DNA sequence encodes the position of DNA supercoils
Figures
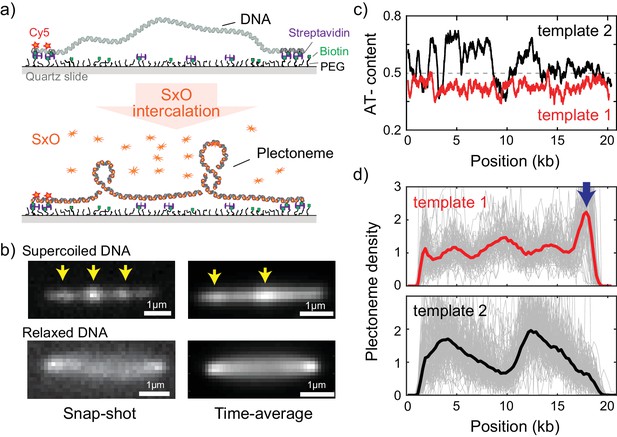
Direct visualization of individual plectonemes on supercoiled DNA.
(a) Schematic of the ISD assay. (top) A flow-stretched DNA is doubly-tethered on a PEG-coated surface via streptavidin-biotin linkage. One-end of the DNA is labeled with Cy5-fluorophores (red stars) for identifying the direction of each DNA molecule. (bottom) Binding of SxO fluorophores induces supercoiling to the torsionally constrained DNA molecule. (b) Representative fluorescence images of a supercoiled DNA molecule. Left: Snap-shot image of a supercoiled DNA with 100 ms exposure. Yellow arrows highlight higher DNA density, that is individual plectonemes. Right: Time-averaged DNA image by stacking 1000 images (of 100 ms exposure each). Arrows indicate peaks in the inhomogeneous average density of plectonemes. (c) AT-contents of two DNA samples: template1 and template2 binned to 300 bp. (d) Plectoneme densities obtained from individual DNA molecules. (top) Plectoneme density on template1 (grey thin lines, n = 70) and their ensemble average (red line). Arrow indicates a strong plectoneme pinning site. (bottom) Plectoneme densities obtained from individual DNA molecules of template2 (grey thin lines, n = 120) and their ensemble average (black line).
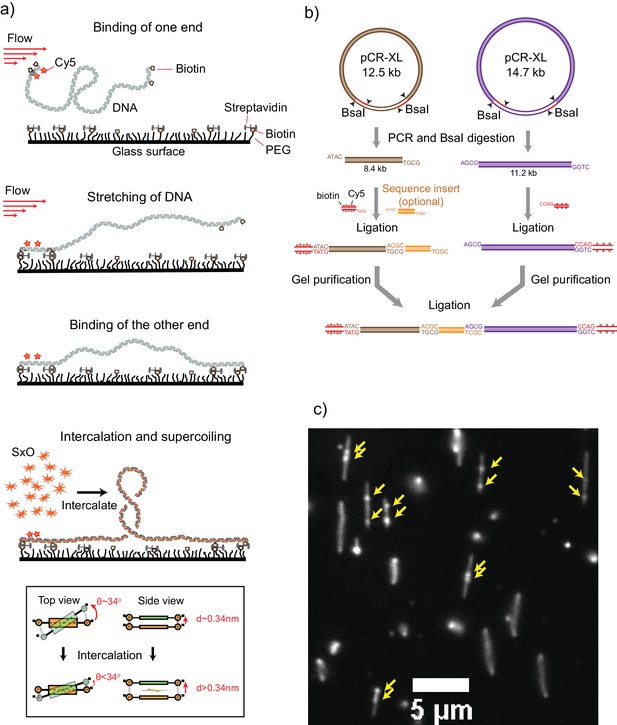
Details of the ISD assay.
