Identification of EOMES-expressing spermatogonial stem cells and their regulation by PLZF
Figures
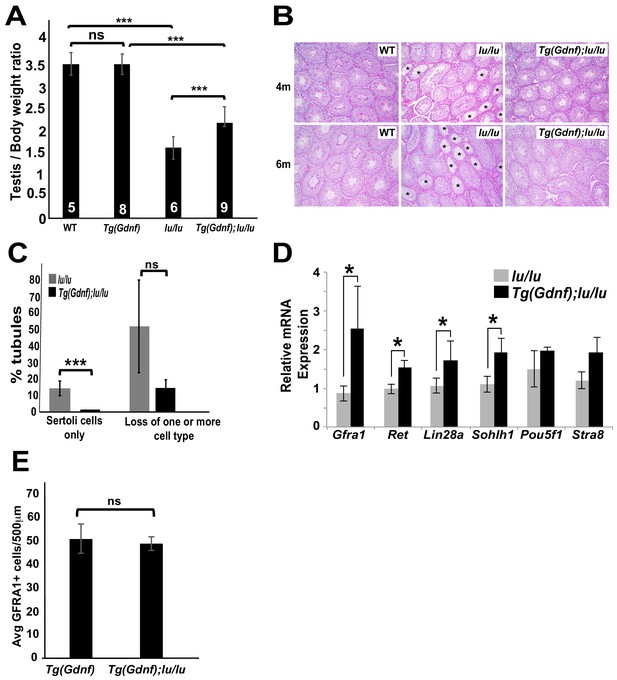
Sertoli-cell overexpression of GDNF partially reverses the loss of undifferentiated spermatogonia in luxoid mice.
(A) Testis/body weight ratios. No significant (ns) difference was observed in testis/body weight between WT and Tg(Gdnf). Significant differences (***) were detected between WT and lu/lu (p=0.0001), Tg(Gdnf) and Tg(Gdnf);lu/lu (p=0.0001) and between lu/lu and Tg(Gdnf);lu/lu (p=0.0005). Number in bar = n animals. (B) Representative images of periodic-acid-Schiff stained cross-sections of 4- and 6 month old testes showing agametic tubules (asterisks) in lu/lu. (C) Fewer tubules with germ cell loss were present at 6 months in Tg(Gdnf):lu/lu. A total of 200–300 tubules were counted for each genotype (n = 3). p<0.005. (D) qRT-PCR on testis RNA from 4 month old mice shows a significant increase in SSC gene expression in lu/lu mice overexpressing GDNF in Sertoli cells. Relative mRNA levels are normalized to a β-actin internal control. *, p<0.01. (n = 3) (E) Average number of GFRA1+ cells per 500 μm length of tubules in Tg(Gdnf) and Tg(Gdnf);lu/lu in 10wk old mice (n = 3).
-
Figure 1—source data 1
Source file for Figure 1A, 1C and 1E.
- https://doi.org/10.7554/eLife.43352.005
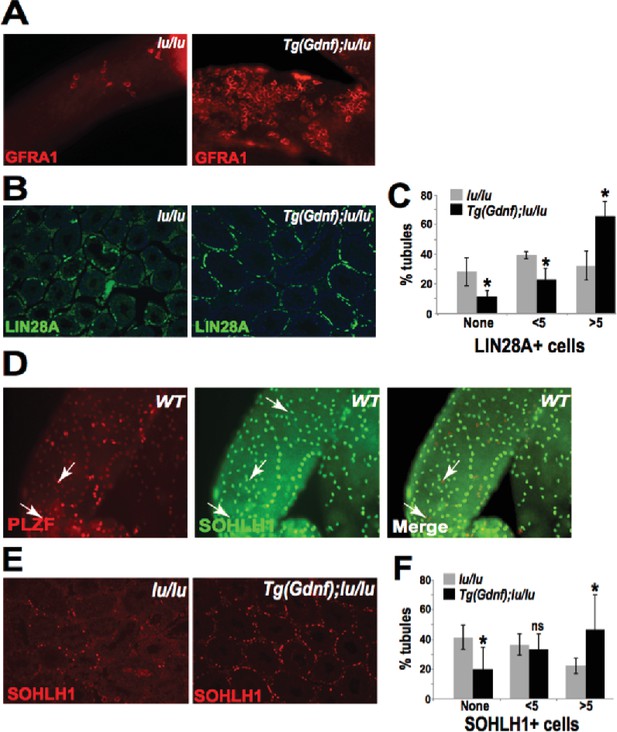
Sertoli-cell overexpression of GDNF increases the SSC population in Plzf-mutant luxoid mice.
