Mycobacterium tuberculosis induces decelerated bioenergetic metabolism in human macrophages
Figures
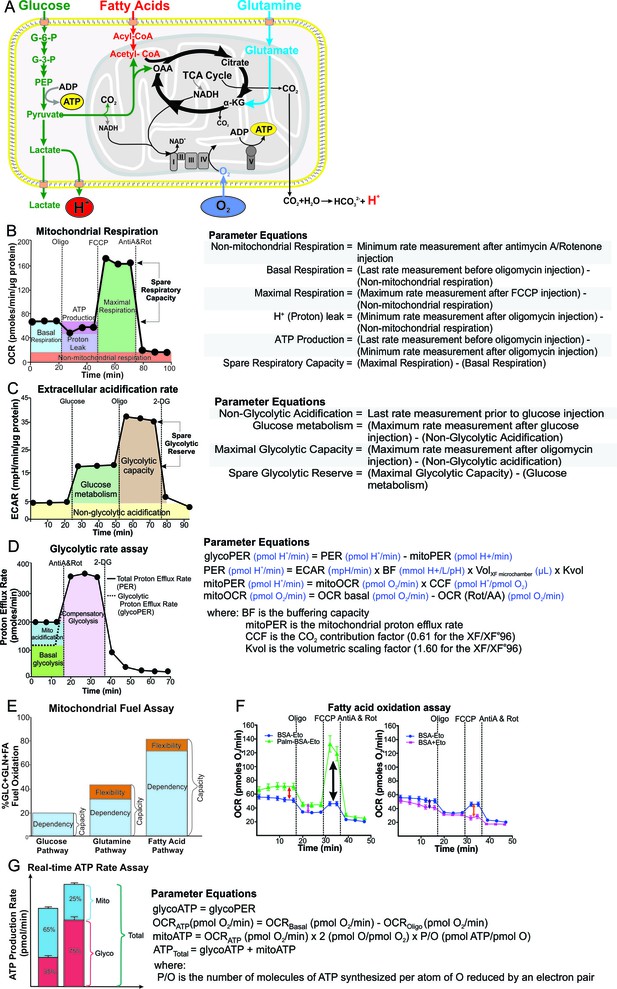
Schematic illustration of cellular metabolism pathways and XF assays used to analyze metabolic pathways.
(A) The XF measures oxygen consumption rate (OCR) of the cell, which is mostly consumed at complex IV of the electron transport chain (ETC) in the mitochondria, and extracellular acidification rate (ECAR), which is generated from lactic acid produced from pyruvate, the end-product of glycolysis, and carbonic acid produced from CO2 released during the TCA cycle. Assays performed on the XF include: (B) mitochondrial respiration test, (C) extracellular acidification test, (D) glycolytic rate assay, (E) mitochondrial fuel test, (F) fatty acid oxidation assay and (G) real-time ATP rate assay. Oligo, oligomycin; FCCP, cyanide-4-[trifluoromethoxy]phenylhydrazone; AntiA and Rot, antimycin A and rotenone; 2-DG, 2-Deoxyglucose; G-6-P, glucose-6-phosphate; G-3-P, glyceraldehyde-3-phosphate; PEP, phosphoenolpyruvate; α-KG, α-ketoglutarate; OAA, oxaloacetate.
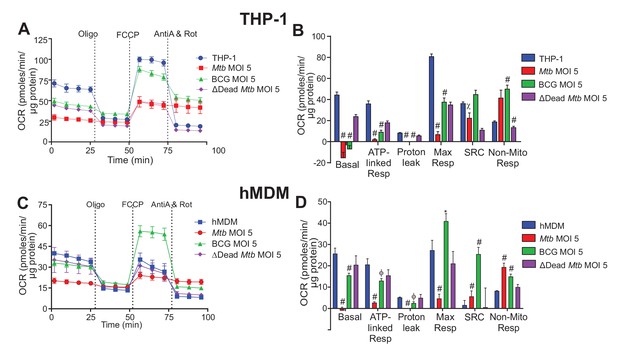
Respiratory profiles and parameters of infected macrophages are dependent on cell type, mycobacterial strain and MOI.
