SUMO peptidase ULP-4 regulates mitochondrial UPR-mediated innate immunity and lifespan extension
Figures
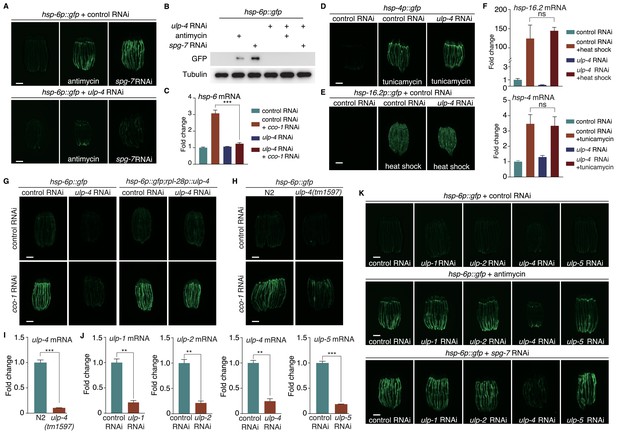
SUMO-specific peptidase ULP-4 is required for the activation of UPRmt.
(A) hsp-6p::gfp animals fed with control RNAi (top) or ulp-4 RNAi (bottom) were untreated, or treated with antimycin or spg-7 RNAi. Scale bar is 200 μm in this study unless otherwise indicated. Secondary RNAi was added 24 hr later after first RNAi unless otherwise indicated. Antimycin was added 48 hr later after first RNAi unless otherwise indicated. (B) Immunoblot of GFP expression in untreated hsp-6p::gfp worms, or hsp-6p::gfp worms treated with antimycin or spg-7 RNAi. (C) Quantitative PCR of endogenous hsp-6 mRNA levels. (D–E) hsp-4p::gfp (D) or hsp-16.2p::gfp (E) animals on control or ulp-4 RNAi were treated with tunicamycin (D) or heat-shocked (E). (F) Quantitative PCR of endogenous hsp-16.2 and hsp-4 mRNA levels. (G) hsp-6p::gfp or hsp-6p::gfp; rpl-28p::ulp-4 (ulp-4 overexpression, OE) animals fed with control or ulp-4 RNAi were untreated or treated with cco-1 RNAi. Overexpressed ulp-4 is codon optimized. (H) hsp-6p::gfp or ulp-4(tm1597); hsp-6p::gfp animals were fed with control or cco-1 RNAi. (I) Quantitative PCR of ulp-4 mRNA level in wild-type or ulp-4(tm1597) animals. (J) Quantitative PCR of endogenous ulp-1,2,4,5 mRNA levels under each respective RNAi. (K) hsp-6p::gfp animals fed with control or ulp-1,2,4,5 RNAi were treated with antimycin or spg-7 RNAi. Error bars show standard deviation. Student’s t-test, ns not significant, **p<0.002 and ***p<0.0002. All experiments in this paper, if not specifically indicated, have been repeated for at least three times.
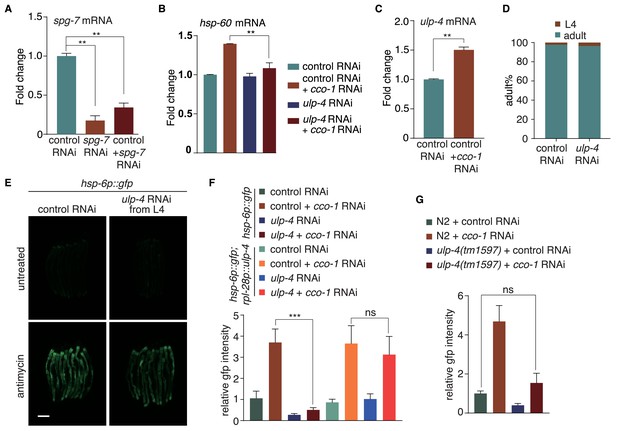
SUMO-specific peptidase ULP-4 is required for the activation of UPRmt.
(A–C) Quantitative PCR of spg-7 (A), hsp-60 (B) and ulp-4 (C) mRNA levels under indicated RNAi. (D) Statistic results for developmental stages of worm fed with control or ulp-4 RNAi at 20°C for 60 hr, n > 80. (E) L4 stage hsp-6p::gfp animals were fed with control or ulp-4 RNAi for 24 hr before antimycin treatment. Representative fluorescent images were taken 24 hr after antimycin treatment. (F–G) Statistic result of relative gfp intensity in Figure 1C (F) and 1D (G). Student’s t-test. Error bars indicate standard deviation. ns not significant, *p<0.05, **p<0.002 and ***p<0.0002.

Sequence alignment of worm ULP proteins.
The alignment was performed by MAFFT (https://www.ebi.ac.uk/Tools/msa/mafft/). The catalytic domain of ULP-4 is highlighted in red.
