T-ALL leukemia stem cell 'stemness' is epigenetically controlled by the master regulator SPI1
Figures
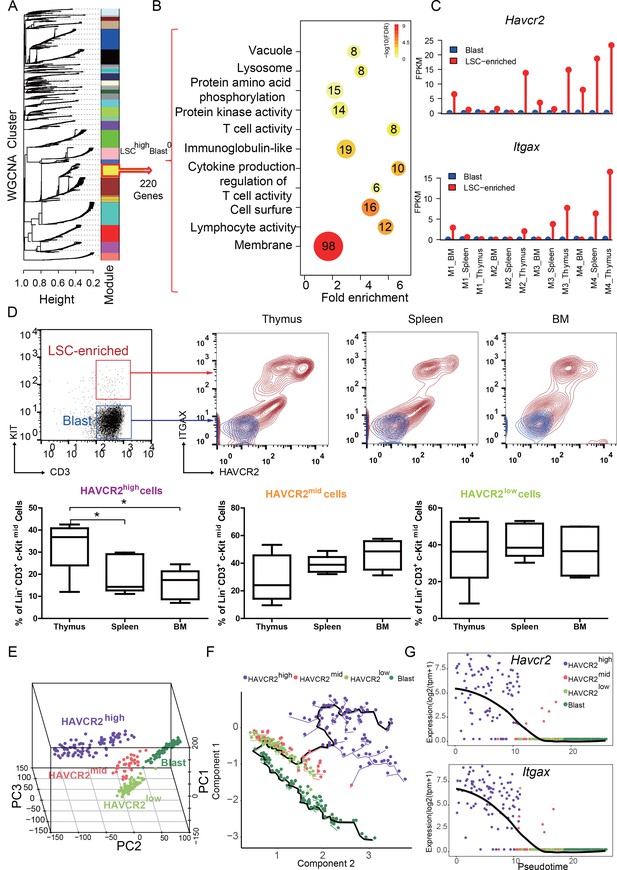
HAVCR2 redefines a heterogeneous LSC-enriched subpopulation at single-cell resolution (A) WGCNA analysis for the bulk RNA-seq of LSC-enriched and leukemic blast subpopulations.
The yellow module contains 220 genes that are preferentially expressed in the LSC-enriched subpopulation (LSChigh-Blast0); (B) Gene Ontology (GO) analysis of LSC-enriched genes in the yellow module; (C) Havcr2 and Itgax are specifically expressed in LSC-enriched (red) but not in leukemic blast (blue) subpopulations isolated from the indicated hematopoietic organs of M1-M4 Pten-null T-ALL mice; (D) Upper panel: FACS plots are overlaid to show the differential expression of HAVCR2 and ITGAX in the LSC and blast subpopulations. The previously defined Lin-CD3+KITmid LSC-enriched subpopulation (in the red box in the left panel) can be further separated into several subgroups based on the expression of the cell-surface markers HAVCR2 and ITGAX. The Lin-CD3+KIT- leukemic blast subpopulation (in the blue box in the left panel) does not express HAVCR2 or ITGAX. Lower panel: Quantitative measurement of the HAVCR2high, HAVCR2mid and HAVCR2low subgroups in different hematopoietic organs from Pten-null T-ALL mice (n = 5; *, p<0.05). The HAVCR2high subgroup is enriched in the thymus; (E) PCA analysis of the single-cell transcriptome shows four subgroups, labeled in different colors. Cells from two independent mice are indicated by different shapes; (F) Pseudotime analysis shows the expression profiles of T-ALL cells in 2-D component space. The solid black line shows the main differentiation path from HAVCR2high (purple) to blasts (dark green); (G) Pseudotemporal ordering of single cells based on Havcr2 or Itgax expression.BM: bone marrow.
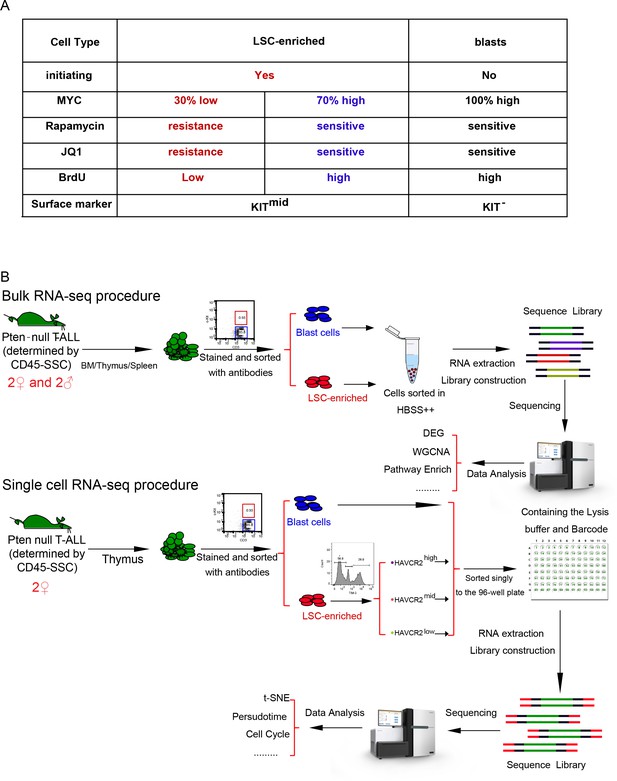
A schematic illustration of procedures used for Bulk and single cell RNAseq analysis.