Respiratory profiles (OCR) and respiratory parameters of (A–B) PMA differentiated THP-1 macrophages, and (C–D) hMDMs infected with Mtb, BCG and ∆Dead Mtb (heat-killed Mtb) at MOIs of 5 for 24 h. Refer to Figure 2—figure supplement 1 for profiles of lower MOIs. After obtaining basal respiration, cells were subjected to oligomycin (Oligo, 1.5 µM), which inhibits ATP synthase and demonstrates the mitochondrial ATP-linked OCR, followed by FCCP (cyanide-4-[trifluoromethoxy]phenylhydrazone), which uncouples mitochondrial respiration and maximizes OCR (1 µM for THP-1 and hMDMs), and finally antimycin A and rotenone (AntiA and Rot), which inhibit complex III and I in the ETC, respectively, and shut down respiration (0.5 µM of each for THP-1; 2.5 µM of each for hMDMs). Profiles and respiratory parameters are representative of three independent experiments. Data shown are the mean ± SD (n = 6 biological replicates). Student’s t test relative to uninfected cells; #, p < 0.0001; χ, p < 0.0005; ϕ, p < 0.001; *p < 0.005; +, p < 0.05.
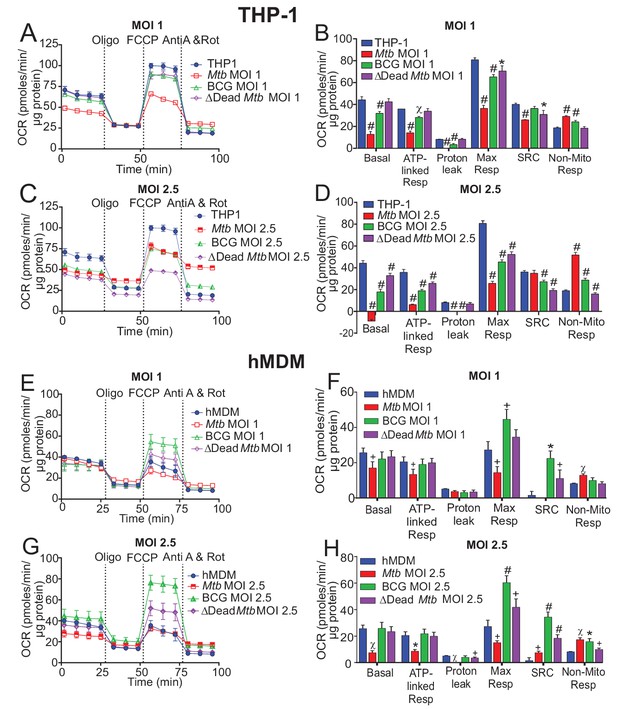
Respiratory profiles and parameters of infected macrophages are dependent on cell type, mycobacterial strain and MOI.
Respiratory profiles (OCR) and respiratory parameters of (A–D) PMA differentiated THP-1 macrophages, and (E–H) hMDMs infected with Mtb, BCG and dead Mtb at MOIs of 1 or 2.5 for 24 h. Profiles and respiratory parameters are representative of three independent experiments. Data shown are the mean ± SD (n = 6 biological replicates). Student’s t test relative to uninfected cells; #, p < 0.0001; χ, p < 0.0005; ϕ, p < 0.001; *p < 0.005; +, p < 0.05.
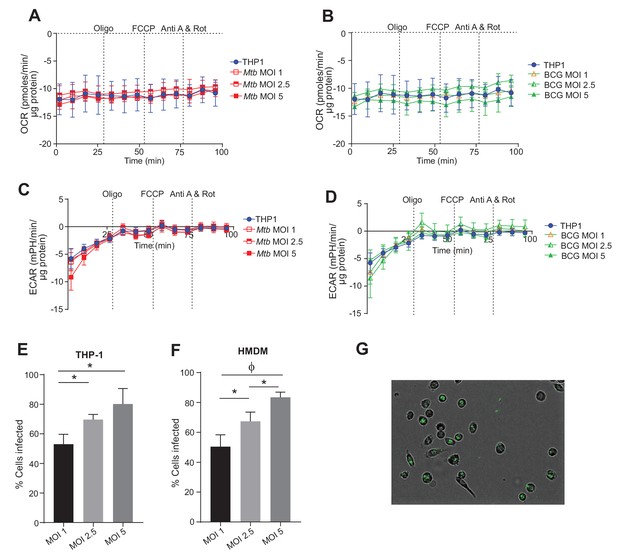
Contribution of extracellular mycobacteria to OCR and ECAR of infected macrophages and percentage of Mtb-infected macrophages.
