accumulation. Data shown are representative BM flow profiles of Ctrl mice and those with Rptor deletion in the CD11b+Gr-1+ compartment, depicting distributions of CD11b and Gr-1 markers in adult mice. Similar data have been obtained in three experiments, involving a total of 5 mice per group. (E, F) Rptor-/- LSK, CLP and CMP cells differentiate into IMLECs in vitro. (E) Diagram of experimental design. 2 × 103 LSK cells or 5 × 104 CLP or 5 × 104 CMP cells were co-cultured with the OP9 cells for 10 days. (F) Rptor deletion promoted generation of IMLEC. The CD45.2+ leukocytes were gated and analyzed for their expression of CD11b, Gr-1 and PD-L1. Data shown are representative of three independent experiments. (G, H) Both LSK and CMP cells differentiate into IMLECs in vivo. (G) Diagram of BM cells transplantation. FACS-sorted, Ctrl or cKO LSK cells (5 × 104/mouse) or CMP cells (1.2 × 105/mouse) were injected i.v. to CD45.1 recipient mice which were immediately administrated with five daily pIpC treatments. BM cells were harvested on day seven for FACS analyses. (H) Rptor deletion promoted differentiation of progenitor cells into IMLEC. Donor-derived CD45.2+ BM cells were gated to analyze surface markers CD11b, Gr-1 and PD-L1. Data shown are representative flow profiles from one of 3 independent experiments.
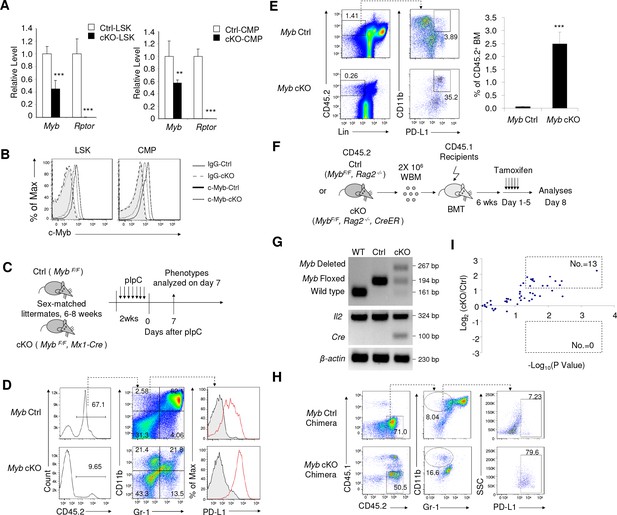
Inactivation of Myb is an underlying cause for accumulation of IMLEC.
(A) Reduction of Myb mRNA in the Rptor-deficient HSPCs. BM LSK and CMP populations were FACS-sorted from Ctrl and cKO mice at 1–3 weeks after pIpC treatment. Quantitation of Myb mRNA was performed by qPCR. n = 5 for Ctrl-LSK; n = 4 for cKO-LSK (left). n = 4 for Ctrl-CMP; n = 3 for cKO-CMP (right). (B) Detection of c-Myb protein by intracellular staining. Data represent one of three independent experiments with similar results. (C–E) Deletion of c-Myb in mice with homozygous floxed Myb resulted in enhanced generation of IMLECs. (C) Schematic of experimental design. Sex-matched 6–8 weeks old c-Myb Ctrl (Myb F/F) and cKO (Myb F/F, Mx1-Cre) mice were treated with pIpC for seven times. The phenotypes were analyzed on day seven after the complement of pIpC treatment. Inducible deletion of c-Myb showed obvious increase of PD-L1 expression on CD11b+Gr-1- BM cells (D) and production of IMLECs (E). n = 3 for c-Myb Ctrl mice; n = 3 for c-Myb cKO mice. (F–H) Deletion of Myb enhances generation of IMLEC. (F) Diagram of experimental design. Whole BM cells (2 × 106/mice) of given genotypes were used for transplantation. Once the chimera mice were established, deletion of Myb was induced by five daily injection of tamoxifen. (G) Detection of Myb deletion in the whole BM cells after tamoxifen treatments. Data are representative of two independent experiments. (H) Generation of IMLEC is promoted by inactivation of Myb. BM cells were harvested at 7 days after first tamoxifen treatment and analyzed for IMLECs based on surface markers CD11b, Gr-1 and PD-L1 within the donor-derived CD45.2+ BM cells. Data shown represent one of three experiments using either first or second generation of BM chimeras. (I) Rptor deletion broadly increases miRNAs targeting Myb. Lin- c-Kit+ HSPCs were isolated from Raptor Ctrl and cKO mice at 10 days after pIpC treatment. miRNA levels were measured by miRNA microarray. The y-axis shows the log2 ratio of signal, while the x-axis shows the -log10P value. The doted boxes show the numbers of significantly (p<0.05) up-regulated (fold change >2) or down-regulated (fold change <0.5) miRNAs among 50 miRNAs with mirSVR score <−1.0. Each dot represents the mean value of a unique miRNA from three independent samples.
