ZMYND10 functions in a chaperone relay during axonemal dynein assembly
Figures
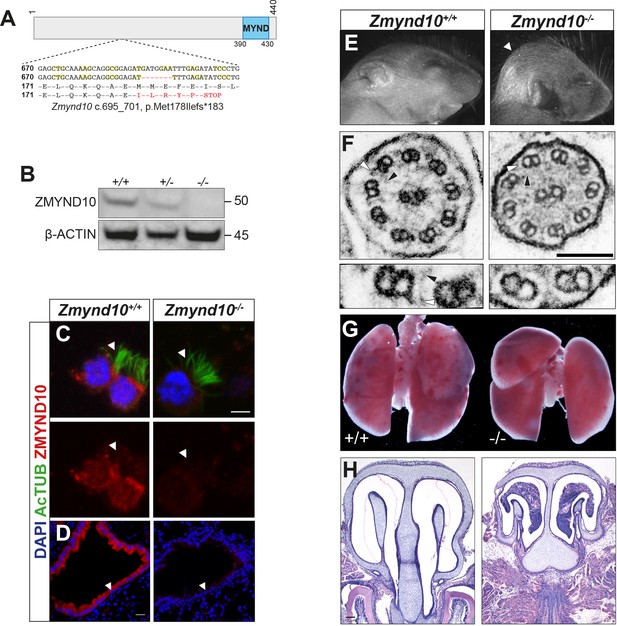
Loss of Zmynd10 in mice results in a PCD phenotype.
(A) Schematic illustrating the null allele generated by a −7 bp CRISPR deletion in Zmynd10 exon 6. (B) Immunoblots from testes extracts from postnatal day 26 (P26) control and mutant male mice show loss of ZMYND10. (C, D) Immunostaining for ZMYND10 reveals a complete loss of signal in multiciliated ependymal cells (C) and lung cryo-sections (D). Multicilia are marked with acetylated α- tubulin (C). (E) Neonatal Zmynd10 mutants display hydrocephaly; the white arrowhead points to doming of the head. See also Figure 1—figure supplement 2E. (F) TEM of tracheal ciliary axoneme cross-sections shows a lack of axonemal outer (ODA: white arrowhead) and inner (IDA: black arrowhead) dynein arms in mutants. (G) Representative image of a gross dissection of lungs shows situs inversus totalis in mutants. See also Figure 1—figure supplement 1F. (H) H&E staining of coronal sections of nasal turbinates reveals mucopurulent plugs in mutants. Scale bars in (C) = 5 μm, in (D) = 100 μm, in (F) = 100 nm.
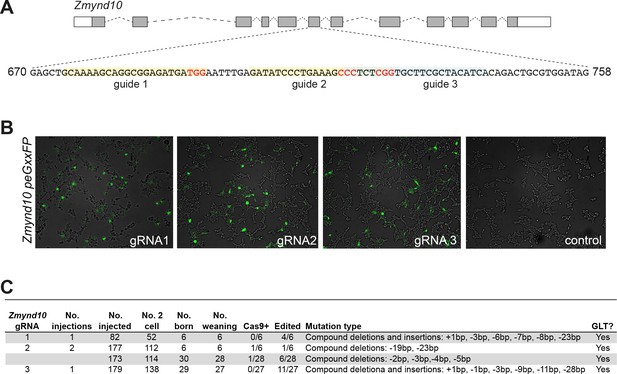
Generation of Zmynd10 mutant mice by CRISPR genome editing.
(A) Schematic of Zmynd10 mouse locus (ENSEMBL: ENSMUSG00000010044) and guide design targeting critical exon six which were cloned into px330 (Addgene:#42230) (Cong et al., 2013). (B) Validation of activities of gRNAs to generate double-stranded breaks were initially tested in HEK293 cells by using a reporter to assay reconstitution of EGFP upon cleavage (Addgene: #50716) (Mashiko et al., 2014). (C) Table summarizing pronuclear microinjection rounds for generating Zmynd10 mutants.
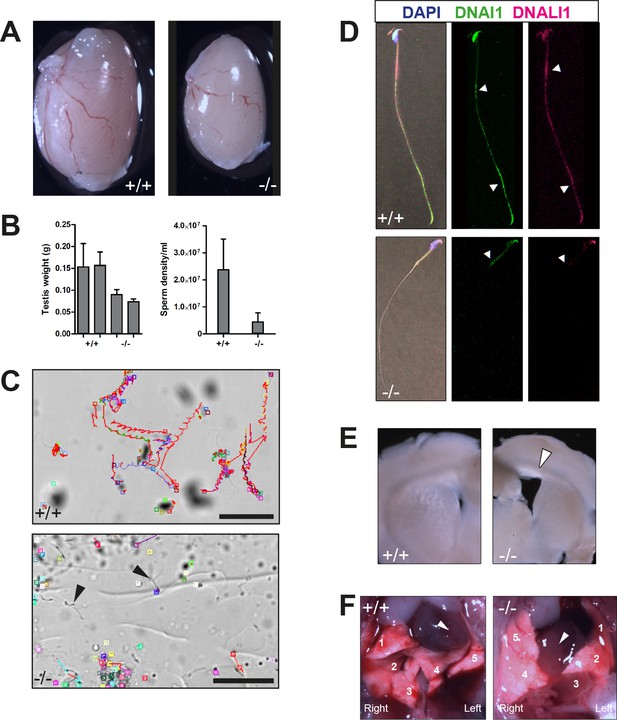
Detailed phenotypic analysis of Zmynd10 mutants.
(A) Representative image of a testis from Zmynd10 mutant and wild type control (P150). (B) As a result of defects in flagellar development, mutants have smaller testes (mean ± s.e.m, n = 3 males/genotype). Zmynd10 mutants have lower sperm density (mean ± s.e.m, n = 3 males/genotype). (C) Zmynd10 mutant sperm have reduced motility. Arrows point to static/paralyzed sperm in mutants. Scale bars = 50 μm. (D) By immunofluorescence, staining of DNAI2 and DNALI1 is absent along Zmynd10-/- flagella Arrowhead in wild type point to the annulus region and flagellar tip. In the mutant, the arrowhead points to the annulus. (E) Coronal vibratome sections of brains show Zmynd10 mutants display dilated lateral ventricles (arrowhead) due to hydrocephaly. (F) Zmynd10 mutants display heterotaxy defects. Arrowheads denote direction of the heart’s position, which is reversed in the mutants; numbers denote lobes of the lungs (1,2 and 3 correspond to upper, mid and lower lobes of the right lung; 4 and 5 correspond to the upper and lower lobes of the left lung).