OCR (A, B) and ECAR (C, D) of the remaining extracellular Mtb (A, C) and BCG (B,D) in the final wash of infected THP-1 macrophages that were adhered to wells of an XF96 culture plate using Cell-TakTM prior to a mitochondrial respiration assay. The percentage of THP-1 (E) and hMDM cells (F) that were infected with Mtb GFP-reporter strain at MOI 1, 2.5 or 5 after 16 h. Data shown are the mean ± SD (n = 5 biological replicates) Student’s t test, ϕ, p < 0.001; *p < 0.005.
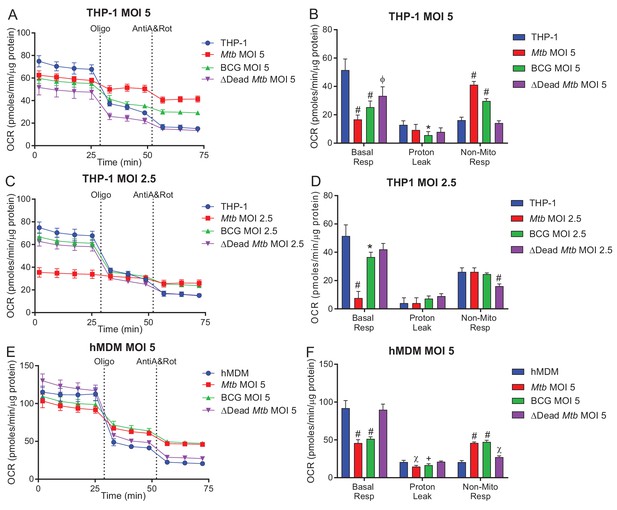
Mitochondrial respiration assays without FCCP.
Mitochondrial respiratory profiles of THP-1 cells (A–D) and hMDMs (E, F) without the addition of FCCP were generated to measure the non-mitochondrial respiration. The non-mitochondrial respiration was then used to calculate the basal respiration and proton leak in scenarios when the addition of FCCP induces oxidative bursts in infected cells at a high MOI. Profiles and respiratory parameters are representative of three independent experiments. Data shown are the mean ± SD (n = 6 biological replicates). Student’s t test relative to uninfected cells; #, p < 0.0001; χ, p < 0.0005; ϕ, p < 0.001; *p < 0.005; +, p < 0.05.
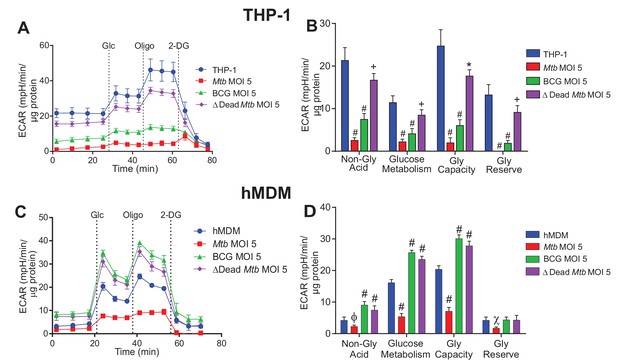
Extracellular acidification profiles and glycolytic parameters of THP-1 and hMDMs are affected by macrophage type, mycobacterial strain and MOI.
ECAR profiles and glycolytic parameters of (A–B) PMA differentiated THP-1 macrophages, and (C–D) hMDMs infected with Mtb, BCG and dead Mtb at MOI of 5 for 24 h. Refer to Figure 3—figure supplement 1 for profiles at lower MOIs. After obtaining non-glycolytic acidification, glucose (Glc, 10 mM) was added to the cells, followed by oligomycin (1.5 µM), which inhibits ATP synthase inducing maximal glycolysis to compensate for loss of mitochondrial generated ATP, and finally 2-deoxyglucose (2-DG, 100 mM) to inhibit glycolysis and demonstrate that the prior acidification was generated by glycolysis. Profiles and glycolytic parameters are representative of three independent experiments. Data shown are the mean ± SD (n = 6 biological replicates). Student’s t test relative to uninfected cells; #, p < 0.0001; χ, p < 0.0005; ϕ, p < 0.001; *p < 0.005; +, p < 0.05.