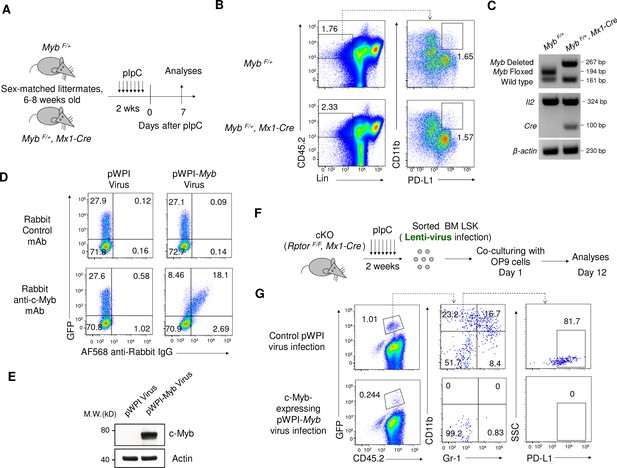
Myb expression and accumulation of IMLECs.
(A–C) Induced deletion of c-Myb in mice with heterozygous floxed c-Myb (Myb F/+, Mx1-Cre) did not show significant accumulation of IMLECS in mouse BM. (A) Schematic of experimental design. Sex-matched 6–8 weeks old Ctrl (Myb F/+) and heterozygous floxed c-Myb (Myb F/+, Mx1-Cre) mice were treated with pIpC for seven times. The phenotypes were analyzed on day seven after the complement of pIpC treatment. (B) Representative flow profile of BM IMLECs. (C) Detection of Myb deletion in the whole BM cells after pIpC treatment. Data in (B) and (C) are representative results of independent experiments with 3 groups of paired mice. (D–G) The provision of heterologous c-Myb significantly diminished the generation of IMLECs from Raptor-deficient LSK cells. (D, E) Myb-expression lenti-virus (pWPI-Myb) was validated by intracellular staining (D) and western blot (E) of CHO cells 36 hr after virus infection. Lenti-virus with GFP co-expression (pWPI) was used as control. AF568, Alexa Fluor 568 dye. (F) Diagram of experimental design. 1 × 104 LSK cells sorted from pIpC treated Raptor cKO mice were infected with indicated lenti-virus and subsequently co-cultured with the OP9 cells for 12 days. (G) GFP+ cells with heterologous c-Myb expression did not give rise to CD11b+ Gr-1- PD-L1+ IMLECs. Data represent the results of two independent experiments.
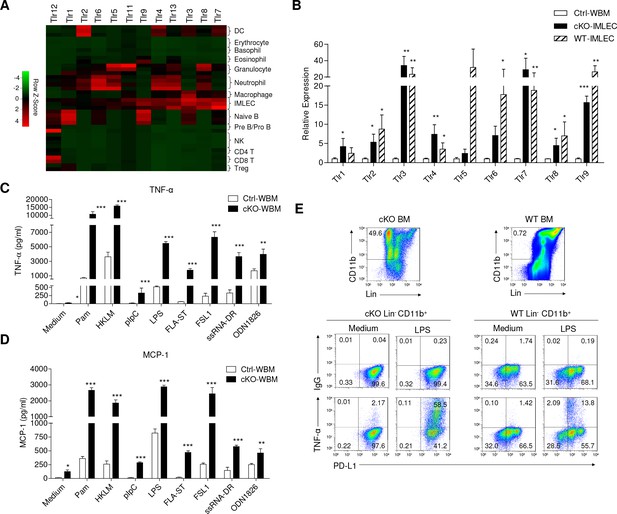
IMLECs broadly express PPRs and produce large amounts of inflammatory cytokines upon stimulation by various TLR ligands.
(A) Heat map showing the relative transcript levels of TLRs among the indicated populations. (B) Both WT and Raptor-cKO IMLECs have greatly elevated expression of multiple TLRs genes in comparison to Ctrl BM. q-PCR was performed to determine transcript levels of Tlr1-9 genes. After normalizing for cDNA input based on Hprt mRNA in each sample, the Tlr1-9 levels in the FACS-sorted cKO CD11b+ Gr-1- BM IMLEC were compared with Ctrl BM (artificially defined as 1.0). n = 3 for Ctrl WBM; n = 5 for cKO- IMLEC; n = 5 for WT-IMLEC. Similar results were obtained using mice sacrificed at 2 weeks ~2 months after pIpC treatment. (C, D) In responses to various TLR ligands, Rptor cKO BM cells produced greatly elevated amounts of TNF-α (C) and MCP-1 (D) than the Ctrl BM. Data shown are from one experiment involving three repeats per group and have been reproduced in five independent experiments. (E) Lin- (B220-CD3-Ter119-Gr-1-NK1.1-F4/80-CD115-) CD11b+ PD-L1+ IMLECs from both cKO and WT BM were robust TNF-α producers after stimulation with LPS. BM cells (1 × 107 cells / well) were stimulated with LPS (1 μg/ml) for 16 hr with the presence of Golgi blocker in the last 4 hr. Data shown are representative profiles from one experiment and have been reproduced in three independent experiments.
