Multivalency of NDC80 in the outer kinetochore is essential to track shortening microtubules and generate forces
Figures
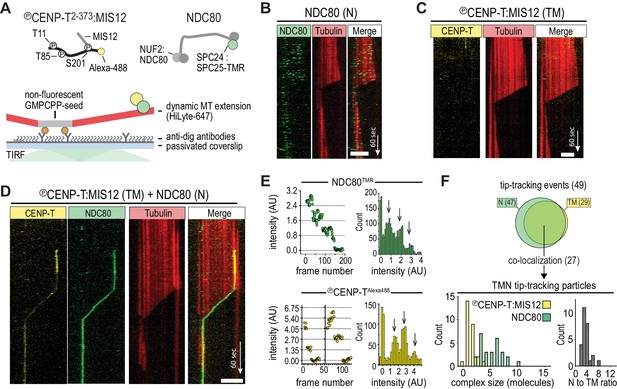
CENP-T-mediated oligomerization of NDC80 produces particles that follow microtubule disassembly.
(a) Schematic representation of Alexa-488-labeled, phosphorylated CENP-T:MIS12 and TMR-labeled NDC80 in a single-molecule TIRF setup with dynamic microtubules. (b) Monomeric NDC80 at 200 pM does not follow microtubule shortening. (c) MIS12-CENP-T at 2 nM does not bind to microtubules and does not follow microtubule shortening. (d) MIS12:CENP-TAlexa488 (2 nM, yellow) and NDC80TMR (20 pM, green) mixed in the flow-chamber, co-localize in a particle that follows the shortening end of a microtubule. (e) Left: time traces showing photobleaching of NDC80TMR (green) and CENP-TAlexa488 particles (yellow). Note that the CENP-T particle initially bleaches two fluorophores simultaneously. Right: distributions containing all bleaching traces (n = 13 for TMR and n = 20 for Alexa488). Arrows show intensities of 1, 2 and 3 fluorophores. See Materials and methods for details. (f) Euler diagram of tip-tracking particles produced by mixing NDC80 and MIS12:CENP-T (top). Distributions of initial, unbleached fluorescence intensities (expressed as the number of fluorophores) of NDC80TMR (green) and MIS12:CENP-TAlexa488 (yellow) in the tip-tracking particles containing both signals (bottom left). Black bars show ratio of NDC80 to CENP-T (bottom right). Scale bars: 5 µm (horizontal), 60 s (vertical).
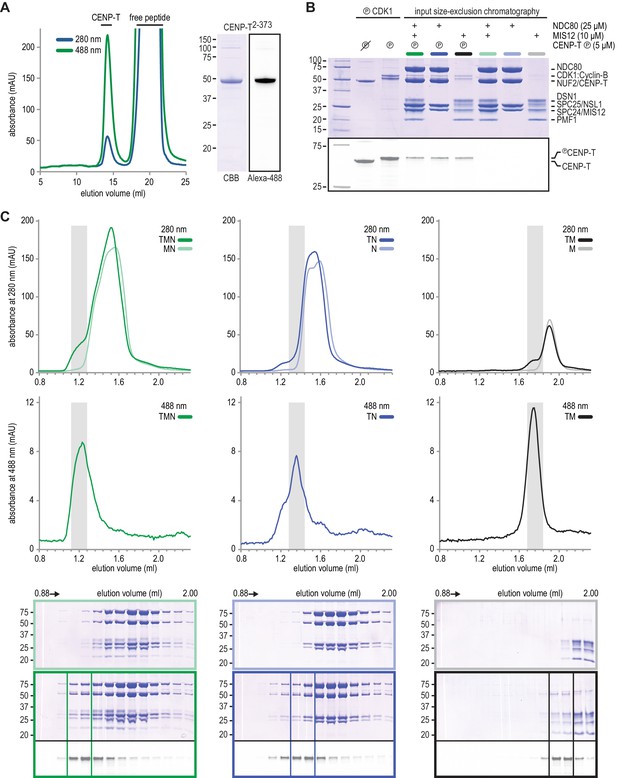
Reconstitution of NDC80 and MIS12 on CENP-TAlexa-488 phosphorylated by CDK1:Cyclin-B.
