Crk proteins transduce FGF signaling to promote lens fiber cell elongation
Figures
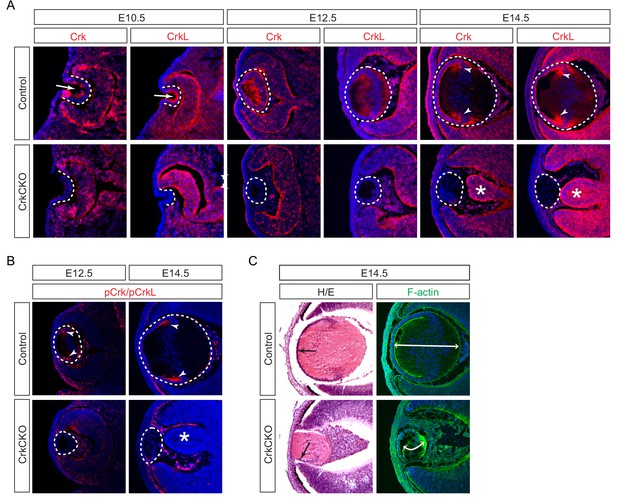
Crk and Crkl are essential for lens development.
(A) Crk and Crkl immunostaining were localized to the invaginating lens vesicle at E10.5 (arrows) and to the elongating lens fiber cells near the transitional zone of the lens at E14.5 (arrowheads). These staining patterns were specifically lost in the CrkCKO lens. The dotted lines enclose the region of the lens and the disorganization of the retina was marked with asterisks (B) The phosphorylation of both Crk and Crkl was noticeably absent in the CrkCKO lens (arrowheads). (C) The CrkCKO lens size was significantly reduced with the anterior lens epithelium rotated sideways (arrows) and the disorganized lens fiber cells markedly shortened (double headed arrows).
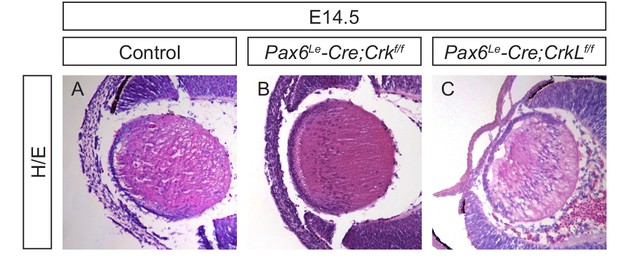
Crk and Crkl single mutants did not display lens phenotype.
(A–C) H/E staining showed that the individual deletion of Crk and Crkl did not affect aspects of lens development.

Molecular defects in the Crk and Crkl double mutant lens.
(A) There were no significant changes in the staining intensity of the lens determinant markers Prox1 and Pax6. (B) Further, none of the three forms of Crystallins (α, β, γ) displayed any changes in staining intensity in the CrkCKO lens. (C) The polarity of the CrkCKO lens fiber cells was maintained as indicated by both the apical expression of Zo-1 and the basal expression of β1 integrin. (D) The number of Ki67-expressing proliferative cells was significantly decreased and the number of TUNEL-positive apoptotic cells was increased (arrowheads). (E) Quantification of proliferation and apoptosis in wild type and CrkCKO lens. Student’s t test, *p<0.01, **p<0.001, n = 4.
-
Figure 2—source data 1
Source data for Figure 2E.
- https://doi.org/10.7554/eLife.32586.006
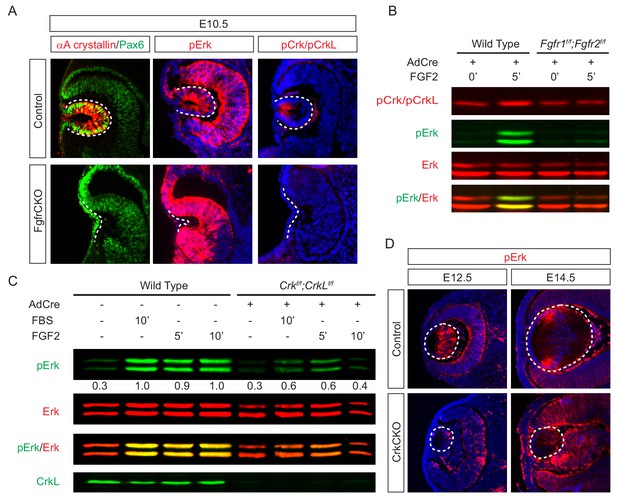
Crk proteins mediate FGF signaling in Erk phosphorylation.