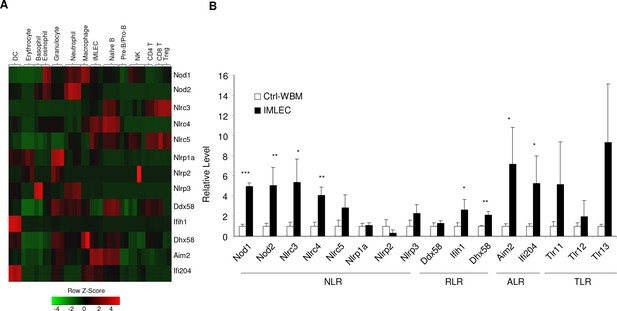
IMLECs broadly over-express multiple families of pattern recognition receptors (PRRs) when compared with other blood cells.
(A) Heat map showing the relative expression levels of genes encoding key PRRs. TLRs, Toll-like receptors; NLR, Nod-like receptors; RLRs, RIG-I-like receptors; ALRs, Aim2-like receptors. (B) mRNA levels of the indicated PPRs genes were measured by q-PCR in WBM cells from Ctrl mice and CD11b+ Gr-1- BM IMLECs from cKO mice. n = 3 for Ctrl WBM; n = 5 for cKO BM IMLECs.
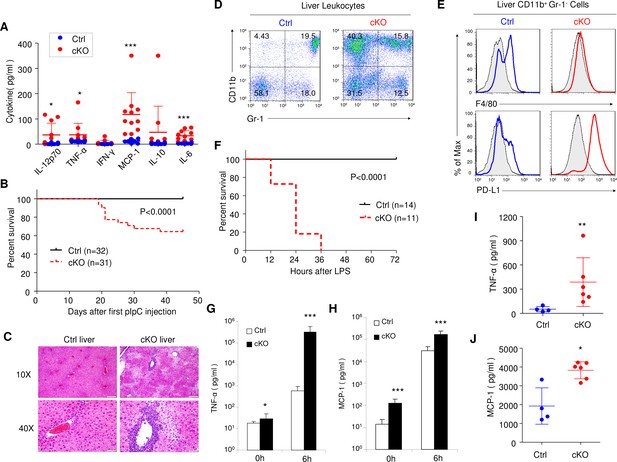
Rptor cKO mice are hyper sensitive to challenges by TLR ligands.
(A) High blood levels of inflammatory cytokines in cKO mice that have received 7 pIpC treatments. Serum samples collected at 7 days after the last pIpC treatment were tested for levels of IL-12p70, TNF-α, IFN-γ, MCP-1, IL-10 and IL-6. Dot plots depict cytokine levels in individual cKO (red dots) or Ctrl (blue dots) mice, and line drawings depict means and SD. n = 12 for Ctrl mice and n = 11 for cKO mice. (B) High proportions of Rptor cKO mice that received 7 pIpC injections died within 7 weeks after first pIpC treatment. Data shown are Kaplan-Meier survival curve, and the statistical significance is determined by log-rank test. n = 32 for Ctrl mice and n = 31 for cKO mice. (C) Representative H and E staining of liver sections from Rptor Ctrl and cKO mice at 2 weeks post pIpC treatment. Scale bars represent 200 μm (10X) and 50 μm (40X), respectively. (D, E) Massive accumulation of IMLEC in cKO liver. (D) Accumulation of CD11b+ Gr-1- cells in the cKO liver. (E) CD11b+ Gr-1- cells in cKO livers demonstrated IMLEC surface markers, including PD-L1high and F4/80low/-. Gray lines indicate the staining for isotype control antibodies. Results are representative of at least five independent analyses. (F–H) cKO mice are vulnerable to low doses of LPS challenges. At 2–3 months after pIpC treatments, cKO and Ctrl mice were challenged with 5 mg/kg body weight of LPS and observed survival. (F) Kaplan- Meier survival analysis. Data are pooled from three independent experiments. n = 14 for Ctrl mice; n = 11 for cKO mice. Inflammatory cytokines TNF-α (G) and MCP-1 (H) levels from mice serum before (0 h) and at 6 hr after (6 h) LPS injection are shown. n = 10 for Ctrl mice; n = 9 for cKO mice. Note greater than 500-fold increase in plasma TNF-α levels. (I, J) cKO mice mounted enhanced inflammatory response to acetaminophen-triggered liver necrosis. Serum TNF-α (I) and MCP-1(J) levels at 6 hr upon acetaminophen (3.2 mg/mouse) treatment are shown. n = 4 for Ctrl mice; n = 6 for cKO mice. Mann-Whitney test was used for statistics analysis, and lines indicate Mean ±SD. Similar trends were observed in another independent experiment.
