The cellular basis of mechanosensory Merkel-cell innervation during development
Figures
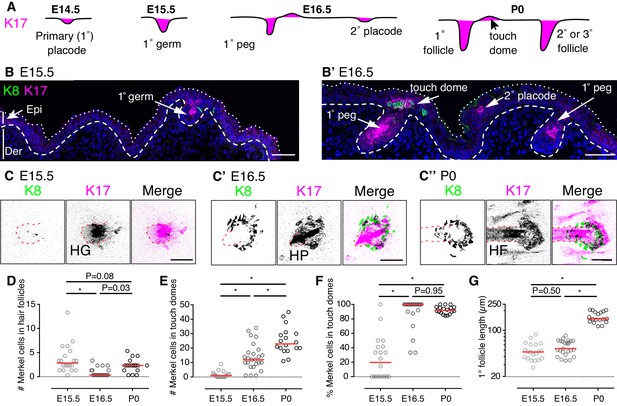
Touch domes emerge at E16.5.
(A) Stages of hair-follicle and touch-dome morphogenesis. (B) Sagittal cryosections of dorsal skin at E15.5 and E16.5. Merkel cells are labeled with antibodies against K8 (green) and hair follicle and touch-dome keratinocytes are stained for K17 protein (magenta). Nuclei are labeled with DAPI (blue). Dotted and dashed lines outline the skin surface and basal epidermis, respectively. (C) Confocal axial projections show full-thickness cleared skin specimens at E15.5 (left trio of panels), E16.5 (middle trio), and P0 (right trio). K8 immunoreactivity: left panels and green in merged images; K17 immunoreactivity: middle panels and magenta in merged images. In the inverted lookup table (LUT) applied to merged images here and in Figure 2,3,4,5,7 and Figure 5—figure supplement 1, black denotes co-localization of green and magenta pixels. Hair follicle structures (hair germ, HG, and hair peg, HP) are indicated by red dashed lines. (D–G) Quantification of Merkel-cell distributions and follicle lengths for primary hair follicles and touch domes at E15.5 (N = 20), E16.5 (N = 25) and P0 (N = 18). Red lines denote medians. Scatter plots show the number of Merkel cells present within each primary hair follicle (D) or adjacent touch domes (E), the corresponding percentage of Merkel cells in touch domes (F), and the lengths of reconstructed primary follicles (G). One-way ANOVA with Tukey’s multiple comparisons test. *p<0.0001. Primary follicles associated with at least one Merkel cell were quantified from three mice per stage. Scale bars: 50 µm. See also Figure 1—figure supplement 1 and Figure 1–video 1.
-
Figure 1—source data 1
Numerical values for data plotted in Figure 1.
- https://doi.org/10.7554/eLife.42633.005
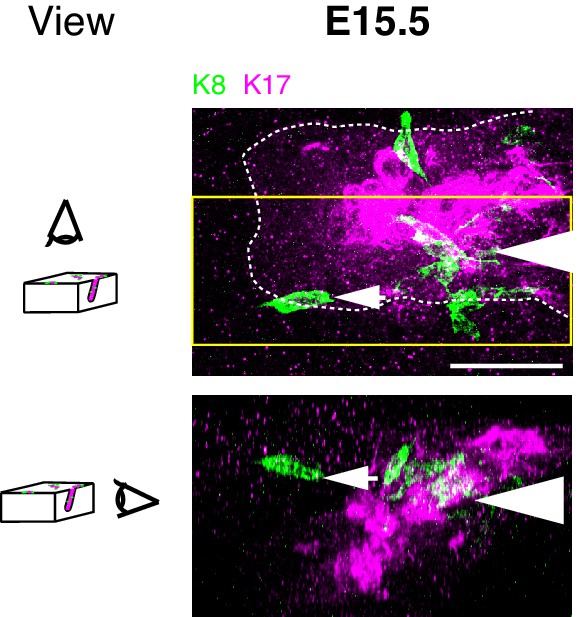
Three-dimensional projections in different planes show that Merkel cells are located in both primary hair follicles and touch dome epidermis.