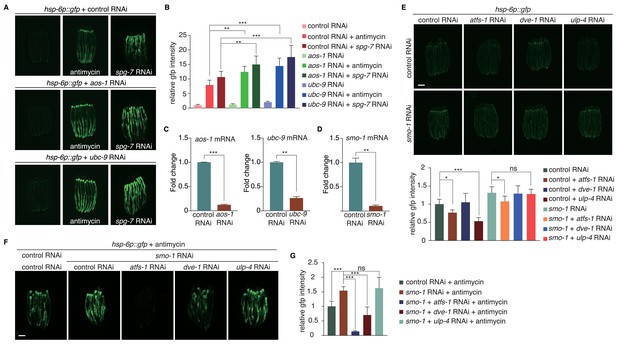
Deficiency of smo-1 or SUMO conjugating enzymes enhances UPRmt.
(A) hsp-6p::gfp animals on control, aos-1 or ubc-9 RNAi were untreated, or treated with antimycin or spg-7 RNAi.(B) Statistic result of relative gfp intensity in (A). (C–D) Quantitative PCR testing RNAi efficiency of aos-1, ubc-9 (B) and smo-1 (C) RNAi. (E) Representative fluorescent images (upper panel) and statistic result of relative gfp intensity (lower panel) for hsp-6::gfp animals fed with indicated RNAi. (F) Representative fluorescent images of hsp-6p::gfp animals fed with indicated RNAi and then treated with antimycin. (G) Statistic result of relative gfp intensity in (F). Student’s t-test. Error bars indicate standard deviation. ns not significant, **p<0.002 and ***p<0.0002.
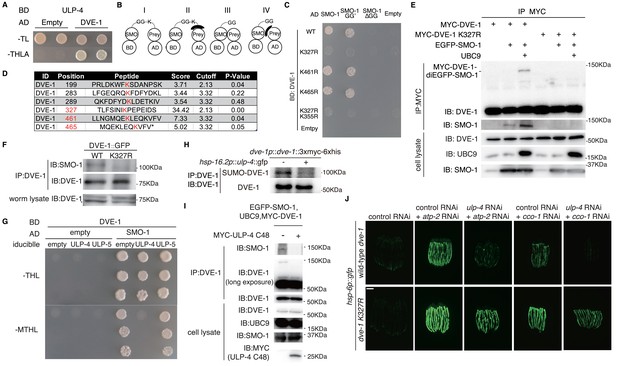
ULP-4 deSUMOylates DVE-1 at K327 residue during mitochondrial stress.
(A) Yeast two-hybrid result of ULP-4 and DVE-1 interaction. AH109 strain was transformed with Gal4-BD-ULP-4 and empty Gal4-AD or Gal4-AD-DVE-1 plasmids. –TL, tryptophan and leucine dropout agar media plates. –THLA, tryptophan, leucine, histidine and adenine dropout agar media plates. (B) Diagrams showing possible mechanisms for SMO-1 and prey protein interaction in yeast two-hybrid assay. (C) Yeast two-hybrid assay of the interaction between wild-type, K327R, K461R, K465R or K355R DVE-1 and SMO-1. SMO-1 GG’, residues after C-terminal di-glycine were deleted. SMO-1ΔGG, di-glycine residues in SMO-1 were deleted. (D) Predicted SUMOylation sites of DVE-1 using GPS-SUMO web service (http://sumosp.biocuckoo.org/online.php). (E) In vivo SUMOylation assay for DVE-1. Myc-tagged wild-type DVE-1 or DVE-1 K327R, SMO-1 and E2 SUMO conjugating enzyme UBC9 were expressed in 293 T cells. (F) DVE-1 SUMOylation in worms. DVE-1::GFP or DVE-1 K327R::GFP driven by dve-1 promotor were expressed in worms. (G) Yeast three-hybrid assay of DVE-1 deSUMOylation. Constitutive Gal4-BD-DVE-1 expression and inducible ULP-4/ULP-5 expression elements were cloned into pBridge. ULP-4/5 was induced when methionine was dropout. –MTHL, methionine, tryptophan, leucine and histidine dropout agar plates. (H) Immunoblot of DVE-1 SUMOylation. L4 stage dve-1p::dve-1::3xmyc-6xhis worms or dve-1p::dve-1:: 3xmyc-6xhis; hsp-16.2p::ulp-4::gfp worms were heat shocked at 37°C for 1 hr, and cultured at 20°C for 8 hr before immunoprecipitation. (I) In vivo deSUMOylation assay for DVE-1. Myc-tagged ULP-4 C48 domain, SMO-1 and E2 SUMO conjugating enzyme UBC9 were expressed in 293 T cells. (J) hsp-6p::gfp or dve-1 K327R; hsp-6p::gfp animals fed with control or ulp-4 RNAi were treated with control, atp-2 or cco-1 RNAi.
