Multisite dependency of an E3 ligase controls monoubiquitylation-dependent cell fate decisions
Figures
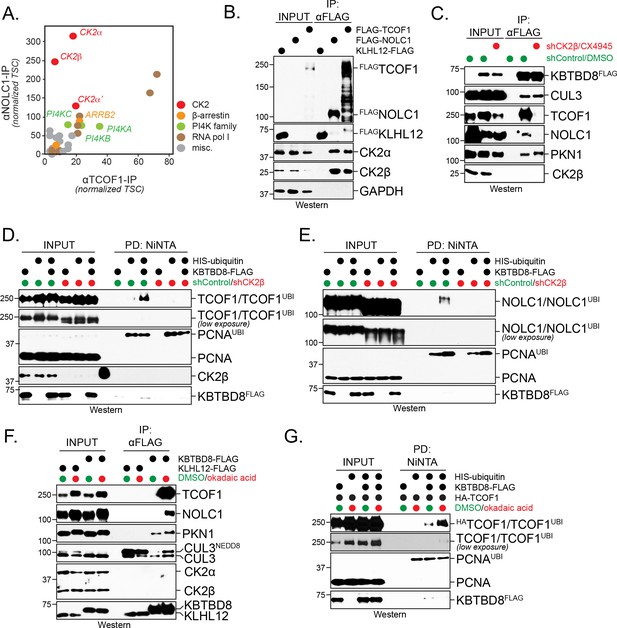
CK2 kinase is required for CUL3KBTBD8-substrate binding and ubiquitylation in cells.
(A) Both NOLC1 and TCOF1 associate with the CK2 kinase. FLAGNOLC1 and FLAGTCOF1 were affinity-purified from 293 T cells and specific binding partners were determined by CompPASS mass spectrometry. Total spectral counts (TSCs) of specific interactors were normalized to 1000 TSCs for each bait protein and plotted against each other. For each CUL3KBTBD8-substrate, three independent affinity-purification and mass spectrometry experiments were performed. The results of each affinity experiments were compared to a database, which included ~100 unrelated affinity-purifications performed with the same antibody and in the same cell line. (B) TCOF1 and NOLC1 associate with endogenous CK2 in cells. FLAGNOLC1 and FLAGTCOF1 were affinity-purified from 293 T cells, and co-purifying proteins were identified by western blotting using the indicated antibodies. KLHL12FLAG was used as a negative control. The low molecular weight species seen in TCOF1 samples are proteolytic degradation products caused by partial cleavage of the large and intrinsically disordered TCOF1 during the affinity-purification. (C) CK2 is required for substrate recognition by CUL3KBTBD8 in cells. 293 T cells were depleted of CK2β using shRNAs and the activity of the remaining CK2 kinase was simultaneously inhibited using the small molecule CX4945. KBTBD8FLAG, the substrate adaptor of CUL3, was affinity-purified and bound endogenous proteins were detected by western blotting using the indicated antibodies. PKN1 is a specific binding partner of KBTBD8 that is not known to be ubiquitylated by CUL3KBTBD8 and that is not important for neural crest specification. (D) CK2 is required for monoubiquitylation of TCOF1 in cells. Ubiquitylated proteins were purified under denaturing conditions from 293 T cells expressing HISubiquitin. Ubiquitylated proteins were detected by western blotting using the indicated antibodies. PCNA is ubiquitylated by a distinct E3 ligase and hence is used as a control for general ubiquitylation efficiency; unmodified PCNA is not detected in NiNTA pulldowns, demonstrating the specificity of this assay. Input: 3% of lysate used for denaturing purification. (E) CK2 is required for monoubiquitylation of NOLC1 in cells. Ubiquitylated NOLC1 was purified both in the absence or presence of CK2 inhibitors, as described for TCOF1 above. Input: 3% of lysate used for denaturing purification. (F) Inhibition of PP1 and PP2A phosphatases, which counteract CK2 in cells, promotes substrate binding by CUL3KBTBD8. KBTBD8FLAG or the negative control KLHL12FLAG were affinity-purified from cells treated with the PP1/PP2A-inhibitor okadaic acid. Co-purifying proteins, including endogenous TCOF1, NOLC1, and the control KBTBD8-interactor PKN1, were detected by western blotting. This experiment was performed at smaller scale than Figure 1C, explaining the lack of TCOF1 binding to KBTBD8 under conditions without phosphatase inhibitors. (G) Inhibition of PP1/PP2A phosphatases increases TCOF1 ubiquitylation in cells. Ubiquitylated HATCOF1 was purified under denaturing conditions from 293 T cells expressing HISubiquitin and treated with or without okadaic acid. Ubiquitylated proteins were detected by Western blotting. Input: 3% of lysate used for denaturing purification.
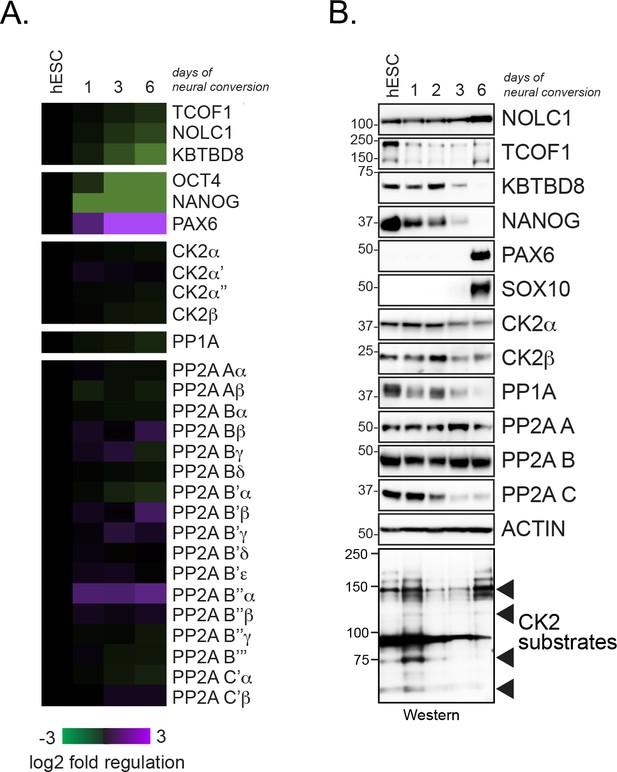
Expression analysis of CK2 and PP1/PP2A in differentiating hESCs.