(A) Genetic ablation of Fgfr1 and Fgfr2 disrupted the proper formation of the lens vesicle with the phosphorylation of Erk and Crk/Crkl proteins being noticeably absent. (B) Mouse Embryonic fibroblast (MEF) cells treated with FGF2 displayed an increase in pCrk/Crkl and pErk levels, which were abrogated by the removal of Fgfr1 and Fgfr2 using a Cre-expressing adenovirus. (C) Ablation of Crk proteins in MEF cells reduced FGF2-induced Erk phosphorylation. The pErk/Erk ratios were noted below the pERK blot. (D) The CrkCKO lens displayed a significant decrease in pERK staining compared to the wild type lens.
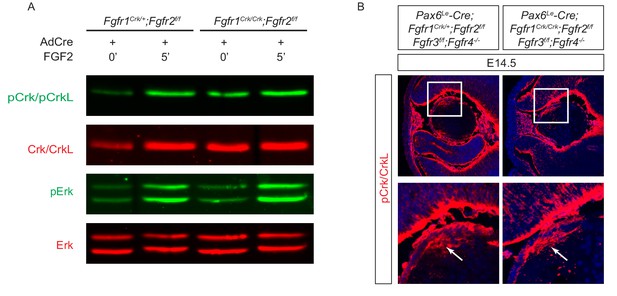
The Y463F mutation in Fgfr1 (Fgfr1Crk) did not affect the phosphorylation of Crk and Erk proteins that is essential for lens development.
(A) FGF2 was still able to induce an upregulation of pCrk/Crkl and pERK in Fgfr1Crk/Crk;Fgfr2flox/flox MEF cells after treatment with a Cre-expressing adenovirus. (B) A homozygous Fgfr1Crk/Crk mutation did not affect pCrk/Crkl staining (arrows) in the lens even after the genetic removal of other Fgf receptors (Fgfr2-4).
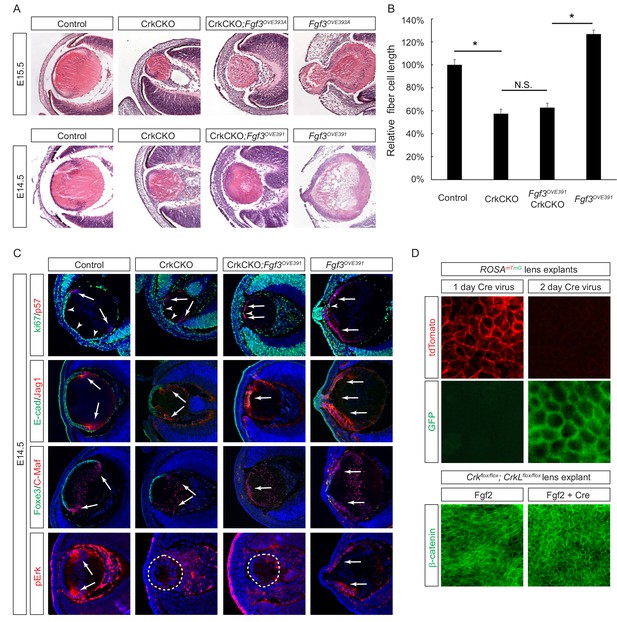
Crk/Crkl deletion prevented FGF-induced lens cell elongation without affecting differentiation.
(A) Overexpression of Fgf3 in Fgf3OVE391 and Fgf3OVE393A strains resulted in increased lens fiber cell elongation and overall lens sizes. However, both of these phenotypes were suppressed after being crossed with the CrkCKO mutant, indicating a genetic epistasis interaction between the FGF and Crk signaling pathways. (B) Quantification of lens fiber cell length. One-way ANOVA test followed by Tukey's multiple comparisons test, *p<0.01, n = 3. (C) Deletion of the Crk genes did not prevent the premature differentiation phenotype observed in the Fgf3 overexpressing lens, as indicated by a reduction of lens progenitor cell markers (Ki67, E-cad and Foxe3) and an increase of differentiation cell markers (p57, Jag1, and C-Maf) within the presumptive lens epithelial layer (arrows). Nonetheless, pERK staining was significantly reduced in the CrkCKO;Fgf3OVE391 lens. (D) The Cre-expressing adenovirus induces efficient genetic recombination in lens explant cultures as indicated by the ROSAmTmG reporter. In Crkflox/flox;Crklfloxflox explants, the Cre-mediated deletion of Crk proteins prevented Fgf2 from inducing cell shape changes.