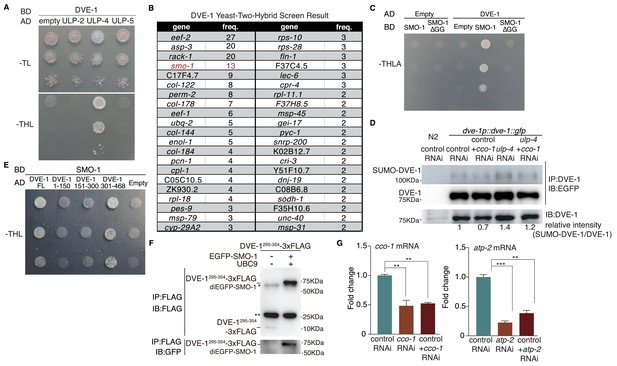
DVE-1 interacts with ULP-4 and SMO-1 in yeast two-hybrid assay.
(A) Yeast two-hybrid result of DVE-1 and ULPs. (B) Screen hits of yeast two-hybrid experiment with dve-1 as bait. Gal4-BD-dve-1 vector and Gal4-AD vector containing worm cDNAs were transformed in yeast. Clones grew on –THLA plates were picked and sequenced. Genes that have been identified more than twice were listed. (C) SMO-1 covalently modifies DVE-1. SMO-1 Δ GG: SMO-1 deleted with C-terminal di-glycine. (D) Immunoblot of SUMOylated DVE-1 in N2 or DVE-1::GFP worms with indicated RNAi. (E) SMO-1 interacts with DVE-1 301–468. Full-length (FL) DVE-1 or DVE-1 fragments 1–150, 151–300 and 301–468 are tested in yeast two-hybrid assay. (F) Immunoprecipitation and western blotting to detect SUMOylation of DVE-1 295–354. * indicates nonspecific band. ** indicates Igg light chain. (G) Quantitative PCR testing RNAi efficiency of cco-1 or atp-2 RNAi. Student’s t-test. Error bars indicate standard deviation. ns not significant, **p<0.002 and ***p<0.0002.
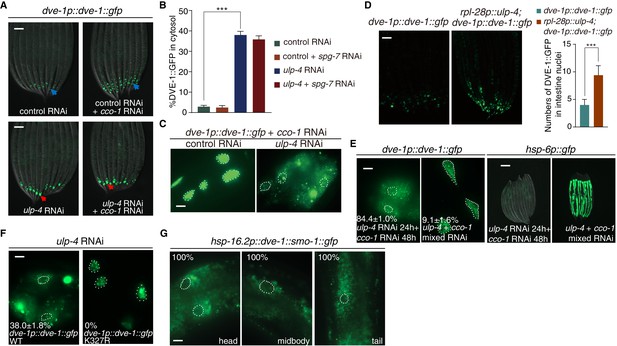
SUMOylation affects the subcellular localization of DVE-1.
(A) Representative fluorescent images of dve-1p::dve-1::gfp animals fed with indicated RNAi. Blue arrows indicate that DVE-1::GFP remains in the nucleus. Red arrows indicate that DVE-1::GFP localizes in the cytosol. Scale bar is 100 μm. (B) Percentage of DVE-1::GFP in cytosol with indicated RNAi. Student’s t-test, ***p<0.0002. Error bars indicate standard deviation. (C) Representative fluorescent images of dve-1p::dve-1::gfp animals fed with control or ulp-4 RNAi. Scale bar is 10 μm. (D) Representative fluorescent images (left) and statistic result (right) for nuclear accumulation of DVE-1 in dve-1p::dve-1::gfp animals with or without ulp-4 overexpression. (E) Induction of UPRmt correlates with nuclear accumulation of DVE-1. dve-1p::dve-1::gfp and hsp-6p::gfp P0 animals were fed with ulp-4 RNAi. dve-1p::dve-1::gfp (left) or hsp-6p::gfp (right) animals were cultured with ulp-4 RNAi for one generation, and F1s were then treated with indicated RNAi starting at L1 stage. Scale bar is 10 μm (left) and 200 μm (right). The number indicates the proportion of animals with DVE-1::GFP in cytosol. Numbers indicate Mean ±standard deviation. N = 3 biological replicates, n > 100 worms each replicate. (F) DVE-1 K327R constitutively localizes in the nucleus, even under ulp-4 RNAi. The number indicates the proportion of animals with DVE-1::GFP in cytosol. Numbers indicate Mean ±standard deviation. N = 3 biological replicates, n > 100 worms per replicate. (G) Fusion of SMO-1 to DVE-1 to mimic its SUMOylated form results in DVE-1 expression in the cytosol. Worms were observed 2 hr after 1 hr heat shock at 37°C. The number indicates the proportion of animals with DVE-1::GFP in cytosol.