(a) Intercalation induced Supercoiling of DNA- Schematics for the preparation of the intercalation-induced supercoiled DNA. A doubly biotinylated DNA at the ends is flowed along a streptavidin-coated surface at a constant flow velocity. One end of the DNA first binds to the surface via biotin-streptavidin interaction, which is followed by stretching of the molecule along the flow. The other end of the DNA then binds to surface resulting in a torsionally constrained DNA. Upon addition of an intercalating dye (Sytox Orange), DNA becomes supercoiled due to local unwinding induced by intercalation. Inset at the bottom panel, Schematics showing local unwinding of stacked base pairs due to intercalation of a dye molecule. In the B-form DNA structure, a pair (orange) of stacked bases make an angle of 34° with the next pair (green) and is separated by 0.34 nm. Intercalation of a dye molecule between the stacked bases increases the separation and decreases the angle, resulting in local unwinding of the DNA, which adds positive supercoiling to the rotationally constrained DNA molecule. (b) Schematics for DNA template preparation. Two plasmids (pCR-XL 12.5 kb and 14.7 kb) were expressed in methylation-free E. coli cells using a midiprep kit. Each plasmid contains two BsaI sites. The plasmids were first PCR-amplified and then digested with BsaI endonuclease to obtain the required sticky ends. An 8.4 kb DNA fragment was obtained from the 12.5 kb plasmid and a 11.2 kb DNA fragment from the 14.7 kb plasmid. The 8.4 kb fragment, a sequence of interest, and biotin-Cy5-DNA handle were ligated together. At the same time, the 11.2 kb fragment is ligated with biotin-DNA handle. The ligated fragments were then agarose gel-purified and ligated together. The final ligation product is purified by agarose gel electrophoresis. In the case of template 1, we skipped the sequence of interest and the 8.4 kb and 11.2 kb fragments were directly ligated together. Template two was obtained by digesting the single 21 kb plasmid and ligating with biotin handle at one end and biotin/Cy5 handle at the other end. (c) Fluorescence snap-shot showing several supercoiled DNA molecules. Yellow arrows point to the plectonemes on supercoiled DNA.
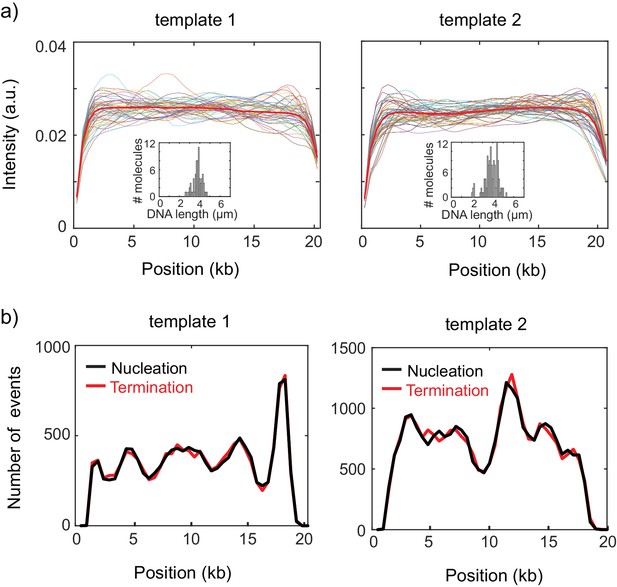
Characterization of the supercoiled and torsionally relaxed DNA.
(a) Intensity profiles of a torsionally relaxed DNA molecules. Intensity profiles of individual DNA molecules (thin lines) from template 1 (left) and template 2 (right) after nicking. Thick red lines are the ensemble average of the intensity profiles. Insets show the distribution of the end-to-end lengths of the surface immobilized DNA molecules. (b) Total number of observed plectoneme nucleation and termination events for template 1 (left) and template 2 (right). The near-equal number of nucleation and termination events at every position on both templates implies that DNA- SxO complexes are in equilibrium.
