eIF2B activator prevents neurological defects caused by a chronic integrated stress response
Figures
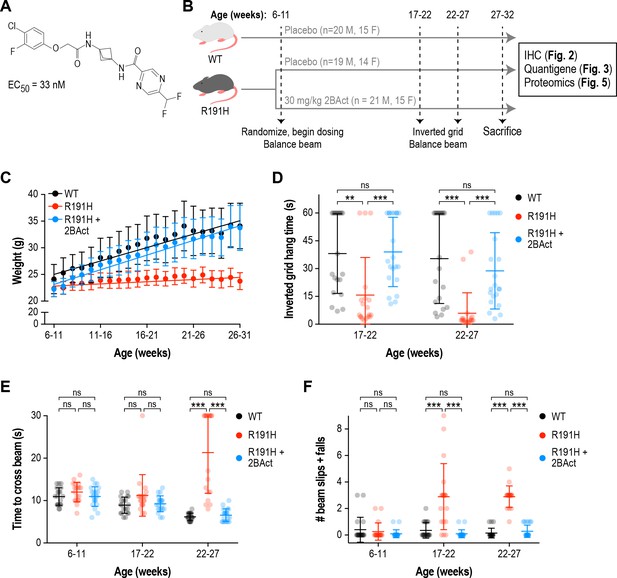
2BAct normalized body weight gain and prevented motor deficits in male R191H mice.
(A) Chemical structure of 2BAct and ATF4-luciferase reporter assay EC50. (B) Schematic of the 2BAct treatment experiment. Body weights were measured weekly for the duration of the experiment. (C) Body weight measurements of male mice along the study. Lines are linear regressions. At the 6–11 week time point when 2BAct treatment was initiated, body weights were not significantly different among the three conditions (p>0.05; two-way ANOVA with Holm-Sidak pairwise comparisons). R191H body weight was significantly lower at all subsequent time points. 2BAct-treated animals caught up to WT animals at the 8–13 week time point, and their weights were not significantly different thereafter. (D) Inverted grid test of muscle strength. Time spent hanging was measured up to a maximum of 60 s. (E–F) Beam-crossing assay to measure balance and motor coordination. Time to cross the beam was measured up to a maximum of 30 s (E), and the number of foot slips/falls was counted (F). For (C)-(F), N = 20 (WT), 19 (R191H) and 21 (R191H + 2 BAct) males were analyzed. Error bars are SD. For (D)-(F), *p<0.05; **p<0.01; ***p<10-3; nsp>0.05 by Mann-Whitney test with Bonferroni correction.
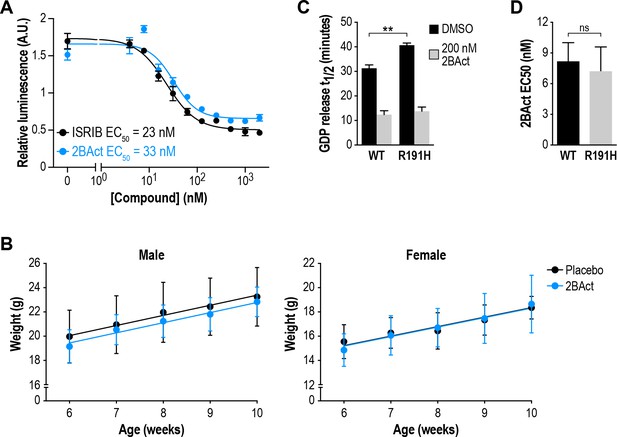
2BAct is an eIF2B activator with similar potency to ISRIB.
(A) Dose response curves of 2BAct and ISRIB in an ATF4-luciferase reporter assay of ISR activation in HEK293T cells. Cells were treated with 100 nM thapsigargin and increasing doses of 2BAct or ISRIB for 7 hr (N = 4, mean ±SD). (B) Body weight measurements of WT mice treated with placebo or 2BAct for 4 weeks, beginning at 6 weeks of age (N = 6/condition/sex). Lines are linear regressions. Weight gain was not significantly different between the treatment conditions (p>0.48; treatment factor in two-way ANOVA). (C) Half-lives of GDP release from eIF2 stimulated by WT and R191H mouse embryonic fibroblast lysates. R191H had significantly reduced GEF activity (p<0.01; two-tailed t-test) that was enhanced by 2BAct. (D) 2BAct EC50 values calculated from dose-response measurements in GEF assays. WT and R191H were not significantly different (p>0.05; two-tailed t-test). For (C) and (D), each bar represents 12 measurements from four independent experiments (mean ±SD).
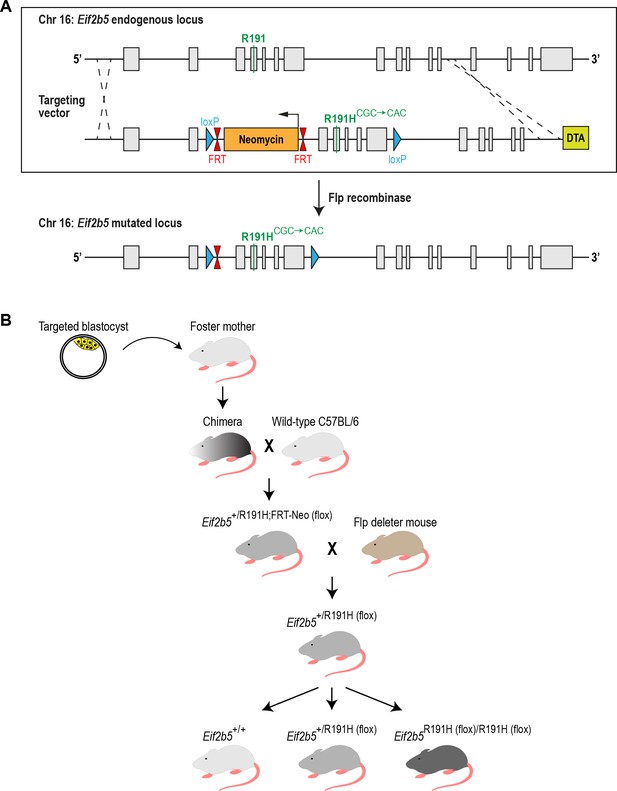
Generation of the R191H (Eif2b5R191H (flox)/R191H (flox)) mouse model.