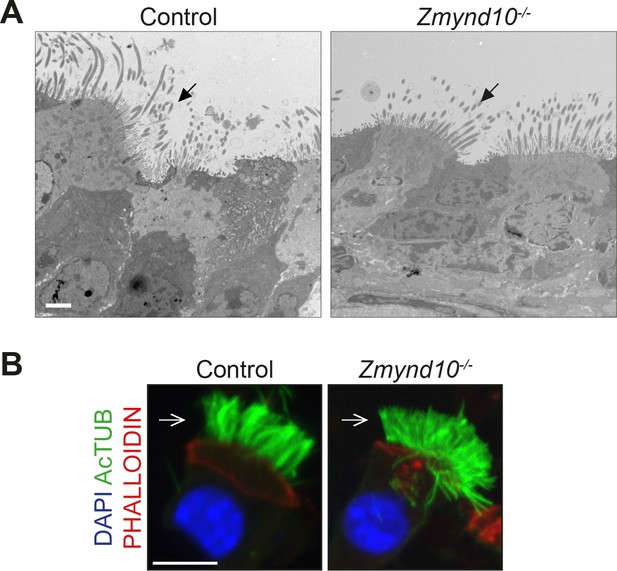
No gross ciliary defects are seen in Zmynd10 mutants.
(A) Transmission electron micrographs (TEM) of tracheal multiciliated cells from wild type and mutant female mice (P9). Arrows point to cilia on luminal surface of the cells; basal bodies appear correctly docked in the mutant cells similar to controls. Scale bar = 2 μm. (B) Immunostaining for cilia (acetylated α-tubulin) and apical F-actin (phalloidin) reveals no cytoskeletal gross defects in Zmynd10 mutants. Scale bars = 10 μm.
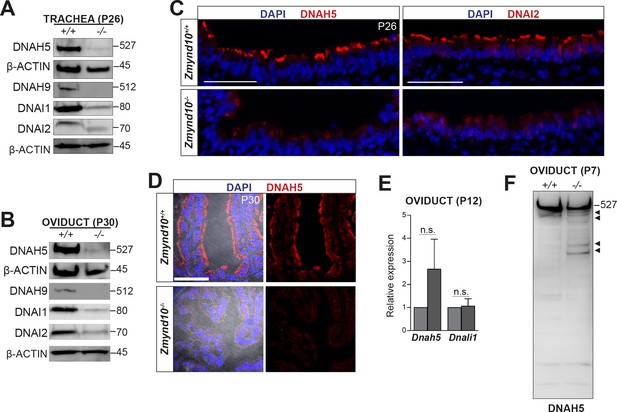
Global post-transcriptional destabilization of dyneins occurs in Zmynd10 mutant motile ciliated tissues.
Immunoblots of whole protein extracts from trachea P26 (A) and P30 oviducts (B) show reduced abundance of ODA subunits HC-γ, HC-β (DNAH5 and DNAH9), IC1 and IC2 (DNAI1 and DNAI2). Immunofluorescence of trachea (C) and oviduct (D) tissue sections show loss of axonemal DNAH5 and DNAI2 staining as well as reduced total abundance in Zmynd10 mutants compared to controls. Brightfield is included in oviduct merge panel (D) to highlight absence of staining in cilia in Zmynd10 mutants. Scale bars in (C) and (D) = 50 μm. (E) No significant changes are detected in levels of dynein transcripts by quantitative RT-PCR of (Dnah5, Dnali1) normalized to (Tbp) in P12 Zmynd10 mutant oviducts (n = 3/genotype, dark grey Zmynd10 mutants). (F) During early motile ciliogenesis, mildly reduced levels and laddering consistent with degradative, misfolded intermediates (arrowheads) of ODA HC-γ DNAH5 are detected in Zmynd10 mutant oviducts (P7). These will be subsequently cleared as tissue differentiation proceeds.
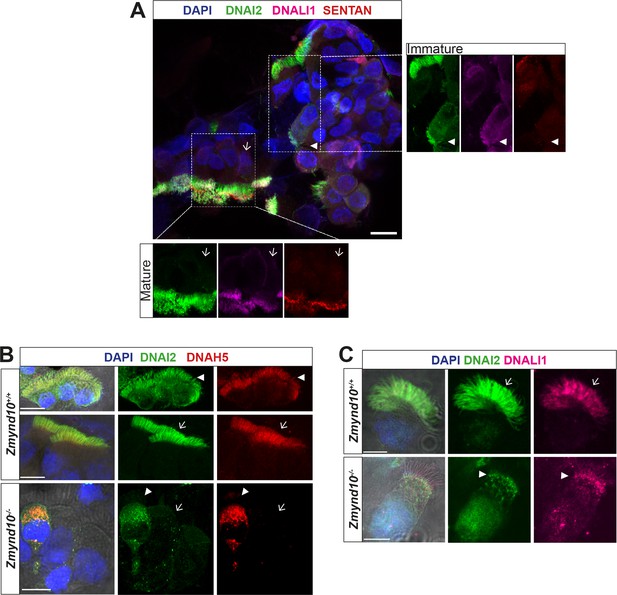
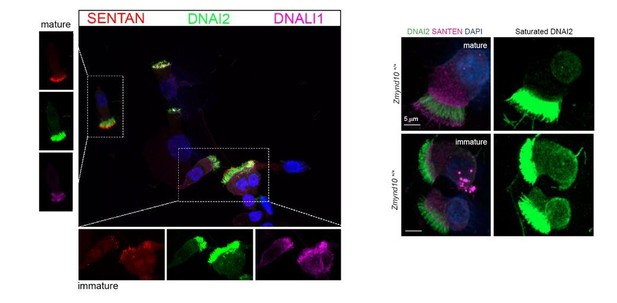
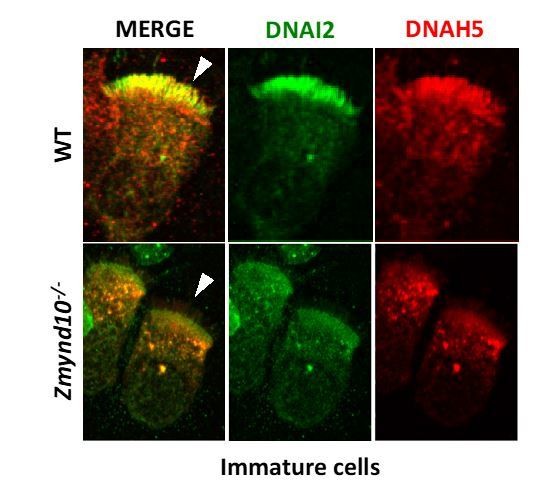
Loss of ZMYND10 perturbs sub-cellular distribution and levels of dynein complexes during pre-assembly.