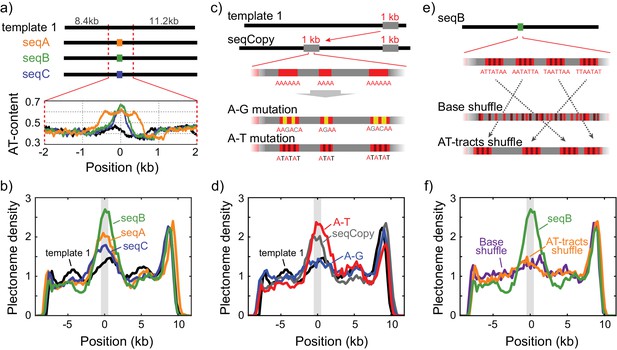
Sequence-dependent pinning of DNA plectonemes.
(a) Top: Schematics showing DNA constructs with AT-rich fragments inserted in template1. Three different AT-rich segments, SeqA (400 bp), SeqB (500 bp), and SeqC (1 kb), are inserted at 8.8 kb from Cy5-end in template1. Bottom: AT-contents of these DNA constructs zoomed in at the position of insertion. (b) Averaged plectoneme densities measured for the AT-rich fragments denoted in (A). The insertion region is highlighted with a gray box. (n = 43, 31, and 42 for SeqA, SeqB, and SeqC, respectively) (c) Schematics of DNA constructs with a copy of the 1 kb region near the right end of template1 where strong plectoneme pinning is observed (seqCopy). Poly(A)-tracts within the copied region are then mutated either by replacing A bases with G or C (A-G mutation), or with T (A-T mutation). (d) Plectoneme densities measured for the sequences denoted in (c). Plectoneme density of template1 is shown in black, seqCopy in green, A-G mutation in blue, and A-T mutation in red. (n = 45, 34, and 42 for seqCopy, A-G mutation, and A-T mutation, respectively) (e) Schematics of DNA constructs with mixed A/T stretches modified from seqB. The insert is modified either by shuffling nucleotides within the insert to destroy all the poly(A) and poly(A/T)-tracts (Base shuffle), or by re-positioning the poly(A) or poly(A/T)-tracts (AT-tracts shuffle) – both while maintaining the exact same AT content across the insert. (f) Plectoneme densities measured for the sequences denoted in (e). seqB from panel (b) is plotted in green; base shuffle data are denoted in purple; AT-tracts shuffle in orange. (n = 24, and 26 for Base shuffle, and AT-tracts shuffle, respectively).
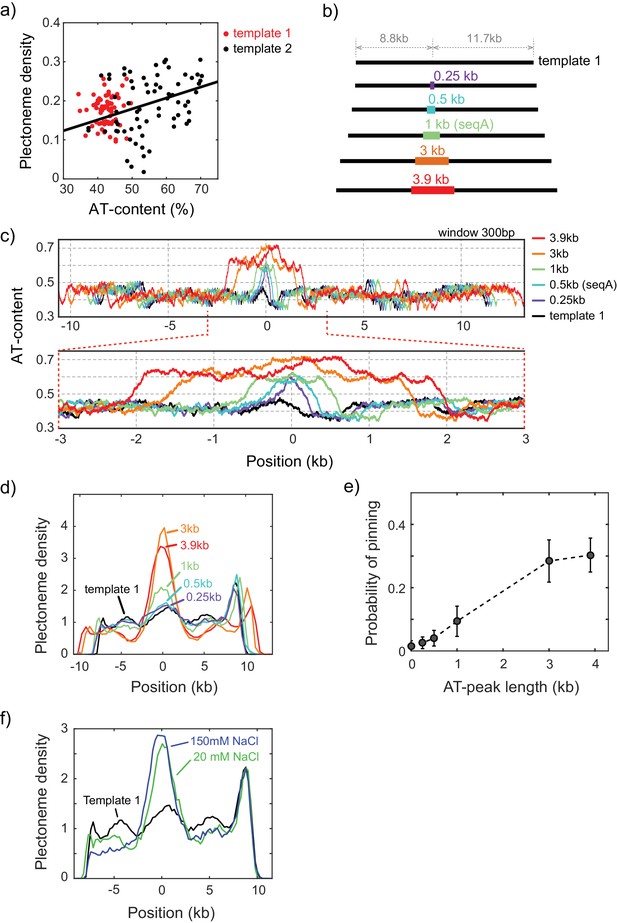
Plectoneme pinning at AT-rich inserts of various lengths.