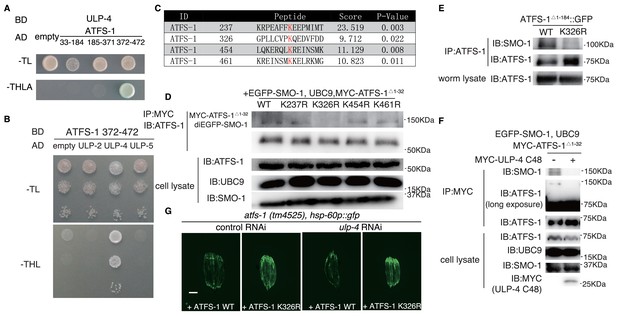
ULP-4 deSUMOylates ATFS-1 at K326 residue upon mitochondrial stress.
(A) Yeast two-hybrid assay for ATFS-1 and ULP-4 interaction. ATFS-1 was truncated into 33–174, 175-371and 372–472 fragments. (B) Yeast two-hybrid assay for ATFS-1 372–472 and ULPs. (C) Predicted SUMOylation sites of ATFS-1 using GPS-SUMO web service. (D) In vivo SUMOylation assay for ATFS-1. Myc-tagged wild-type ATFS-1Δ1-32 or ATFS-1Δ 1-32 lysine mutants, SMO-1 and E2 SUMO conjugating enzyme UBC9 were expressed in 293 T cells. (E) ATFS-1 SUMOylation in worms. Wild-type or ATFS-1Δ 1-184 K326R was expressed in worms. (F) ATFS-1 Δ 1-32 deSUMOylation in 293 T cells. SMO-1, UBC9, ATFS-1 Δ 1-32 and ULP-4 C48 domain were expressed in 293 T cells. (G) Representative fluorescent images of atfs-1(tm4525); hsp-60p::gfp with wild-type ATFS-1 or ATFS-1 K326R animals fed on control or ulp-4 RNAi.
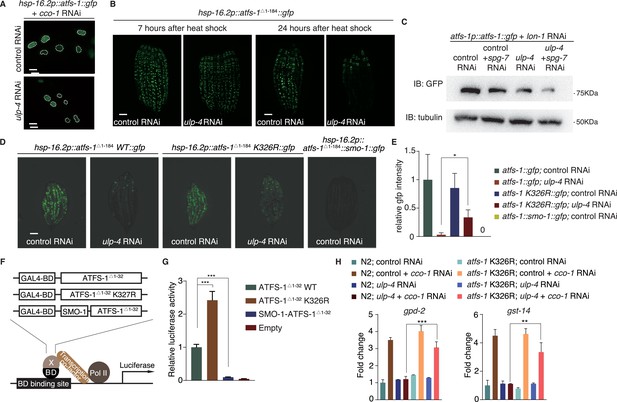
SUMOylation affects the stability and transcriptional activity of ATFS-1.
(A) Representative fluorescent images of ATFS-1::GFP localization with indicated treatment. (B) Representative fluorescent images of ATFS-1 Δ 1-184::GFP. Worms were heat-shocked at 37°C for 1 hr and photographed at indicated time points. (C) Immunoblot of ATFS-1::GFP expression in atfs-1p::atfs-1::gfp worms fed with indicated RNAi. (D) Representative fluorescent images of wild-type, SUMOylation mutant or SUMOylation mimetic ATFS-1 Δ 1-184::GFP with indicated RNAi. (E) Statistic result of (D). (F) Diagram of transcriptional activity assay. ATFS-1 Δ 1-32 was cloned into a vector containing Gal4-BD at N-terminus to induce luciferase expression. (G) Transcriptional activity assay of ATFS-1 Δ 1-32, ATFS-1 Δ 1-32 K326R or SMO-1-ATFS-1 Δ 1-32. Student’s t-test. ***p<0.0002. Error bars indicate standard deviation. (H) Quantitative PCR of god-2 and gst-14 mRNA levels in wild-type or atfs-1 K326R worms fed with indicated RNAi. .
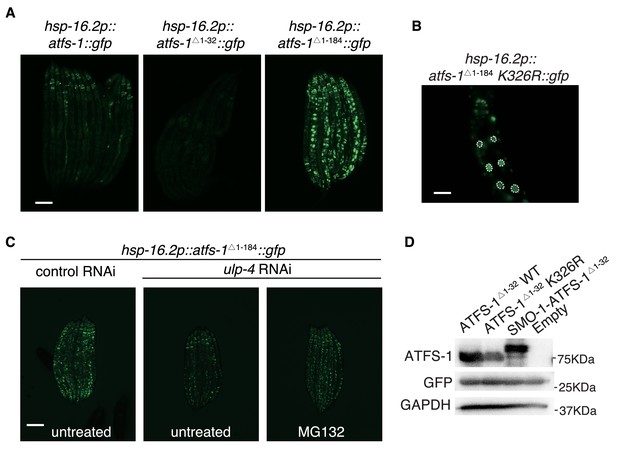
SUMOylation affects the stability and transcriptional activity of ATFS-1.