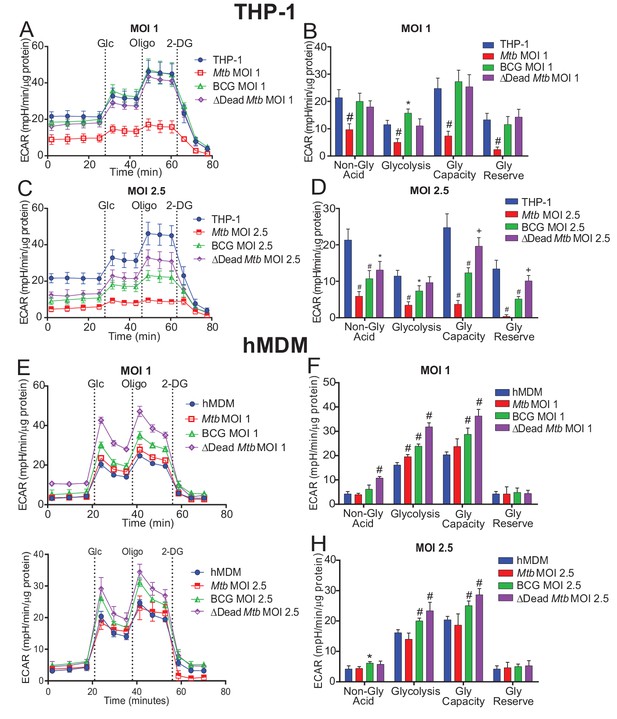
Extracellular acidification profiles and glycolytic parameters of THP-1 and hMDMs are affected by macrophage type, mycobacterial strain and MOI.
ECAR profiles and glycolytic parameters of (A–D) PMA differentiated THP-1 macrophages, and (E–H) hMDMs infected with Mtb, BCG and dead Mtb at MOIs of 1 or 2.5 for 24 h. Profiles and glycolytic parameters are representative of three independent experiments. Data shown are the mean ± SD (n = 6 biological replicates). Student’s t test relative to uninfected cells; #, p < 0.0001; χ, p < 0.0005; ϕ, p < 0.001; *p < 0.005; +, p < 0.05.
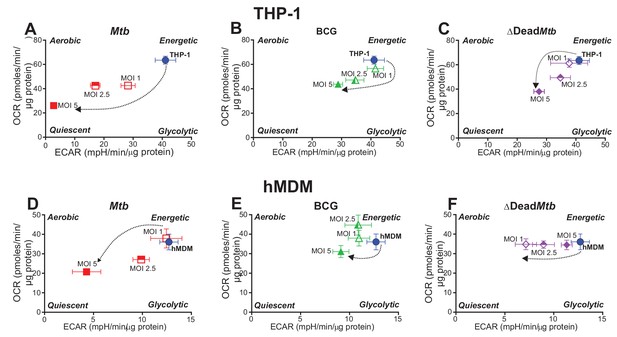
Phenograms demonstrate that increasing MOI of Mtb shifts macrophages towards quiescent energy phenotypes.
Basal OCR and ECAR measurements from the respiratory assay (Figure 2) before addition of oligomycin were plotted to generate phenograms of (A–C) PMA-differentiated THP-1 cells and (D–F) hMDMs infected with Mtb, BCG and ∆Dead Mtb at MOIs of 1, 2.5 and 5. Data are representative of three independent experiments. Data shown are the mean ± SD (n = 6 biological replicates). Student’s t test relative to uninfected cells; #, p < 0.0001; χ, p < 0.0005; ϕ, p < 0.001; *p < 0.005; +, p < 0.05.
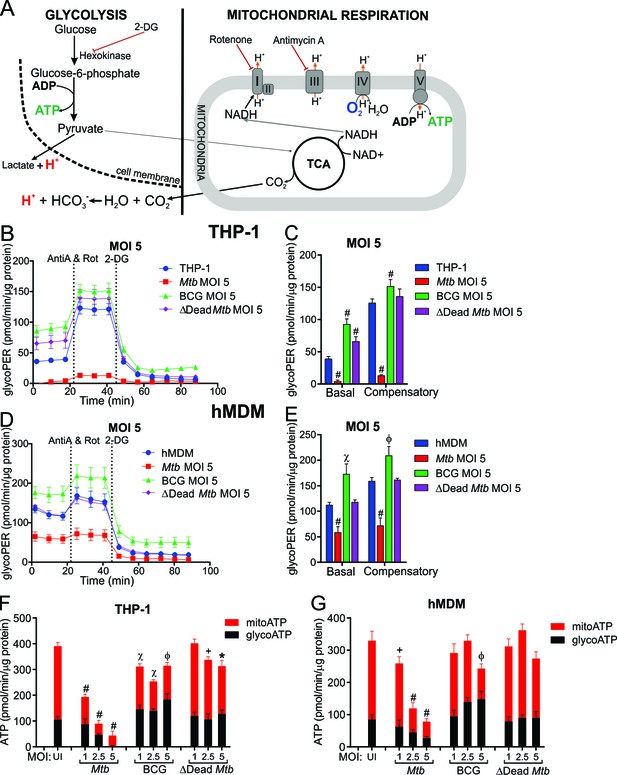
Mtb infection reduces the glycolytic proton efflux rate of macrophages.