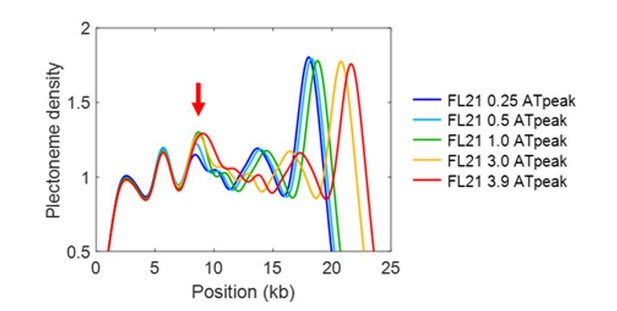
(a) Correlation between plectoneme density and AT-content. Plectoneme densities at each position in Figure 1d were plotted against local AT-contents for template 1 (red dots) and template 2 (black dots). Pearson correlation coefficient calculated for template 2 is 0.33. (b–e) Plectoneme pinning at AT-rich regions with different lengths. Schematics of DNA constructs carrying an AT-rich fragment (B) and their AT contents (C). The AT-rich fragments with different length are indicated by different colors both in the schematics and the plots. The AT percentage were calculated with 300 bp windows. (d) Plectoneme densities measured with the AT-rich fragments denoted in (b). Position 0 kb indicates the center location of the inserted AT-rich sequences. (n = 15, 21, 43, 21, and 22 for 0.25, 0.5, 1, 3, 3.9 kb, respectively). (e) Increase in the probability of plectoneme pinning plotted against the length of the AT region. The probability is calculated from the area below the center peak (with respect to the base line) in the plectoneme density curves in (d). (f) Effect of ionic strength on the plectoneme pinning. Plectoneme density measured for seqB at 150 mM NaCl (blue), as compared to the data at 20 mM NaCl (green). (n = 18 for 150 mM NaCl).
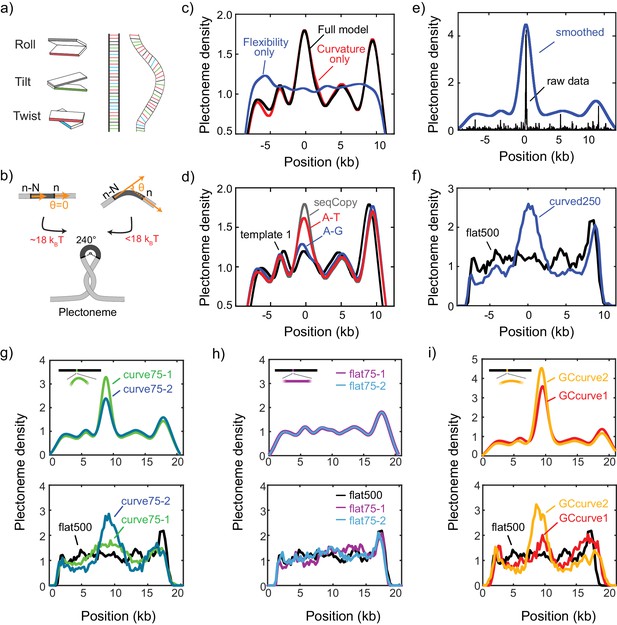
DNA plectonemes pin to sequences that exhibit local curvature.