(A) Z-projection through healthy human donor nasal brush immunofluorescence shows a mix of ‘mature’ motile ciliated cells (lower inset, arrow) with exclusively axonemal staining of dynein subunits (DNAI2 green, DNALI1 magenta) and strong foci of SENTAN (red) at cilial tips, as well as ‘immature’ cells having cytoplasmic staining of dynein subunits and no SENTAN (right inset, arrowheads). Scale bar = 10 μm. (B) Nasal brush immunofluorescence from Zmynd10 mice shows components of outer arm dyneins (DNAH5, DNAI2) are initially expressed in apical cytoplasm of immature mutant cells (arrowheads) but subsequently undergo ‘clearance’ in mature cells (lower panels, arrows), whilst all complexes exclusively translocate into cilia in control mature cells (middle panels, arrows). (C) Inner arm dynein component DNALI1 is expressed and apically arrested in immature mutant cells (arrowhead) and unlike controls, will get subsequentlycleared along with the ODA IC, DNAI2, in mature mutant cells. Scale bars = 5 μm.
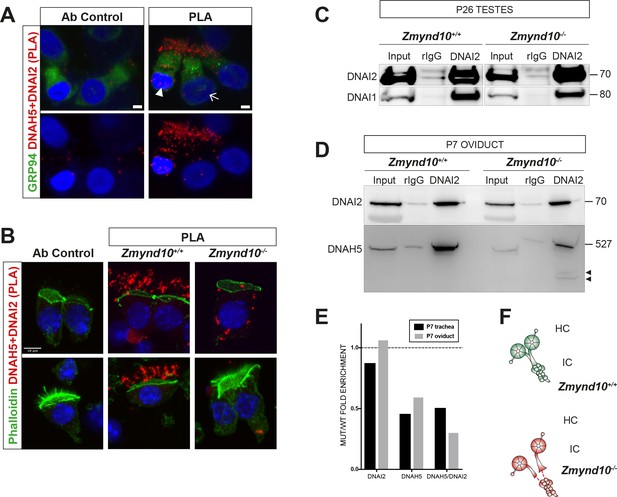
Sequential cytosolic assembly of outer arm dynein components occurs in mammalian motile ciliogenesis in a process requiring ZMYND10.
(A,B) Z-projections of Proximity Ligation Assay (PLA) on human donor (A) or mouse P7 (B) nasal brush biopsies confirming ODA subunits (mouse IC2/DNAI2 and HC-γ/DNAH5) are pre-assembled in the cytoplasm of mammalian multiciliated cells. Control (antibody only control, Ab control) sections incubated with only DNAH5 show no PLA signal. Red spots denote individual ODA complexes (<40 nm) that appear as peri-nuclear foci in immature cells (arrowhead) and translocate to cilia in mature cells (arrow). (B) In Zmynd10 mutant cells, reduced number of foci are observed and restricted only to the cytoplasm, highlighting defects in cytoplasmic pre-assembly. GRP94 was used as a pan-cytosolic marker (A) or phalloidin for apical actin ring (B). Nuclei are stained with DAPI (blue). See also Figure 4—figure supplement 1A for mouse IC2/DNAI2 and HC-β/DNAH9 PLA. Scale bars in (A) = 5 μm and (B) = 10 μm. (C) Endogenous Immunoprecipitation of DNAI2 from P26 Testes extracts reveals no defects in DNAI2 association with its heterodimeric partner DNAI1 in Zmynd10 mutants. (D) Endogenous Immunoprecipitation of DNAI2 from P7 oviduct extracts show disruption in subsequent association between DNAI2 and DNAH5 in mutants compared to controls as quantified by intensities of the DNAH5 pull-down bands (E). Arrowheads show predicted degradative or misfolded intermediates of DNAH5 polypeptide in the mutants only. Numbers to the right of panels denote protein molecular weight in kDa. See also Figure 4—figure supplement 1B,C for analysis in P7 and P90 trachea. (E) Ratio of mutant versus wild type fold enrichment (from 4D, Figure 4—figure supplement 1B: IP/input) for DNAI2 and DNAH5, normalized for differences in stability in input, as well as amount of DNAH5/DNAI2 complexes. (F) Schematic of axonemal ODA showing the intermediate chain heterodimers (IC) bind normally to heavy chains (HC) to form the entire motor complex in controls (green) and that this association is perturbed in mutants (red).
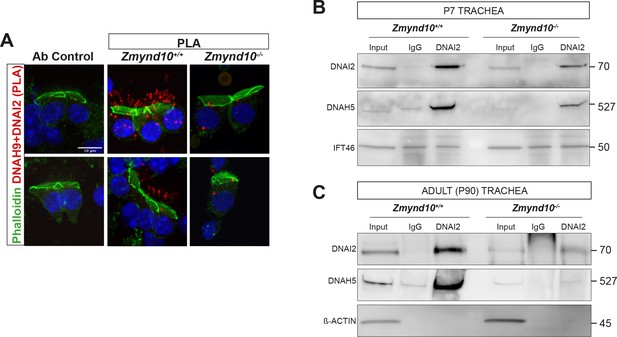
Sequential cytosolic assembly of outer arm dynein components occurs during mammalian motile ciliogenesis in a process requiring ZMYND10.