(A) Extracellular acidification can be caused by both lactate and protons produced from pyruvate, the final product of glycolysis, in addition to carbonic acid generated from CO2 from pyruvate oxidation in the mitochondria. Calculating proton efflux rate (PER) enables the glycolytic PER to be elucidated separately from the mitochondrial PER (Figure 5—figure supplement 1A–D). (B–E) Basal and compensatory glycolytic PER of THP-1 cells (B–C) and hMDMs (D–E) infected with Mtb, BCG and ∆Dead Mtb at MOI of 5 for 18 h. Refer to Figure 5—figure supplement 1E–L for profiles at lower MOIs. Following basal measurement of ECAR and OCR, to determine basal glycolytic PER, rotenone and antimycin A were added to determine compensatory PER. This was followed by addition of 2-DG to ensure that the PER observed was caused by glycolysis. Profiles and PER are representative of two independent experiments. Data shown are the mean ± SD (n = 6). Student’s t test relative to the uninfected cells; #, p < 0.0001; χ, p < 0.0005; ϕ, p < 0.001; *p < 0.005; +, p < 0.05. (F–G) Total rate of ATP production was calculated as the sum of glycolytic ATP rate formation (equivalent to glycoPER) and mitochondrial-derived ATP rate formation that was estimated from the ATP-linked OCR, assuming a P/O ratio of 2.79. Rate of ATP formation in (F) THP-1 cells and (G) hMDM cells infected with Mtb, BCG or ∆Dead Mtb at indicated MOI for 18 h. Refer to Figure 5—figure supplement 1M–N for % contribution of glycolysis and OXPHOS to the total rate of ATP production. Error bars are SD (n = 6 biological replicates). Student’s t test; #, p < 0.0001; χ, p < 0.0005; ϕ, p < 0.001; *p < 0.005; +, p < 0.05.
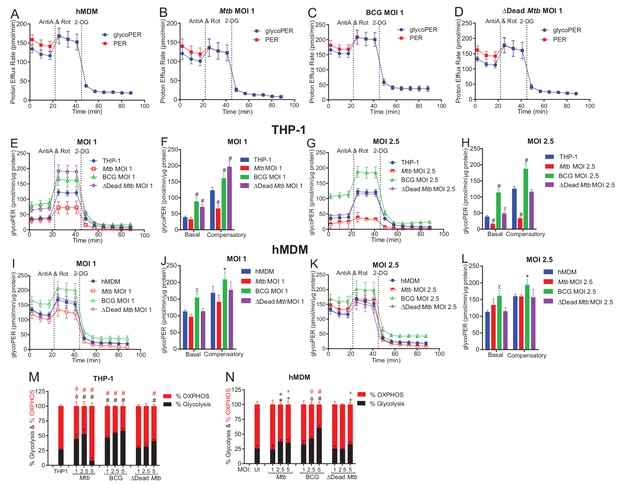
Mtb infection reduces the glycolytic proton efflux rate of macrophages.
(A–D) Profiles of total proton efflux rate (PER) in red and the glycolytic proton efflux rate (glycoPER) in blue of hMDMs infected with Mtb, BCG and dead Mtb at MOI of 1 for 18 h. The difference between the total PER and glycoPER will give the PER caused by mitochondrial respiration. (E–L) Basal and compensatory glycolytic PER of THP-1 cells (E–H) and hMDMs (I–L) infected with Mtb, BCG and dead-Mtb at MOIs of 1 and 2.5 for 18 h. (M, N) % Contribution of glycolysis and OXPHOS to the total rate of ATP production in (M) THP-1 cells and (N) hMDM cells infected with Mtb, BCG or dead Mtb at indicated MOI for 18 h. Profiles, PER and % contribution of glycolysis and OXPHOS to total ATP production are representative of two independent experiments. Data shown are the mean ± SD (n = 6 biological replicates). Student’s t test relative to the uninfected cells; #, p < 0.0001; χ, p < 0.0005; ϕ, p < 0.001; *p < 0.005; +, p < 0.05.