(a) CENP-T2-373 labelled with Alexa-488 was separated from the excess of GGGGC-Alexa488 peptide by size-exclusion chromatography. (b) SDS-PAGE analysis to demonstrate the phosphorylation of CENP-T by CDK1:Cyclin-B and to compare the input for size-exclusion chromatography shown in panel C. (c) MIS12 and NDC80 bind phosphorylated CENP-T. Size-exclusion chromatography traces (top) and SDS-PAGE analysis (bottom) of TMN (green), TN (blue) and TM (black). The two shaded fractions were pooled for single-molecule TIRF experiments.
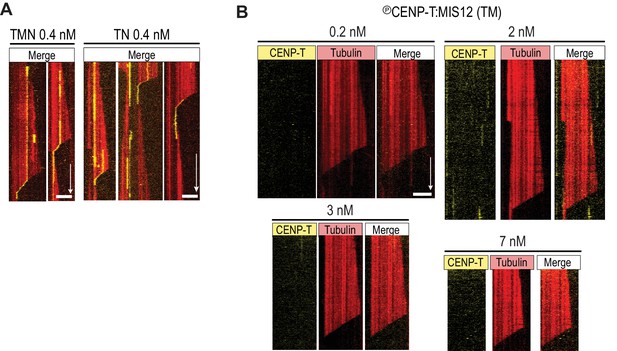
NDC80 promotes tip-tracking of reconstituted TN and TMN.
(a) TMN and TN (both at 0.4 nM) follow shortening microtubules in a single-molecule TIRF experiment. (b) TM does not bind to microtubules and does not follow microtubule shortening in a concentration range from 0.2 to 7 nM. Scale bars: 5 µm (horizontal), 60 s (vertical).
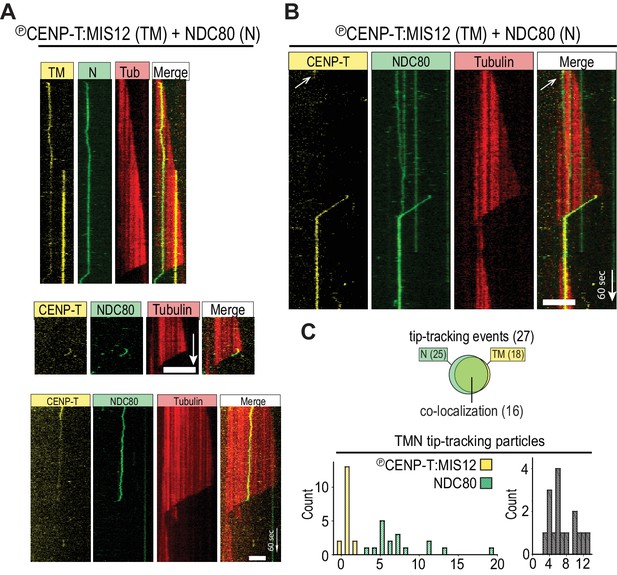
Tip-tracking by NDC80 oligomerized on CENP-T:MIS12.
(a) Two example kymographs as in Figure 1D. (b) Example kymograph of co-localizing NDC80TMR and CENP-TAlexa488:MIS12 on the microtubule lattice and at the shortening microtubule end in a flow chamber with a 1:1 N/TM (both added at 200 pM). The white arrow indicates a CENP-T bleaching event. (c) Quantification of tip-tracking events as in Figure 1F for particles containing initially both TMR and Alxa488 signal at 200 pM of both NDC80TMR and CENP-TAlexa488:MIS12. 5 µm (horizontal), 60 s (vertical).
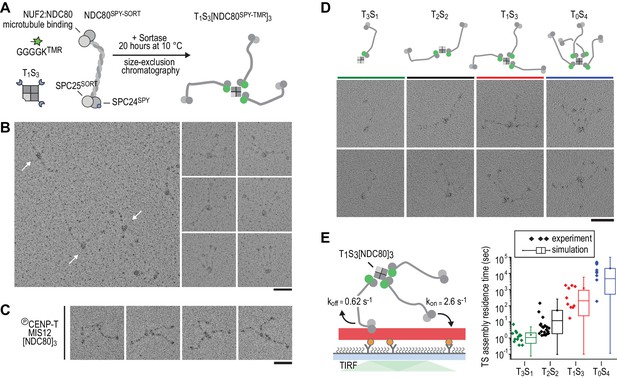
Incremental addition of NDC80 results in hyperstable microtubule binding.