(A) Schematic of the mouse Eif2b5 locus and the targeting vector used. Gray bars, exons; black lines, introns; DTA, diphtheria toxin negative selection cassette. A single base point mutation (CGC to CAC) in exon four changed Arg191 to His. Excision of the neomycin selection cassette by Flp recombinase left behind a single FRT site and two loxP sites flanking exons 3–7. (B) Breeding scheme used to generate homozygous R191H mice. Experiments were performed with homozygous mice from heterozygous parents, and their WT littermates as controls.
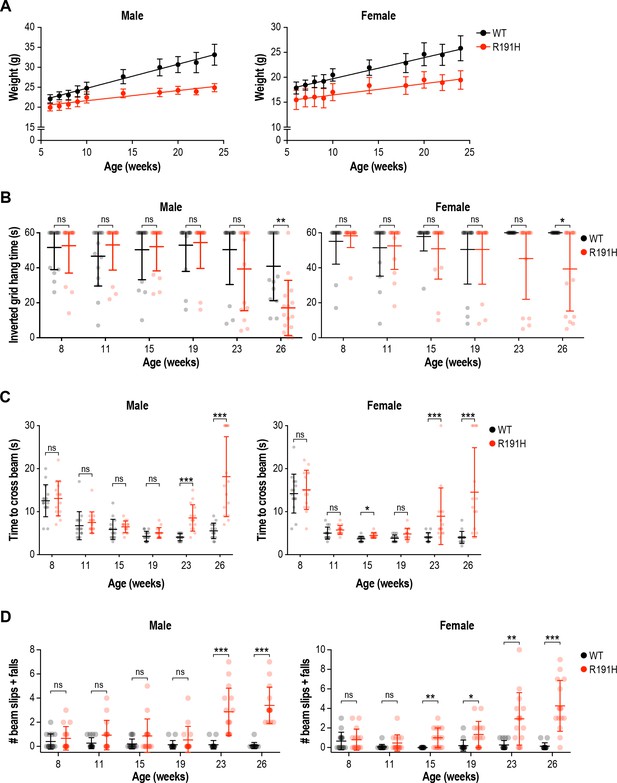
R191H mice exhibited reduced body weight and age-dependent performance impairment in motor assays.
(A) Body weight measurements along the study. Lines are linear regressions. 5-week-old R191H mice had significantly lower body weight compared to WT littermates, and the weight difference persisted as animals aged. In both males and females, weights were significantly different at all time points (p<0.01; two-way ANOVA with Holm-Sidak pairwise comparisons). (B) Inverted grid test of grip strength. At week 26, the difference in hang time between WT and R191H was statistically significant. (C–D) Beam-crossing assay to measure motor coordination. Motor deficits were not evident in R191H mice at early time points (8–19 weeks), but became apparent in older animals (23 and 26 weeks). At the later time points, R191H animals took significantly longer to cross the beam (C) and suffered more slips and falls while crossing (D). For all panels, n = 15/genotype were analyzed. Error bars are SD. *p<0.05; **p<0.01; ***p<10-3; nsp>0.05 by Mann-Whitney test with Bonferroni correction.
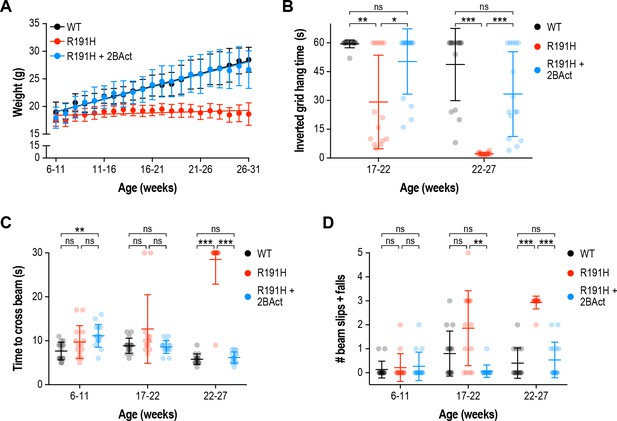
2BAct normalized body weight gain and prevented motor deficits in female R191H mice.
(A) Body weight measurements along the study. Lines are linear regressions. At the 6–11 week time point when 2BAct treatment was initiated, body weights were not significantly different between the three conditions (p>0.05; two-way ANOVA with Holm-Sidak pairwise comparisons). R191H body weight was significantly lower at all subsequent time points. The body weights of 2BAct-treated R191H females were not significantly different from WT at any time point. (B) Inverted grid test of muscle strength. Time spent hanging was measured up to a maximum of 60 s. (C–D) Beam-crossing assay to measure balance and motor coordination. Time to cross the beam was measured up to a maximum of 30 s (C), and the number of foot slips/falls was counted (D). For all panels, N = 15 (WT), 14 (R191H) and 15 (R191H + 2 BAct) females were analyzed. Error bars are SD. For (B)-(D), *p<0.05; **p<0.01; ***p<10-3; nsp>0.05 by Mann-Whitney test with Bonferroni correction.
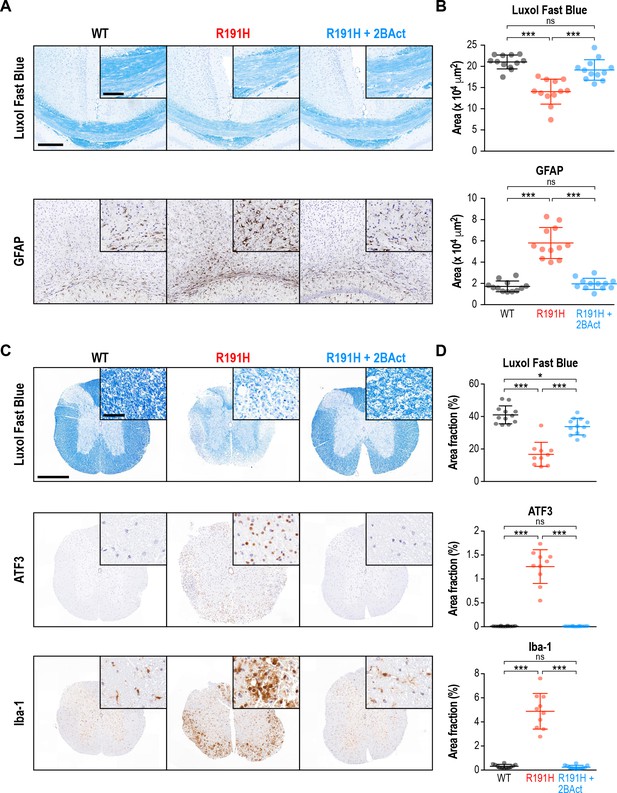
2BAct prevented myelin loss and reactive gliosis in the brain and spinal cord of R191H mice.