-
Figure 4—source data 1
- https://doi.org/10.7554/eLife.32586.010
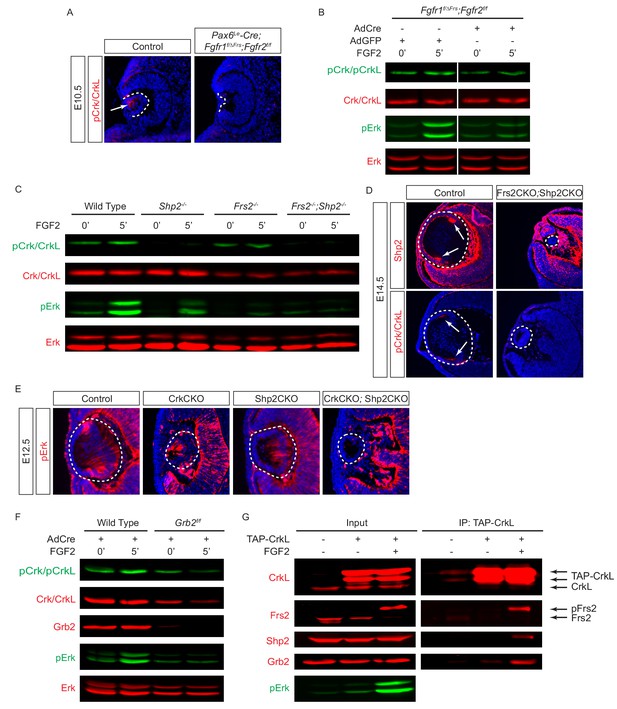
Crk proteins are recruited to the Frs2-Shp2-Grb2 complex in FGF signaling.
(A–B) Mutating the Frs2-bindng site in Fgfr1ΔFrs resulted in the loss of pCrk/Crkl in the Pax6Le-Cre;Fgfr1flox/ΔFrs;Fgfr2floxflox mutant lens. (B) FGF2 was unable to induce the phosphorylation of Crk proteins in Fgfr1flox/ΔFrs;Fgfr2floxflox MEF cells after treatment with the Cre expressing adenovirus. (C) FGF2-induced pCrk/Crkl and pErk were significantly downregulated in both Frs2 and Shp2 null MEF cells. (D) Shp2 was successfully depleted in the Frs2CKO;Shp2CKO lens, which resulted in the loss of pCrk/Crkl staining. (E) pERK was downregulated in both CrkCKO and Shp2CKO lenses and was further reduced in CrkCKO;Shp2CKO mutants. (F) FGF2-induced pCrk/Crkl and pErk were down regulated in Grb2 deficient MEF cells. (G) TAP-taged Crkl pulled down Frs2, Shp2 and Grb2 after FGF2 stimulation. Note that only the slower moving phosphorylated form of Frs2 successfully interacted with Crkl.