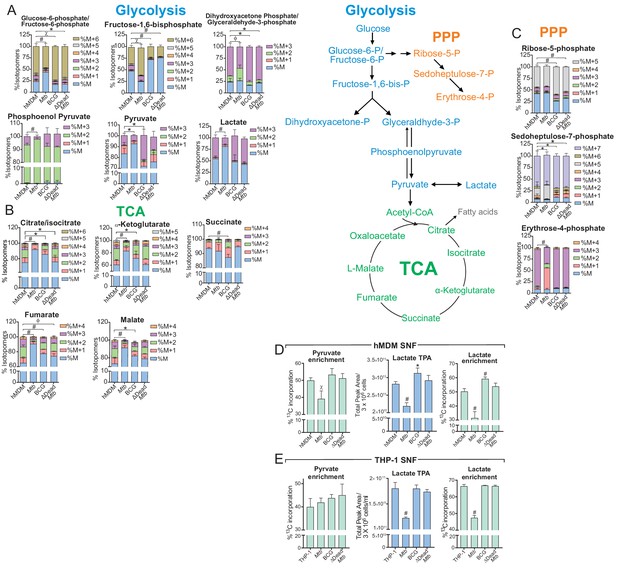
Mtb infection decelerates flux through glycolysis and the TCA cycle of the macrophage.
(A–C) 13C-tracing of metabolites extracted from hMDM cells infected with Mtb, BCG and ∆Dead Mtb at MOI 5 (hMDM) for 10 h followed by incubation with [U-13C]glucose for 8 h. The stacked mass isotopomer distributions of the intracellular metabolites depict the contribution of glucose to (A) glycolysis, (B) the tricarboxylic acid (TCA) cycle and (C) the pentose phosphate pathway (PPP). Figure 6—figure supplement 1 demonstrates the isotopomer distributions of the intracellular metabolites of THP-1 cells infected at a MOI of 2.5. (D–E) 13C enrichment of pyruvate and lactate, including total peak areas of lactate in the supernatant of the (D) hMDM cells and (E) THP-1 cells used for metabolite analysis. Data are representative of two independent experiments and shown as the mean ± SD (n = 6 biological replicates). Student’s t test relative to uninfected cells; #, p < 0.0001; χ, p < 0.0005; ϕ, p < 0.001; *p < 0.005; +, p < 0.05.
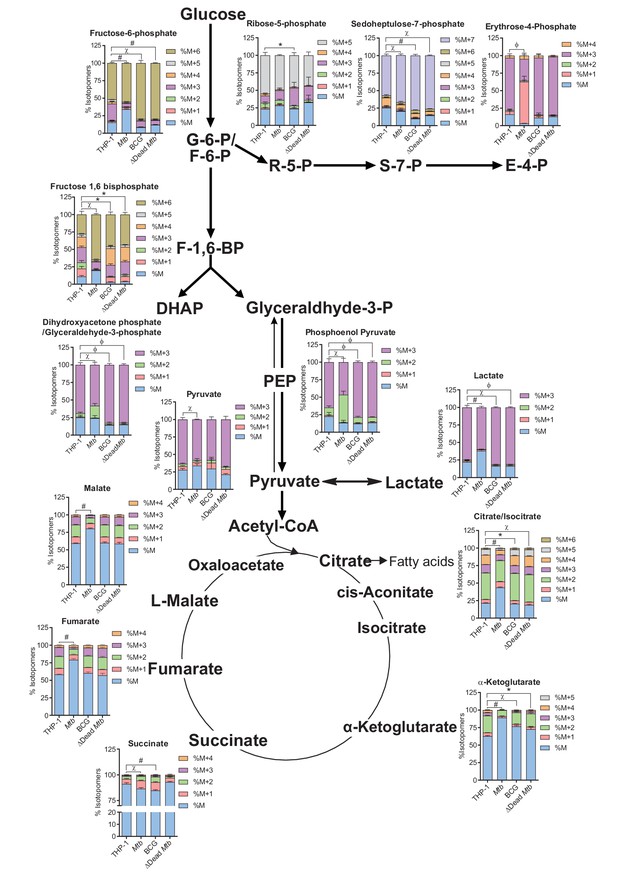
Mtb at a MOI of 2.5 decelerates flux through glycolysis in THP-1 cells and induces breaks in the TCA cycle at citrate and succinate.
13C-tracing of extracted metabolites of THP-1 cells infected with Mtb, BCG and ∆Dead Mtb at a MOI of 2.5 for 10 h followed by incubation with [U-13C] glucose for 8 h. The stacked mass isotopomer distributions of the intracellular metabolites depict the contribution of glucose to the TCA cycle and the PPP and some amino acids associated with the TCA cycle.