(A) Heterogenous properties of LSC-enriched and leukemic blast subpopulations. (B) Schematic illustrations of the procedures used for the isolation of LSC-enriched and leukemic blast subpopulations and the bulk (upper panel) and single-cell (lower panel) RNA-seq analyses.
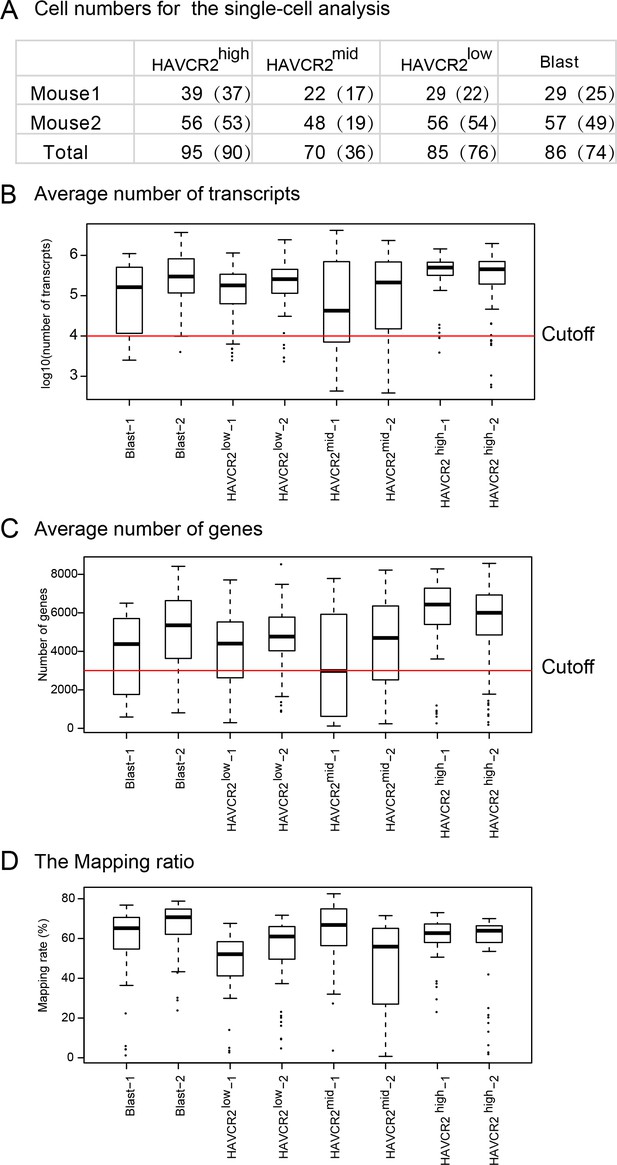
Quality control of single cell RNAseq analysis.
(A) Cell numbers used for single-cell RNA-seq. The numbers in parenthesis represent the number of cells that passed the quality control and were used for further analyses; (B–C) Boxplots of the average numbers of transcripts (B) and genes (C) detected in each subgroup; (D) Mapping ratio of the raw reads in each subgroup.
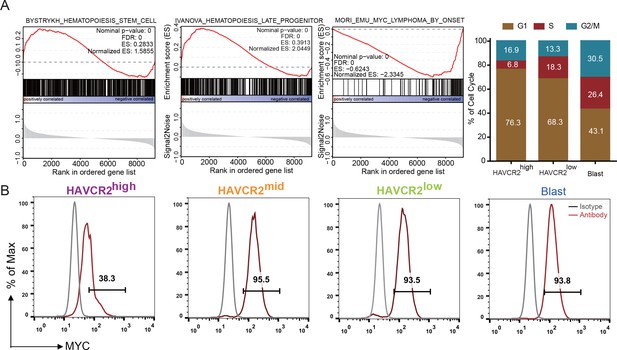
Cells in the HAVCR2high subgroup are in a quiescent cell cycle state.
(A) Left panel: GSEA analysis shows signaling pathways enriched in the HAVCR2high and blast subpopulations. Right panel: Percentage of cells in each phase of the cell cycle based on single-cell RNA-seq; (B) Intracellular FACS analyses of MYC levels in the HAVCR2high,HAVCR2mid, HAVCR2low and blast subgroups. Gray line: isotype control.
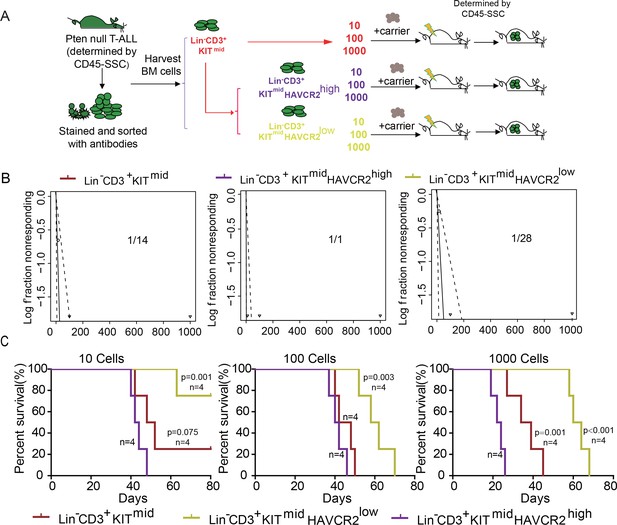
The HAVCR2high subgroup contains the vast majority of LSC activity.