(a) NDC80SPY-SORT was fluorescently labelled and covalently bound to TS assemblies. The cartoon shows the formation of T1S3[NDC80]3 assemblies. Size-exclusion chromatography and SDS-PAGE analysis are shown in Figure 1—figure supplement 1. (b) T1S3[NDC80]3 assemblies were analysed by electron microscopy after low-angle rotary shadowing. Three flexible NDC80 complexes of approximately 60 nm originate from central T1S3 densities (white arrows in the field of view) Scale bar 50 nm. (c) Representative micrographs of CENP-T:MIS12:[NDC80]3 as analysed previously (Huis In 't Veld et al., 2016). (d) Side-by-side comparison of NDC80 coupled to T3S1, T2S2, T1S3, and T0S4. Cartoons represent the approximate orientation of assemblies in the upper row of micrographs. Scale bar 50 nm. (e) Residence time of quantized NDC80 assemblies on taxol-stabilized microtubules as determined experimentally (dots) and as predicted by a series of 1000 simulations (box and whiskers plot; box: 25–75%, horizontal line: median, whiskers: 5–95%). NDC80 complexes of a microtubule-bound TS-NDC80 assembly attach to and detach from microtubules with rates of kon and koff, respectively. The residence time of an oligomer is defined as the time between the association of its first NDC80 tether and the detachment of all NDC80 tethers.
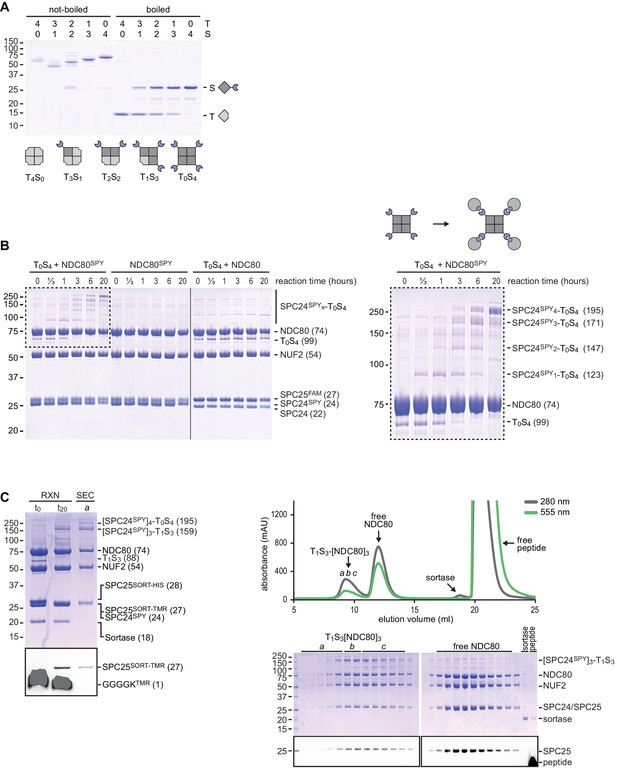
A reconstituted system to precisely control NDC80 stoichiometry (Part I).
(a) TySx variants were separated by ion-exchange chromatography based on the pI difference of T (5.1) and S (4.5). Collected assemblies were analyzed by SDS-PAGE as tetramers (not-boiled) or in a denatured form (boiled). (b) SDS-PAGE analysis (samples not-boiled) to monitor the formation of S-SPC24SPY complexes. NDC80 complexes without added T0S4 or without a SPY-tag were analysed as a control. The boxed area is shown at larger magnification on the right. (c) Samples before (t0) and after (t20) the reaction were analysed by SDS-PAGE (samples not-boiled) to monitor coupling of SPC24SPY to T1S3 tetramers and fluorescent labelling of SPC25SORT. Size-exclusion chromatography was used to separate T1S3[NDC80]3 assemblies from sortase and from the excess of unreacted NDC80 and free peptide. The lower panels show the GGGGKTMR and SPC25TMR in-gel fluorescence.