(A) Representative IHC images of the corpus callosum. Scale bar, 250 μm. Inset is magnified 2X. Inset scale bar, 100 μm. (B) Quantification of staining in (A). Area of positive staining expressed as μm2. (C) Representative IHC images of the lower cervical/upper thoracic region of the spinal cord. Scale bar, 500 μm. Inset is magnified 6.8X. Inset scale bar, 50 μm. (D) Quantification of staining in (C). For (B) and (D), N = 12 mice/condition (6 males and six females combined; no significant sex differences were detected). Error bars are SD. *p<0.05; ***p<10-4; nsp>0.05 by 1-way ANOVA with Holm-Sidak pairwise comparisons.
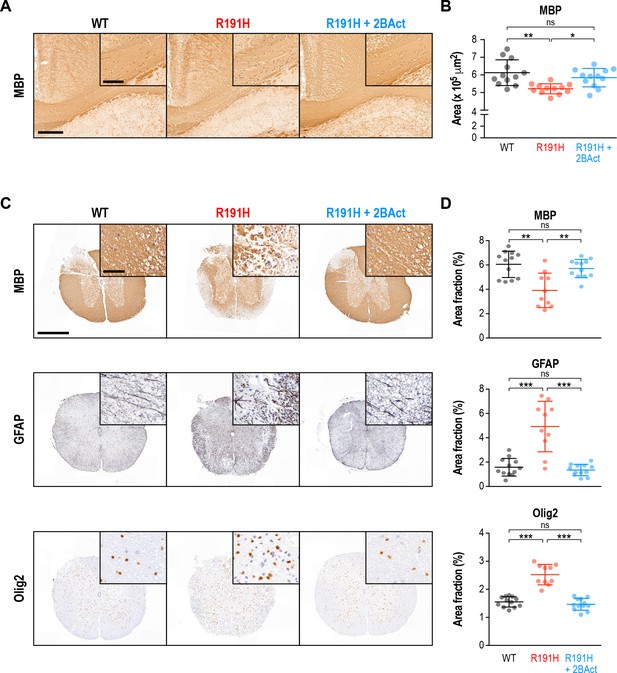
2BAct prevented myelin loss and reactive gliosis in the brain and spinal cord of R191H mice.
(A) Representative MBP staining in the corpus callosum. Scale bar, 250 μm. Inset is magnified 2X. Inset scale bar, 100 μm. (B) Quantification of staining in (A). Area of positive staining expressed as μm2. (C) Representative IHC images of the lower cervical/upper thoracic region of the spinal cord. Scale bar, 500 μm. Inset is magnified 6.8X. Inset scale bar, 50 μm. (D) Quantification of staining in (C). For (B) and (D), N = 12 mice/condition (6 males and six females combined; no significant sex differences were observed). Error bars are SD. *p<0.05; **p<10−3; ***p<10-4; nsp>0.05 by 1-way ANOVA with Holm-Sidak pairwise comparisons.
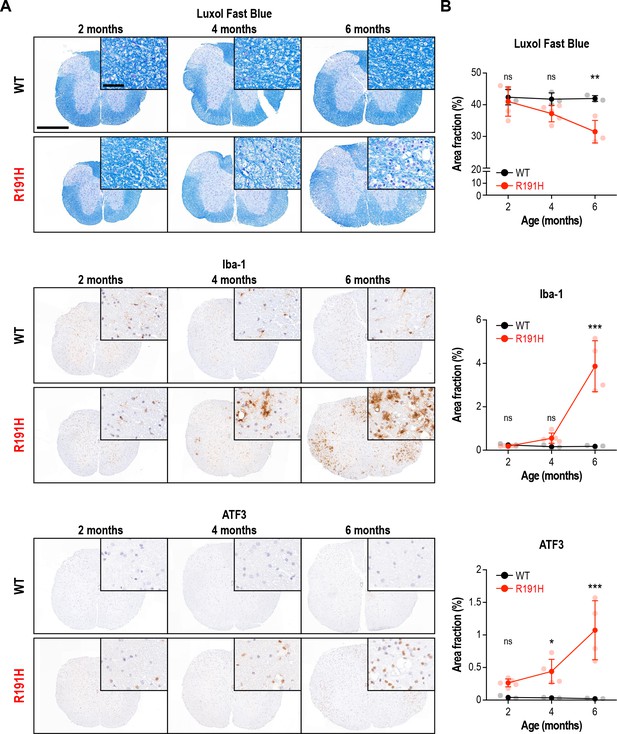
Age-dependent myelin loss and inflammation in the spinal cord of R191H mice.
(A) Representative IHC images of the lower cervical/upper thoracic region of the spinal cord of female mice at 2, 4 and 6 months of age. Scale bar, 500 μm. Inset is magnified 6.8X. Inset scale bar, 50 μm. (B) Quantification of staining in (A). N = 3 (2 mo WT), 5 (2 mo R191H), 3 (4 mo WT), 5 (4 mo R191H), 3 (6 mo WT), 4 (6 mo R191H) females. Error bars are SD. *p<0.05; ***p<10-4; nsp>0.05 by 2-way ANOVA with Holm-Sidak correction.