(A) Expression analysis of CK2 and PP1/PP2A subunits in self-renewing hESC and hESCs undergoing neural conversion for the indicated time periods using previously published RNAseq data sets (Werner et al., 2015). (B) Expression analysis of CK2 and PP1/PP2A subunits during neural conversion of hESCs by Western blotting using specific antibodies.
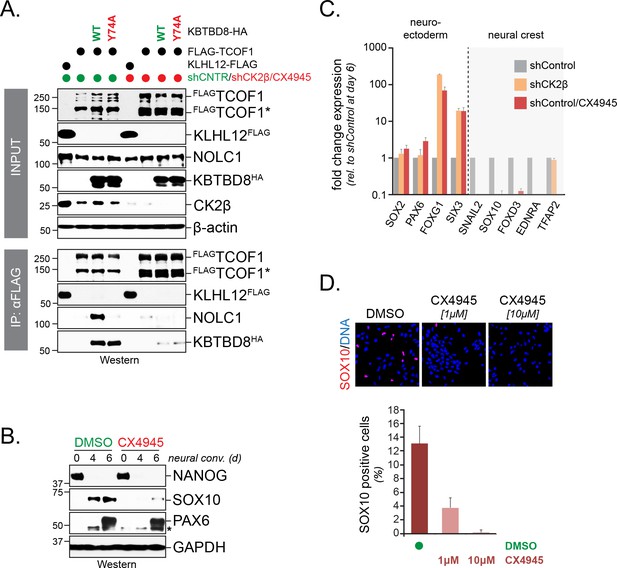
CK2 is required for neural crest specification.
(A) CK2 is required for CUL3KBTBD8- and ubiquitylation-dependent formation of TCOF1-NOLC1 complexes. FLAGTCOF1 was affinity-purified from control- or CK2 activity-depleted cells (shCK2β/CX4945) that expressed WT-KBTBD8 or mutant KBTBD8Y74A, which is unable to bind CUL3 and thus cannot catalyze monoubiquitylation. Bound proteins, including endogenous NOLC1, were detected by western blotting. KLHL12FLAG was used as a control protein. (B) CK2 activity is required for neural crest formation from hESCs. H1 hESCs were subjected to neural conversion in the absence or presence of 1 μM CK2 inhibitor CX4945. Expression of markers for central nervous system precursor (PAX6) or the neural crest (SOX10) was analyzed by Western blotting. Differentiation was monitored by following the stem cell-specific transcription factor NANOG, using specific antibodies. (C) CK2 controls a switch between CNS and neural crest precursors. Neural conversion of H1 hESCs was induced under control conditions (shControl), in the presence of an shRNA to deplete endogenous CK2β (shCK2β), or in the presence of 1 μM CK2 inhibitor CX4945 (shControl/CX4549). Differentiation was followed by qRT-PCR analysis of markers of the CNS (SOX2, PAX6, FOXG1, SIX3) or the neural crest (SNAIL2, SOX10, FOXD3, EDNRA, TFAP2). Fold change expression is relative to shControl at day 6 of differentiation. Error bars denote s.e.m. of three technical replicates. (D) CK2 is required for neural crest specification, as detected at the single cell level. H1 hESCs were subjected to neural conversion in the absence of presence of indicated concentrations of CK2 inhibitor CX4945 for 7d. Formation of neural crest cells was assessed by indirect immunofluorescence microscopy using antibodies specific to SOX10. Representative images are shown. SOX10 positive cells were quantified from three independent experiments and expressed as percentage of total cells imaged. Error bars denote standard deviation.
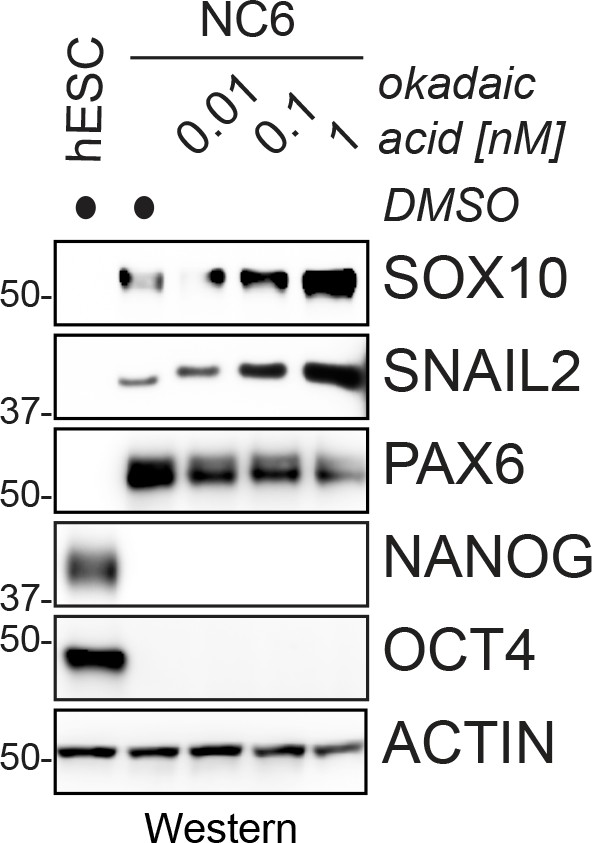
Phosphatase inhibition increases the efficiency of neural crest specification.
hESCs were subjected to neural conversion in the presence of increasing concentrations of the PP1/PP2A inhibitor okadaic acid, and neural crest specification was monitored by following the expression of the neural crest markers SOX10 or SNAIL2 using western blotting. Okadaic acid is toxic to hESCs at concentrations above 1 nM.