(A) Representative fluorescent images showing expression level of full-length ATFS-1, ATFS-1Δ 1-32, and ATFS-1Δ 1-184. (B) Representative fluorescent images showing subcellular localization of ATFS-1Δ 1-184 K326R. (C) ulp-4 RNAi resulted in ATFS-1 degradation, which was partially prevented by treating with protease inhibitor MG132. (D) Immunoblot of wild-type, K326R or SMO-1-conjugated ATFS-1 ATFS-1Δ1-32 in 293 T cells. GFP was co-transfected as a control of transfection efficiency.
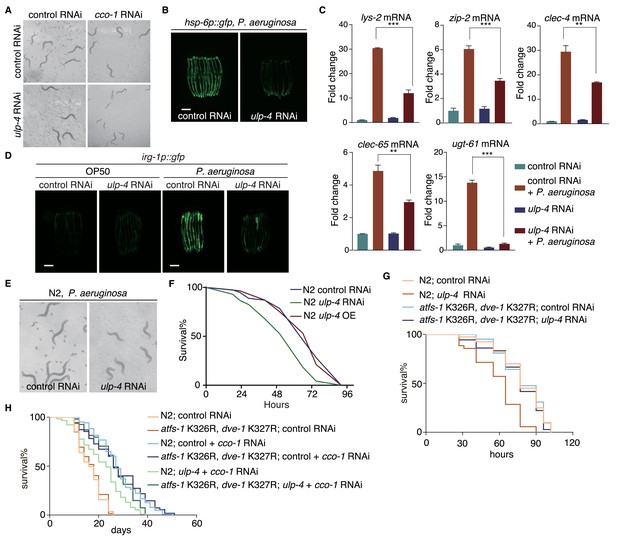
ulp-4 is essential for UPRmt-regulated innate immunity and lifespan extension.
(A) Representative images of wild-type worms raised on indicated RNAi. cco-1 RNAi was treated 16 hr later after control or ulp-4 RNAi. (B) Representative fluorescent images of hsp-6p::gfp worms. Animals were pre-treated with control or ulp-4 RNAi for 24 hr, and then fed with P. aeruginosa for additional 48 hr. Student’s t-test. Error bars indicate standard deviation. **p<0.002 and ***p<0.0002. (C) Fold changes of immune response gene lys-2, zip-2, clec-4, clec-65 and ugt-61 mRNA levels in control or ulp-4 RNAi animals after exposure to P. aeruginosa. (D) Immune response reporter strain irg-1p::gfp fed with control or ulp-4 RNAi was infected with P. aeruginosa strain PA14. (E) Representative images of wild-type worms fed with control or ulp-4 RNAi for 24 hr and then exposed to P. aeruginosa. (F) PA14 survival assay in wild-type, ulp-4 knockdown or overexpression animals. N = 3 biological replicates, n > 40 worms per replicate. Analyzed using Log-Rank method and p<0.05 (N2 control RNAi vs N2 ulp-4 RNAi). (G) PA14 survival assay in N2 or atfs-1 K326R; dve-1 K327R animals with indicated RNAi. N = 2 biological replicates, n > 35 worms per replicate. Analyzed using Log-Rank method, p<0.05 (N2 control RNAi vs N2 ulp-4 RNAi), p>0.05(atfs-1 K326R; dve-1 K327R control RNAi vs atfs-1 K326R; dve-1 K327R ulp-4 RNAi). (H) Representative lifespan result of N2 or atfs-1 K326R; dve-1 K327R animals fed with indicated RNAi. n > 80 worms per condition. Analyzed using Log-Rank method, p<0.0001 (N2 control +cco-1 RNAi vs N2 ulp-4 RNAi + cco-1 RNAi), p<0.01(N2 ulp-4 +cco-1 RNAi vs atfs-1 K326R; dve-1 K327R ulp-4 +cco-1 RNAi).
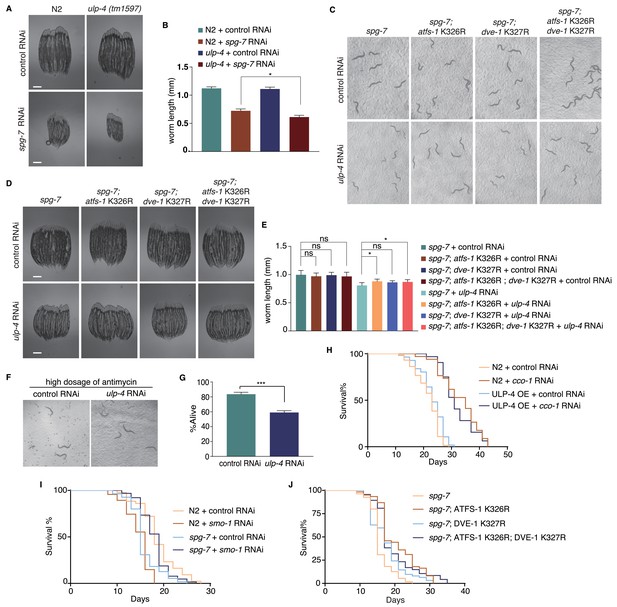
ulp-4 is essential for UPRmt-regulated innate immunity and lifespan extension.