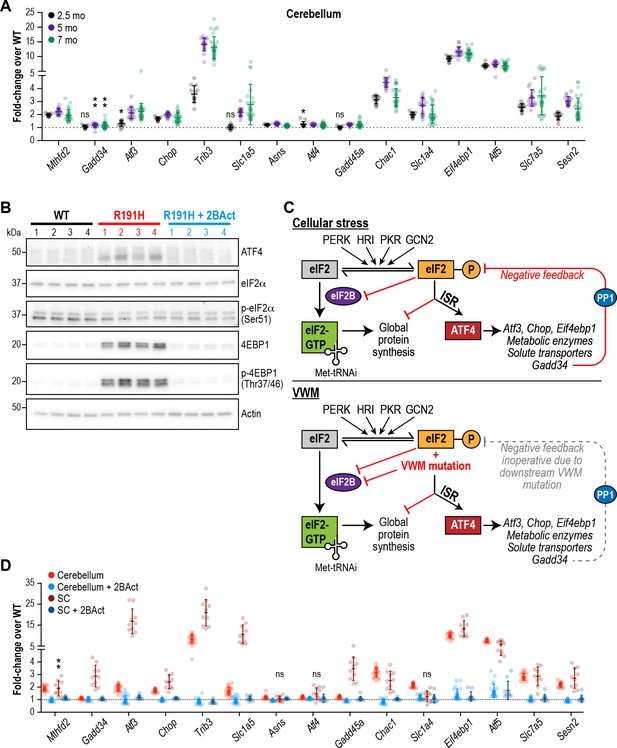
The ISR is activated in the brain of R191H mice and its induction is prevented by 2BAct.
(A) mRNA expression in R191H cerebellum at 2.5 (N = 13/genotype), 5 (N = 20 WT, 19 R191H) and 7 (N = 30/genotype) months of age. (B) Western Blots of the indicated proteins from 7-month-old male cerebellum. Actin was used as a loading control. Each lane represents an individual animal. (C) Schematic of ISR activation in the context of external stressors or VWM. PP1, protein phosphatase 1. Gadd34, an ATF4-induced regulatory subunit of PP1 that targets it to eIF2. (D) mRNA expression in R191H cerebellum (N = 23 WT, 21 R191H, 24 R191H + 2 BAct) and spinal cord (N = 10/condition) at 27–32 weeks of age from the 2BAct treatment study (Figure 1B). For (A) and (D), males and females were combined as there was no significant difference between sexes. Data are shown normalized to WT transcript levels. Bars, mean ±SD. *p<0.01; **p<10-3; nsp>0.05 by Student’s t-test with Holm-Sidak correction (compared to WT). Transcripts without symbols were highly significant with p<10−4. 2BAct treatment was highly significant for all transcripts (p<0.01 vs. placebo treatment). A table of p-values from tests is available in Figure 3—source data 1.
-
Figure 3—source data 1
Adjusted p-values from t-tests of multiplex transcript expression quantification.
- https://doi.org/10.7554/eLife.42940.015
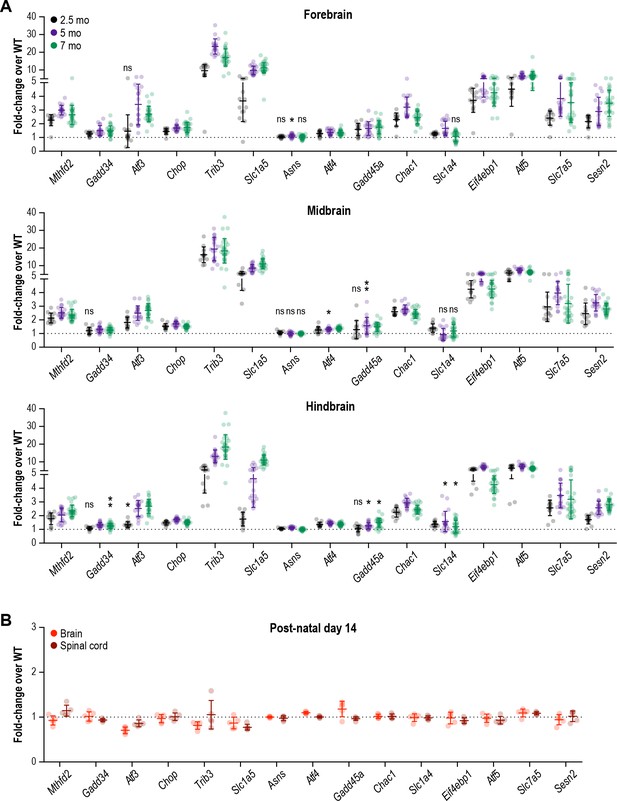
A robust and chronic ISR is triggered in all regions of the R191H mouse brain by 2.5 months.
(A) mRNA expression in R191H brain regions at 2.5 (N = 13/genotype), 5 (N = 20 WT, 19 R191H) and 7 (N = 30/genotype) months of age. *p<0.01; **p<10-3; nsp>0.05 by Student’s t-test with Holm-Sidak correction (compared to WT). Transcripts without symbols were highly significant with p<10−4. (B) mRNA expression in R191H whole brain and spinal cord at postnatal day 14. N = 5/genotype. None of the transcripts were significantly different from WT (p>0.1 by Student’s t-test with Holm-Sidak correction). For (A) and (B), males and females were combined as there was no significant difference between sexes. Data are shown normalized to WT transcript levels. Bars, mean ±SD. A table of p-values from tests is available in Figure 3—source data 1.
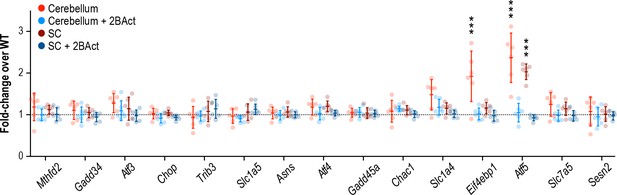
2BAct prevents ISR induction in the cerebellum and spinal cord of Eif2b5R132H/R132H mice.
mRNA expression in Eif2b5R132H/R132H cerebellum and spinal cord (N = 6/condition) at 12–14 weeks of age. 2BAct treatment was initiated at 8–10 weeks of age. Data are shown normalized to WT transcript levels. Bars, mean ±SD. ***p<10−5 by Student’s t-test with Holm-Sidak correction (compared to WT). Transcripts without symbols were not significant (p>0.05). A table of p-values from tests is available in Figure 3—source data 1.