(A) Schematic illustrating the cell isolation, limiting dilution and transplantation procedures used for testing LSC activity as described in Guo et al. (Guo et al., 2008); (B) LSC frequencies were calculated for each subgroup according to Hu et al. (Hu and Smyth, 2009);(C) Survival curves showing LSC activity in each of the sorted subgroups upon transplantation (n = 4). Student’s t-test was used to calculate the p-value.
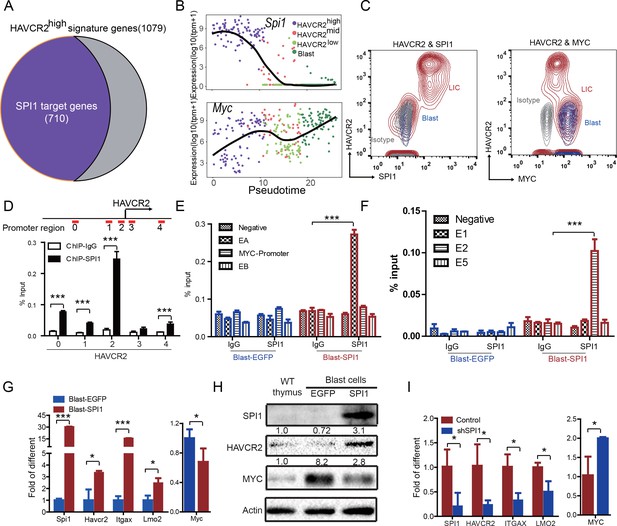
SPI1 is the master regulator of LSC signature genes and controls HAVCR2 and c-MYC expression.
(A) Nearly 70% of the genes highly expressed in the HAVCR2high subgroup—the LSC signature genes—are potential SPI1 target genes (purple);(B) Pseudotemporal ordering of single cells based on Spi1 or Myc expression; (C) FACS analysis shows the correlation of HAVCR2 cell surface expression and intracellular SPI1 and c-MYC levels in the LSC-enriched (Lin-CD3+KITmid; red) and blast (Lin-CD3+KIT-; blue) subpopulations. Gray, isotype control; (D–E) ChIP-qPCR analysis identifies SPI1 binding regions in the HAVCR2 promoter (left) and Tcra/d enhancer A(EA) region (right), using Blast-SPI1 cells; (F) ChIP analysis identifies a SPI1 binding site in the endogenous Myc enhancer;(G) q-PCR shows the fold changes in Havcr2, Itgax, Lmo2 and Myc expression between Blast-SPI1 cells (red) and Blast-EGFP cells (blue); (H) Western blotting shows the SPI1, HAVCR2 and c-Myc protein levels in WT thymus, Blast-EGFP and Blast-SPI1 cells. The fold changes relative to expression in the WT thymus are shown above each lane; (I) q-PCR analysis shows the fold changes in HAVCR2, ITGAX, LMO2 and MYC expression in control shRNA (blue) and shSPI1 knockdown human T-ALL KE-37 cells (red); (D–I) All experiments were performed at least three independent times, and the data in D, E, F, G, and I are the means ± S.Ds; *p≤0.05; **p≤0.01; ***p≤0.001.
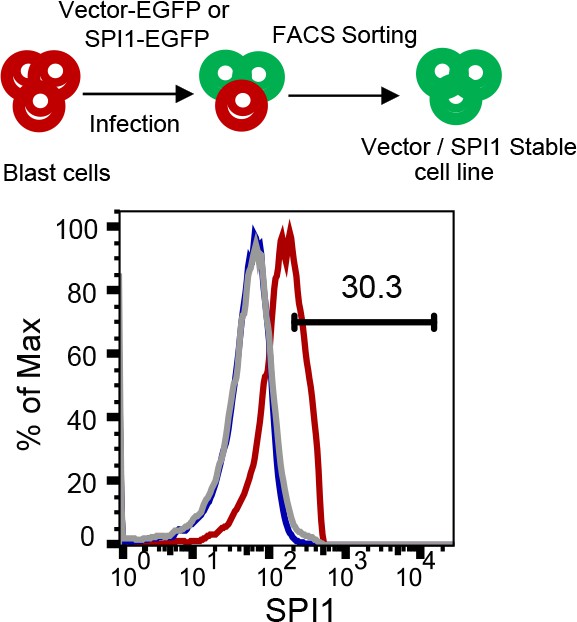
A schematic illustration of establishing lines expressing EGFP vector or EGFP-PU.1.
Upper, a schematic illustration for establishing blast lines expressing the EGFP or EGFP-Spi1 vector; lower, FACS for intracellular SPI1 expression in blast cells infected with the empty EGFP retroviral vector (Blast-EGFP; blue) or the vector with EGFP-Spi1 cDNA (Blast –SPI1; red).