(A) Z-projections of proximity ligation assay (PLA) on mouse P7 nasal brush biopsies confirming that ODA subunits (mouse IC2/DNAI2 and HC-β/DNAH9) are pre-assembled in the cytoplasm. Control (antibody only control, Ab control) sections were incubated with only DNAH9 antibody and show no PLA signal. Red spots denote individual ODA complexes (<40 nm) in control cells and cilia. In Zmynd10 mutant cells, reduced number of foci are observed and restricted only to the cytoplasm, highlighting defects in cytoplasmic assembly. Phalloidin was used to mark the apical actin ring. Nuclei are stained with DAPI (blue). Scale bar = 10 μm. (B, C) Immunoprecipitation of DNAI2 from P7 trachea extracts show disruption in subsequent association between DNAI2 and DNAH5 in mutants compared to controls, as quantified by intensities of the DNAH5 pull-down bands (Figure 4E). In contrast, in P90 adult Zmynd10 mutant trachea (C), levels of DNAI2 are profoundly affected (input), and negligible complexes can be purified.
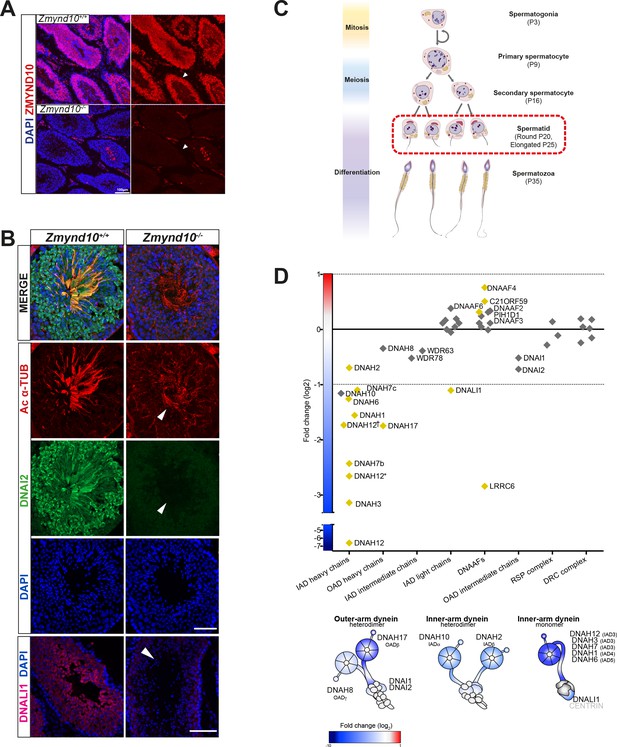
Early and specific defects in axonemal dynein heavy chain stability are observed in Zmynd10 mutants during cytoplasmic assembly.
(A) Immunofluorescence of ZMYND10 in control and mutant adult testes (P150) in asynchronous seminiferous tubules (arrowhead). (A) ZMYND10 is strongly expressed in primary spermatocytes and spermatids, where it is restricted to the cytoplasm and never in developing sperm tails. This staining is lost in mutants. Nuclei are stained with DAPI. Scale bar = 100 μm. (B) Cross-sections of similarly staged seminiferous tubules reveal similar developmental staging of sperm between adult control and mutants, but loss of DNAI2 (ODA) and DNALI1 (IDA) proteins from cytoplasm and axonemes (arrowhead) of mutant sperm. Nuclei are stained with DAPI. Scale bars = 50, 100 μm. (C) Schematic summarizing mouse spermatogenesis which is initially synchronized postnatally (stages shown to right), before occurring in asynchronous waves across seminiferous tubules. Axonemal dynein pre-assembly in the cytoplasm occurs from spermatid stage around P25 and continues till flagellogenesis at P30. (D) Unbiased quantitative proteomics of control and mutant testes (elongated spermatid: P25) reveal that loss of ZMYND10 leads to a primary reduction in abundance of all dynein HC subunits during cytoplasmic pre-assembly, whilst other components remain initially unaffected at this stage. Yellow diamonds highlight significantly (p<0.05) changed hits based on LFQ intensity between Zmynd10 mutants and wild type littermates (n = 3/genotype). Schematic below highlights fold change of specific subunits on given dynein arms. Some heavy chains had specific multiple isoforms detected (DNAH7b/c and DNAH12 (E9QPU2*, F6QA95, Q3V0Q1†). See also Supplementary file 1. Key: IAD: inner arm dynein; OAD: outer arm dynein; DNAAFs: dynein axonemal assembly factors; RSP: radial spoke protein; DRC: dynein regulatory complex.
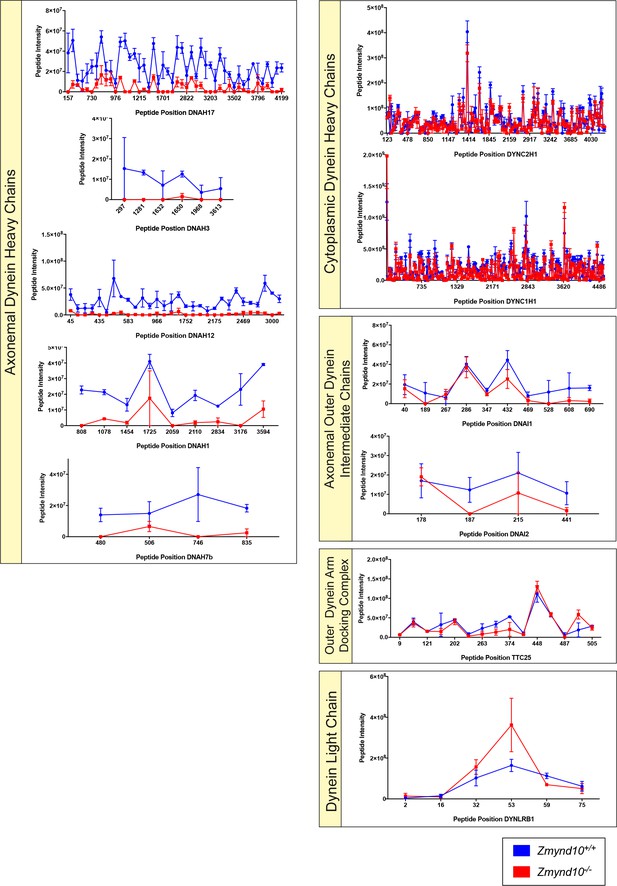
Peptide intensities across the axonemal dynein heavy chains reveal profound and significant decreases, whilst other dynein subunits are not affected.