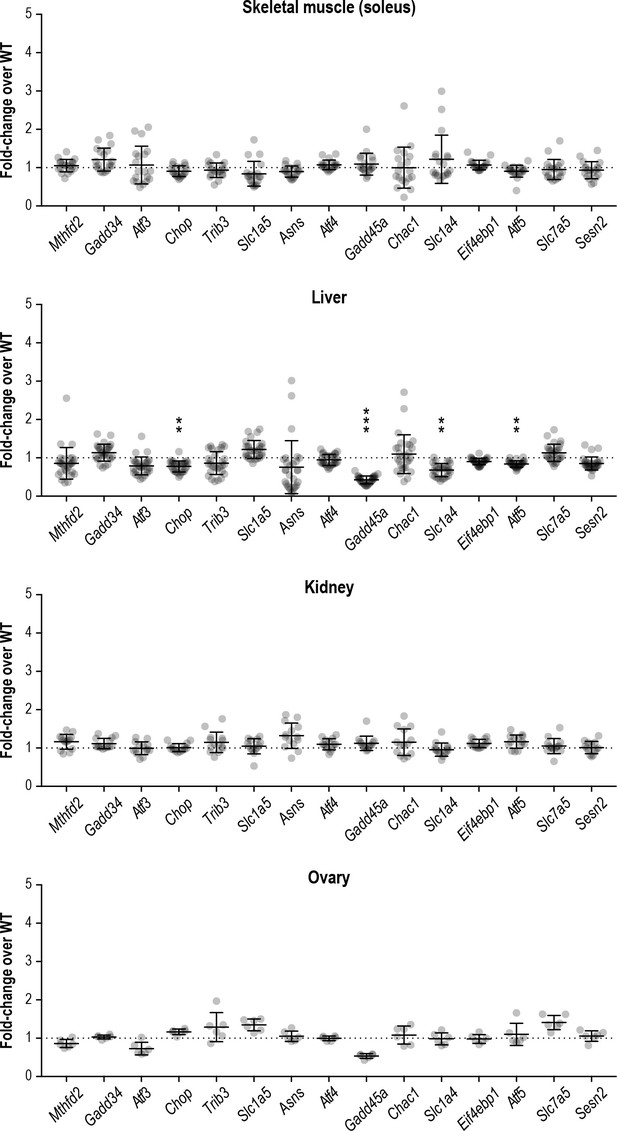
The ISR is not induced in peripheral organs of R191H mice.
mRNA expression in R191H soleus muscle (N = 20 WT, 19 R191H), liver (N = 28 WT, 30 R191H), ovary (N = 6/genotype) and kidney (N = 14/genotype) at 7, 7, 10 and 2.5 months of age, respectively. Males and females were combined as there was no significant difference between sexes. Data are shown normalized to WT transcript levels. Bars, mean ±SD. **p<10−3; ***p<10−5 by Student’s t-test with Holm-Sidak correction (compared to WT). Transcripts without symbols were not significant (p>0.05). A table of p-values from tests is available in Figure 3—source data 1.
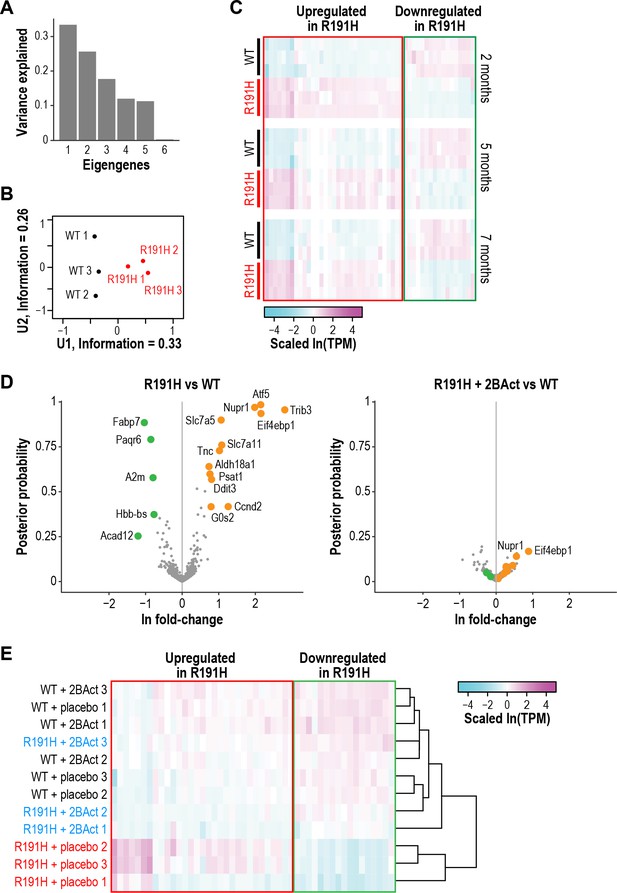
R191H mice have an abnormal brain transcriptome at 2 months of age that is normalized by 2BAct treatment.
(A) Scree plot showing the variance explained by each component of the SVD analysis of 2-month-old WT and R191H cerebellum. (B) Individual 2-month-old cerebellum samples plotted along the first and second components of SVD analysis. (C) Heatmap of gene expression changes in WT and R191H cerebellum at 2, 5, and 7 months of age (N = 3/genotype/time point). Shown are the 50 genes with the largest absolute loadings in the first eigengene from SVD analysis of 2 month samples. Source data for (A) -(C) are available in Figure 4—source data 1. (D) Volcano plots showing gene expression changes between R191H and WT (left) and R191H + 2 BAct and WT (right). Orange and green dots indicate transcripts that were more than 2X increased or decreased, respectively, in the R191H vs. WT plot. These dots are replicated on the R191H + 2 BAct vs. WT plot for comparison. (E) Heatmap of gene expression changes in WT and R191H cerebellum treated with placebo or 2BAct for 4 weeks. Genes are the same set plotted in Figure 4C. Colors indicate the scaled ln(TPM) from the mean abundance of the gene across all samples. For (D) and (E), N = 3/condition. Source data for (D) and (E) are available in Figure 4—source data 2.
-
Figure 4—source data 1
Fold-changes of transcripts identified in RNA-seq of 2-, 5- and 7-month-old cerebellum.