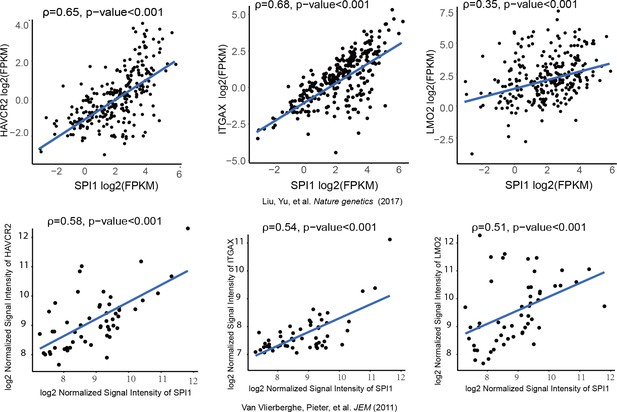
SPI1 expression is positively correlated with HAVCR2, ITGAX and LMO2 expression in human T-ALL.
Correlation of SPI1 expression with HAVCR2, ITGAX and LMO2 expression in two different cohorts of human T-ALL samples, ρ: Spearman’s rank correlation coefficient, p-value: p-value of Spearman’s rank correlation test.
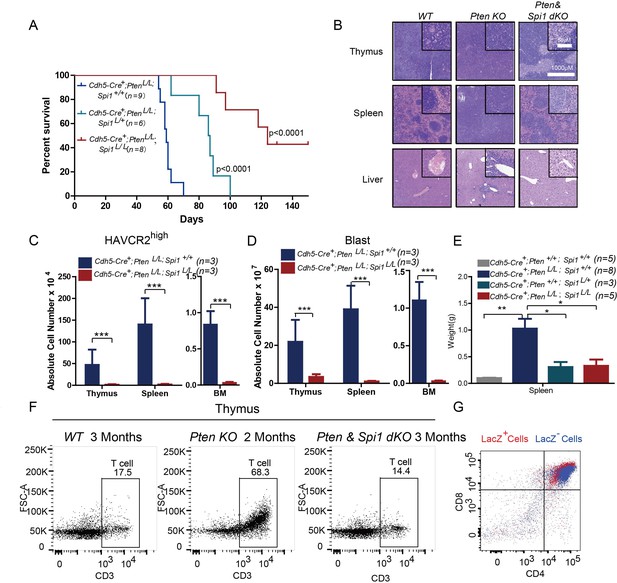
SPI1 is essential for LSC formation and T-ALL development.
(A) Survival curves for Cdh5-Cre+;PtenL/L T-ALL model mice (blue line) with heterozygous (Cdh5-Cre+;PtenL/L;Spi1L/+; green line) or homozygous (Cdh5-Cre+;PtenL/L;Spi1L/L; red line) Spi1 conditional deletion; (B) HE-stained images of thymus, spleen and liver tissue from 2-month-old mice with the indicated genotypes;(C–D) Comparison of the absolute number of HAVCR2high and blast cells in each organ in 2-month-old Cdh5-Cre+;PtenL/L (blue bars) and Cdh5-Cre+;PtenL/L;Spi11L/L (red bars) mice;(E) Comparison of spleen weights in the mice in B-C; (F) Representative FACS plots show CD3-positive T cells in the thymus of WT, Pten-null T-ALL and Pten/Spi1 double knockout mice. WT and Pten/Spi1 double knockout mice were 3 months old, and Pten-null T-ALL mice were 2 months old. n = 3; (G) FACS-Gal analysis of T cell development in the thymus of Pten/Spi1 double knockout mice. LacZ+ cells (red dots) and LacZ−cells (blue dots) from the same sample are overlaid. C-D, the data are presented as the means ± S.Ds; *p≤0.05; **p≤0.01; ***p≤0. 001.The bars in the HE images and inserts represent 1000 μM and 50 μM, respectively.
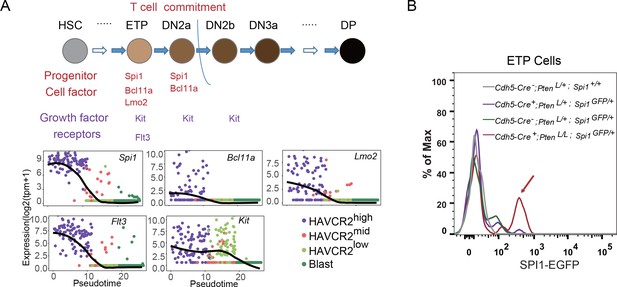
Spi1 is upregulated at the ETP/DN1 stage during T cell development.
(A) Upper panel: Diagram of progenitor cell factors and growth factor receptors involved in early T cell development, modified from (Rothenberg et al., 2016); lower panels: pseudotemporal ordering of single cells based on Spi1, Bcl11a, Lmo2, Flt3 and Kit expression; (B) Spi1-GFP expression is upregulated in ETP/DN1progenitor cells from Cdh5-Cre+;PtenL/L;Spi1GFP/+Pten null (red line), compared to that in Cdh5-Cre-;Pten+/L;Spi1+/+ WT (gray line), Cdh5-Cre+;Pten+/L;Spi1GFP/+Pten heterozygous (purple line) and Cdh5-Cre-;Pten+/L;Spi1GFP/+ WT GFP+ (green line) mice.