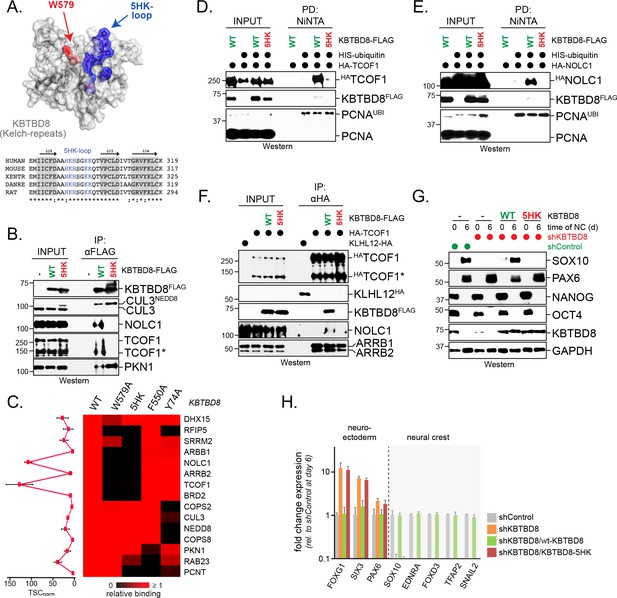
A conserved loop in KBTBD8 is required for CK2-substrate recognition.
(A) KBTBD8 contains a loop with five His- or Lys-residues that is close to W579, a KBTBD8-residue known to be important for substrate recognition. Depicted is a structural model of the Kelch-domain of KBTBD8; W579 is shown in red, whereas the 5HK-loop is shown in blue. (B) The 5HK-loop in KBTBD8 is required for substrate recognition. KBTBD8FLAG or KBTBD85HK-FLAG (i.e. a KBTBD8-mutant in which all five His- or Lys-residues in the conserved loop were exchanged to Ala) were affinity-purified from cells. Co-purifying TCOF1, NOLC1, or PKN1 were detected by western blotting. (C) The 5HK-loop of KBTBD8 is essential for substrate recognition. The indicated KBTBD8 variants (wt; W579A; 5HK-Ala; F550A; Y74A) were affinity-purified from 293 T cells and analyzed for binding partners by CompPASS mass spectrometry. The heatmap depicts the relative binding of interactors identified for wild-type KBTBD8 in the respective KBTBD8 variant (black = no interaction, red = equal or more interaction). A quantification of normalized total spectral counts (TSCs) of three biological replicates is shown for the wild-type KBTBD8 affinity-purification is shown on the left. (D) The 5HK-loop of KBTBD8 is required for TCOF1 ubiquitylation in cells. Ubiquitylated proteins were purified under denaturing conditions from 293 T cells expressing HISubiquitin. Modified proteins were detected by western blotting, using the indicated antibodies. (E) The 5HK-loop of KBTBD8 is required for NOLC1 ubiquitylation in cells. Ubiquitylated proteins were purified from 293 T cells under denaturing conditions and analyzed as described above. (F) The 5HK loop of KBTBD8 is required for TCOF1-NOLC1 complex formation. Cells expressing HATCOF1 and either KBTBD8FLAG, KBTBD85HK-FLAG, or the control protein KLHL12FLAG were subjected to anti-HA affinity-purification and analyzed for co-purifying proteins by western blotting. ARBB1 and ARBB2 binding to TCOF1 shows that the efficiency of affinity-purification was identical under all conditions. (G) The 5HK loop in KBTBD8 is required for neural crest specification. H1 hESCs were depleted of endogenous KBTBD8 using stably expressed shRNAs. Where indicated, hESCs were reconstituted with shRNA-resistant wild-type (WT) or mutant (5HK) KBTBD8. Differentiation was followed be western blotting using markers of the neural crest (SOX10), CNS precursors (PAX6), or pluripotency (NANOG; OCT4). (H) The 5HK loop of KBTBD8 is required for neural crest specification. H1 hESCs were depleted of KBTBD8 and reconstituted with shRNA-resistant wild-type KBTBD8 or mutant KBTBD85HK. Expression of markers of CNS/forebrain (FOXG1, SIX3, PAX6) or the neural crest (SOX10, EDNRA, FOXD3, TFAP2, SNAIL2) was followed by qRT-PCR. Fold change expression is relative to shControl at day 6 of differentiation. Error bars denote s.e.m. of three technical replicates.
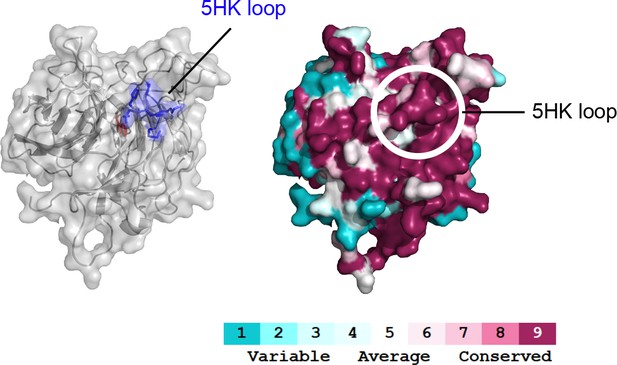
Conservation analysis of KBTBD8.