- https://doi.org/10.7554/eLife.42940.019
-
Figure 4—source data 2
Fold-changes of transcripts identified in RNA-seq in the 4-week 2BAct treatment experiment.
- https://doi.org/10.7554/eLife.42940.020
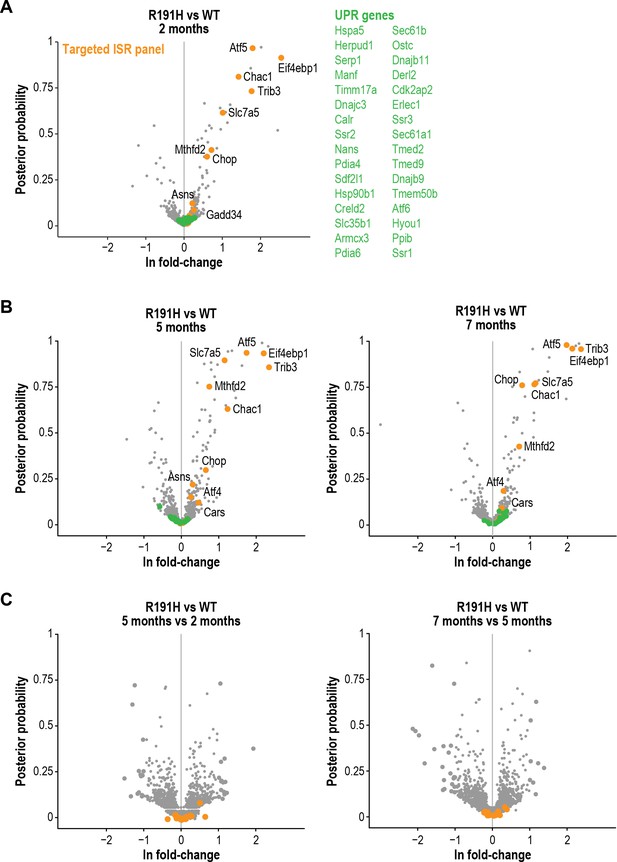
Sustained ISR induction is a feature of R191H cerebellum across different ages.
(A) Volcano plot showing gene expression changes between R191H and WT cerebellum at 2 months of age. Orange dots indicate genes from the targeted ISR panel. Green dots indicate targets of the IRE1α and ATF6 branches of the UPR. (B) Volcano plots showing gene expression changes between R191H and WT cerebellum samples at 5 (left) and 7 (right) months of age. Orange dots indicate genes from the targeted ISR panel. Green dots indicate targets of the IRE1α and ATF6 branches of the UPR. (C) Volcano plots showing gene expression changes between the R191H vs. WT cerebellum comparisons at different ages (5 mo vs. two mo, left; 7 mo vs. five mo, right). Orange dots indicate genes from the targeted ISR panel. For all panels, N = 3/genotype/time point. Source data are available in Figure 4—source data 1.
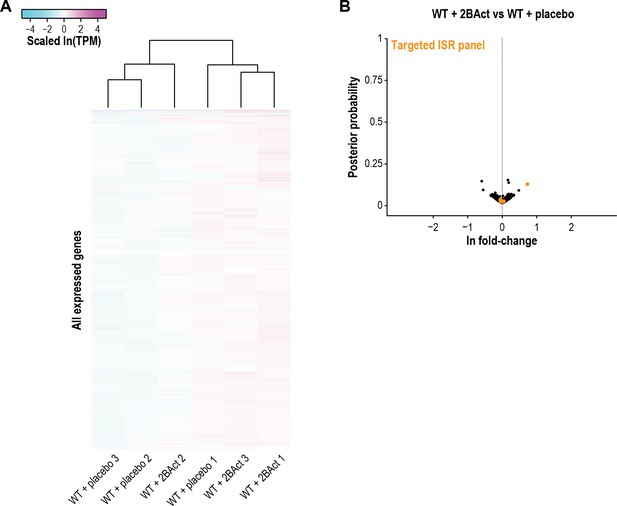
2BAct does not elicit spurious gene changes in WT mice.
(A) Heatmap of gene expression changes in WT cerebellum from mice treated with placebo or 2BAct for 4 weeks. The entire analyzed transcriptome is shown. Placebo- and 2BAct-treated samples did not cluster away from each other. Colors indicate the scaled ln(TPM) from the mean abundance of the gene across all samples. Scale limits are the same as shown in Figure 4C and E. (B) Volcano plot showing gene expression changes between 2BAct-treated and placebo-treated WT cerebellum. Orange dots indicate genes from the targeted ISR panel. For (A) and (B), N = 3/condition. Source data are available in Figure 4—source data 2.
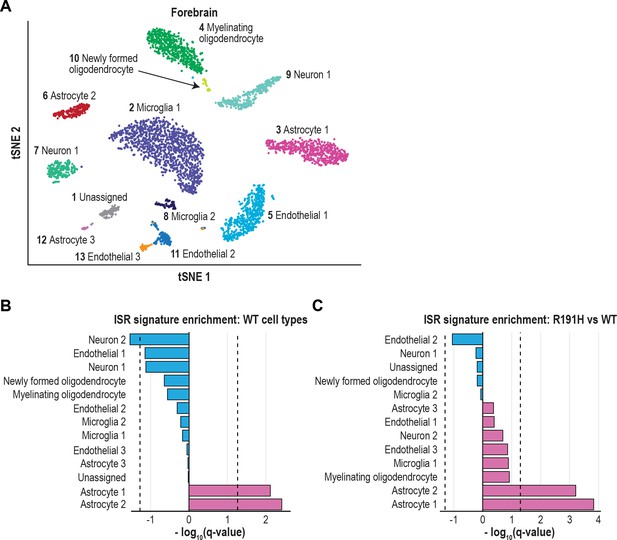
The ISR is strongly activated in astrocytes of VWM forebrain.
(A) tSNE plot showing the 13 transcriptionally defined clusters identified from single-cell analysis of WT and R191H forebrain. (B) Q-values from GSEA based on the differential expression analysis of each WT cluster versus all other WT clusters, using the ISR gene expression signature derived from CLIC. (C) Q-values from the differential expression analysis of R191H vs WT cells for each cluster, using the same ISR signature as in (B). In (B) and (C), dotted lines indicate Q-value thresholds of 0.05.