(A) Immunostained whole-mount tubules from 12-week old Tg(Gdnf);lu/lu mice show large clusters of GFRA1+ cells compared to lu/lu. (B) Representative LIN28A immunostaining of lu/lu and Tg(Gdnf);lu/lu tubule paraffin sections. (C) Quantification of immunostained sections as in (B) show 70% more tubules with 5 or more LIN28A+ cells in Tg(Gdnf);lu/lu compared to lu/lu (n=4). *, p < 0.05. (D) In WT, SOHOLH1 expression is detected in Aal cells transitioning to A1 and differentiating cells A2-A4, absent from As , Apr , and Aal cells. (E) Representative SOHLH1 immunostaining of lu/lu and Tg(Gdnf);lu/lu tubule paraffin sections. (F) Quantification of immunostained sections as in (E) show a significant increase in the number of tubules with 5 or more SOHLH1+ cells in Tg(Gdnf);lu/lu compared to lu/lu. (n=4). * p<0.05, ns, not significant.
-
Figure 1—figure supplement 1—source data 1
Source file for Figure 1—figure supplement 1C and 1F.
- https://doi.org/10.7554/eLife.43352.004
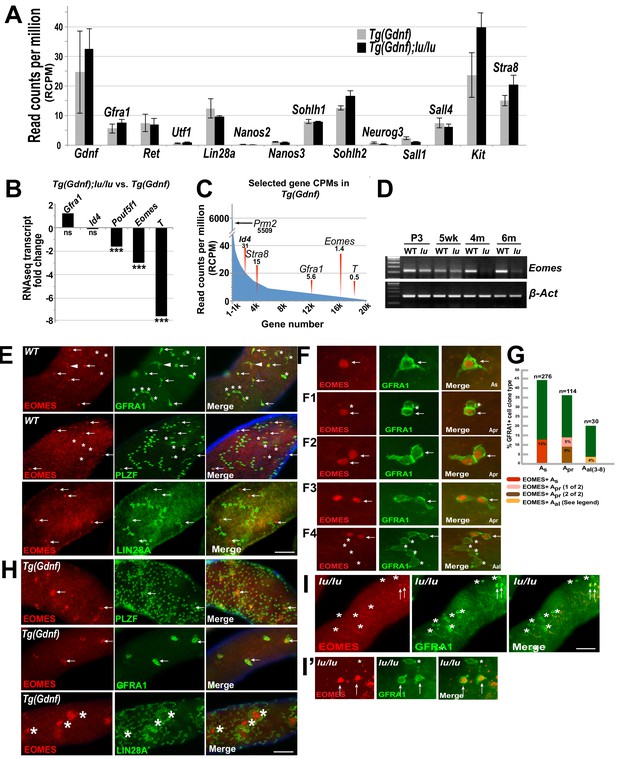
EOMES marks a subpopulation of SSCs.