KBTBD8 variants from 54 vertebrate species were subjected to conservation analysis, and conserved residues were mapped onto a structural model of the KELCH-repeats of human KBTBD8. The 5HK loop is highly conserved.
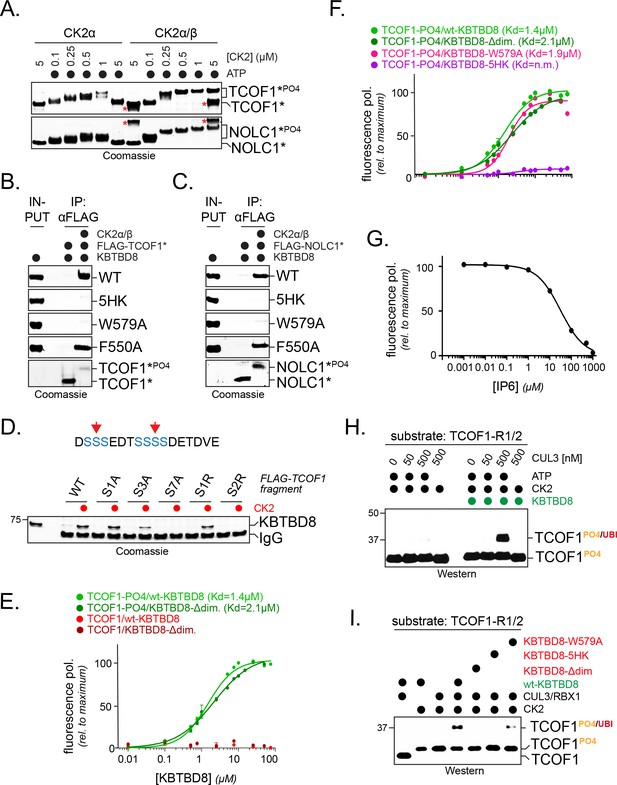
CK2 marks CUL3KBTBD8-substrates for monoubiquitylation in vitro.
(A) TCOF1 and NOLC1 are phosphorylated by recombinant CK2 in vitro. Recombinant Flag-tagged TCOF1190-364 or NOLC176-187 fragments with two CK2 motifs (Flag-TCOF1*, Flag-NOLC1*) were incubated with either the catalytic subunit CK2α or complexes between CK2α and the regulatory subunit CK2β. ATP was added as indicated, and proteins were detected by Coomassie staining. The molecular weight shift observed in the presence of both CK2 and ATP is indicative of phosphorylation. The red asterisk marks CK2β, as detected at its highest concentration. In the absence of the stabilizing CK2β-subunit, high concentrations of CK2α likely lead to aggregation and reduced TCOF1 phosphorylation. (B) KBTBD8 selectively binds phosphorylated TCOF1. The binding of recombinant Flag-TCOF1* (containing two CK2 motifs) to KBTBD8 was analyzed in the absence or presence of CK2α/β. TCOF1 was immobilized on beads and bound KBTBD8 was detected by Coomassie staining. The KBTBD8 variants tested (WT, 5HK, W579A, F550A) are indicated on the right. (C) KBTBD8 selectively binds phosphorylated NOLC1. The binding of recombinant Flag-NOLC1* (containing two CK2 motifs) to KBTBD8 was analyzed as described above for TCOF1. (D) Mutation of a CK2 motif obliterates its recognition by KBTBD8. The indicated CK2 motif of TCOF1197-213 was altered by introduction of one or two Arg residues (sites indicated by red arrows in scheme above) or by mutation of one (S1A), three (S3A) or all seven (S7A) phosphoacceptor Ser residues. Binding to immobilized KBTBD8 was analyzed either in the absence or presence of CK2, using Coomassie staining. (E) KBTBD8 binds a CK2 motif in TCOF1 in a phosphorylation-dependent manner. A single N-terminally TAMRA-labeled CK2 motif of TCOF1194-216 was either incubated with CK2 (‘-PO4’) or kept unmodified and incubated with increasing concentrations of recombinant KBTBD8. Binding of KBTBD8 to the CK2 motif was measured as change in fluorescence polarization, and the dissociation constant of the binding reaction was determined by fitting to a two-state binding curve. In addition to wild-type KBTBD8, the ability of a monomeric variant of KBTBD8 (KBTBD8-Δdim) was monitored (green curves). In the absence of CK2, no measurable binding was detected (red curves). (F) Mutation of the 5HK loop, but not of W579, obliterates binding of KBTBD8 to a single CK2 motif. Binding of distinct KBTBD8 variants (WT, Δdim, W579A, 5HK) to a phosphorylated CK2 motif of TCOF1 was measured by fluorescence polarization. (G) Substrate binding by KBTBD8 is likely electrostatic in nature. Binding of a single phosphorylated CK2 motif from TCOF1 to recombinant KBTBD8 was measured by fluorescence polarization in the presence of increasing concentrations of the highly negatively charged molecule inositolehexakisphosphate (IP6). (H) TCOF1 is monoubiquitylated in vitro. A TCOF1 fragment containing two CK2 motifs (‘R1/2’) was first phosphorylated by CK2α/β and then immobilized on FLAG-agarose followed by incubation with KBTBD8; immobilization interfered with non-specific ubiquitylation of the substrate peptide. Next, recombinant neddylated CUL3/Rbx1, E1, UBE2D3, ubiquitin, and ATP were added as indicated, and ubiquitylation of TCOF1 was followed by Western blotting. (I) Monoubiquitylation of TCOF1-R1/2 by CUL3KBTBD8 was analyzed as described above, using the indicated KBTBD8 mutants.