(A,B) Representative images (A) and statistical results of worm lengths (B) for wild-type or ulp-4(tm1597) fed with control or spg-7 RNAi, n > 15 worms. (C–E) Representative images (C,D) and statistical results of worm lengths (E) for worms with indicated genotype and RNAi treatment, n > 15 worms. (F,G) Representative images (F) and statistical results of survival rates for control or ulp-4 RNAi animals treated with high dosage of antimcyin (20 µg antimycin in a 6-well plate), N = 3 biological replicates, n > 40 worms per replicate. (H–J) Lifespan of worms with indicated genotype and RNAi treatment. Analyzed using Log-Rank method (n > 60 worms). (H) p=0.2254 (N2 +control RNAi vs ULP-4 OE +control RNAi), p=0.6672 (N2 +cco-1 RNAi vs ULP-4 OE +cco-1 RNAi). (I) p<0.0001 (N2 +control RNAi vs N2 +smo-1 RNAi), p=0.0006 (spg-7 +control RNAi vs spg-7 +smo-1 RNAi) (J) p<0.0001 (spg-7 vs spg-7; ATFS-1 K326R), p=0.0047 (spg-7 vs spg-7; DVE-1 K327R) and p<0.0001 (spg-7 vs spg-7; ATFS-1 K326R; DVE-1 K327R). Student’s t-test. Error bars indicate standard deviation. ns not significant, *p<0.05 and ***p<0.0002.
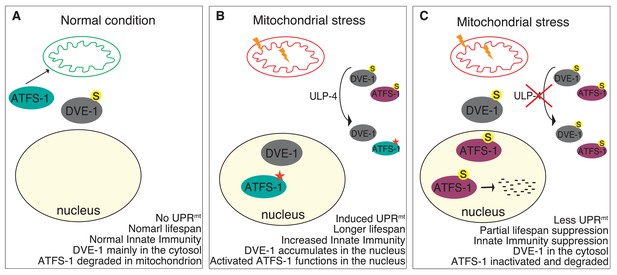
Model for ULP-4-mediated UPRmt signaling.
(A) Under normal condition, ATFS-1 is imported into mitochondria and degraded by mitochondrial protease. DVE-1 is mainly localized in the cytosol. No UPRmt is induced. The animal is normal lived and with normal innate immunity. (B) Upon mitochondrial stress, DVE-1 is deSUMOylated by ULP-4 and translocated to the nucleus. Mitochondrial importer efficiency is compromised, leading to ATFS-1 nuclear accumulation. ULP-4 deSUMOylates ATFS-1, leading to increased stability and transcriptional activity (active form). UPRmt is induced. Animals are long-lived, with increased innate immunity. (C) Reduction of ULP-4 activity during mitochondrial stress results in SUMOylation of DVE-1 and ATFS-1. SUMOylated of DVE-1 localizes in the cytosol; whereas SUMOylated ATFS-1, which has lower transcriptional activity, is prone to degradation. Induction of UPRmt is impaired. UPRmt-regulated innate immunity and lifespan extension is partially suppressed.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (C. elegans) | hsp-6p::gfp | Caenorhabditis Genetics Center | WB Strain: SJ4100 | |
| Genetic reagent (C. elegans) | hsp-16.2p::gfp | Caenorhabditis Genetics Center | WB Strain: CL2070 | |
| Genetic reagent (C. elegans) | hsp-4p::gfp | Caenorhabditis Genetics Center | WB Strain: SJ4005 | |
| Genetic reagent (C. elegans) | dve-1p::dve-1::gfp | Caenorhabditis Genetics Center | WB Strain: SJ4198 | |
| Genetic reagent (C. elegans) | hsp-60p::gfp | Caenorhabditis Genetics Center | WB Strain: SJ4058 | |
| Genetic reagent (C. elegans) | N2 | Caenorhabditis Genetics Center | WB Strain: N2 | |
| Genetic reagent (C. elegans) | DA2249 (spg-7 mutant) | Caenorhabditis Genetics Center | WB Strain: DA2249 | |
| Genetic reagent (C. elegans) | tm1597 (ulp-4 mutant) | National Bioresource Project | WB Variation: tm1597 | |
| Genetic reagent (C. elegans) | hsp-60p::gfp; atfs-1 (tm4525) | PMID:22700657 | Dr. Cole Haynes (University of Massachusetts Medical School) | |
| Genetic reagent (C. elegans) | hsp-60p::gfp; tm1597 | This paper | N/A | |
| Genetic reagent (C. elegans) | dve-1p::dve-1 K327R::gfp | This paper | N/A | |
| Genetic reagent (C. elegans) | hsp-16.2p::dve-1::smo- 1::gfp | This paper | N/A | |
| Genetic reagent (C. elegans) | hsp-16.2p::atfs-1::gfp | This paper | N/A | |
| Genetic reagent (C. elegans) | hsp-16.2p::atfs-1Δ1–32::gfp | This paper | N/A | |
| Genetic reagent (C. elegans) | hsp-16.2p::atfs-1Δ1– 184::gfp | This paper | N/A | |
| Genetic reagent (C. elegans) | hsp-16.2p::atfs-1Δ1– 184::smo-1::gfp | This paper | N/A | |