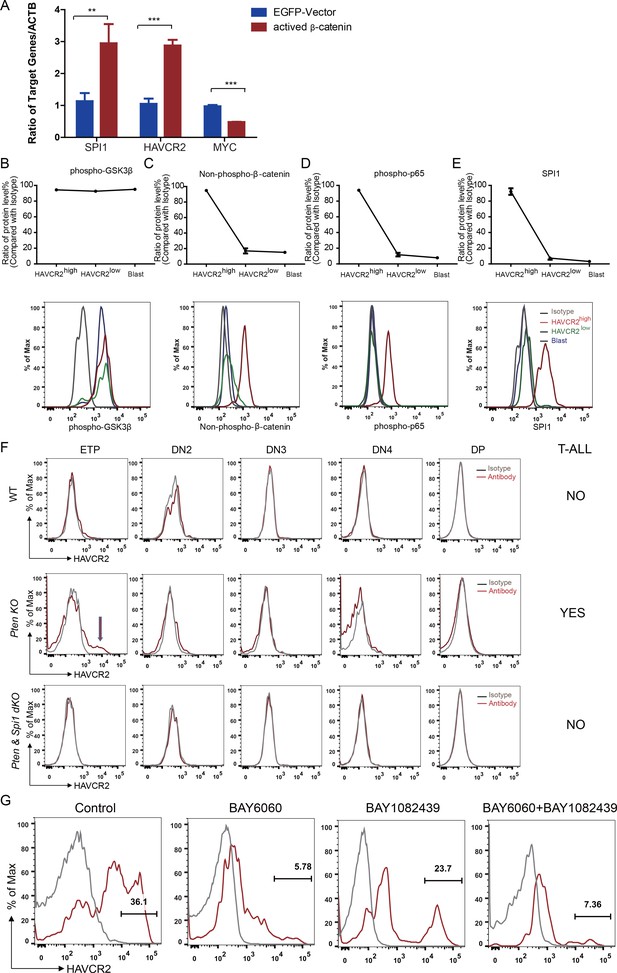
Spi1 expression is maintained by β-catenin-SPI1-HAVCR2 regulatory circuit.
(A) q-PCR analysis of SPI1 and SPI1-regulated HAVCR2 and c-MYC expressions after the overexpression of active β-catenin in the Jurkat T-ALL cell line (red bars). The data are normalized to that of empty plasmid controls (blue bars); (B–E) Upper panels: quantitative intracellular FACS analyses of P-GSK-3β, non-phospho-β-catenin, P-p65 and SPI1 levels in the HAVCR2high, HAVCR2low and blast subgroups; lower panels: representative intracellular FACS analysis of P-GSK-3β, non-phospho-β-catenin, P-p65 and SPI1 levels in the HAVCR2high, HAVCR2low and blast subgroups. Gray line, isotype control;(F) FACS analysis shows cells in the HAVCR2high subgroup at the ETP/DN1 stage, which are absent in WT and dKO mice; (G) Representative FACS plots show the number of cells in the HAVCR2high subgroup in the different drug treatment groups. The data in A, B, C, D and E are the means ± S.Ds of 3 independent tests; *p≤0.05; **p≤0.01; ***p≤0.001.
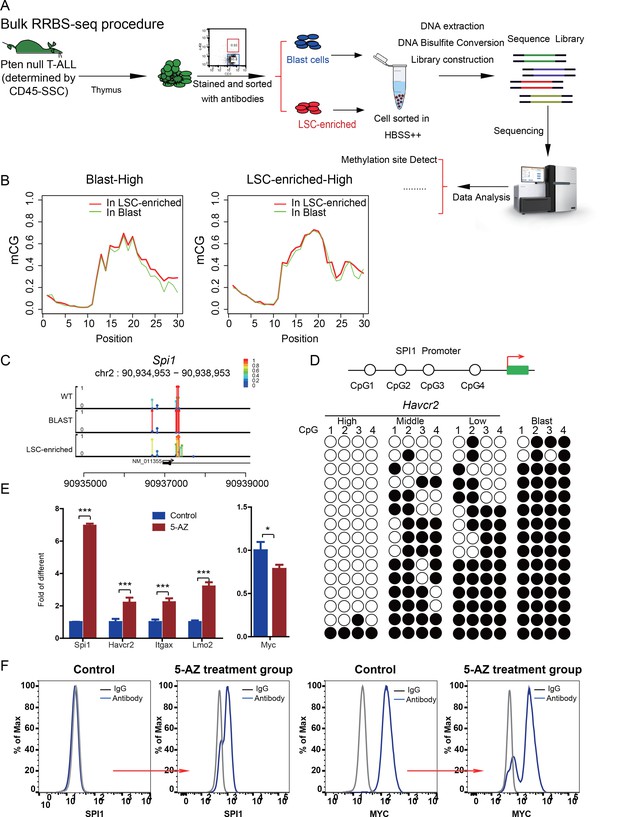
Spi1 expression is controlled by DNA methylation
(A) Schematic illustrating the procedures involved in cell isolation and RRBS analysis; (B) DNA methylation status of genes specifically expressed in the leukemic blast (left) and LSC-enriched (right) subpopulations; (C) Spi1 promoter methylation status in normal T cells, LSC-enriched cells and blast-enriched cells.