Intensities of peptides detected by mass spectrometry from P25 testes whole proteome analysis (Figure 5D) are plotted across the polypeptide lengths for individual axonemal dynein subunits (ODA and IDA-HCs, ODA-ICs, ODA-LC). TTC25 (ODA-docking complex) and cytoplasmic dynein 1 and 2 heavy chain polypeptides are used as controls. Axonemal dynein heavy chains specifically show reduced peptide intensities in Zmynd10 mutant testes samples (red) compared to controls (blue). All the other proteins tested have comparable peptide intensities between mutant and control samples.
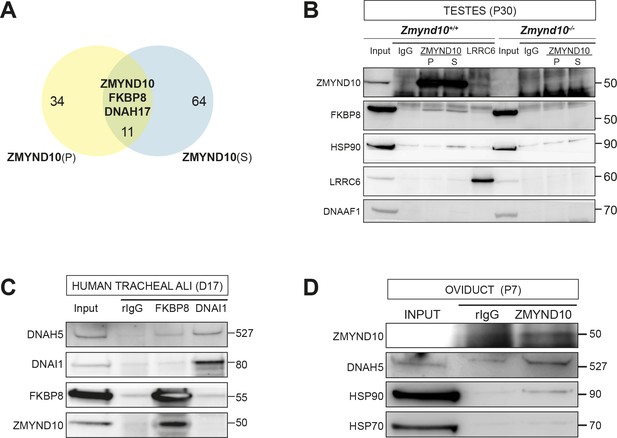
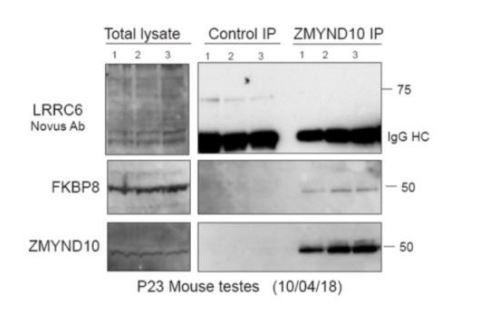
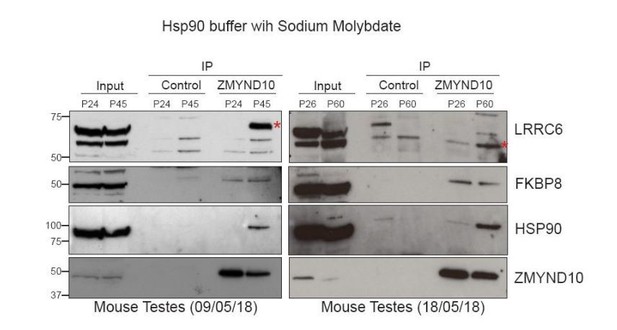
ZMYND10 interacts with a novel chaperone relay at a distinct stage of dynein heavy chain stability during cytoplasmic assembly.
(A) Summary schematic of affinity purification-mass spectrometry (AP-MS) analysis of endogenous ZMYND10 interactomes from P30 mouse testes, overlapping between two commercial polyclonal Sigma (S) and Proteintech (P) antibodies. See also Supplementary file 2. (B) Endogenous ZMYND10 affinity purification with two validated ZMYND10 antibodies from Zmynd10 control and mutant P30 testes extracts confirm the ZMYND10 interaction with FKBP8 and HSP90AB1 (S only) in control samples. These interactions are not found in mutant samples lacking ZMYND10, serving as specificity controls. An in vivo interaction between ZMYND10 and LRRC6, as well as with DNAAF1, was not detected in reciprocal endogenous immunoprecipitations in P30 control testes extracts, using rIgG as a control. (C) Endogenous FKBP8 and DNAI1 immunoprecipitations in differentiating healthy human donor tracheal epithelial cultures (D17 ALI) both show binding of client DNAH5, the rest of the complexes are distinct, suggesting they act at sequential steps of assembly. (D) Endogenous ZMYND10 immunoprecipitation of client DNAH5 and chaperone HSP90, but not HSP70 from differentiating oviduct epithelial tissue (P7) using rIgG as a control. Protein molecular weights displayed in KDa.
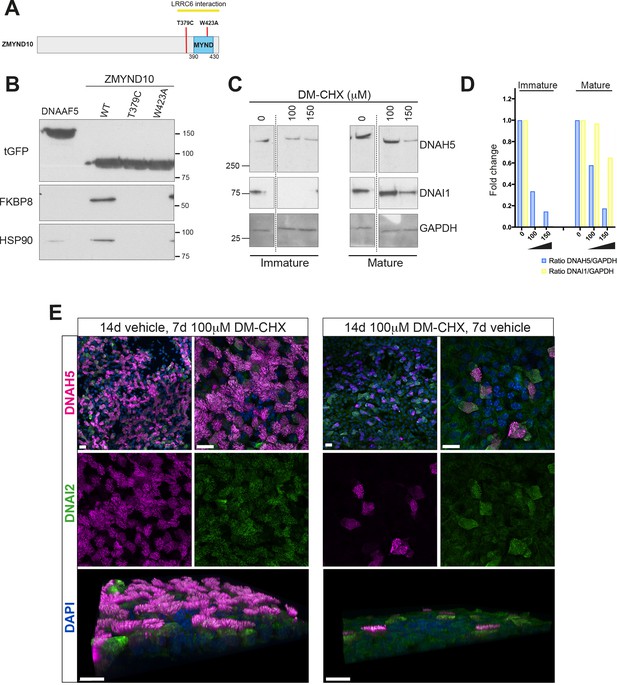
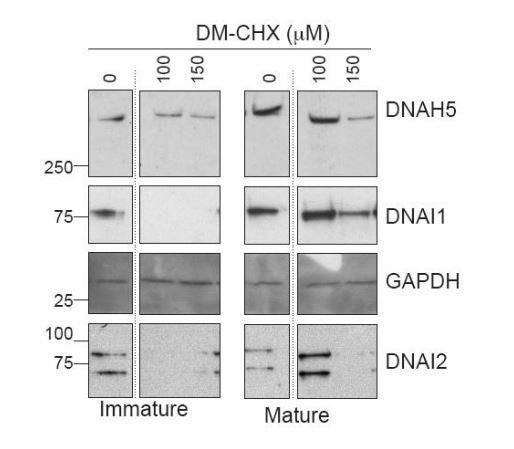
Ubiquitous FKBP8 actively participates in axonemal dynein heavy chain stability via its interaction with ZMYND10.