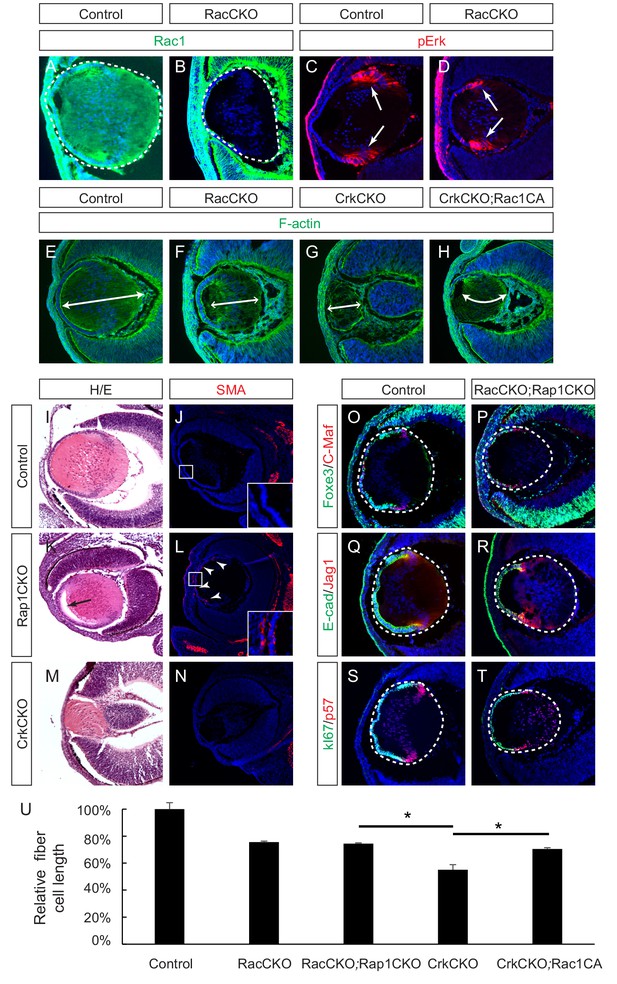
Rac proteins are downstream effectors of Crk signaling.
(A–D) Staining of Rac-depleted lenses with a pErk antibody showed no significant difference in staining intensity. (E–H) Phalloidin (F-actin) staining revealed that the length of lens fiber cells (indicated by arrows) was significantly reduced in CrkCKO and RacCKO lenses, which was partially reversed after the activation of Rac signaling in CrkCKO;Rac1CA lenses. (I–N) The Rap1 depleted mutants displayed a detachment of the lens fiber cells from the anterior side of the lens epithelial cells (arrow) and Smooth Muscle Actin (SMA) staining within the lens epithelial layer itself (arrowheads). These phenotypes were absent both in the control and the Crk/Crkl depleted lenses. (O–T) Immunstaining of the Rac and Rap1 depleted lenses with progentior (Foxe3, E-cad, Ki67) and differentiation (C-Maf, Jag1,p57) markers did not reveal any defects in differentiation or any further shortening of the fiber cells. (I) Quantification of fiber cell lengths. One-way ANOVA test followed by Tukey's multiple comparisons test, *p<0.01, n = 3.
-
Figure 6—source data 1
Source data for Figure 6U.
- https://doi.org/10.7554/eLife.32586.013
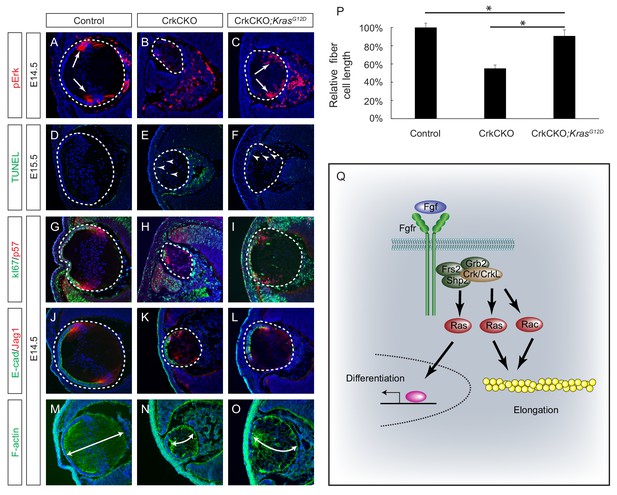
Constitutive Kras signaling can compensate for the loss of Crk and Crkl in lens development.