(A) Relative gene expression (read counts per million, RCPM) of selected transcripts from RNAseq performed on testis RNA from 4 month old Tg(Gdnf) and Tg(Gdnf);lu/lu mice. No significant change is detected in known spermatogonial marker genes. (B) Fold change in RNAseq transcripts for a subset of pluripotency and developmental genes in Tg(Gdnf);lu/lu compared to Tg(Gdnf) testes. ***, p<0.0001. (C) Relatively low RCPM of the highest-fold change transcripts shown in (B), compared to the spermatid-expressed protamine two gene Prm2, and differentiating spermatogonia marker Stra8. X-axis represents 20,000 genes numbered in descending order by RCPM. (D) Semi-quantitative RT-PCR of total testis RNA showing decreasing Eomes expression over time in lu/lu testes. (E) Immunostaining for EOMES and undifferentiated spermatogonial markers in WT whole-mount testes. EOMES marks some (arrows), but not all (asterisks), GFRA1+ PLZF+ As cells. Weak EOMES staining can occasionally be detected in chains of GFRA1+ Aal cells (arrowhead). EOMES+ As cells do not express LIN28A (bottom row, arrows). (F) Examples of EOMES+ subset of GFRA1+ cells. F1 shows an example of two adjacent cells where one is EOMES+ (arrow) and one is EOMES- (asterisk). F2 and F2 illustrate two examples of two juxtaposed cells that are both EOMES+. F4 illustrates an example of a chain of 6 Aal cells with two EOMES+ (arrow) and four EOMES– (asterisks) cells. (G) Graph showing the percent of EOMES+ cell in the GFRA1+ population clone types in WT adult mice (631 GFRA1+ cells from three mice). n = number of clone types counted (e.g. Apr = 114 pairs of cells = 228 GFRA1+ total cells). Of the total GFRA1+ Apr population, in 9% of cases both cells were positive for EOMES, while 5% had only one cell positive for EOMES. Within the Aal fraction, 4% of the clones express EOMES in a heterogeneous pattern (two Aal chains where 3 of 3 were EOMES+; two Aal chains of 4 where all were EOMES+; one Aal chain of 4 where only one was EOMES+; one Aal chain of 6 where only 2 of 6 cells were EOMES+). (H) EOMES is expressed in tightly packed GFRA1+ PLZF+ clusters (arrows) found in Tg(Gdnf) whole-mount tubules. EOMES is not expressed in the LIN28A+ cells in the cortical region of the clusters (bottom row, asterisks). (I) Detection of EOMES+ GFRA1+ cells (arrows) in Plzf lu/u mutants. Asterisks mark an EOMES- GFRA1+ cell.
-
Figure 2—source data 1
Source file for Figure 2G.
- https://doi.org/10.7554/eLife.43352.007
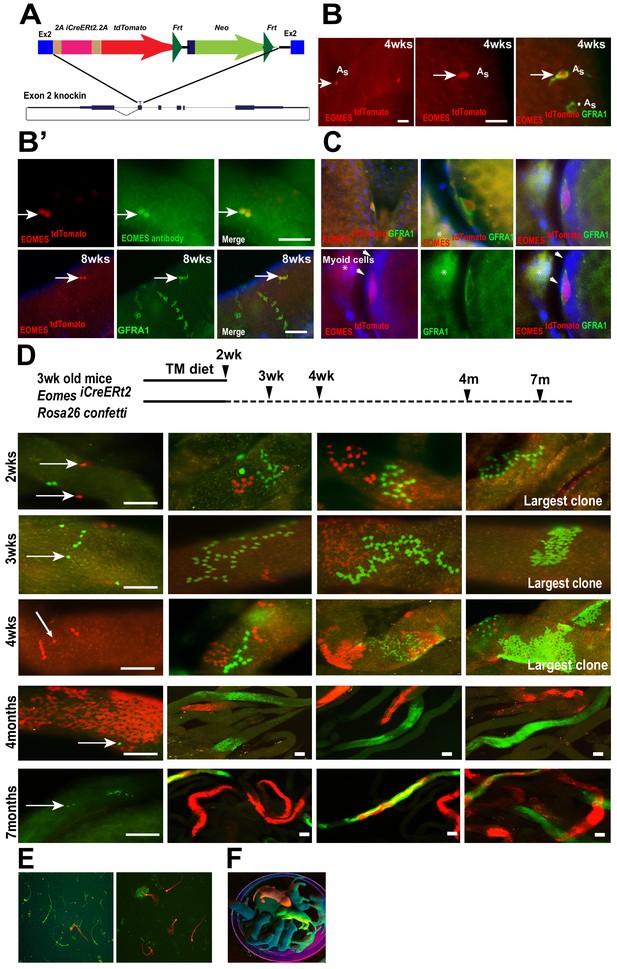
EOMES+ cells are long-term SSCs.