(A) Summary of mutations generated in ZMYND10-tGFP by site-directed mutagenesis to disrupt the binding interface for FKBP8, including the W423A mutation predicted to functionally disrupt one of two Zn2+-fingers in the MYND domain and the T379C PCD patient mutation (Zariwala et al., 2013), lying just before the MYND domain. (B) Extracts from transiently transfected HEK293 cells affinity purified against turboGFP shows C-terminal mutations interrupt endogenous FKBP8 and HSP90 binding to ZMYND10. (C) Healthy human donor tracheal epithelial cultures (MucilAir) before ciliation (D17, post-ALI) or fully ciliated (D60, post-ALI), were cultured for 24 hr in control (vehicle only) or DM-CHX (concentrations indicated in μM) before harvesting protein extracts. Immature cultures (D17) were more sensitive to effects of specific PPIase inhibitor DM-CHX, destabilizing dynein components, whilst mature cultures were minimally affected. (D) Quantification of band intensities for DNAH5 (blue) or DNAI1 (yellow) from (C) were normalized to loading control, and plotted as a fold change after 24 hr. (E) Z projections of whole-mount immunofluorescence of mouse tracheal epithelial cultures (MTECs) stained with DNAH5 (magenta), DNAI2 (green) and DAPI (blue) after treatment of three days post-airlift (ALI) with either 100 μm DM-CHX or vehicle control for 14 days in culture to differentiate, followed by switching cultures from control to 100 μm DM-CHX (mature: cilial beat visualized) or from DM-CHX into vehicle control (no cilia beat) and culturing an additional 7 days. Treatment of mature ciliated cultures had little effect on DNAH5 and DNAI2 levels, whilst treatment during differentiation disrupted expression of dynein subunits, as evidenced by lack of axonemal staining in most cells (right panels) but release allows recovery of dynein pre-assembly, mostly cytoplasmic after 7 days.
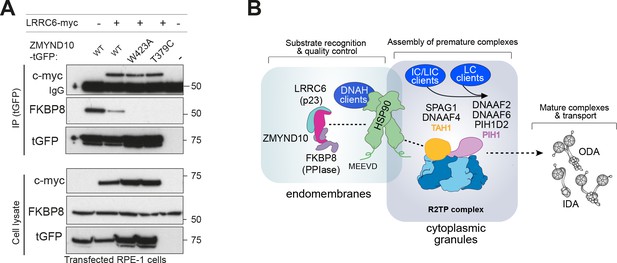
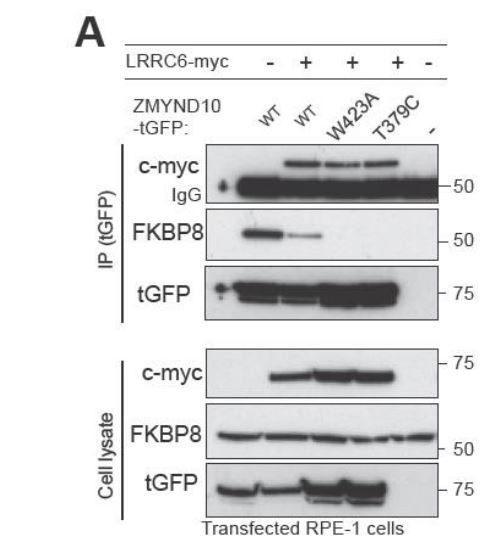
ZMYND10 specifies dynein heavy chains as clients in a chaperone relay during dynein pre-assembly.
(A) Extracts from transiently transfected RPE-1 cells with ZMYND10-turboGFP variants and human LRRC6-myc were affinity purified against turboGFP (upper panels). Inclusion of LRRC6 destabilizes binding of wild type ZMYND10 to endogenous FKBP8. C-terminal ZMYND10 mutations do not affect its interaction with LRRC6. Expression of interactors is confirmed in input cell lysates (lower panels). (B) During dynein arm assembly in the cytoplasm, ZMYND10 interaction with co-chaperone PPIase FKBP8 and HSP90 is required for the stabilization and folding of dynein heavy chains (DNAH). Competitive binding of LRRC6 to ZMYND10 may advance the HSP90-FKBP8 chaperone cycle taking the client DNAHs to the next stage of assembly. We propose either R2TP or a specialized R2TP-like complex may function in parallel to assemble, scaffold or tether the DNAI1-DNAI2 heterodimeric complex via the PIH-domain proteins, such as DNAAF6, DNAAF2 and PIH1D2. The TPR-domain containing SPAG1 or DNAAF4 could recruit HSP90 via its MEEVD domain and together with RUVBL1/RUVBL2 these assembly factors could form a stable platform to promote HC-IC subunit interactions. This R2TP or R2TP-like complex likely operates distinctly to ZMYND10 as DNAI1/2 heterodimers are detected in Zmynd10 mutants, however, these are likely degraded if fully functional mature complexes (with heavy chains) cannot be assembled. This chaperone-relay system comprises several discrete chaperone complexes overseeing the folding and stability of discrete dynein subunits. Folding intermediates are handed off to successive complexes to promote stable interactions between subunits all the while preventing spurious interactions. Stable dynein complexes, once formed are targeted to cilia via transport adaptors and intraflagellar transport (IFT).