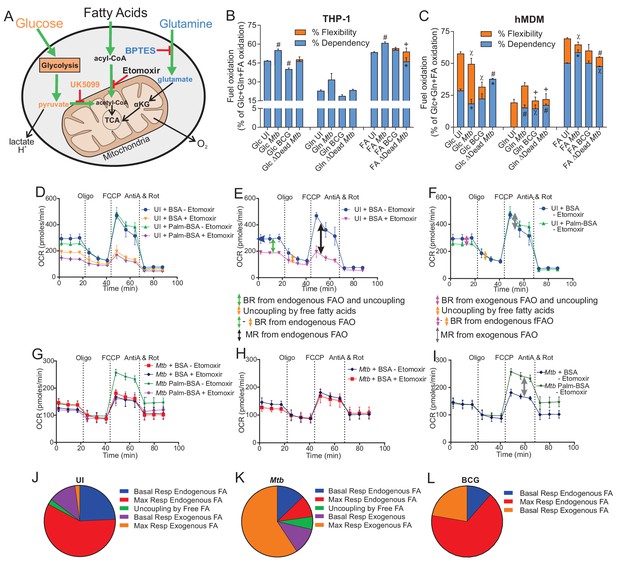
Mtb infection alters the mitochondrial substrate preference of macrophages.
(A–C) UK5099, etomoxir and BPTES were used to assess the mitochondrial flexibility and dependency on glucose (Glc), fatty acids (FA) and glutamine (Gln) in (B) THP-1 and (C) hMDM cells infected with Mtb, BCG and ∆Dead Mtb at MOIs of 1 and 5, respectively, for 18 h. Data shown are the mean ±SEM of five independent experiments. Student’s t test relative to uninfected cells (UI); #, p < 0.0001; χ, p < 0.0005; ϕ, p < 0.001; *p < 0.005; +, p < 0.05. (D–I) The oxidation of endogenous or exogenous fatty acids in (D–F) uninfected hMDM cells in addition to (G–I) Mtb-infected macrophages at a MOI of 5 were assessed by adding a palmitate-BSA (palm-BSA) conjugate and BSA controls before analysis on the XF with the respiration test. Etomoxir was used to assess inhibition of the transport of long-chain fatty acids into the mitochondria. BR, basal respiration; MR, maximal respiration (which gives a measure of respiration under conditions of stress); FAO, fatty acid oxidation; FA, fatty acids. (J–L) Pie charts illustrating the mitochondrial fatty acid preferences of (J) uninfected, (K) Mtb- and (L) BCG-infected hMDMs under basal and stressful conditions. Profiles and pie charts are representative of two independent experiments (n = 6 biological replicates).
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Mycobacterium Tuberculosis) | Mtb | BEI resources | NR-123 | |
| Strain, strain background (Mycobacterium bovis) | BCG | Stratens Serum Institut | Danish BCG vaccine strain 1331 | |
| Strain, strain background (Mycobacterium Tuberculosis) H37Rv AYs330 | Mtb-GFP | Steyn Laboratory | GFP Reporter strain | |
| Cell line (Homo sapiens) | THP-1 | ATCC Cat# TIB-202 | RRID:CVCL_0006 | |
| Biological sample (Homo sapiens) | Buffy coats | South African National Blood Service | ||
| Biological sample (Homo sapiens) | Human plasma | South African National Blood Service | ||
| Antibody | CD14 microbeads, Human | Miltenyi Biotec | Cat# 130- 050-201 | MACS (1:25) |
| Commercial assay or kit | Seahorse XF Mito Fuel Flex Test Kit | Agilent | Cat# 103260–100 | |
| Commercial assay or kit | Lookout Mycoplasma PCR detection kit | Sigma | Cat# MO0035 | |
| Chemical compound, drug | Seahorse XF Palmitate- BSA FAO Substrate | Agilent | Cat# 102720–100 | |
| Chemical compound, drug | Oligomycin (from Streptomyces diastatochromogenes) | Sigma | Cat# O4876 | |
| Chemical compound, drug | Carbonilcyanide p-triflouromethoxyphenylhydrazone (FCCP) | Sigma | Cat# C2920 | |
| Chemical compound, drug | Antimycin A | Sigma | Cat# A8674 | |
| Chemical compound, drug | Rotenone | Sigma | Cat# R8875 | |
| Chemical compound, drug | D-(+)- Glucose | Sigma | G8270 | |
| Chemical compound, drug | GlutaMAXTM | Gibco | Cat# 35050–038 | |
| Chemical compound, drug | DMEM | Lonza | Cat# BE12-604F | |
| Chemical compound, drug | L-Carnitine hydrochloride | Sigma | Cat# C0283 | |
| Chemical compound, drug | NaCl | Sigma | Cat# S3014 | |
| Chemical compound, drug | KCl | Sigma | Cat# SAAR5042020EM | |
| Chemical compound, drug | CaCl2 | Sigma | Cat# 1023780500 | |