Projections of a confocal z-stack of full-thickness skin at E15.5. Merkel cells, labeled with K8 antibodies (green) are present both in the primary hair peg (arrowhead) and in the surrounding interfollicular epidermis that makes up the touch dome (arrow). Hair follicles and touch-dome keratinocytes were labeled with K17 antibodies (magenta). Yellow box indicates the slice of the axial view (top) that is projected in the sagittal plane (bottom). Dashed lines denote the hair peg. Scale bar: 25 µm.
Three-dimensional image stack of whole-mount immunofluorescence at E15.5.
The movie progresses from epidermal to deep dermal levels. Touch-dome keratinocytes (superficial optical sections) and hair follicle keratinocytes (deeper optical sections) were labeled with K17 antibodies (magenta). Merkel cells, labeled with K8 antibodies (green) are present both in the touch-dome epidermis and embedded in the primary hair peg. The image contrast was linearly adjusted to facilitate visualizing immunoreactive cells in the movie file.
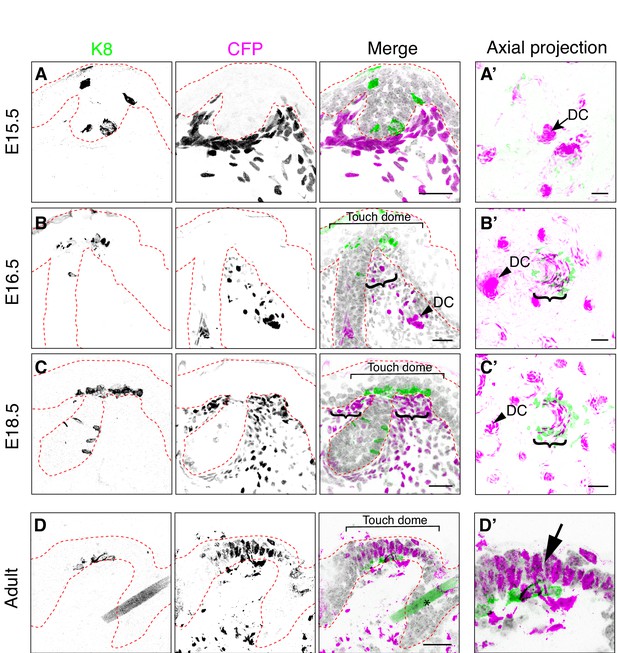
Bmp4-expressing cells form mesenchymal condensates under touch-dome placodes.
Representative images of immunohistochemistry in Bmp4CFP mice. (A–D, D’) Sagittal cryosections of dorsal skin at E15.5 (A), E16.5 (B), E18.5 (C) and adult (9 month old female; D, D’). Anti-K8 antibodies labeled Merkel cells (left panels and green in merged images). Anti-GFP antibodies labeled CFP-expressing cells (middle panels and magenta in merged images). DAPI is shown in grayscale. Red dashed lines outline epidermis. Asterisk (D) indicates an autofluorescent hair shaft in adult skin. In (B–C), brackets denote touch domes, curly brackets indicate clusters of CFPhigh cells beneath touch domes, and arrowheads indicate CFPhigh cells in dermal condensates (DC). In (D’), arrow indicates epidermal expression of CFP driven from the Bmp4 locus in adult epidermis. (A’–C’) Confocal axial projections of full-thickness dorsal skin at E15.5 (A’), E16.5 (B’) and E18.5 (C’). Pseudocolor in merged images indicates K8 (green) and K17 staining (magenta). In axial projections, DCs and dermal papilla (DP) appear as intensely stained dots and touch dome mesenchymal clusters appear as crescents. Images are representative of touch domes from 2–4 animals per stage. Scale bars: 25 µm.