(A–C) Despite the loss of Crk and Crkl, Erk phosphorylation was partially recovered in CrkCKO;KrasG12D lenses. (D–F) A signficant amount of TUNEL positive cells remained in both CrkCKO and CrkCKO;KrasG12D lenses. (G–L) Cell proliferation indicated by Ki57 increased in the CrkCKO;KrasG12D lens as compared to the CrkCKO lens, but there was no significant difference in staining intensity for the differentiation markers E-cad and Jag1. (M–P) Lens fiber cell length increased significantly in CrkCKO;KrasG12D lenses as compared to CrkCKO ones. Fiber cell length was measured based on F-actin staining and statistical analysis was performed using the one-way ANOVA test followed by Tukey's multiple comparisons test (*p<0.01, n = 3). (Q) Model of Crk function in FGF signaling. The binding of FGF to its receptor induces the assembly of the Frs2-Shp2-Grb2 complex, which subsequently activates Ras signaling to promote lens differentiation. When FGF signaling is further elevated at the transitional zone of the lens, Crk proteins were additionally recruited by the Frs2-Shp2-Grb2 complex to further promote Ras and Rac signaling, resulting in actin cytoskeletal rearrangement and cell shape changes.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (M. musculus) | Crkflox, CrkLflox | PMID: 19074029 | RRID:MGI:3830069 | Dr. Tom Curran (The Children's Research Institute, Children's Mercy Kansas City) |
| Genetic reagent (M. musculus) | Fgfr3flox | PMID: 20582225 | RRID:MGI:4459834 | Dr. Xin Sun (University of California San Diego) |
| Genetic reagent (M. musculus) | Frs2αflox | PMID: 17868091 | RRID:MGI:3768915 | Dr. Feng Wang (Texas A and M) |
| Genetic reagent (M. musculus) | Rap1aflox, Rap1bflox | PMID: 18305243 | RRID:MGI:3777607 | Alexei Morozov (National Institutes of Health) |
| Genetic reagent (M. musculus) | Shp2flox | PMID: 15520383 | RRID:MGI:3522138 | Gen-Sheng Feng (UCSD) |
| Genetic reagent (M. musculus) | Fgf3OVE391 | PMID: 7539358 | Dr. Michael Robinson (Miami University) | |
| Genetic reagent (M. musculus) | Fgf3OVE393A | PMID: 9640329 | Dr. Michael Robinson (Miami University) | |
| Genetic reagent (M. musculus) | Fgfr1ΔFrs | PMID: 16421190 | RRID:MGI:3620075 | Dr. Raj Ladher (RIKEN Kobe Institute-Center for Developmental Biology) |
| Genetic reagent (M. musculus) | Fgfr1Crk | PMID: 26341559 | RRID:MGI:5882534 | Dr. Philipo Soriano (Washington University Medical School) |
| Genetic reagent (M. musculus) | Fgfr2flox | PMID: 12756187 | RRID:MGI:3044690 | Dr. David Ornitz (Washington University Medical School) |
| Genetic reagent (M. musculus) | Fgfr4-/- | PMID: 9716527 | RRID:MGI:3653043 | Dr. Chu-Xia Deng (National Institute of Health) |
| Genetic reagent (M. musculus) | Grb2flox | PMID: 21427701 | RRID:MGI:4949890 | Dr. Lars Nitschke (University of Erlangen-Nürnberg) |
| Genetic reagent (M. musculus) | Pax6Le-Cre (Le-Cre) | PMID: 11069887 | RRID:MGI:3045795 | Dr. Ruth Ashery-Padan (Tel Aviv University) |
| Genetic reagent (M. musculus) | Rac1flox | PMID: 12759446 | RRID:MGI:2663672 | Dr. Feng-Chun Yang (Indiana University School of Medicine) |
| Genetic reagent (M. musculus) | Rac2-/- | PMID: 10072071 | RRID:MGI:3840460 | Dr. Feng-Chun Yang (Indiana University School of Medicine) |
| Genetic reagent (M. musculus) | KrasLSL-G12D | PMID: 15093544 | RRID:MGI:3044567 | Mouse Models of Human Cancers Consortium Repository at National Cancer Institute |
| Genetic reagent (M. musculus) | Fgfr1flox | Jackson Laboratory | Stock #: 007671 RRID:MGI:3713779 | PMID:16421190 |