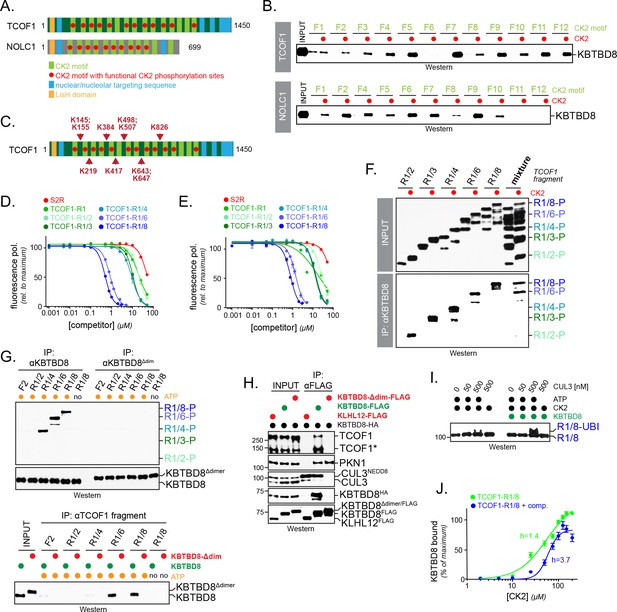
Multisite dependency of substrate recognition by CUL3KBTBD8 in vitro.
(A) TCOF1 and NOLC1 contain multiple CK2 motifs (light green). Functional motifs are marked with a red dot (non-functional motifs typically contain Arg substitutions that are incompatible with recognition by CK2). The LisH domain is a protein interaction module found in many proteins, but often with unknown function. (B) Multiple CK2 motifs are recognized by KBTBD8. TCOF1 or NOLC1 fragments containing single CK2 motifs (number according to position in sequence) were incubated with buffer or with CK2/ATP, immobilized on beads, and incubated with recombinant KBTBD8. Bound KBTBD8 was visualized by western blotting using a specific antibody. (C) TCOF1 is ubiquitylated in vivo on multiple Lys residues that are all in close proximity to CK2 motifs. Ubiquitylated TCOF1 was purified under denaturing conditions from 293 T cells and ubiquitylated Lys residues were determined by mass spectrometry. The figure depicts the location of ubiquitylated lysine residues identified in individual peptides. (D) The more CK2 motifs are in a substrate, the better it binds to KBTBD8. A single phosphorylated and fluorescently labeled CK2 motif of TCOF1 was bound to KBTBD8. Unlabeled TCOF1 competitor fragments containing either two (R1/2), three (R1/3), four (R1/4), six (R1/6) or eight (R1/8) CK2 motifs were phosphorylated and titrated into the binding reaction, and dissociation of the reporter peptide was monitored by loss of fluorescence polarization. (E) The increase in affinity afforded by multiple CK2 motifs also applies to monomeric KBTBD8. A monomeric variant of KBTBD8 was produced by mutation of conserved residues at the BTB dimer interface, as described for KLHL12 (McGourty et al., 2016). Binding of a single phosphorylated CK2 motif of TCOF1 to monomeric KBTBD8 was then monitored in the presence of competitor peptides by fluorescence polarization, as described above. (F) Multiple CK2 motifs increase substrate affinity for KBTBD8. TCOF1 fragments containing either two (R1/2), three (R1/3), four (R1/4), six (R1/6) or eight (R1/8) CK2 motifs were incubated with immobilized KBTBD8 either in the presence or absence of CK2. In the last two lanes, all TCOF1 fragments were mixed and incubated at the same time with KBTBD8. Binding was monitored by western botting (top lanes: input; bottom lanes: KBTBD8-immunoprecipitation). (G) Multiple CK2 motifs increase substrate affinity towards KBTBD8 at low substrate concentrations. Upper panel: 450 nM of TCOF1 fragments with a single (F2) or multiple (R1/2, R1/4, R1/8) CK2 motifs were incubated with either wild-type KBTBD8 or monomeric KBTBD8Δdim, and KBTBD8 was then affinity-purified using a specific antibody. Lower panel: Same binding reaction as above, but TCOF1 was immobilized on beads. Complex formation was analyzed by western blotting. (H) Dimerization of KBTBD8 is required for TCOF1, but not PKN1, binding in cells. Cells expressing indicated KBTBD8 variants were subjected to KBTBD8FLAG affinity-purification and analyzed for dimerization (KBTBD8HA), CUL3-binding, or substrate recognition (TCOF1, PKN1) using Western blotting with specific antibodies. (I) CUL3KBTBD8-substrates with multiple CK2 motifs are monoubiquitylated. Ubiquitylation of TCOF1-R1/8, a fragment containing eight CK2 motifs, was analyzed in vitro using purified neddylated CUL3-RBX1, KBTBD8, E1, UBE2D3, ubiquitin and ATP, as described above. (J) Ultrasensitivity of the KBTBD8-TCOF1 interaction. Binding of TCOF1-R1/8 to KBTBD8 was analyzed in the presence of increasing CK2 concentrations, as described in the Figure 5F. Western blots of three independent experiments were quantified by Image J, and error bars denote standard deviation. Green curve: binding reaction in the absence of competitor; blue curve: binding of TCOF1-R1/8 in the presence of a ten-fold excess of TCOF1-R1/2, which mimics endogenous CK2 substrates that typically have one or two CK2 motifs.