(A) Schematic diagram of the Eomes locus on chromosome nine and insertion cassette used to generate an Eomes/2AiCreERT2/2A/tdTomato knock-in allele. (B) GFRA1+ tdTomato+ As cell in testis of 4wk old mouse (arrow). GFRA1+ tdTOMATO As cell (asterick). (B’) tdTOMATO is co-expressed with EOMES (top row, arrow). tdTOMATO is expressed in GFRA1+ As cells but not in chains of GFRA1+ Aal cells (bottom row). (C) tdTOMATO cells (Red), positive for GFRA1 (Green), are present within the seminiferous tubules. Top - An optical section of confocal image showing the localization of EOMES+ cell along the basement membrane. Bottom – Magnified view of GFRA1+ EOMES tdTOMATO+ cells shown above. Arrowheads mark myoid cell (M) nucleus, which are negative for EOMES tdTOMATO. * Background fluorescence in the interstitial space associated with GFRA1 staining. (D) Experimental flow chart showing the induction times and clonal analysis of Eomes iCreERt2 targeted cells labeled in mice crossed to Rosa26 Confetti reporter and fed a tamoxifen-enriched diet. Fluorescence images of whole-mount seminiferous tubules showing the different size of labeled clones. Representative clones at 2 (n = 5, animals), 3 (n = 4), and 4wks (n = 3), and 4 (n = 3) and 7 (n = 3) months after tamoxifen induction. (E) Images of labeled sperm from the epididymis of mice 7 months after tamoxifen induced recombination. (F) Fluorescence positive offspring born to WT female mated with ROSA26confetti: Eomes-iCreERt2 4 months after tamoxifen-induced recombination at Confetti locus by Eomes-iCreERt2.
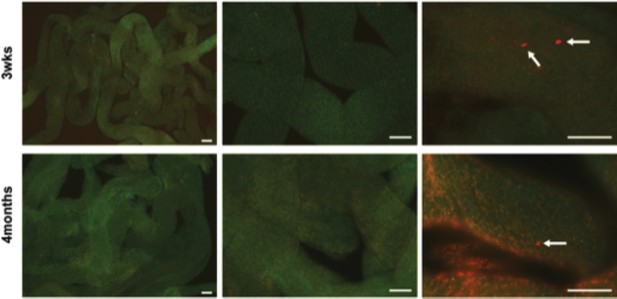
No TAM controls for lineage tracing of Eomes iCreERt2 targeted cells in mice crossed to Rosa26 Confetti reporter on a normal (no tamoxifen) diet.
Representative images at 3wks and 4mos post completion of TM diet fed to experimental cohorts shown in Figure 3. Arrows indicate tdTOMATO+ cells. Scale bar = 100 μm.
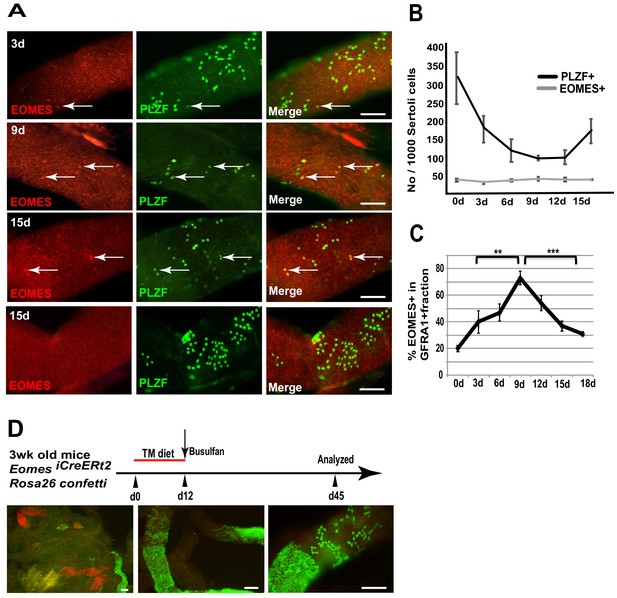
EOMES+ cells are resistant to busulfan and contribute in regeneration.