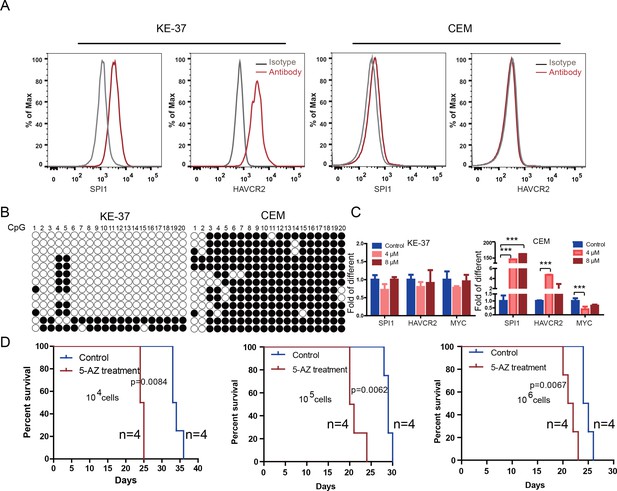
Human SPI1 expression is silenced by DNA methylation
(A) FACS analysis shows the surface expression of HAVCR2 and the intracellular level of SPI1 in the human T-ALL cell lines KE-37 and CEM.(B) Methylation status of CpG islands in the SPI1 promoter in the human T-ALL cell lines KE-37 and CEM; (C) q-PCR analysis of SPI1, HAVCR2 and Myc expression in KE-37 and CEM cells without (blue) and with(pink and red) 5-AZ treatment in vitro; (D) Survival curves show T-ALL development by CEM cells without (blue) and with (red) 5-AZ treatment upon transplantation (n = 4; t-test). The data in Care the means ± S.Ds of 3 independent tests; *p≤0.05; **p≤0.01; ***p≤0.001.
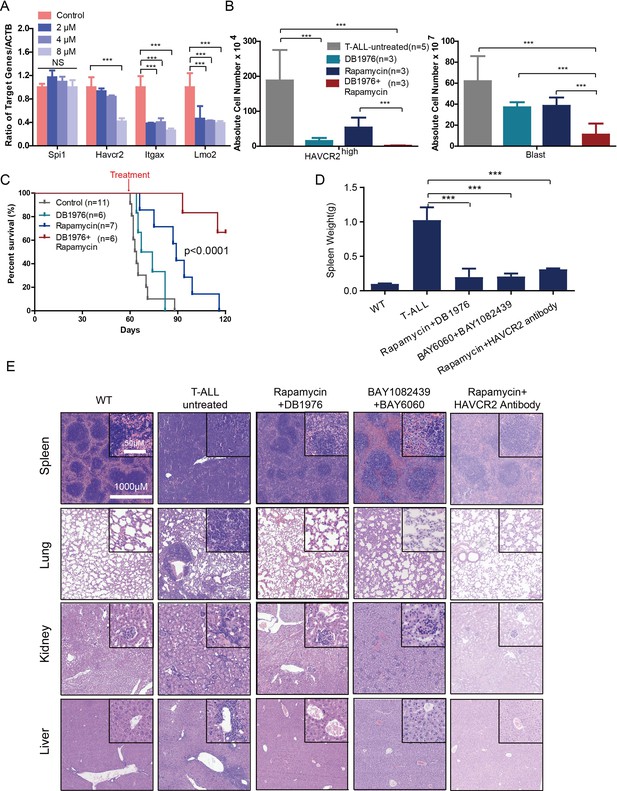
Cotargeting oncogenic driver mutations and the LSC ‘stemness’ maintenance circuit eliminated LSC and T-ALL cells
(A) q-PCR analysis of Spi1 and Spi1-regulated Havcr2, Itgax and Lmo2 expression after 24 hr of DB1976 treatment (blue bars). The data are normalized to that of the untreated controls (red bars); (B) A comparison of the absolute number of HAVCR2high and blast cells in the untreated (gray bars) and differently treated groups; (C) Survival curve of Cdh5-Cre+;PtenL/L model mice treated with DB1976 and rapamycin alone and in combination; (D) A comparison of the spleen weights of 2-month-old WT mice, untreated Cdh5-Cre+;PtenL/L mice, and combination-treated mice upon euthanasia; (E) HE-stained images of spleen, lung, kidney and liver tissue from2-month-old WT, untreated and combination-treated mice. A, B and D: the data are presented as the means ± S.Ds; ***p≤0.001; the bars in the HE images and inserts represent 1000 μM and 50 μM, respectively.