N- and C-terminal TCOF1 fragments both require multiple CK2 motifs for recognition.
TCOF1 fragments containing different numbers of CK2 motifs were generated from a more C-terminal part of the protein and compared for their affinity to KBTBD8 to N-terminal fragments described in Figure 5. Note that the R3/8 fragment contains an inactive motif in between two and three active CK2 motifs, respectively, and it accordingly behaves like a TCOF1 fragment with three consecutive CK2 motifs (R1/3).
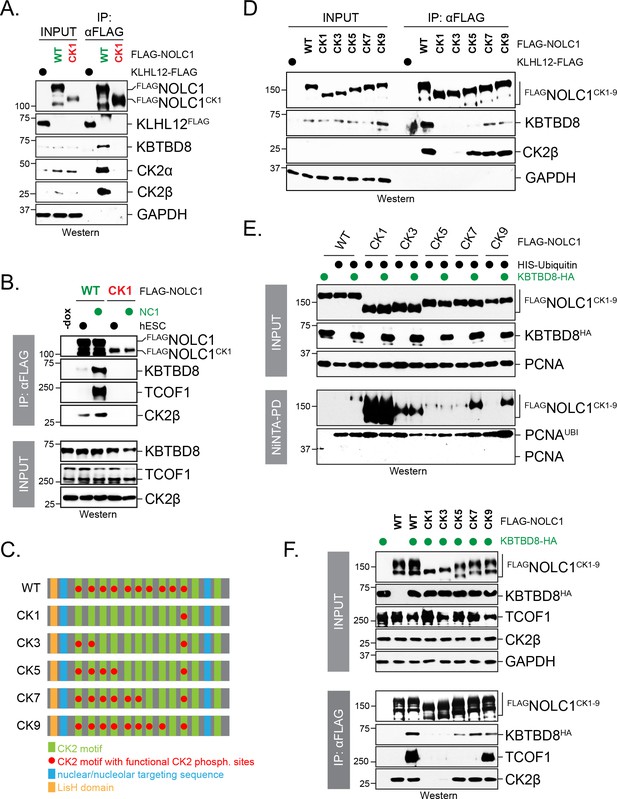
Multisite dependency of CUL3KBTBD8 substrate recognition in cells.
(A) A single CK2 motif is not sufficient for substrate recognition by KBTBD8 in cells. Wild-type FLAGNOLC1 or a mutant containing only a single CK2 motif (FLAGNOLC1CK1) were affinity-purified from 293 T cells and analyzed for binding to endogenous KBTBD8 and CK2 by western blotting using specific antibodies. (B) A single CK2 motif is not sufficient for KBTBD8 and TCOF1 recognition by NOLC1 in differentiating hESCs. FLAGNOLC1 was affinity-purified from hESCs or hESCs subjected to 1 day of neural conversion (NC1), and bound endogenous KBTBD8, TCOF1, or CK2 were detected by western blotting using specific antibodies. (C) Scheme of CK2-mutants of NOLC1. CK2 motifs that remained functional are labeled with a red dot; the last two CK2 motifs of NOLC1 do not mediate KBTBD8-binding in vitro, as shown above. (D) KBTBD8-recognition of NOLC1 depends on multiple CK2 motifs. The indicated NOLC1 variants were affinity-purified from 293 T cells and analyzed for binding to endogenous KBTBD8 or CK2β by western blotting using specific antibodies. Note that wild-type NOLC1, which contains all 10 functional CK2 motifs is the most efficient KBTBD8-recruiter. (E) Multisite dependency of NOLC1 ubiquitylation in cells. Ubiquitylated NOLC1 mutants were purified under denaturing conditions from 293 T cells that expressed HISubiquitin and, where indicated, KBTBD8HA. Ubiquitylation of FLAGNOLC1 was analyzed by αFLAG-Western; note that NOLC1 mutants containing up to five CK2 motifs are ubiquitylated by an E3 ligase distinct from KBTBD8 (i.e. ubiquitylation is observed in the absence of KBTBD8 expression in these cells). Ubiquitylation of endogenous PCNA was monitored by western blotting using specific antibodies. (F) Multisite dependency of TCOF1-NOLC1 complex formation. FLAGNOLC1 variants with increasing numbers of CK2 motifs were affinity-purified from 293 T cells that expressed KBTBD8 as indicated. Complex formation with endogenous TCOF1 was monitored by western blotting using specific antibodies. In addition, overexpression of KBTBD8 allowed for KBTBD8 recognition already in the presence of five CK2 motifs, consistent with multisite dependency providing an increase in affinity (i.e. that can in part be overcome by overexpression), rather than creating a specific complex E3 recognition element.
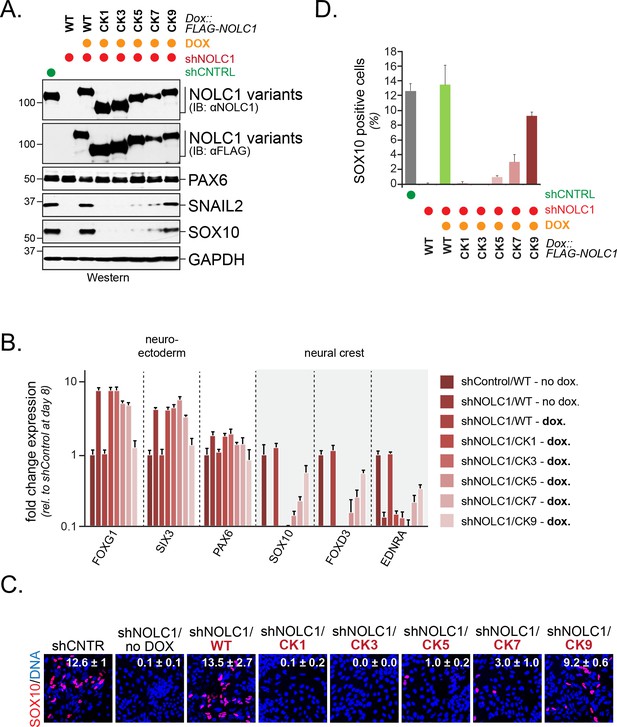
Multisite dependency of CUL3KBTBD8 during neural crest specification.