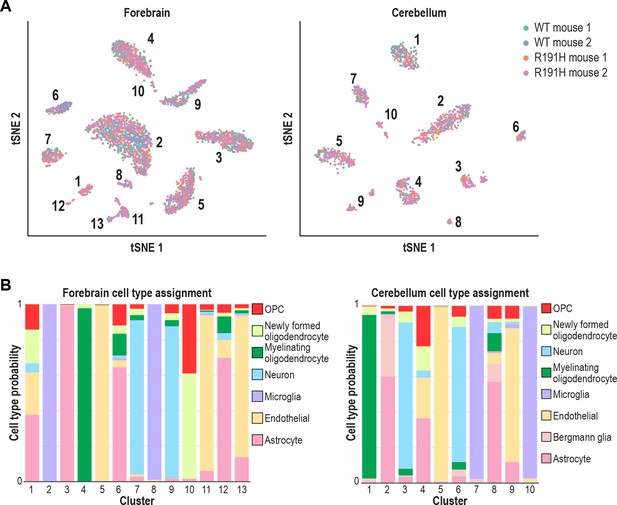
scRNA-seq of WT and R191H forebrain and cerebellum yields distinct transcriptionally defined clusters that do not depend on genotype.
(A) tSNE plots showing the cell type clusters identified from scRNA-seq analysis of WT and R191H forebrain and cerebellum. Points represent individual cells color-coded by source animal. (B) Probability of each cluster corresponding to a cell type of defined identity based on published gene signatures (Koirala and Corfas, 2010; Zhang et al., 2014). Each cluster was assigned a single cell type label if any individual probability exceeded 0.5.
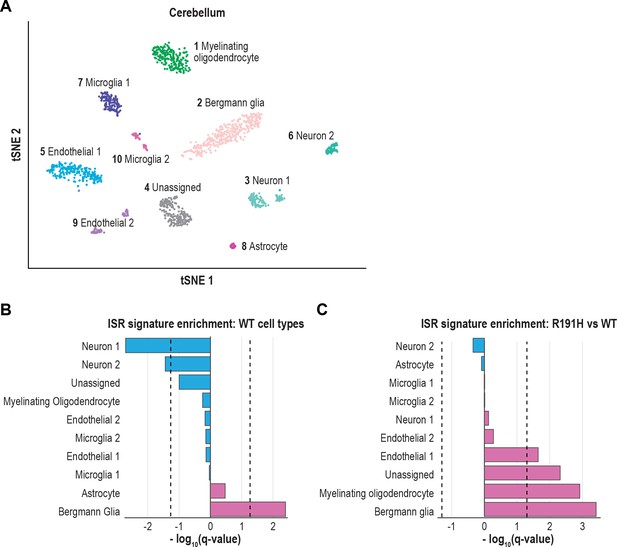
The ISR is strongly activated in Bergmann glia of VWM cerebellum.
(A) tSNE plot showing the 10 cell type clusters identified from scRNA-seq analysis of WT and R191H cerebellum. (B) Q-values from GSEA based on the differential expression analysis of each WT cluster versus all other WT clusters, using the ISR gene expression signature derived from CLIC. (C) Q-values from the differential expression analysis of R191H vs WT cells for each cluster, using the same ISR signature as in (B). In (B) and (C), dotted lines indicate Q-value thresholds of 0.05.
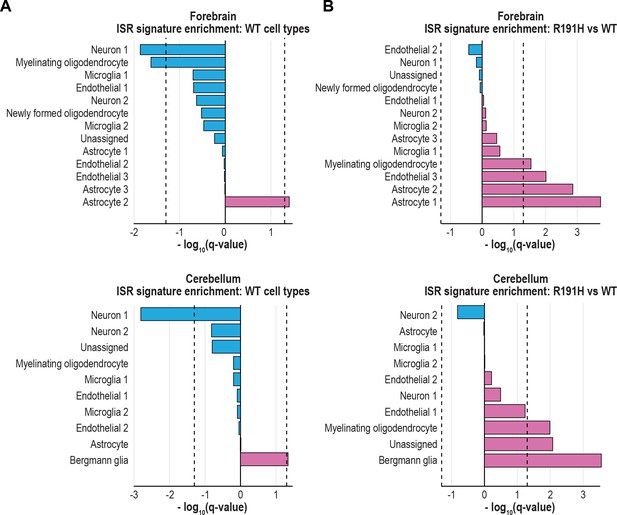
A curated list of ISR targets reveals activation in astrocytes and Bergmann glia of VWM brain.
(A) Q-values from GSEA based on the differential expression analysis of each WT cluster versus all other WT clusters, using the ISR gene expression signature derived from the curated ISR target list. (B) Q-values from the differential expression analysis of R191H vs WT cells for each cluster, using the same ISR signature as in (A). In (A) and (B), dotted lines indicate Q-value thresholds of 0.05.
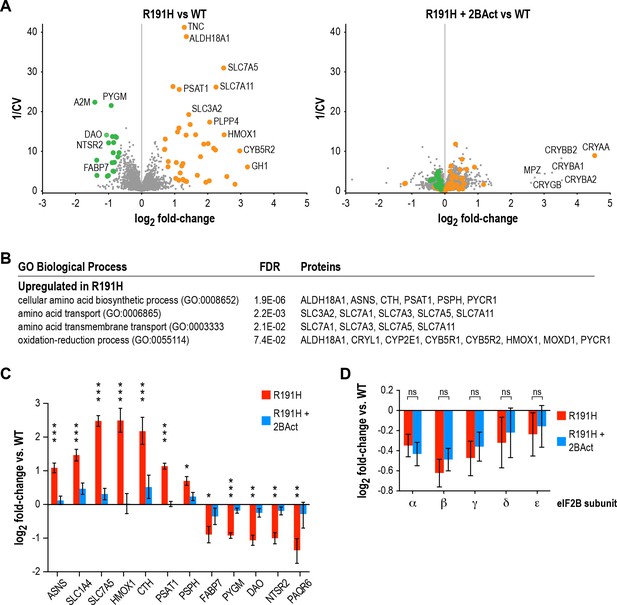
2BAct normalizes the R191H brain proteome without affecting eIF2B subunit levels.