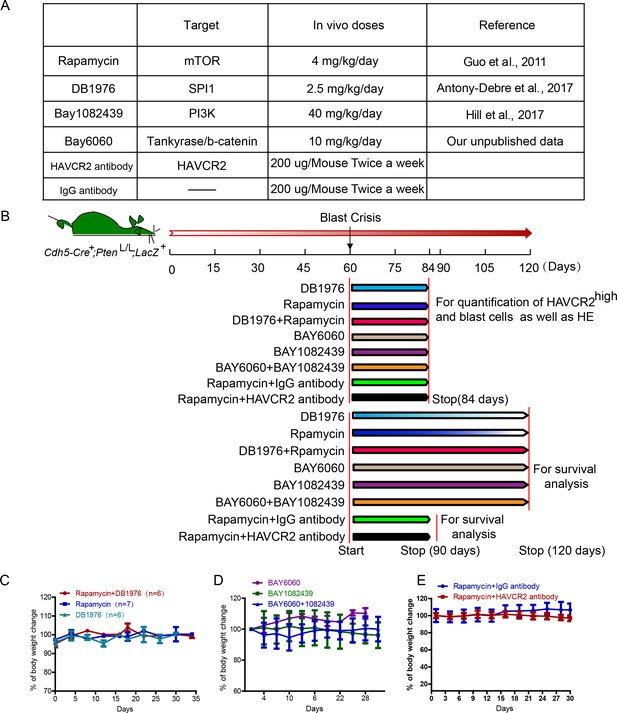
A schematic illustration of dosing schedules and treatment cohorts.
(A) Table showing all inhibitors used in this study, including their targets, in vivo dosages and references. (B) Schematic illustration of the different treatment cohorts and analyses; (C– E) Lack of general body weight loss in each treatment cohort.
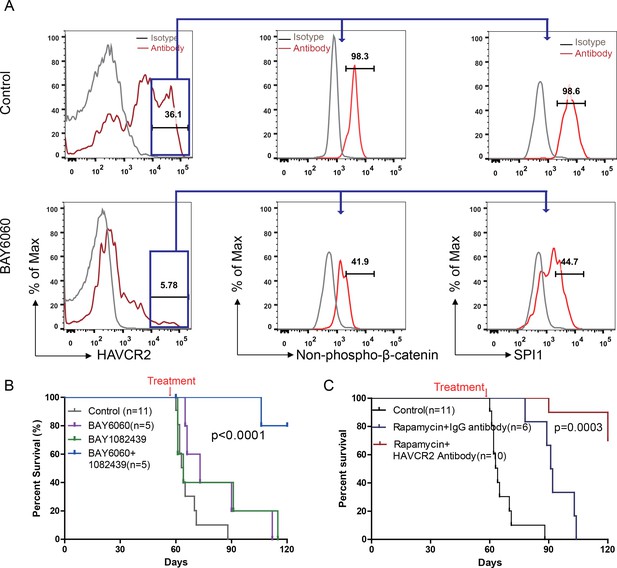
Cotargeting oncogenic driver mutations and LSC ‘stemness’ maintenance circuit.
(A) Comparison of the HAVCR2high subgroup population (left panel) and of the levels of non-phosphorylated β-catenin (middle panel) and SPI1 (right panel) within the HAVCR2high subgroup without (upper panels) and with BAY6060 treatment (low panels); (B) Survival curve for Cdh5-Cre+;PtenL/L mice treated with BAY6060 and BAY1082439 alone and in combination; (C) Survival curve for Cdh5-Cre+;PtenL/L mice treated with rapamycin in combination with either an IgG control antibody or an anti-HAVCR2 antibody.
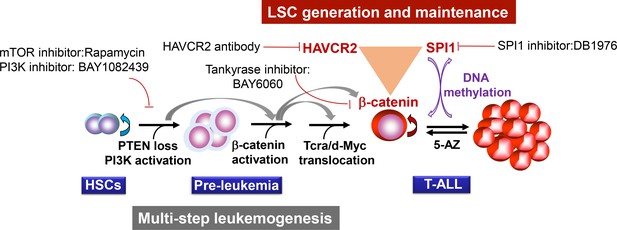
Two-layer control mechanisms for leukemogenesis and LSC maintenance.
https://doi.org/10.7554/eLife.38314.019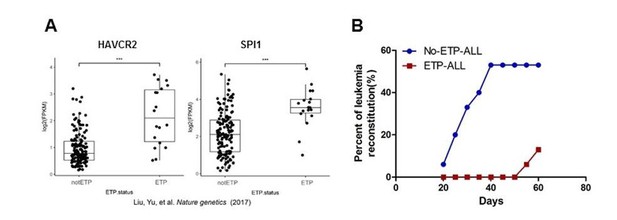
ETP and non-ETP T-ALL have different HAVCR2 and SPI1 expression levels and leukemia reconstitution abilities.
(A). The HAVCR2 and SPI1 expression levels are significant higher in ETP T-ALL samples as compared to non-ETP T-ALL samples. ***, p<0.001. (B). Bone marrow cells (1X106) from human T-ALL patients were i.v. injected into the NSG mice. Leukemia reconstitution was determined when >25% of the cells in the peripheral blood were human CD45+;CD7+ leukemic cells.
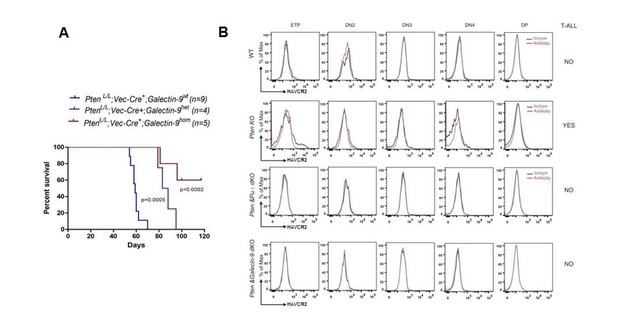
Gal-9 is essential for the Pten null T-ALL LSC formation and leukemia development.