(a) Ingredients for an intrinsic-curvature model that is strictly based on dinucleotide stacking. (Left) Cartoons showing the relative alignment between the stacked bases which are characterized by three modes: roll, tilt, and twist. (Middle) In the absence of variations in the roll, tilt, and twist, a DNA molecule adopts a strictly linear conformation in 3D space. (Right) Example of a curved free path of DNA that is determined by the slightly different values for intrinsic roll, tilt, and twist angles for every dinucleotide. (b) Schematics showing the energy required to bend a rigid elastic rod as a simple model for the tip of a DNA plectoneme. (c) Plectoneme density prediction based on intrinsic curvature and/or flexibility for seqCopy. Predicted plectoneme densities calculated based on either DNA flexibility (blue), only curvature (red), or both (black). Combining flexibility and curvature did not significantly improve the prediction comparing to that solely based on DNA curvature. (d) Predicted plectoneme densities for the DNA constructs carrying a copy of the end peak and its mutations, as in Figure 2b. Note the excellent correspondence to the experimental data in Figure 2b. (e–f) Predicted (e) and measured (f) plectoneme density of a synthetic sequence (250 bp) that is designed to strongly pin a plectoneme. Raw data from the model are shown in black and its Gaussian-smoothed (FWHM = 1600 bp) is shown in blue in the left panel. Plectoneme densities measured from individual DNA molecules carrying the synthetic sequence (thin grey lines) and their averages (thick blue line) are shown in the right panel. (n = 37, and 21 for curved250, and flat500, respectively) (g–h) Model-predicted (upper panels) and experimentally measured (bottom panels) plectoneme densities of 75 bp-long highly curved (g) and flat (h) inserts. (i) Model-predicted (upper panels) and experimentally measured (bottom panels) plectoneme densities of curved GC-rich sequences. (n = 36, 26, 21, 20, 52, and 29 for curve75-1, curve75-2, flat75-1, flat75-2, GCcurve1, and GCcurve2, respectively).
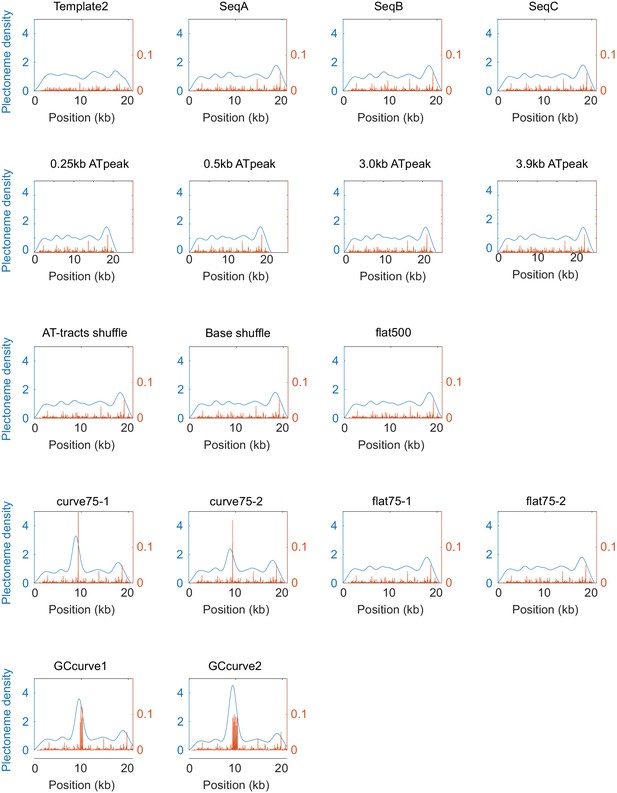
Model-predicted plectoneme density of various sequences.
Raw data from the model are shown in orange, and their Gaussian-smoothed profiles (FWHM = 1600 bp) are shown in blue.