Videos
Rapid ependymal ciliary motility in lateral ventricles of a wild type mouse.
High-speed video microscopy on a coronal brain vibratome section (postnatal day 11 mouse, littermate control) shows ependymal cilia lining lateral ventricles beating with high frequency in a wild type mouse.
Immotile ependymal cilia lining lateral ventricles of Zmynd10 mutant mouse.
High-speed video microscopy on a coronal vibratome section of a brain from a Zmynd10 mutant mouse (postnatal day 11) shows complete loss of ependymal cilia motility.
Control murine ependymal cilia with metachronal waveform.
High-speed video microscopy on a coronal vibratome section of a Zmynd10 mild hypomorphic mutant mouse brain (p. M179del; postnatal day 24) shows arrays of cilia beating in a metachronal waveform and actively generating fluid flow to move particulates over the ventricle tissue.
Tufts of immotile ependymal cilia in Zmynd10 mutant murine brain.
High-speed video microscopy on a coronal vibratome section of a Zmynd10 null mutant mouse brain (p. L188del; postnatal day 29) shows arrays of immotile cilia lining the ventricle tissue with no active fluid flow noticeable.
Aberrant flagellar motility in Zmynd10 mutant murine epididymal spermatozoa.
High-speed video microscopy on mature spermatozoa extracted from the epididymis of a 5 month old Zmynd10 CRISPR founder mutant mouse and slowed-down in methylcellulose. The majority of spermatozoa were completely immotile but rarely displayed highly aberrant flagellar movements as observed in the video.
Sinusoidal flagellar motility in wild type murine epididymal spermatozoa.
High-speed video microscopy on mature epididymal spermatozoa extracted from a 5 month-old wild type mouse (littermate control) and slowed-down in methylcellulose. Virtually all spermatozoa underwent forward motion with the flagella displaying a sinusoidal beat pattern.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Mus musculus) | Zmynd10 | NA | MGI:2387863; ENSMUSG00000010044; | Synonym: Blu; Dnaaf7 |
| Strain (M. musculus) | C57BL/6J | JAX | 664 | |
| Strain (M. musculus) | C3H/HeJ | JAX | 659 | |
| Strain (M. musculus) | CD1 (ICR) | Charles River | 022 | Outbred background |
| Genetic reagent (M. musculus) | Zmynd10em1Pmi | This paper | Allele symbol: Zmynd10em1Pmi; Allele synonym: Zmynd10-; Accession ID: MGI:6159883 | CRISPR null allele of Zmynd10; Zmynd10 c.695_701 p.Met178Ilefs*183 |
| Cell line (H. sapiens) | HEK293 | ATCC | CRL-1573 | Human embryonic kidney cell line. |
| Cell line (H. sapiens) | RPE-1 | ATCC | CRL-4000 | Human retinal pigmented epithelial cell line immortalized with hTERT. |
| Biological sample (M. musculus) | mouse tracheal epithelial cells (mTECs) | This paper | NA | See Vladar and Brody, 2013 for protocol. |
| Biological sample (H. sapiens) | MucilAir tracheal epithelial cell cultures | Epithelix Sarl | EP01MD | |
| Antibody | Acetylated α-tubulin | Sigma | 6-11B-1; T6793, RRID:AB_477585 | IF (1:500–2000) |
| Antibody | β-actin | Sigma | AC-15; A1978, RRID:AB_476692 | WB (1:1000) |
| Antibody | DNAAF1/LRRC50 | Novus Biologicals | NBP2-01936; RRID: AB_2732031 | WB (1:5000) |
| Antibody | DNAH5 | PMID: 23525783 | Custom made | IF (1:100), PLA; WB (1:5000) |
| Antibody | DNAH5 | Sigma | HPA037470, RRID:AB_10672348 | IF (1:100), PLA; WB (1:5000) |
| Antibody | DNAH9 | PMID: 24421334 | Custom made | IF (1:100), PLA; WB (1:5000) |
| Antibody | DNAH5 | Sigma | HPA037470, RRID:AB_10672348 | WB (1:5000) |
| Antibody | DNAI1 | Abcam | ab171964; RRID: AB_2732030 | WB (1:5000) |
| Antibody | DNAI2 | Abnova | M01 clone IC8; H00064446-M01, RRID:AB_426059 | IF (1:100), PLA; WB (1:5000) |
| Antibody | DNAI2 | Proteintech | 17533–1-AP; 17533–1-AP, RRID:AB_2096670 | IF (1:100); WB (1:5000); IP (1.5 μg-3μg/IP) |
| Antibody | DNALI1 | Santa Cruz | N-13; sc-160296, RRID:AB_2246230 | IF (1:75); WB (1:1000) |
| Antibody | FKBP8 | Proteintech | 11173–1-AP, RRID:AB_10597097 | WB (1:5000); IP (1.5 μg-3μg/IP) |
| Antibody | FKBP8 | R and D Systems | MAB3580, RRID:AB_2262675 | WB (1:5000) |
| Antibody | γ tubulin | Abcam | GTU-88; ab11316, RRID:AB_297920 | IF (1:500) |
| Antibody | GAPDH | Abcam | ab8245, RRID:AB_2107448 | WB (1:5000) |
| Antibody | tGFP | Origene | TA150041, RRID:AB_2622256 | IF (1:200); WB (1:5000) |
| Antibody | GFP | Santa Cruz | FL; sc-8334, RRID:AB_641123 | WB (1:5000); IP (1.5 μg-3μg/IP) |
| Antibody | GRP-94/HSP90B1 | Thermo Scientific | clone 9G10; MA3-016, RRID:AB_2248666 | IF (1:100); WB (1:5000) |
| Antibody | HSP70 | Santa Cruz | K-20; sc-1060, RRID:AB_631685 | WB (1:5000) |
| Antibody | HSP90AB1 | R and D Systems | MAB32861, RRID:AB_2121071 | WB (1:5000) |
| Antibody | HSP90 | Santa Cruz | Clone F-8; sc-13119, RRID:AB_675659 | WB (1:5000) |
| Antibody | LRRC6 (Hiroshi Hamada) | PMID:27353389 | Custom made | WB (1:5000), a gift from Hiroshi Hamada |