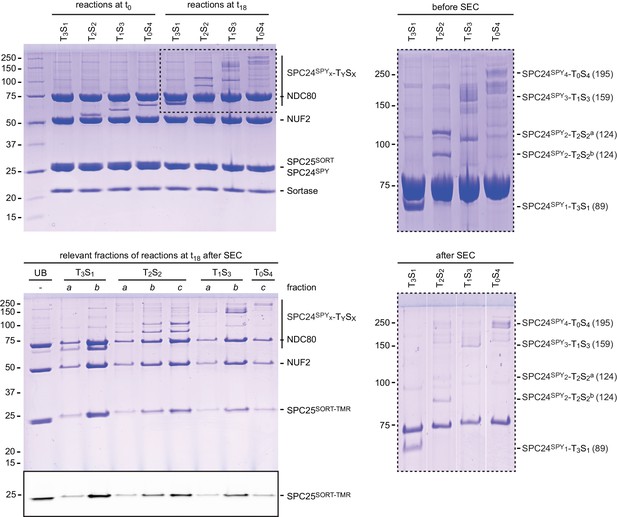
A reconstituted system to precisely control NDC80 stoichiometry (Part II).
Samples before (t0) and after (t18) the reaction were analysed by SDS-PAGE (samples not-boiled) to monitor coupling of SPC24SPY to TySx tetramers and fluorescent labelling of SPC25SORT. Size-exclusion chromatography was used to separate TySx[NDC80]x assemblies from sortase and from the excess of unreacted NDC80 and free peptide. Fractions a, b, and c, indicate the front, middle, and tail of the peak as in panel C.
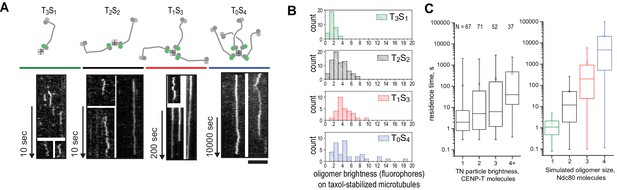
Characterization of oligomerized NDC80 on taxol-stabilized microtubules.
(a) Kymographs of mono-, di-, tri-, and tetravalent NDC80 complexes binding to taxol stabilized microtubules. Scale bar 5 µm. (b) Brightness distribution of TS-NDC80 assemblies on taxol-stabilized microtubules. (c) Residence time of co-purified CENP-T:NDC80 on taxol-stabilized microtubules, grouped by initial intensity of CENP-TAlexa488 fluorescence (left). Box: 25–75%, whiskers: min-max. Right: residence time of simulated NDC80 monomers, dimers, trimers and tetramers (see also Figure 2E). Box: 25–75%, whiskers: 5–95%.
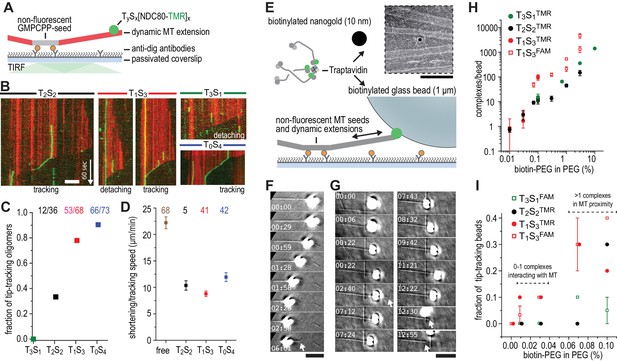
Trivalent TS-NDC80 efficiently tracks depolymerizing microtubules and transports cargo.
(a) Schematic representation of the experimental setup. (b) Kymographs showing NDC80 (green) assembled on T2S2, T1S3, or T0S4 tracking a depolymerizing microtubule (red). An example of a T1S3[NDC80]3 complex that detached from the tip of shortening microtubule is also included. Scale bar 5 µm. See Figure 3—figure supplement 1 and Video 1. (c) The fraction of NDC80 assemblies that track depolymerizing microtubules. (d) Comparison between microtubule depolymerization in the presence and absence of TS-NDC80 following the shortening tips. Data are shown as mean ± SEM. e) Biotinylated glass beads or nanogold particles can be conjugated to traptavidin in TS-NDC80C assemblies. Nanogold particles coated with T1S3[NDC80]3 bound to microtubules as observed by negative-staining EM (see also Figure 3—figure supplement 2). Scale bar 100 nm. (f–g) Examples of glass beads coated with T1S3[NDC80]3 tracking depolymerizing microtubules. The bead in panel g follows the growing microtubule after a rescue event until it detaches during a second depolymerization phase. White arrows indicate the dynamic microtubule tips. Scale bar 5 μm. (h) Fluorescence-based quantification of the number of complexes on glass beads coated with increasing amounts of PLL-PEG-biotin. (i) The fraction of beads coated with various TS-NDC80 assemblies that track depolymerizing microtubules as a function of the amount of biotin-PEG added to the beads.