(A) Volcano plots showing protein abundance changes between R191H and WT (left) and R191H + 2 BAct and WT (right). The y-axis is the inverse of the coefficient of variation. Orange and green dots indicate proteins that were more than 1.5X increased or decreased, respectively, in the R191H vs. WT plot at a posterior probability of >90%. These dots are replicated on the R191H + 2 BAct vs. WT plot for comparison. (B) GO-term enrichment analysis of proteins meeting the threshold for increase in (A). Categories shown have an FDR cutoff smaller than 10−2. Downregulated proteins did not show enrichment for any categories. (C) Quantification of selected ISR, metabolic and neural targets relative to WT levels, showing rescue by 2BAct treatment. Posterior probability *<0.05; **<10−5; ***<10−10 of a <50% difference compared to WT. All targets in the 2BAct-treated condition had posterior probability >0.5 of a <50% difference compared to WT. (D) Quantification of eIF2B subunits relative to WT levels. Posterior probability >0.95 of a <25% difference between placebo-treated and 2BAct-treated conditions. For all panels, N = 6/condition. For (B) and (C), bars are mean ±95% credible intervals. Source data are available in Figure 6—source data 1.
-
Figure 6—source data 1
Fold-changes of proteins identified in TMT-MS proteomics experiment.
- https://doi.org/10.7554/eLife.42940.026
Tables
Measured parameters in R191H mice and effect of 2BAct.
https://doi.org/10.7554/eLife.42940.027| Parameter | R191H phenotype | Effect of 2BAct |
|---|---|---|
| Physiological | ||
| Body weight | Reduced | Normalized |
| Inverted grid test | Reduced hang time | Normalized |
| Balance beam test | Longer crossing time, more errors | Normalized |
| Histological | ||
| Myelin levels | Reduced | Normalized |
| GFAP staining | Increased | Normalized |
| Iba-1 staining | Increased | Normalized |
| ATF3 staining | Increased | Normalized |
| Olig2 staining | Increased | Normalized |
| Molecular | ||
| ATF4 expression | Increased | Normalized |
| ISR target genes expression | Increased | Normalized |
| Transcriptome | Deregulated | Normalized |
| Proteome | Deregulated | Normalized |
| eIF2B complex levels | Reduced | No effect |
| eIF2B specific activity | Reduced | Increased |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (M. musculus) | R191H VWM mouse model | this paper | Eif2b5R191H/R191H mutation in C57BL/6J background | |
| Genetic reagent (M. musculus) | R132H VWM mouse model | this paper | Eif2b5R132H/R132H mutation in C57BL/6J background | |
| Chemical compound, drug | 2BAct | this paper | Synthesized in-house | |
| Cell line (H.sapiens) | HEK293T with ATF4-Luc reporter | PMID: 23741617 | ||
| Commercial assay or kit | Quantigene Plex 2.0 assay | Thermo Fisher Scientific | Custom gene panel | |
| Commercial assay or kit | ONE-GLO luciferase assay system | Promega | #E6120 | |
| Antibody | Rabbit polyclonal anti-MBP | abcam | #ab40390 (RRID:AB_11141521) | IHC 5 ug/ml; epitope retrieval with pepsin pH 2.3, 10–20 min |
| Antibody | Mouse anti-GFAP | Millipore | #MAB3402 (RRID:AB_94844) | IHC 1 ug/ml; epitope retrieval with citrate pH 6, 95C, 30 min |
| Antibody | Rabbit polyclonal anti-Iba1 | Wako Chemicals | #019–19741 (RRID:AB_839504) | IHC 1 ug/ml; epitope retrieval with citrate pH 6, 95C, 30 min |
| Antibody | Rabbit monoclonal anti-ATF3 | abcam | #ab207434 (RRID:AB_2734728) | IHC 4 ug/ml; epitope retrieval with EDTA pH 9, 95C, 30 min |
| Antibody | Rabbit monoclonal anti-Olig2 | abcam | #ab109186 (RRID:AB_10861310) | IHC 0.3 ug/ml; epitope retrieval with EDTA pH 9, 95C, 30 min |
| Antibody | Rabbit monoclonal anti-ATF4 | Cell Signaling Technology | #11815 (RRID:AB_2616025) | Western Blot (1:1000 dilution) |
| Antibody | Rabbit polyclonal anti-eIF2α | Cell Signaling Technology | #9722 (RRID:AB_2230924) | Western Blot (1:1000 dilution) |
| Antibody | Rabbit monoclonal anti-phospho-eIF2α | Cell Signaling Technology | #3398 (RRID:AB_2096481) | Western Blot (1:1000 dilution) |
| Antibody | Rabbit monoclonal anti-4EBP1 | Cell Signaling Technology | #9644 (RRID:AB_2097841) | Western Blot (1:1000 dilution) |
| Antibody | Rabbit monoclonal anti-phospho- 4EBP1 | Cell Signaling Technology | #2855 (RRID:AB_560835) | Western Blot (1:1000 dilution) |
| Antibody | Mouse monoclonal anti-actin | Cell Signaling Technology | #3700 (RRID:AB_2242334) | Western Blot (1:5000 dilution) |
| Antibody | HRP-conjugated goat anti-rabbit | Promega | #W401B | Western Blot (1:5000 dilution) |
| Antibody | HRP-conjugated goat anti-mouse | Promega | #W402B | Western Blot (1:5000 dilution) |
| Other | Luxol Fast Blue | Electron Microscopy Sciences | #26516–01 | Histological stain for myelin |
Additional files
-
Supplementary file 1
(A) Binding of 2BAct at 10 μM to the CEREP panel of receptors/enzymes/ion channels. Compound binding was calculated as a % inhibition of the binding of a radioactively labeled ligand specific for each target. Results showing inhibition or stimulation higher than 50% are considered to represent significant effects of the test compound. (B) Pharmacokinetic data from 2BAct dosing in mice. (C) GO Biological Processes identified by enrichment analysis of RNA-seq data. (D) List of genes used for enrichment analysis of scRNA-seq data.
- https://doi.org/10.7554/eLife.42940.028
-
Transparent reporting form
- https://doi.org/10.7554/eLife.42940.029