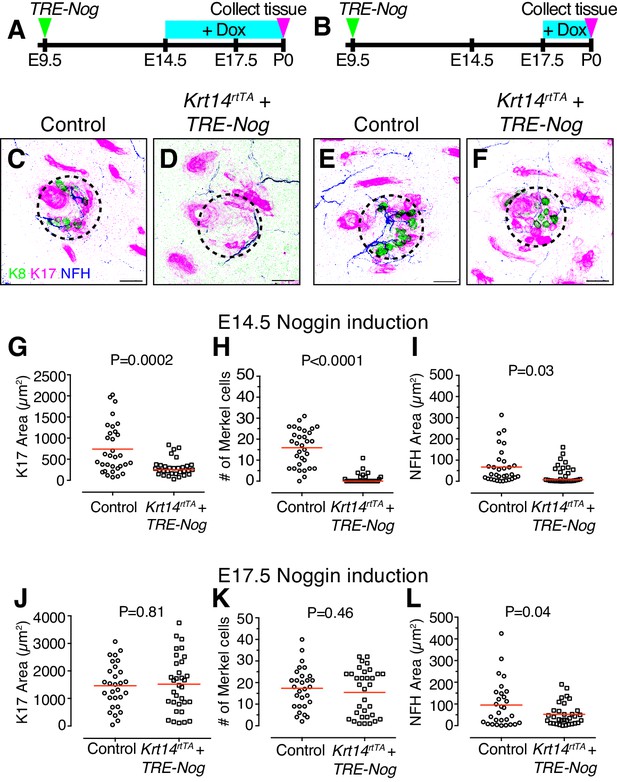
BMP signaling is required for touch-dome development.
(A–B) Experimental design. (C–F) Representative axial projections of P0 dorsal skin labeled with K8 (green), K17 (magenta) and NFH (blue) after doxycycline (Dox)-induction of Noggin expression from E14.5–P0 (C–D) or E17.5–P0 (E–F). Dashed circles denote touch domes. (C, E) Control mice either lacked the Krt14-driven Tet activator transgene (TRE-Nog + Dox) or the virally induced Tet responsive Noggin transgene (Krt14rtTA + Dox). (D, F) Experimental mice overexpressed Noggin in epidermis (Krt14rtTA + TRE-Nog + Dox). (G–I) Quantification of K17 area (G) number of Merkel cells per touch dome (H), and area of NFH immunoreactivity (I) for induction from E14.5–P0 (control, N = 32 touch domes; Noggin overexpression, N = 31 touch domes). (J–L) The same measures from litters induced with Dox at E17.5–P0 (control, N = 29 touch domes; Noggin overexpression, N = 33 touch domes). Red lines denote means. Statistical significance was assessed with Student’s t tests (two-tailed). Three animals per group were analyzed. Scale bars: 25 µm.
-
Figure 3—source data 1
Numerical values for data plotted in Figure 3.
- https://doi.org/10.7554/eLife.42633.008
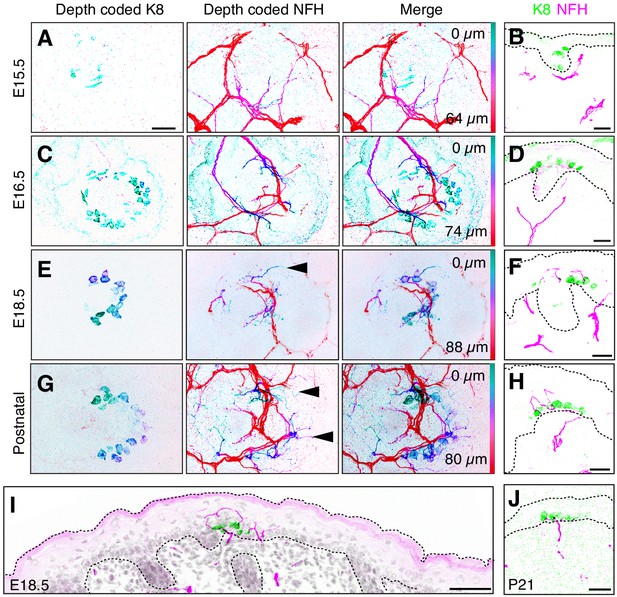
NFH-expressing afferents initiate Merkel-cell contacts at E16.5 and overshoot Merkel cells at E18.5.