-
Figure 3—source data 1
Tip-tracking events for differently coated beads.
Accompanying Figure 3I.
- https://doi.org/10.7554/eLife.36764.014
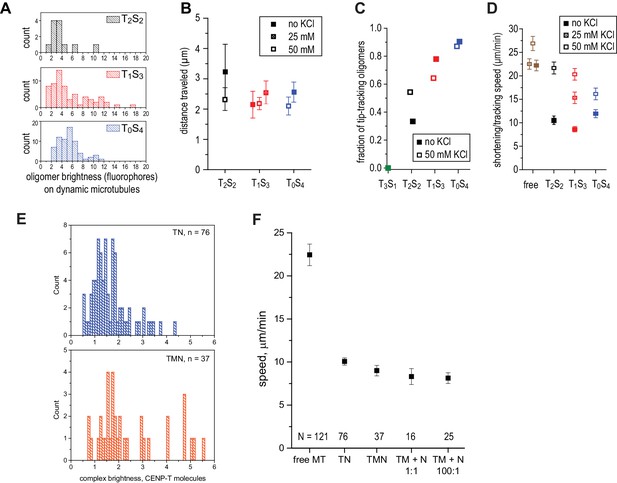
Characterization of oligomerized NDC80 on dynamic microtubules.
(a) Brightness distribution of TS-NDC80 assemblies on dynamic microtubules. b) Distance travelled by TS-NDC80 modules moving with the tips of the shortening microtubules. (c–d) Presence of 25–50 mM KCl in the motility buffer does not prevent TS-NDC80 assemblies from tip-tracking and slowing down the microtubule shortening. (e) Histograms of initial intensities of TN and TMN spots following the shortening ends of the microtubules. (f) Pre-assembled TN and TMN, as well as TM mixed with N in the flow-chamber, slow down microtubule shortening (left, see also Figure 1D,F) to the same extent as TS-NDC80 modules (see also Figure 3B,D). Data are shown as mean ± SEM.
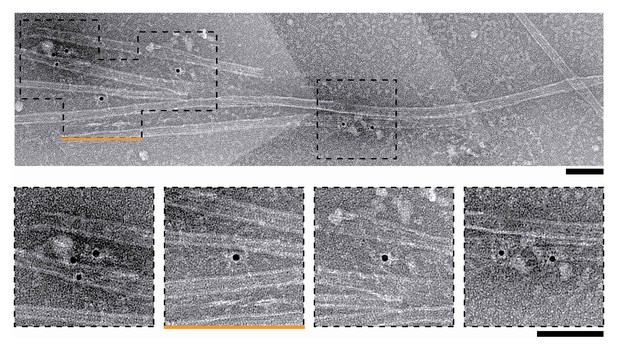
Negative-stain EM of microtubules and nanogold particles coated with T1S3[NDC80]3.
Boxed areas in the upper micrograph are shown below at a higher magnification. The orange line marks the micrograph shown in main Figure 2E. Scale bars 100 nm.
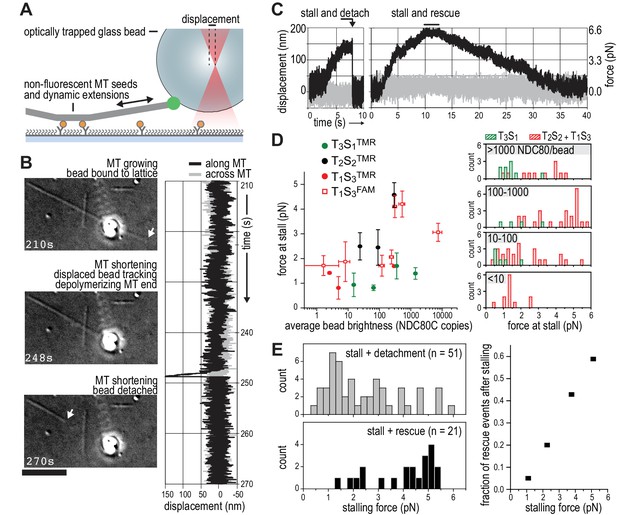
TS-NDC80 modules stall and rescue microtubule depolymerization.