| Antibody | SENTAN | Sigma | HPA043322 , RRID: AB_10793945 | IF (1:150) |
| Antibody | ZMYND10 | Proteintech | 14431–1-AP, RRID:AB_2218002 | WB (1:5000); IF (1:100); IP (1.5 μg-3μg/IP) |
| Antibody | ZMYND10 | Sigma | HPA035255, RRID:AB_10601928 | WB (1:5000); IF (1:100); IP (1.5 μg-3μg/IP) |
| Recombinant DNA reagent | pCMV6-Zmynd10-tGFP | Origene | MG207003 | Mouse Zmynd10 ORF with C-terminal turbo-GFP tag under CMV promoter in plasmid with ampicillin resistance gene |
| Recombinant DNA reagent | pCMV6-DNAAF5-tGFP | Origene | MR221395 | Mouse Dnaaf5 ORF with C-terminal turbo-GFP tag under CMV promoter in plasmid with ampicillin resistance gene |
| Recombinant DNA reagent | pRK5-Myc-LRRC6 | PMID:23891469 | NA | Human LRRC6 ORF with myc tag; gift from the Hildebrandt and Gee labs |
| Recombinant DNA reagent | pX330-U6-Chimeric _BB-CBh-hSpCas9 | PMID: 23287718 | Addgene:#42230 | A human codon-optimized SpCas9 and chimeric guide RNA expression plasmid. pX330-U6-Chimeric_BB-CBh-hSpCas9 was a gift from Feng Zhang. |
| Recombinant DNA reagent | pCAG-EGxxFP | PMID: 24284873 | Addgene:#50716 | 5’ and 3’ EGFP fragments that shares 482 bp were placed under ubiquitous CAG promoter. Used for validation of gRNA sequences by DSB mediated EGFP reconstitution. pCAG-EGxxFP was a gift from Masahito Ikawa |
| Sequence-based reagent | mouse Dnahc5 qRT-PCR primers | This paper | AAGCTGTTGCACCAGACCAT/ CCCAGGTGGCAGTTCTGTAG; Probe:88 | |
| Sequence-based reagent | mouse Dnali1 qRT-PCR primers | This paper | AGTTCCTGAAACGGACCAAC/ TGAGACCCATGTGGAAATGA; Probe:97 | |
| Ssequence-based reagent | mouse Zmynd10 qRT-PCR primers | This paper | GCCATCCTTGATGCAACTATC/ CAATCAGCTCCTCCACCAG; Probe:64 | |
| Sequence-based reagent | mouse Tbp qRT-PCR primers | This paper | GGGGAGCTGTGATGTGAAGT/ CCAGGAAATAATTCTGGCTCA; Probe:97 | |
| Chemical compound, drug | N-(N’N’-Dimethyl carboxamidomethyl) cycloheximide (DM-CHX) | PMID:16547004 | FKBP8 inhibitor, 1 mM stock in sterile PBS | |
| Software, algorithm | Fiji | PMID: 22743772 | ||
| Software, algorithm | Nis-Elements AR V4.6 | Nikon Instruments | ||
| Software, algorithm | Imaris V9.1 | Bitplane | ||
| Software, algorithm | MaxQuant | PMID: 19029910 | ||
| Software, algorithm | Andromeda | PMID: 21254760 | ||
| Software, algorithm | Perseus | PMID: 27348712 | ||
| Software, algorithm | Crapome | PMID: 23921808 |
Additional files
-
Source data 1
Figures 1–8 Uncropped immunoblots
Images show un-cropped versions of immunoblots. Figure panels used are demarcated with a box and the detected proteins are labeled. Asterisks are used to denote either non-specific bands (6C, 6D and 7B) or putative degradation products (7E). HE = High exposure, LE = Low exposure
- https://doi.org/10.7554/eLife.34389.021
-
Supplementary file 1
Summary of label-free quantitative global proteomics of P25 testes from Zmynd10+/+ and Zmynd10-/- mice.
Label free quantitative (LFQ) proteomics were performed on whole testes extracts to compare differential protein expression profiles between Zmynd10-/-mutants and age matched (P25) wild type littermate controls. LFQ intensities from three sets of biological replicates were averaged per genotype and ratios of enrichment or depletion in mutant versus wild type were calculated. Significance values were calculated using a Student's paired 2-tailed t-test. For ease of interpretation, the proteins were grouped into their respective complexes (i.e. Inner Dynein Arm Heavy Chains, Outer Dynein Arm Heavy Chains etc.)
- https://doi.org/10.7554/eLife.34389.022
-
Supplementary file 2
Endogenous ZMYND10 immunoprecipitation protein interactor profiling.
Filtered by significance. Endogenous ZMYND10 was immunoprecipitated from total extracts of control wild type testes (P30) with two commercial polyclonal antibodies (Proteintech and Sigma). Proteins that co-precipitated with ZMYND10 in triplicate experiments were identified by tandem Mass Spectrometry and ranked according to fold-change (FC Ratio) enrichment and significance (t-test p-value) in the ZMYND10 specific pull-downs versus control immunoglobulin pull-downs. Proteins that commonly precipitated with ZMYND10 using the two separate polyclonal antibodies were considered putative interactors. A ranked list of these common hits is provided with ZMYND10 and the interactors investigated in this study that is FKBP8, HSP90AB1 and DNAH17 highlighted in green.
- https://doi.org/10.7554/eLife.34389.023
-
Supplementary file 3
Table of antibodies, qRT-PCR primers and plasmids used in this study.
- https://doi.org/10.7554/eLife.34389.024
-
Supplementary file 4
Table of additional mouse primers used in this study.
- https://doi.org/10.7554/eLife.34389.025
-
Transparent reporting form
- https://doi.org/10.7554/eLife.34389.026