(A–D) Depth-coded axial projections of full-thickness dorsal skin. Teal-green structures are located near the skin surface whereas red structures are embedded deep in the dermis. For each stage, the specific Z-depth code is indicated by color scale bars at right. Black scale bar in (A) (25 µm) applies also to (B–D). (A’–D’), (E–F) Sagittal cryosections of developing skin demonstrate the time course of Merkel-cell innervation. K8: green, NFH: magenta. Scale bars: 50 µm. Dashed lines outline the epidermis. All images are representative of data from 2 to 4 animals per stage.
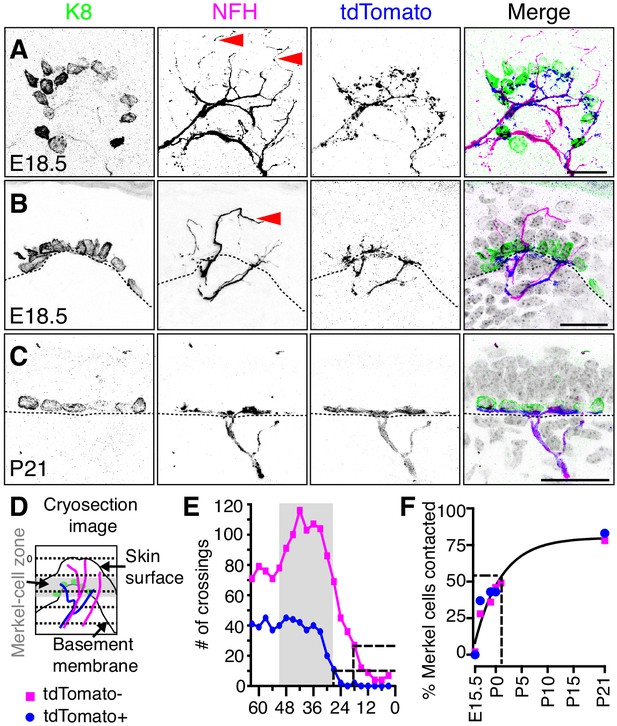
Two molecularly distinct populations of NFH-positive neurons transiently innervate touch domes during development.
(A) Axial projections of full-thickness skin specimens from a Ntrk3tdTomato transgenic reporter mouse at E18.5 reveal that Merkel-cell clusters (left panel and green in merge) are contacted by both tdTomato-negative afferents (arrowheads; second panel and magenta in merge) and tdTomato-positive afferents (third panel and blue in merge). (B) Sagittal sections also show that tdTomato-negative afferents (arrow) innervate superficial layers of touch domes at E18.5. (C) TdTomato and NFH immunoreactivity overlaps at P21. Color scheme in (B–C) is as in A). Dashed lines (B, C) denote the dermal-epidermal border. (D) Schematic of line crossing analysis. A series of lines was overlaid on each image starting from the skin surface and continuing toward the dermis at 3 µm intervals. The number of times each marker (tdTomato or NFH) crossed a line was counted. (E) Quantification of line crossings. Gray box indicates the span of the Merkel-cell bearing epidermal zone. Dashed lines indicate the distance from the surface at which the line crossing curve decays by 75% (N = 50 touch domes from three animals). (F) The proportion of Merkel cells contacted by tdTomato-positive afferent branches (N = 35–387 Merkel cells from 2 to 3 mice per stage) and tdTomato-negative afferent branches (N = 87–521 Merkel cells from 2 to 6 mice per stage) were plotted versus developmental stage. Both datasets were well fit by a single exponential (black line; τ = 5.1 d; R2 = 0.9), which indicates that most Merkel cells are innervated by P1. See also Figure 5—figure supplement 1.
-
Figure 5—source data 1
Numerical values for data plotted in Figure 5.