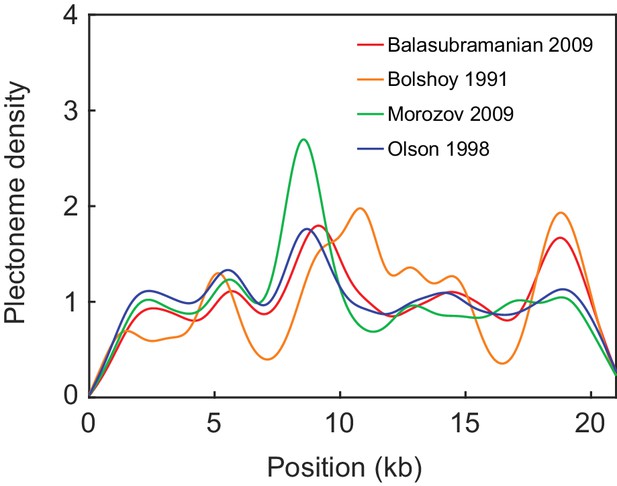
Comparison of the model predictions on seqCopy for various sets of model parameters.
The dinucleotide parameters are taken from four different sources (Balasubramanian et al., 2009; Bolshoy et al., 1991; Morozov et al., 2009; Olson et al., 1998).
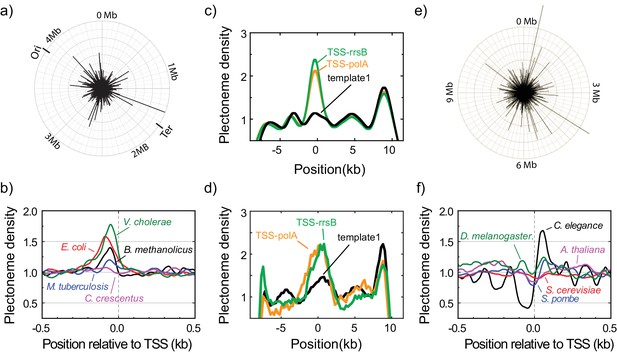
Plectonemes are enriched at prokaryotic transcription start sites.
(a) The strength of plectoneme pinning calculated for the entire E. coli genome (4,639,221 bp; NC_000913). (b) Mean predicted plectoneme densities around transcription start sites (TSS) in prokaryotic genomes. The density profiles were smoothed over a 51 bp window. (c) Model-predicted and (d) experimentally measured plectoneme densities obtained for two selected TSS sites, TSS-rrsB and TSS-polA, which are E. coli transcription start sites encoding for 16S ribosomal RNA and DNA polymerase I, respectively. For comparison to experimental data, we smoothed the predicted plectoneme densities using a Gaussian filter (FWHM = 1600 bp) that approximates our spatial resolution. (n = 26, and 17 for TSS-rrsB, and TSS-polA, respectively) (e) Strength of plectoneme pinning calculated for the entire 12.1 Mb genome (i.e. all 16 chromosomes placed in sequential order) of S. cerevisiae (NC_001134). For quantitative comparison, we kept the radius of the outer circle the same as in (a). (f) Mean predicted plectoneme densities around the most representative TSS for each gene in several eukaryotic genomes. The density profiles are smoothed over a 51 bp window.
Videos
A representative real-time fluorescence image of a supercoiled DNA molecule that shows dynamic bright spots upon plectoneme formation.
At 20 s after acquisition, the DNA was torsionally relaxed due to photo-induced nicking.
Additional files
-
Supplementary file 1
Dinucleotide parameters used in the physical model.
- https://doi.org/10.7554/eLife.36557.012
-
Supplementary file 2
DNA sequences of the inserts
- https://doi.org/10.7554/eLife.36557.013
-
Supplementary file 3
Materials used for DNA template one with and without inserts
- https://doi.org/10.7554/eLife.36557.014
-
Supplementary file 4
Materials used for DNA template 2
- https://doi.org/10.7554/eLife.36557.015
-
Transparent reporting form
- https://doi.org/10.7554/eLife.36557.016