(A) Multiple CK2 motifs in NOLC1 are required for its role in neural crest specification. Control or NOLC1-depleted H1 hESCs were generated by stable expression of shRNAs. Cells were then reconstituted with doxycycline-inducible wildtype (WT) or CK-mutants of NOLC1 and subjected to neural conversion for 6 days. The outcome of cell differentiation was monitored by western blotting using specific antibodies against PAX6 (CNS precursor marker), SNAIL2 and SOX10 (neural crest markers), and GAPDH (control). (B) Multiple CK2 motifs in NOLC1 are required for its role in neural crest specification. H1 hESCs expressing wild type or NOLC1CK variants were generated and subjected to neural conversion as described above, and the outcome of differentiation was monitored using qRT-PCR of markers of the CNS (FOXG1, SIX3, PAX6) or the neural crest (SOX10, SNAIL2, FOXD3). (C) Multiple CK2 motifs in NOLC1 are required for its role in neural crest specification, as seen at single-cell resolution by fluorescence microscopy analysis. H1 hESCs expressing wild type or specific NOLC1CK mutants were generated and subjected to neural conversion as described above, and neural crest differentiation was monitored by immunofluorescence microscopy using anti-SOX10 antibodies. (D) Quantification of SOX10-positive cells in Figure 7C (n = 3, two biological and one technical replicate; error bars denote s.e.m).
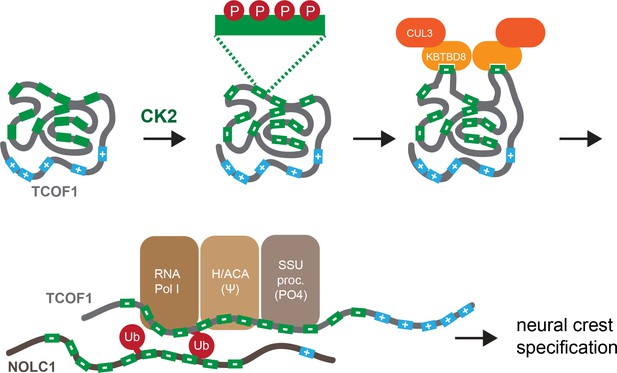
Model of multisite-dependent substrate recognition by CUL3KBTBD8.
CUL3KBTBD8-substrates TCOF1 and NOLC1 are first phosphorylated by CK2 in conserved CK2 motifs (12 motifs in TCOF1; 10 motifs in NOLC1), which increases the negative charge of the central intrinsically disordered region (IDR). Phosphorylation likely occurs on multiple Ser residues within a CK2 motif (see insert). While the phosphorylated substrate likely adopts a compact conformation, CK2 motifs are stochastically exposed, allowing for their recognition by KBTBD8. We speculate that the resulting monoubiquitylation of TCOF1 and NOLC1 opens up their conformations to allow for recognition of ribosome biogenesis factors, including RNA polymerase I, the pseudouridylation machinery, and the SSU processome. As shown previously (Werner et al., 2015), the subsequent production of new, and likely modified, ribosomes then triggers neural crest specification.
Tables
Mutations introduced in different NOLC1-CK2 mutants.
https://doi.org/10.7554/eLife.35407.014| NOLC1 CK2 mutant | Amino acids mutated to R |
|---|---|
| CK1 | CK2 motif 1: D86, S91 CK2 motif 2: S128, S133 CK2 motif 3: D168, D172, E176 CK2 motif 4: S227, S230 CK2 motif 5: S266, S271 CK2 motif 6: S322, S326, S332 CK2 motif 7: S362, S366, S370 CK2 motif 8: S425, S432 CK2 motif 9: S470, S47 |
| CK3 | CK2 motif 3: D168, D172, E176 CK2 motif 4: S227, S230 CK2 motif 5: S266, S271 CK2 motif 6: S322, S326, S332 CK2 motif 7: S362, S366, S370 CK2 motif 8: S425, S432 CK2 motif 9: S470, S477 |
| CK5 | CK2 motif 5: S266, S271 CK2 motif 6: S322, S326, S332 CK2 motif 7: S362, S366, S370 CK2 motif 8: S425, S432 CK2 motif 9: S470, S477 |
| CK7 | CK2 motif 7: S362, S366, S370 CK2 motif 8: S425, S432 CK2 motif 9: S470, S477 |
| CK9 | CK2 motif 9: S470, S477 |
TCOF1 and NOLC1 fragments cloned with N-terminal His6x-FLAG-tag in pET28a.