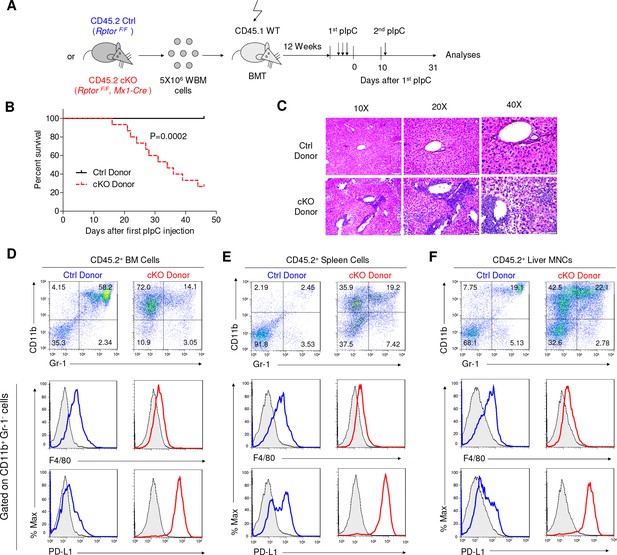
Rptor deletion in hematopoietic cells greatly increases vulnerability of mice to pIpC.
(A) Diagram of experimental design. WT CD45.1+ recipient mice were irradiated and transplanted with 5 × 106 BM cells from RptorF/F and RptorF/F, Mx1-Cre BM. After the fully reconstitution, recipients were treated with pIpC three times every other day to induce gene deletion in chimera mice with cKO BM. 10 days after the last injection of pIpC, recipients were challenged with another pIpC injection and the survival of mice were followed for four more weeks. Arrows denote the injections of pIpC at indicated time points. (B) Kaplan-Meier survival analysis after pIpC treatments. n = 12 for Ctrl chimera mice; n = 15 for cKO chimera mice. (C) Histological analysis of liver sections after H&E staining. Note extensive inflammation and liver damage in cKO chimera mice. Scale bars represent 200 μm (10X), 100 μm (20X) and 50 μm (40X). (D–F) Identification of IMLEC in BM (D), spleen (E) and livers (F) by flow cytometry. Mice were euthanized 45 days after the first pIpC injection. The distinct CD11b+ Gr-1- PD-L1+ F4/80low/- IMLECs greatly enriched in BM, spleen and liver of recipients of Rptor cKO BM. FACS profiles shown represent one of five independent experiments.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| strain, strain background (Mus musculus,C57BL/6) | RptorF/F | PMID: 19046572 | ||
| strain, strain background (Mus musculus,C57BL/6) | MybF/F; MybF/F,Rag2-/-,CreER | PMID: 15195090; PMID: 16169500 | ||
| strain, strain background (Mus musculus,C57BL/6) | Mx1-Cre | The Jackson Laboratory | Stock No: 003556; RRID:IMSR_JAX:003556 | |
| strain, strain background (Mus musculus,C57BL/6) | CreER | The Jackson Laboratory | Stock No: 007001; RRID:IMSR_JAX:007001 | |
| strain, strain background (Mus musculus,C57BL/6) | Lyz2-Cre | The Jackson Laboratory | Stock No: 004781; RRID:IMSR_JAX:004781 |
Additional files
-
Supplementary file 1
Transcript levels of CD markers analyzed from RNA-seq datasets.
The FPKM values of 3 mice samples for cKO-IMLEC group (Rptor cKO CD11b+Gr-1- BM IMLECs) and Ctrl-WBM group (Ctrl whole BM cells) are shown. Fold change indicates the ratio of average FPKM values.
- https://doi.org/10.7554/eLife.32497.022
-
Supplementary file 2
Expression levels of miRNAs targeting Myb in Raptor Ctrl and cKO BM HSPCs.
The normalized signal values of 3 mice samples for cKO-HSPCs group and Ctrl-HSPCs group are shown. The microarray data have been deposited to NCBI GEO database with accession number GSE64042 as previously reported (Ye et al., 2015).
- https://doi.org/10.7554/eLife.32497.023
-
Supplementary file 3
The list of antibodies used for flow cytometry.
- https://doi.org/10.7554/eLife.32497.024
-
Supplementary file 4
Sequences of DNA primers used in PCR assays.
- https://doi.org/10.7554/eLife.32497.025
-
Transparent reporting form
- https://doi.org/10.7554/eLife.32497.026