(A) Whole-mount tubule immunostaining for EOMES and PLZF after low-dosage busulfan treatment (10 mg/mL). (B) Cell counts from (A) show that EOMES+ cell numbers remain constant while PLZF+ cell numbers drop after busulfan treatment. (n = 3 per time point, 8 weeks old) (C) The EOMES+ fraction of GFRA1+ cells increased following busulfan treatment. **, p<0.005; ***, p<0.0001. (D) Schedule to assay the regeneration properties of EOMES+ cells. Three-week old R26R-confetti: Eomes iCreERt2 mice were put on a tamoxifen-enriched diet for 1 week, treated with busulfan, and analyzed on day 45. Images of whole mount seminiferous tubules (n = 5 mice) show labeled clones at different developmental stages during regeneration.
-
Figure 4—source data 1
Source file for Figure 4B and 4C.
- https://doi.org/10.7554/eLife.43352.011
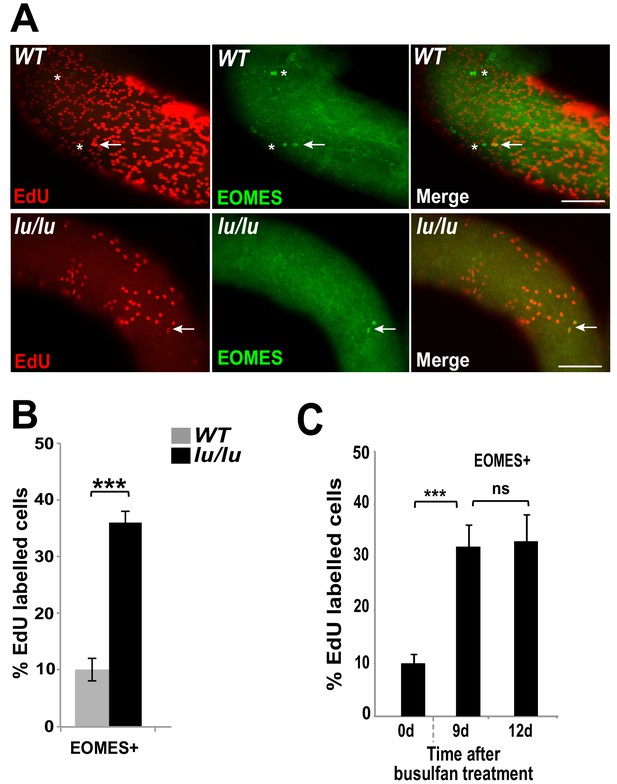
Loss of PLZF increases the proliferative index of EOMES+ cells.
(A) Whole-mount seminiferous tubules from 8 week old WT and lu/lu mice injected with EdU and immunostained 24 hr after injection. EdU+ EOMES+ cells (arrows); EdU- EOMES+ cells (asterisks). (B) Quantification of the EdU labeling index in WT (n = 4 mice) and lu/lu (n = 5 mice) EOMES+ cells. EOMES+ cells have a higher labeling index in lu/lu than in WT (p=0.001). (C) Increase in proliferation rate of EOMES+ cells during the recovery phase after busulfan treatment. EdU incorporation was assayed 24 hr post EdU injection (n = 3 mice per timepoint; ***p=0.001).