- https://doi.org/10.7554/eLife.42633.014
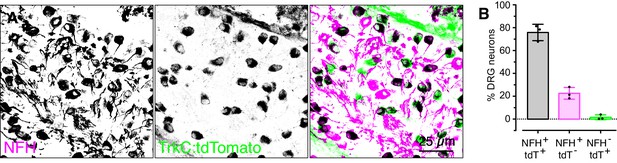
Few tdTomato-positive DRG cell bodies lack NFH expression in E16.5Ntrk3tdTomatomice.
(A) Representative image of E16.5 DRG from Ntrk3tdTomato mice labeled with antibodies against NFH (left panel and magenta in merged images) and tdTomato (middle panel and green in merged images). (B) Quantification of the proportion of NFH-positive, tdTomato-positive and double positive neurons (N = 231–473 neurons quantified per mouse from three mice).
-
Figure 5—figure supplement 1—source data 1
Numerical values for data plotted in Figure 5—figure supplement 1.
- https://doi.org/10.7554/eLife.42633.012
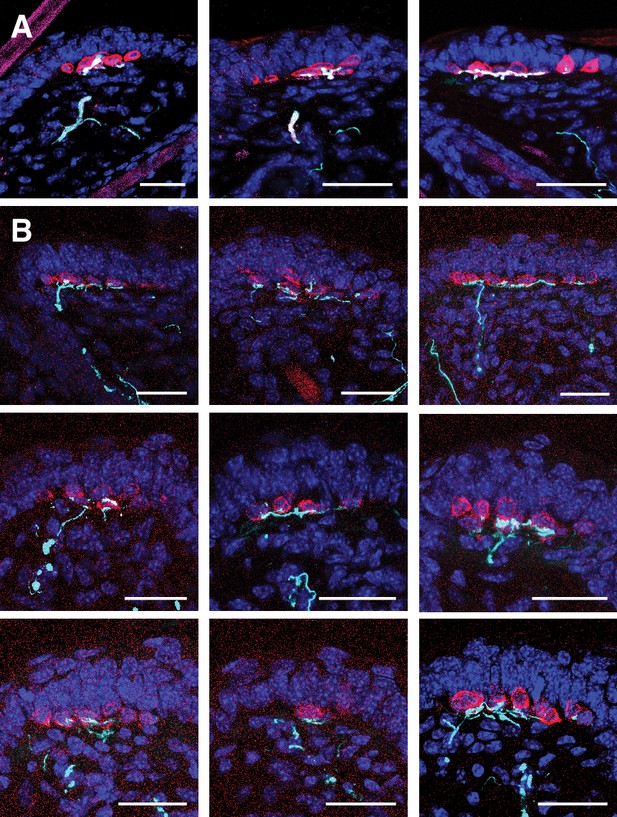
NFH-expressing afferent endings terminate at the basal epidermis at P21.
(A) Images of P21 touch domes (N = 3) from an Ntrk3tdTomato reporter mouse labeled with antibodies against NFH (cyan), K8 (red) and tdTomato (magenta). Another example touch dome from this mouse is shown in Figure 5C. (B) Images of P21 touch domes (N = 6) from a wild-type mouse labeled with antibodies against NFH (cyan) and K8 (red). Another example touch dome from this mouse is shown in Figure 4F.