https://doi.org/10.7554/eLife.35407.015| Fragment | Amino acids | Sequence |
|---|---|---|
| TCOF1-F1 | 72–108 | QAKKTRVSDPI STSESSEEEEEAEAE TAKATPRLAST |
| TCOF1-F2 | 189–225 | GMVSAGQ DSSSEDTSSSSDETDVE GKPSVKPAQVKA |
| TCOF1-F3 | 260–298 | AKRAKKPEEESESSEEGSESEEE APAGTRSQVKASEKIL |
| TCOF1-F4 | 329–364 | QTKAGKP EEDSESSSEESSDSEEE TPAAKALLQAKA |
| TCOF1-F5 | 399–431 | AKAQAGKR EEDSQSSSEESDSEEE APAQAKPSG |
| TCOF1-F6 | 466–499 | VQVGKQ EEDSRSSSEESDSDRE ALAAMNAAQVKP |
| TCOF1-F7 | 536–571 | AQVGKW EEDSESSSEESSDSSDGE VPTAVAPAQEKS |
| TCOF1-F8 | 599–638 | VKAEKPMDNSESSEESSDSADSEEAPAAMTAAQAKPALKI |
| TCOF1-F9 | 683–718 | AGGTQRPA EDSSSSEESDSEEE KTGLAVTVGQAKSV |
| TCOF1-F10 | 754–783 | VKAEKQ EDSESSEEESDSEE AAASPAQVKT |
| TCOF1-F11 | 864–893 | P EEDSGSSEEESDSEEEAE TLAQVKPSGKT |
| TCOF1-F12 | 924–954 | AQAGKQ DDSGSSSEESDSDGE APAAVTSAQV |
| NOLC1-F1 | 76–105 | AKKAKKKA SSSDSEDSSEEEEE VQGPPAKK |
| NOLC1-F2 | 118–149 | GKAAAKA SESSSSEESSDDDDEED QKKQPVQK |
| NOLC1-F4 | 207–242 | APKIANGKAA SSSSSSSSSSSSDDSEEE KAAATPKK |
| NOLC1-F5 | 255–283 | VKAATTPTRK SSSSEDSSSDEEEE QKKPM |
| NOLC1-F6 | 313–346 | AVEKQQPV ESSEDSSDESDSSSEEE KKPPTKAVV |
| NOLC1-F7 | 350–384 | TTKPPPAKKAA ESSSDSSDSDSSEDDE APSKPAGT |
| NOLC1-F8 | 415–448 | QKLLTRKA DSSSSEEESSSSEEE KTKKMVATTKP |
| NOLC1-F9 | 460–494 | AKQAPQGSR DSSSDSDSSSSEEEEE KTSKSAVKKK |
| NOLC1-F10 | 510–544 | AKKGKA ESSNSSSSDDSSEEEEE KLKGKGSPRPQA |
| NOLC1-F11 | 552–586 | ALTAQNGKAAKN SEEEEEE KKKAAVVVSKSGSLKK |
| NOLC1-F12 | 619–648 | EKRASSPFRRVR EEEIEVDS RVADNSFDAK |
| NOLC1-F2 WT | 196–212 | DSSSEDTSSSSDETDVE |
| NOLC1-F2 S1A | 196–212 | DASSEDTSSSSDETDVE |
| NOLC1-F2 S3A | 196–212 | DAAAEDTSSSSDETDVE |
| NOLC1-F2 S7A | 196–212 | DAAAEDTAAAADETDVE |
| NOLC1-F2 S1R | 196–212 | DSRSEDTSSSSDETDVE |
| NOLC1-F2 S2R | 196–212 | DSRSEDTSSRSDETDVE |
| TCOF1* | 190–364 | Contains CK2 motif 2–3 |
| NOLC1* | 76–187 | Contains CK2 motif 2–3 |
| TCOF1-R1/2 | 72–225 | Contains CK2 motif 1–2 |
| TCOF1-R1/3 | 72–298 | Contains CK2 motif 1–3 |
| TCOF1-R1/4 | 72–364 | Contains CK2 motif 1–4 |
| TCOF1-R1/6 | 72–499 | Contains CK2 motif 1–6 |
| TCOF1-R1/8 | 72–638 | Contains CK2 motif 1–8 |
| TCOF1-R3/8 | 260–638 | Contains CK2 motif 3–8 |
| TCOF1-R5/8 | 399–638 | Contains CK2 motif 5–8 |
| TCOF1-R7/8 | 536–638 | Contains CK2 motif 7–8 |
qPCR primers used in this study
https://doi.org/10.7554/eLife.35407.016| qPCR primer | Sequence |
|---|---|
| RPS6 right | TGA TGT CCG CCA GTA TGT TG |
| RPS6 left | TCT TGG TAC GCT GCT TCT TC |
| TFAP2 right | ATT GAC CTA CAG TGC CCA GC |
| TFAP2 left | ATG CTT TGG AAA TTG ACG GA |
| PAX6 right | CAC ATG AAC AGT CAG CCA ATG |
| PAX6 left | GGC CAG TAT TGA GAC ATA TCA GG |
| EDNRA right | CAT GAC TTG TGA GAT GTT GAA CAG |
| EDNRA left | CTG TTT TTG CCA CTT CTC GAC |
| FOXD3 right | TTG ACG AAG CAG TCG TTG AG |
| FOXD3 left | TCT GCG AGT TCA TCA GCA AC |
| SIX3 right | CGA GGG GAG TGG ACA CTT |
| SIX3 left | ATG CCG CTC GGT CCA AT |
| SNAI2 right | TGA CCT GTC TGC AAA TGC TC |
| SNAI2 left | CAG ACC CTG GTT GCT TCA A |
| SOX10 right | CTT TCT TGT GCT GCA TAC GG |
| SOX10 left | AGC TCA GCA AGA CGC TGG |
| FOXG1 right | TGA ATG AAA TGG CAA AGC AG |
| FOXG1 left | TGC AAT GTG GGG AGA ATA CA |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.35407.017