(a) The displacement of an optically trapped glass bead can be used to determine the force exerted by a shortening microtubule on a bead-bound TS-NDC80 oligomer. (b) Example of a trapped glass bead that is displaced along the microtubule axis as it holds on to a depolymerizing microtubule (248 s). Arrows point to the dynamic microtubule tip before (210 s) and after (270 s) the force development. The graph on the right shows unfiltered QPD signal along and across the microtubule axis. (c) Examples of unfiltered QPD signals recorded during microtubule shortening. Stalling of microtubule depolymerization by the coupled bead in the optical trap is followed by detachment of the bead (left) or a rescue of microtubule growth (right). (d) Average forces at which differently coated beads stall shortening microtubules. Data are shown as mean ±SEM. (e) Distribution of stalling forces that were followed by bead detachment from the microtubule (grey bars) or microtubule rescue (black bars). These distributions were used to calculate the fraction of events leading to a force-induced rescue (right).
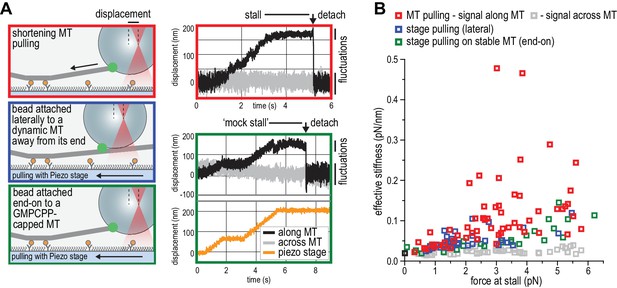
NDC80C oligomers stall microtubules through interaction with the shortening microtubule end.
(a) Experimental setup to compare forces generated by shortening microtubules (red box) with forces generating by a moving stage while a bead with T1S3[NDC80]3 is attached laterally to a dynamic microtubule (blue box), or end-on to a stabilized microtubule (green box). Examples of unfiltered QPD signals recorded during force generation are shown on the right. (b) Effective stiffness of the link between the bead and the microtubule increases with force.
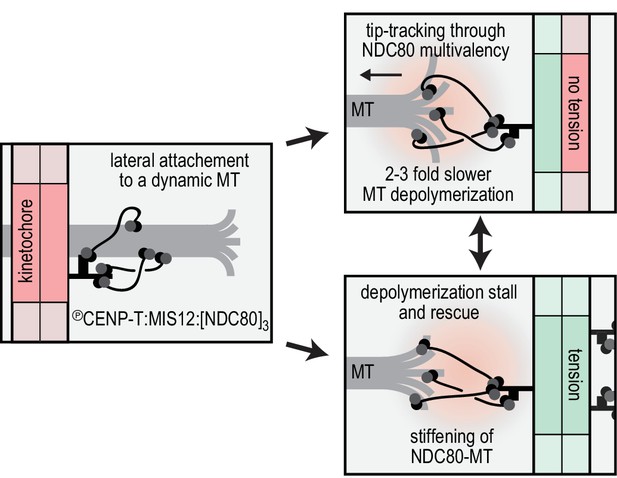
Reconstitution of a dynamic kinetochore-microtubule attachment.
A graphical recapitulation of the kinetochore-microtubule interfaces reconstituted and characterised in this study and their occurrence in vivo.
Videos
Single-molecule TIRF microscopy of TS-NDC80 modules on dynamic microtubules.
35 pM of T0S4[NDC80TMR]4 (green) in the presence of 8 µM tubulin labelled with HiLyte-642 (red) and in the absence of KCl. The two-channel images were acquired every 1.1 s (shown at 30 fps). Top left corner shows time in min:sec. White arrows mark tip-tracking events. Scale bar 5 µm.
A disassembling microtubule tip pulls on a trapped bead.