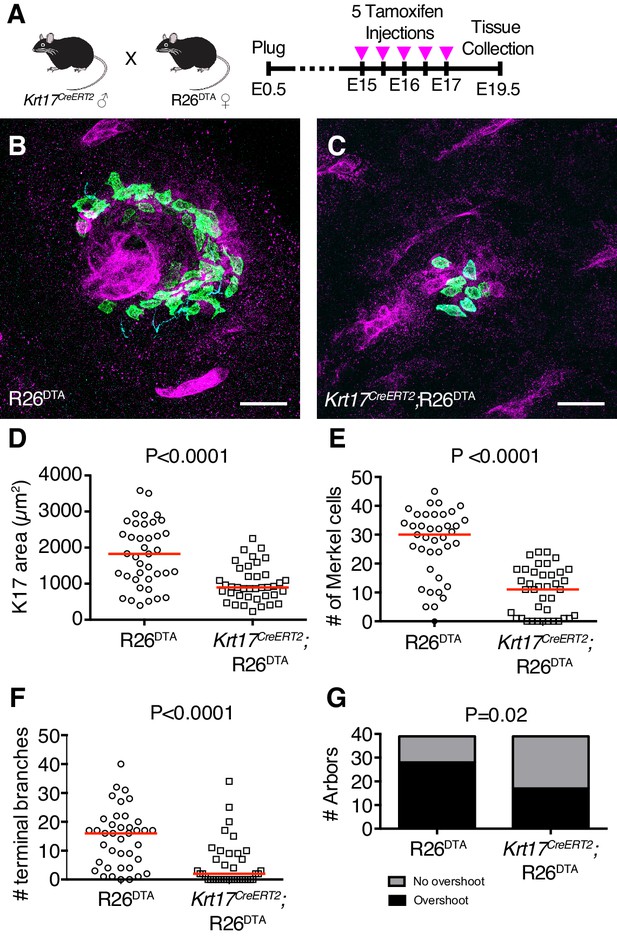
K17-lineage keratinocytes are necessary for patterning touch-dome afferents during embryogenesis.
(A) Experimental design. (B–C) Axial projections of full-thickness E19.5 skin specimens from R26DTA and Krt17CreERT2;R26DTA mice labeled with antibodies against K8 (green), K17 (magenta) and NFH (cyan). (D–G) Quantification of K17 area (D), number of Merkel cells (E), number of terminal branches (F), number of arbors exhibiting sprouting above Merkel cells (G), across genotypes for mice that were administered Tamoxifen from E15–E17. N = 39 touch domes from three animals per group. Red lines denote medians. Scale bars: 25 µm. Statistical significance was assessed with a Student’s t-test in (D), Mann-Whitney tests in (E–F) and a Fisher’s exact test in (G). See also Figure 6—figure supplements 1 and 2.
-
Figure 6—source data 1
Numerical values for data plotted in Figure 6.
- https://doi.org/10.7554/eLife.42633.018
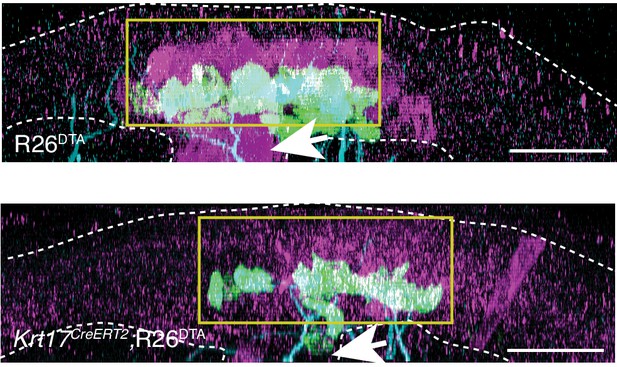
K17-lineage keratinocytes are successfully ablated in embryonic Krt17CreERT2;R26DTA mice.
Sagittal projections of full-thickness E19.5 skin specimens from R26DTA (top) and Krt17CreERT2;R26DTA (bottom) mice labeled with antibodies against K8 (green), K17 (magenta) and NFH (cyan). Yellow boxes outline touch-dome epithelia. Arrows indicate hair follicles. Scale bar :25 µm.
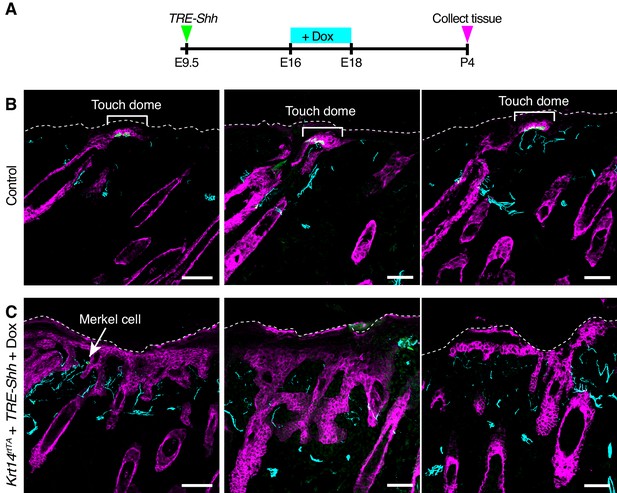
K17-lineage keratinocytes are sufficient to recruit NFH-expressing afferents to the epidermis.