(A). Deletion of the HAVCR2 ligand Gal-9 delays or prevents the Pten null T-ALL development in vivo. (B). HAVCR2high LSCs at ETP stage in the Pten null T-ALL are eliminated by SPI1 or Gal-9 deletion in vivo.
Tables
The biological properties of the newly defined HAVCR2high LSC subgroup in comparison to other subgroups in the Pten-null T-ALL model.
https://doi.org/10.7554/eLife.38314.006| Cell type | HAVCR2High | HAVCR2mid and HAVCR2low | Blasts |
|---|---|---|---|
| MYC | high | ||
| Rapamycin | sensitive | ||
| JQ1 | sensitive | ||
| BrdU | high | ||
| Surface marker | KIT mid | KIT - | |
| | | HAVCR2-/ITGAX- | |
| β-catenin activity | low | ||
| LIC activity | 1/104–105 | ||
| Pathway | Myc/lymphoma | ||
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Mus musculus) | Cdh5-Cre+; PtenloxP/loxP; Rosa26floxedSTOP-LacZ+ | (Guo et al., 2008) | ||
| Strain, strain background (Mus musculus) | Spi1loxP/loxP | (Dakic et al., 2005) | ||
| Strain, strain background (Mus musculus) | Spi1-GFP | (Nutt et al., 2005) | ||
| Strain, strain background (Mus musculus) | Pten/Spi1 double KO | This Paper | ||
| Cell line (Homo sapiens) | KE-37 | Deutsche Sammlung von Mikroorganismen und Zellkulturen (DSMZ) | ACC-46, RRID:CVCL_1327 | |
| Cell line (Homo sapiens) | Jurkat | (Schubbert et al., 2014) | Received Drs. G. Cheng at UCLA | |
| Cell line (Homo sapiens) | CEM | (Schubbert et al., 2014) | Received C. Radu at UCLA | |
| Cell line (Homo sapiens) | HEK 293T | American Type Culture Collection (ATCC) | CRL-3216, RRID:CVCL_0063 | |
| Cell line (Mus musculus) | HE001 | (Schubbert et al., 2014). | ||
| Antibody | TER119- APC-Cy7 | Biolegend | 116223 | |
| Antibody | B220-APC-Cy7 | Biolegend | 103224 | |
| Antibody | CD45-PE | Biolegend | 103108 | |
| Antibody | CD3-PE-Cy7 | Biolegend | 100320 | |
| Antibody | c-Kit-APC | Biolegend | 105812 | |
| Antibody | HAVCR2-PE | Biolegend | 134004 | |
| Antibody | ITGAX-FITC | Biolegend | 117306 | |
| Antibody | Mac-1-PB | Biolegend | 101224 | |
| Antibody | Gr-1-APC | Biolegend | 108412 | |
| Antibody | SPI1-PE | Biolegend | 681308 | |
| Antibody | MYC | Cell Signaling Technology | 5605S | |
| Antibody | Nonphospho (active)-β- catenin | Cell Signaling Technology | 70034 s | |
| Antibody | rabbit IgG | Cell Signaling Technology | 3900 s | |
| Antibody | Phospho- NF-κB p65 | Cell Signaling Technology | 3033 s | |
| Antibody | SPI1 | Cell Signaling Technology | 2258 s | |
| Antibody | Phospho- GSK-3β | Cell Signaling Technology | 5558 s | |
| Antibody | HAVCR2 | Abcam | ab185703 | |
| Antibody | HAVCR2 | BioxCell | RMT3-23 | |
| Antibody | IgG | BioxCell | 2A3 | |
| Antibody | Fluorescein (FITC) AffiniPure Fab Fragment Donkey Anti-Rabbit IgG (H + L) | Jackson Immuno Research | 711-097-003 | |
| Chemical compound, drug | Rapamycin | LC laboratories | R-5000 | |
| Chemical compound, drug | DB1976 | (Belver and Ferrando, 2016) | ||
| Chemical compound, drug | 5-AZ | Selleck | S1782 | |
| Chemical compound, drug | BAY10 82439 | (Hill et al., 2017) | Provided by Bayer Pharmaceuticals | |
| Chemical compound, drug | BAY 6060 | This paper | Provided by Bayer Pharmaceuticals |
Additional files
-
Supplementary File 1
Primers used for genotyping in this study.
- https://doi.org/10.7554/eLife.38314.020
-
Supplementary File 2
Primers used for q-PCR in this study.
- https://doi.org/10.7554/eLife.38314.021
-
Supplementary File 3
List of FPKM for Bulk RNA-seq in T-ALL and genes list of the yellow module reported in the Figure 1A.
- https://doi.org/10.7554/eLife.38314.022
-
Supplementary File 4
The genes list of the HAVCR2high signature genes (1079) and SPI1 target genes (710) reported in the Figure 4A.
- https://doi.org/10.7554/eLife.38314.023
-
Transparent reporting form
- https://doi.org/10.7554/eLife.38314.024