| Genetic reagent (C. elegans) | hsp-16.2p::atfs-1Δ1–184 K326R::gfp | This paper | N/A | |
| Genetic reagent (C. elegans) | hsp-60p::gfp;atfs-1(tm4525);atfs-1p::atfs-1 | This paper | N/A | |
| Genetic reagent (C. elegans) | hsp-60p::gfp;atfs-1(tm4525);atfs-1p::atfs-1 K326R | This paper | N/A | |
| Genetic reagent (C. elegans) | dve-1 K327R | This paper | N/A | Cas9 mutation |
| Genetic reagent (C. elegans) | atfs-1 K326R | This paper | N/A | Cas9 mutation |
| Genetic reagent (C. elegans) | atfs-1 K326R dve-1 K327R | This paper | N/A | Cas9 mutation |
| Genetic reagent (C. elegans) | dve-1 K327R;hsp-6p::gfp | This paper | N/A | Cas9 mutation |
| Genetic reagent (C. elegans) | rpl-28p::ulp-4(opti)::gfp | This paper | N/A | |
| Genetic reagent (C. elegans) | rpl-28p::ulp-4(opti)::gfp; dve-1p::dve-1::gfp | This paper | N/A | |
| Cell line (Homo sapiens) | 293T | ATCC | CRL-3216 | |
| Antibody | Mouse monoclonal anti-GFP | sungen biotech | CAT#KM8009 | (1:1000) |
| Antibody | Rabbit polyclonal anti-GFP | abcam | CAT#ab290 | (1:1000) |
| Antibody | Rabbit polyclonal anti-ATFS-1 | abclonal | N/A | custom made (1:5000) |
| Antibody | Rabbit polyclonal anti-DVE-1 | abclonal | N/A | custom made (1:5000) |
| Antibody | Rabbit polyclonal anti-SMO-1 | abclonal | N/A | custom made (1:5000) |
| Antibody | Mouse monoclonal anti-MYC | CST | CAT#2276 | (1:1000) |
| Antibody | Rat monoclonal anti-TUBULIN | abcam | CAT#6161 | (1:1000) |
| Antibody | Rabbit monoclonal anti-UBC9 | abcam | CAT#ab75854 | (1:1000) |
| Strain (Pseudomonas aeruginosa) | Pseudomonas aeruginosa | PMID:24695221 | N/A | |
| Strain (Pseudomonas aeruginosa) | Pseudomonas aeruginosa | PMID:25274306 | WB Strain: PA14 | |
| Commercial assay or kit | -Trp DO suplement | Coolaber | CAT#PM2140 | |
| Commercial assay or kit | -Leu -Trp DO suplement | Clontech | CAT#630417 | |
| Commercial assay or kit | -Leu -Trp -His DO suplement | Clontech | CAT#630419 | |
| Commercial assay or kit | -Leu -Trp -His -Ade DO suplement | Clontech | CAT#630428 | |
| Commercial assay or kit | -Leu -Trp -His -Met DO suplement | Coolaber | CAT#PM2250 | |
| Commercial assay or kit | Minimal SD Base | Clontech | CAT#630411 | |
| Commercial assay or kit | SYBR Green QPCR mix | Biorad | CAT#172–5122 | |
| Commercial assay or kit | transcript one-step gDNA removal and cDNA synthesis supermix | Transgene | AT311-03 | |
| Commercial assay or kit | Triton X-100 | sigma | T9284 | |
| Commercial assay or kit | Trizol | Cwbio | CW0580A | |
| Commercial assay or kit | Lipofectamine 3000 | life | L3000015 | |
| Commercial assay or kit | Yeast Plasmid Extraction Kit | solarbio | D1160-100 | |
| Commercial assay or kit | dynabeads protei G | life | 10004D | |
| Commercial assay or kit | Protease Inhibitor Cocktail | bimake | B14002 | |
| Commercial assay or kit | Antimycin | sigma | A8674 | |
| Commercial assay or kit | N-Ethylmaleimide | J and K | 128-53-0 | |
| Commercial assay or kit | 5-FLUORO-2'- DEOXYURIDINE | sigma | F0503 | |
| Commercial assay or kit | ECL western blotting kit | Pierce | CAT#32106 | |
| Commercial assay or kit | Dual-Luciferase Reporter Assay Systerm | Promega | CAT#E1910 | |
| Commercial assay or kit | Matchmaker GAL4 Two-Hybrid System 3 | Clontech | CAT# PT3247-1 | |
| Software, algorithm | Graph Pad Prism Software | GraphPad Software | https://www.graphpad.com/scientific-software/prism/ | |
| Software, algorithm | MAFFT | PMID:23329690 | https://www.ebi.ac.uk/Tools/msa/mafft/ | |
| Software, algorithm | SMART | PMID: 29040681 | http://smart.embl-heidelberg.de | |
| Software, algorithm | ImageJ | PMID: 22930834 | https://imagej.net/Downloads | |