(A) Experimental design. (B–C) Sagittal cryosections of P4 skin specimens from Krt14rtTA + TRE-Shh + Doxycycline and control mice labeled with antibodies against K8 (green), K17 (magenta) and NFH (cyan). Scale bars : 50 µm.
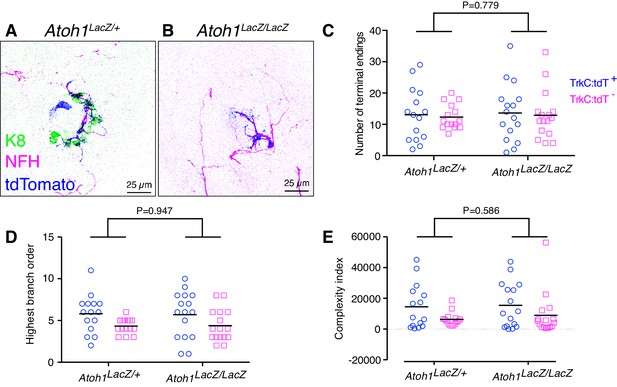
Merkel-cells are not required for embryonic touch-dome innervation.
(A–B) Axial projections of whole-mount immunohistochemistry from control (A) and Atoh1 knockout mice (B). Tissues were labeled with antibodies against K8 (Merkel cells, green), NFH (myelinated neurons, magenta) and tdTomato (Ntrk3-lineage neurons; blue). (C–E) Quantification of terminal branches per touch dome (C), highest branch order (D) and complexity index (E) for tdTomato-expressing and tdTomato-negative afferents (N = 15 control and 16 Atoh1 knockout touch domes from four animals per group). Black lines indicate medians. By two-way ANOVA, the genotype effect was not significantly different for any measure. For the afferent type effect, tdTomato-positive and tdTomato-negative afferents showed significant differences in highest branch order (D, p=0.012) and complexity index (E, p=0.028) but number of terminal endings did not differ (C, p=0.705).
-
Figure 7—source data 1
Numerical values for data plotted in Figure 7.
- https://doi.org/10.7554/eLife.42633.020
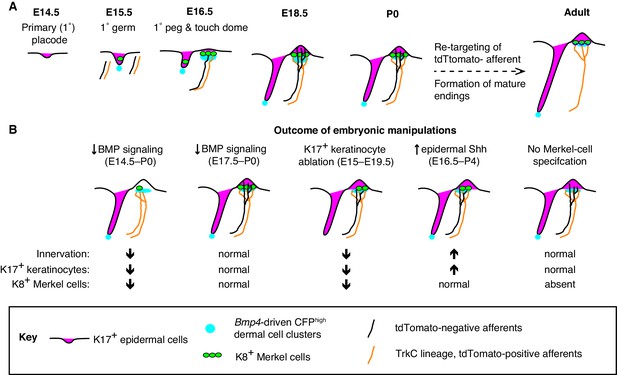
A model of touch-dome innervation during development.