A bead coated with 3% PLL-PEG-biotin and then saturated with T2S2[NDC80TMR]2 was attached to a microtubule with a trap (see also Figure 3B for still images and a complete QPD trace of this signal). Images were acquired at eight fps in DIC, background subtracted, and each 10 consecutive frames averaged (see Materials and methods). The timer at the bottom left corner shows elapsed time in seconds. The microtubule continues growing until about 230 s, when it switches to shortening and pulls on the bead at 248 s (evident from the bead’s brief displacement in the direction of microtubule disassembly). From 265 s till 290 s the microtubule end is seen disassembling to the left of the bead. Scale bar 5 µm.
A microtubule is rescued five times at the bead attachment site.
A bead coated with 0.3% PLL-PEG-biotin and then saturated with T1S3[NDC80FAM]3 was attached to a microtubule with a trap. The microtubule experiences dynamic instability, but its shortening is five times in a row rescued at the attached bead (at 2, 9, 24, 47 and 60 min from the start of the experiment, see timer in the top left corner). Note that each rescue is preceded by a displacement of the bead along the microtubule axis in the direction of disassembly. After 63 min, the trap stiffness is increased from initial 0.03 pN/nm to 0.13 pN/nm to manually remove the bead from the microtubule. The microtubule without the attached bead depolymerizes without stalling or rescue events. Scale bar 5 µm.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Recombinant DNA reagent | pLIB | Peters laboratory,Weissmann et al., 2016 | addgene plasmid 80610 | |
| Recombinant DNA reagent | pBIG1A | Peters laboratory,Weissmann et al., 2016 | addgene plasmid 80611 | |
| Recombinant DNA reagent | pLIB NDC80 | Musacchio laboratory,Huis In 't Veld et al., 2016 | ||
| Recombinant DNA reagent | pLIB NUF2 | Musacchio laboratory,Huis In 't Veld et al., 2016 | ||
| Recombinant DNA reagent | pLIB SPC25-HIS | Musacchio laboratory,Huis In 't Veld et al., 2016 | ||
| Recombinant DNA reagent | pLIB SPC24 | Musacchio laboratory,Huis In 't Veld et al., 2016 | ||
| Recombinant DNA reagent | pBIG1A with NDC80C (SPC25-HIS) | Musacchio laboratory,Huis In 't Veld et al., 2016 | combined by biGBac cloning (Weissmann et al., 2016) | |
| Recombinant DNA reagent | pLIB SPC25-SORT-HIS | Musacchio laboratory, this study | ||
| Recombinant DNA reagent | pLIB SPC24-SPY | Musacchio laboratory, this study | ||
| Recombinant DNA reagent | pBIG1A with NDC80C (SPC25-SORT-HIS) | Musacchio laboratory, this study | combined by biGBac cloning (Weissmann et al., 2016) | |
| Recombinant DNA reagent | pBIG1A with NDC80C (SPC25-SORT-HIS SPC24-SPY) | Musacchio laboratory, this study | combined by biGBac cloning (Weissmann et al., 2016) | |
| Recombinant DNA reagent | pGEX-6P GST-CENP-T2–373 SORT | Musacchio laboratory,Huis In 't Veld et al., 2016 | ||
| Recombinant DNA reagent | pBIG1A CDK1-GST:Cyclin-B1-HIS | Musacchio laboratory,Huis In 't Veld et al., 2016 | ||
| Recombinant DNA reagent | pET21a Core Traptavidin | Howarth laboratory,Chivers et al. (2010) | addgene plasmid 26054 | |
| Recombinant DNA reagent | pET21a Dead Strepatavidin SpyCatcher | Howarth laboratory,Fairhead et al. (2014) | addgene plasmid 59547 | |
| Peptide, recombinant protein | MIS12C (DSN1-d100-109) | Musacchio laboratory,Petrovic et al. (2016) | ||
| Peptide, recombinant protein | Sortase 5M and Sortase 7M | Ploegh laboratory, seeHirakawa et al. (2015) | addgene plasmids 51140 and 51141 | |
| Peptide, recombinant protein | GGGGC-Alexa488 | ThermoFisher | peptide for C-terminal sortase labeling | |
| Peptide, recombinant protein | GGGGK-TMR | GenScript | peptide for C-terminal sortase labeling | |
| Peptide, recombinant protein | GGGGK-FAM | GenScript | peptide for C-terminal sortase labeling | |
| Software, algorithm | Kymo.m | Dogterom laboratory, this study | Matlab script to trace fluorescent particles in kymographs |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.36764.021