-
Figure 5—source data 1
Source file for Figure 5B and 5C.
- https://doi.org/10.7554/eLife.43352.013
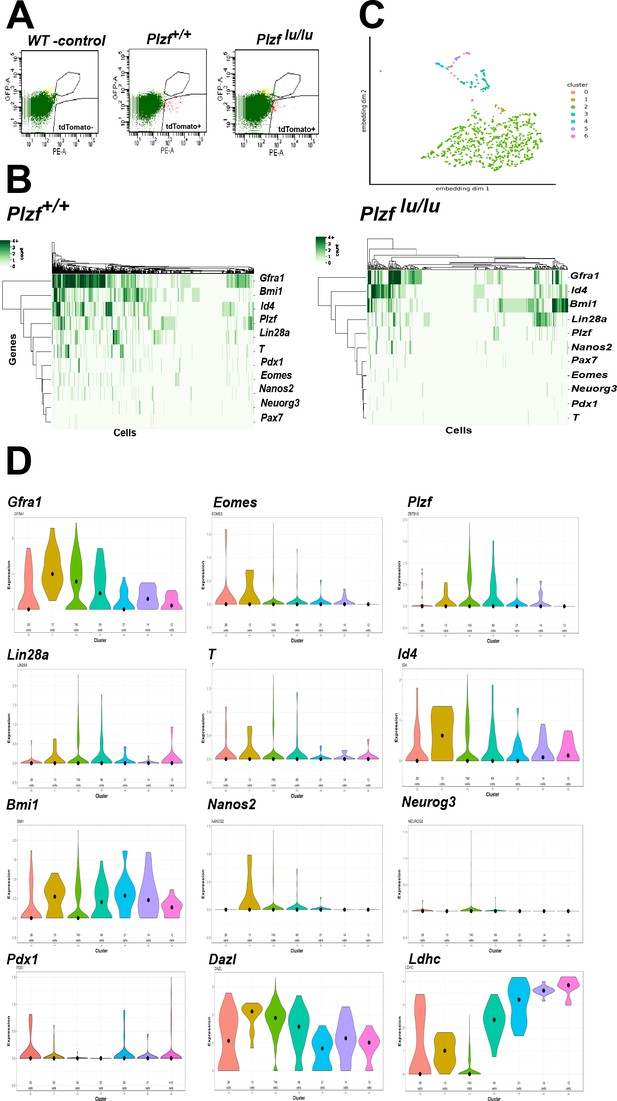
Single cell RNA sequencing of EOMES tdTOMATO cells.
(A) FACscan profile of EOMES tdTOMATO+ cells sorted from Plzf +/+ and Plzf lu/lu mutant testis. The boxed region in the upper right indicated cells with background autofluorescence that were excluded from the analysis. (B) Heat map and dendogram of SSC identity and progenitor genes expressed in EOMES tdTOMATO cells from Plzf +/+ and Plzf lu/lu mutant testis. (C) Clusters generated from WT cells. Cellular expression profiles of 110 genes were embedded into a 2dimensional latent space using UMAP (see supplemental information for details). Cells in cluster 0 could not be confidently placed in any of the other six clusters. (D) Violin plots showing the mean expression of each SSC identity gene per cluster in cells sorted from Plzf +/+ testes.
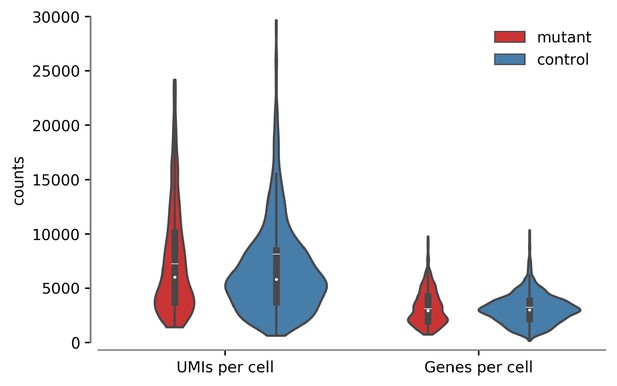
Distribution of unique molecular identifiers (UMIs) and genes per cell.