| Genetic reagent (M. musculus) | R26-Rac1LSL-G12V | Jackson Laboratory | Stock #: 012361 RRID:MGI:4430563 | PMID:19879843 |
| Genetic reagent (M. musculus) | ROSAmTmG | Jackson Laboratory | Stock #: 007676 RRID:MGI:3722405 | PMID:17868096 |
| Cell line(M. musculus) | NIH-3T3 | American Type Culture Collection | Cat# CRL-1658, RRID:CVCL_0594 | |
| Transfected construct (synthesized) | TAP-CrkL | PMID: 26527617 | David J. Glass | |
| Antibody | Rabbit anti-C-maf | Santa Cruz Biotechnology | Cat. #: sc-7866 RRID: AB_638562 | IHC (1:200) |
| Antibody | Rabbit anti-CrkL | Santa Cruz Biotechnology | Cat. #: sc-319 RRID: AB_631320 | IHC (1:100), WB (1:1000) |
| Antibody | Mouse anti-CrkL | Santa Cruz Biotechnology | Cat. #: sc-365471 | WB (1:1000) |
| Antibody | Mouse anti-E-cadherin | Sigma | Cat. #: U3254 | IHC (1:200) |
| Antibody | Rabbit anti-Frs2 | Santa Cruz Biotechnology | Cat. #: sc-8318 RRID: AB_2106228 | WB (1:1000) |
| Antibody | Rabbit anti-Grb2 | Santa Cruz Biotechnology | Cat. #: sc-255 RRID: AB_631602 | WB (1:1000) |
| Antibody | Rabbit anti-Jag1 | Santa Cruz Biotechnology | Cat. #: sc-6011 RRID: AB_649689 | IHC (1:100) |
| Antibody | Mouse anti-Ki-67 | BD Pharmingen | Cat. #: 550609 RRID: AB_393778 | IHC (1:200) |
| Antibody | Rabbit anti-p57 | Abcam | Cat. #: ab75974 | IHC (1:2000) |
| Antibody | Rabbit anti-Pax6 | Covance | Cat. #: PRB-278P RRID: AB_291612 | IHC (1:500) |
| Antibody | Rabbit anti-pCrk (Tyr221) | Cell Signaling | Cat. #: 3491 | IHC (1:200), WB (1:1000) |
| Antibody | Rabbit anti-pCrkL (Tyr207) | Cell Signaling | Cat. #: 3181 | WB (1:1000) |
| Antibody | Rabbit anti-pERK1/2 | Cell Signaling | Cat. #: 4370 | IHC (1:200), WB (1:1000) |
| Antibody | Mouse anti-pERK1/2 | Santa Cruz Biotechnology | Cat. #: sc-7383 | WB (1:1000) |
| Antibody | Rabbit anti-Prox1 | Covance | Cat. No.: PRB-238C | IHC (1:1000) |
| Antibody | Mouse anti-Rac1 | BD Transduction Laboratory | Cat. #: 610652 RRID: AB_397979 | IHC (1:200) |
| Antibody | Rabbit anti-Shp2 | Santa Cruz Biotechnology | Cat. #: sc-280 RRID: AB_632401 | IHC (1:100), WB (1:1000) |
| Antibody | Rabbit anti-α-crystallin | Sam Zigler (National Eye Institute) | IHC (1:5000) | |
| Antibody | Mouse anti-α-SMA | Sigma | Cat. #: C6198 | IHC (1:1000) |
| Antibody | Mouse anti-β-catenin | Sigma | Cat. #: 6F9 | IHC (1:200) |
| Antibody | Rabbit anti-β-crystallin | Sam Zigler (National Eye Institute) | IHC (1:5000) | |
| Antibody | Rabbit anti-γ-crystallin | Sam Zigler (National Eye Institute) | IHC (1:5000) | |
| Recombinant DNA reagent | Ad5CMVCre-eGFP | Gene Transfer Vector Core, University of Iowa, IA | VVC-U of Iowa-1174 | |
| Recombinant DNA reagent | Ad5CMVeGFP | Gene Transfer Vector Core, University of Iowa, IA | VVC-U of Iowa-4 | |
| Peptide, recombinant protein | recombinant human FGF2 | ScienCell | 104–02 | |
| Peptide, recombinant protein | recombinant murine FGF2 | ScienCell | 124–02 | |
| Commercial assay or kit | In situ cell death detection kit, Fluorescein | Sigma | 11684795910 ROCHE | |
| Commercial assay or kit | streptavidin resin | Agilent | 240207 | |
| Chemical compound, drug | Alexa Fluor 488 Phalloidin | ThermoFisher Scientific | A12379 | 1:50 |
| Chemical compound, drug | heparin sodium | Sigma | H3393 |
Additional files
-
Supplementary file 1
List of genetic crosses and experimental progenies.
- https://doi.org/10.7554/eLife.32586.015
-
Transparent reporting form
- https://doi.org/10.7554/eLife.32586.016