| Software, algorithm | SUMO-GPS | PMID: 24880689 | http://sumosp.biocuckoo.org | |
| Recombinant DNA reagent | CMVp-myc-dve-1 | This paper | N/A | |
| Recombinant DNA reagent | CMVp-myc-dve-1 K327R | This paper | N/A | |
| Recombinant DNA reagent | CMVp-myc-33–472 atfs-1 | This paper | N/A | |
| Recombinant DNA reagent | CMVp-myc-33–472 atfs-1 K326R | This paper | N/A | |
| Recombinant DNA reagent | CMVp-gfp-smo-1 | This paper | N/A | |
| Recombinant DNA reagent | CMVp-ubc9 | This paper | N/A | |
| Recombinant DNA reagent | CMVp-ulp-4 C48 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 BD-dve-1 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 BD-dve-1 K327R | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 BD-dve-1 K461R | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 BD-dve-1 K465R | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 BD-dve-1 K327R K355R | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 BD-ulp-4 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 BD-smo-1 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 BD-smo-1 delta GG | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 AD-smo-1 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 AD-smo-1 delta GG | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 AD-smo-1 GG' | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 AD-33–184 atfs-1 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 AD-185–371 atfs-1 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 AD-372–472 atfs-1 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 AD-dve-1 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 AD-1–150 dve-1 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 AD-151–300 dve-1 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 AD-301–468 dve-1 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 AD-ulp-1 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 AD-ulp-2 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 AD-ulp-4 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 AD-ulp-5 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 BD-dve-1 met17p-ulp-4 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 BD-dve-1 met17p-ulp-5 | This paper | N/A | |
| Recombinant DNA reagent | adh1p-gal4 BD-dve-1 in pBridge | This paper | N/A | |
| Recombinant DNA reagent | Adh1p-gal4 AD-33–184 atfs-1 | This paper | N/A | |
| Recombinant DNA reagent | Adh1p-gal4 AD-185–371 atfs-1 | This paper | N/A | |
| Recombinant DNA reagent | Adh1p-gal4 AD-372–472 atfs-1 | This paper | N/A | |
| Recombinant DNA reagent | CMVp-gal4 BD-33–472 atfs-1 | This paper | N/A | |
| Recombinant DNA reagent | CMVp-gal4 BD-33–472 atfs-1 K327R | This paper | N/A | |
| Recombinant DNA reagent | CMVp-gal4 BD-smo- 1-33-472 atfs-1 | This paper | N/A | |
| Recombinant DNA reagent | rpl-28p::ulp-4(opti)::gfp | This paper | N/A | |
| Recombinant DNA reagent | hsp-16.2p::atfs-1::gfp | This paper | N/A | |
| Recombinant DNA reagent | hsp-16.2p::33–472 atfs-1::gfp | This paper | N/A | |
| Recombinant DNA reagent | hsp-16.2p::185–472 atfs-1::gfp | This paper | N/A | |
| Recombinant DNA reagent | hsp-16.2p::185–472 atfs-1 K326R::gfp | This paper | N/A | |
| Recombinant DNA reagent | atfs-1p::atfs-1 | This paper | N/A | |
| Recombinant DNA reagent | atfs-1p::atfs-1 K326R | This paper | N/A | |
| Recombinant DNA reagent | dve-1p::dve-1 K327R::gfp | This paper | N/A | |
| Recombinant DNA reagent | hsp-16.2p::smo-1::dve-1::gfp | This paper | N/A | |
| Recombinant DNA reagent | hsp-16.2p::dve-1::smo-1::gfp | This paper | N/A | |
| Recombinant DNA reagent | hsp-16.2p::atfs-1::smo-1::gfp | This paper | N/A |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.41792.017