| Chemical compound, drug | HEPES | Sigma | Cat# H0887 | |
| Chemical compound, drug | Sodium pyruvate | Sigma | Cat# S8636 | |
| Chemical compound, drug | XF Base medium | Agilent | Cat# 102353–100 | |
| Chemical compound, drug | Etomoxir | Sigma | Cat# E1905 | |
| Chemical compound, drug | Formalin Buffered, Neutral | Sigma | Cat# SAAR2436021EL | |
| Chemical compound, drug | Methanol | Sigma | Cat# 34860 | |
| Chemical compound, drug | Acetonitrile | Sigma | Cat# 34851 | |
| Chemical compound, drug | Human GM-CSF | Celtic Diagnostics | Cat# 300-03-100 | |
| Chemical compound, drug | Histopaque 1077 | Sigma | Cat# 10771 | |
| Chemical compound, drug | DPBS | Lonza | Cat# BE17-512F | |
| Chemical compound, drug | DMSO | Sigma | Cat# 41639 | |
| Chemical compound, drug | Bradford Dye | BIO-RAD | Cat# 500–0205 | |
| Chemical compound, drug | Middlebrook 7H11 Agar | BD | Cat# 283810 | |
| Chemical compound, drug | Polymyxin B | Sigma | Cat# P1004 | |
| Chemical compound, drug | Amphotericin B | Sigma | Cat# A4888 | |
| Chemical compound, drug | Carbenicillin | Sigma | Cat# C1389 | |
| Chemical compound, drug | Trimethoprim | Sigma | Cat# T7883 | |
| Chemical compound, drug | Cell-TakTM | Corning | Cat# 354241 | |
| Chemical compound, drug | Phorbol 12 -myristate 13-acetate | Sigma | Cat# P8139 | |
| Chemical compound, drug | Agarose | Sigma | Cat# A9539 | |
| Chemical compound, drug | SyBr Safe | Invitrogen | Cat# 533102 | |
| Chemical compound, drug | Middlebrook OADC | BD | Cat# 212240 | |
| Chemical compound, drug | RPMI1640 | Lonza | Cat# BE12-167F | |
| Chemical compound, drug | 2-Mercapt oethanol | Gibco | Cat# 21985023 | |
| Chemical compound, drug | 2-Deoxy-D-Glucose | Sigma | Cat# D6134 | |
| Chemical compound, drug | D-Glucose 13C6 | LC Scientific | Cat# GG601L | |
| Chemical compound, drug | Middlebrook 7H9 Broth | BD | Cat# 271310 | |
| Chemical compound, drug | MgSO4 | Sigma | Cat# SAAR4123920EM | |
| Chemical compound, drug | NaH2PO4 | Sigma | Cat# S9638 | |
| Software, algorithm | Wave desktop software | Agilent | Version 2.6 | |
| Software, algorithm | GraphPad Prism | Graphpad Prism | Version 7.04 | |
| Software, algorithm | CorelDRAW | Corel | Version X8 | |
| Software, algorithm | XF Cell Mito Stress Test Report Generator | Agilent | https://www.agilent.com/en/ products/cell-analysis/xf-cell-mito -stress-test-report-generator | |
| Software, algorithm | XF Glycolysis Stress Test Report Generator | Agilent | https://www.agilent.com/en/products/cell-analysis/xf-glycolysis-stress-test-report-generator | |
| Software, algorithm | XF Glycolytic Rate Assay Report Generator | Agilent | https://www.agilent.com/en/products/cell-analysis/xf-glycolytic-rate-assay-report-generator | |
| Software, algorithm | XF Mito Fuel Flex Test Report Generator | Agilent | https://www.agilent.com/en/products/cell-analysis/report-generator-for-the-xf-mito-fuel-flex-test | |
| Software, algorithm | XF Real-Time ATP Rate Assay Report Generator | Agilent | https://www.agilent.com/en/products/cell-analysis/xf-real-time-atp-rate-assay-report- generator |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.39169.015