(A) The progression of developing touch domes, including the localization of epidermal K17-positive cells (magenta) and Merkel cells (green), CFPhigh dermal cells (cyan) and NFH-positive sensory afferents (black, non-TrkC lineage and orange, TrkC lineage). (B) Summary of experimental manipulations of touch-dome cell types and BMP signaling.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (M. musculus) | Bmp4CFP | PMID: 20336610 | MGI: 4460795 | MGI symbol: Bmp4-tm5.1Bhr; R. Berhinger laboratory, MD Anderson |
| Gene (M. musculus) | Ntrk3tdTomato | PMID: 26687362 | MGI: 5707201; Jax: 030292 | MGI symbol: Ntrk3-tm2.1Ddg; D. Ginty laboratory, Harvard Univ. |
| Gene (M. musculus) | Atoh1LacZ | PMID: 10662643 | MGI: 2155983; Jax: 005970 | MGI symbol: Atoh1-tm2Hzo; H. Zoghbi laboratory; Baylor College of Medicine |
| Gene (M. musculus) | Krt14rtTA | PMID: 17018284 | MGI: 3762564; Jax: 008099 | MGI symbol: Tg(KRT14-rtTA)F42Efu; Fuchs laboratory |
| Gene (M. musculus) | Krt17CreERT2 | PMID: 23727240 | MGI:5523315 | MGI symbol: Tg(Krt17-cre/ERT2)1F3Dmo; D. Owens; laboratory, Columbia Univ. |
| Gene (M. musculus) | Krt14Cre | Jackson Labs; PMID: 11044393 | MGI: 2445832; Jax: 018964 | MGI symbol: Tg(KRT14-cre)1Amc |
| Gene (M. musculus) | R26DTA | Jackson Labs; PMID: 16407399 | MGI:4440748; Jax: 010527 | MGI symbol: B6; 129-Gt(ROSA) 26Sortm1(DTA)Mrc/J |
| Antibody | anti-Keratin-8 (K8; rat monoclonal) | Devel. Studies Hybridoma Bank | Troma-I; RRID: AB_531826 | Deposited to DSHB by Brulet, P. and Kemler, R. |
| Antibody | anti-Neurofilament-Heavy (NFH; chicken polyclonal) | Abcam | Cat# 4680; RRID: AB_304560 | (1:3000 in cryosections; 1:500 in whole mount preps) |
| Antibody | anti-dsRed (tdTomato; rabbit polyclonal) | Clontech | Clontech: 632496; RRID:AB_10013483 | (1:1000 in cryosections; 1:500 in whole mount preps) |
| Antibody | anti-Keratin-17 (K17; rabbit monoclonal) | Abcam | Cat# 109725; RRID: AB_10889888 | (1:200 in cryosections; 1:200 in whole mount preps) |
| Antibody | anti-Green Fluorescent Protein (GFP; chicken polyclonal) | Abcam | Cat# 13970; RRID: AB_300798 | (1:1000 in cryosections; 1:500 in whole mount preps) |
| Antibody | Goat polyclonal anti-rat IgG (H + L) Alexa-594 | ThermoFisher | Cat# A-11007; RRID: AB_10561522 | (1:1000 in cryosections; 1:500 in whole mount preps) |
| Antibody | Goat polyclonal anti-rat IgG (H + L) Alexa-488 | ThermoFisher | Cat# A-11006; RRID: AB_2534074 | (1:1000 in cryosections; 1:500 in whole mount preps) |
| Antibody | Goat polyclonal anti-chicken IgY (H + L) Alexa-647 | ThermoFisher | Cat# A-21449; RRID: AB_2535866 | (1:1000 in cryosections; 1:500 in whole mount preps) |
| Antibody | Goat polyclonal anti-chicken IgY (H + L) Alexa-488 | ThermoFisher | Cat# A-11039; RRID: AB_2534096 | (1:1000 in cryosections; 1:500 in whole mount preps) |
| Antibody | Goat polyclonal anti-rabbit IgG (H + L) Alexa-594 | ThermoFisher | Cat# A-11037; RRID: AB_2534095 | (1:1000 in cryosections; 1:500 in whole mount preps) |
| Antibody | Goat oligoclonal anti-rabbit IgG (H + L) Alexa-647 | ThermoFisher | Cat# A-27040; RRID: AB_2536101 | (1:1000 in cryosections; 1:500 in whole mount preps) |
| Genetic reagent | pLKO-TRE-Nog- PGK-H2BGFP | PMID: 28008008 | Fuchs laboratory | |
| Genetic reagent | pLKO-TRE-Shh- PGK-H2BGFP | PMID: 24813615 | Fuchs laboratory |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.42633.022