Median (white dot), mean (white line), and interquartile range (dark box) are shown for each of the four distributions, with control (Plzf +/+; Eomes-iCreERt-tdTomato+) in blue and mutant (Plzf lu/lu; Eomes-iCreERt-tdTomato+) in red.
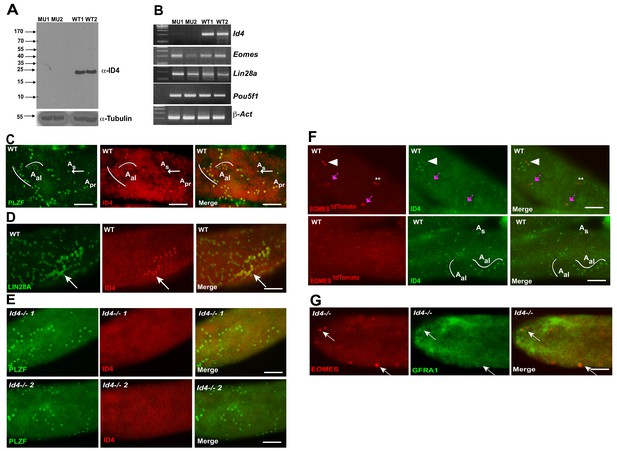
ID4 is expressed in As, Apr and Aal spermatogonia.
(A) Western blot of total testis protein lysate from 2 WT and 2 Id4-/- mutant mice probed with ID4 antibody. For loading control, western blot was probed with α-tubulin antibody. (B) RT-PCR using total RNA from two Id4-/- mutant and 2 WT mice for Eomes, Lin28a, Pou5f1 and β-actin. (C) Detection of PLZF and ID4 by immunofluorescence in whole mount seminiferous tubules. Both ID4 in PLZF+ were detected in As (arrow), Apr, and Aal spermatogonia in WT adult mouse testis. (D) ID4 protein was co-localized in chains of LIN28A+ Aal spermatogonia (arrow). (E) ID4+ cells are not detected in whole mount tubules from prepared from Id4-/- mutant mice. (F) Top row, examples of tdTOMATO/ID4 double-positive cells (arrowhead), tdTOMATO+ ID4- (**), and two adjacent cells where one is tdTOMATO+ and the other is tdTOMATO/ID4 double-positive (magenta arrow). Bottom row, examples of tdTOMATO- ID4+ As and Aal chains of 4 and 8 cells. (G) EOMES+ GFRA1+ cells are present in Id4-/- mutants (arrow).
Tables
Key identity genes expressed in SSCs or progenitor cells.
https://doi.org/10.7554/eLife.43352.016| Gene | Expression | Method | Functional Assay | References |
|---|---|---|---|---|
| Gfra1 | As, Apr, Aal(4-8) | Gfra1-CreERT2 | Lineage tracing Transplant | (Hara et al., 2014) |
| Bmi1 | As | Bmi1CreERT | Lineage tracing | (Komai et al., 2014) |
| Id4 | As | Id4-eGFP | Lineage tracing Transplant | (Oatley et al., 2011; Sun et al., 2015) |
| Plzf | As, Apr, Aal(4-16) | Antibody staining | Lineage tracing Transplant | (La et al., 2018; Buaas et al., 2004) |
| Lin28a | As, Apr, and Aal(4-16) | Antibody staining | Transplant | (Chakraborty et al., 2014) |
| T | As | T(nEGFP-CreERT2) | Lineage tracing | (Tokue et al., 2017) |
| Eomes | As, Apr | Eomes-iCre ERt2 tdTomato | Lineage tracing | This manuscript |
| Nanos2 | As, Apr | Antibody staining | Lineage tracing Knockout | (Sada et al., 2009) |
| Neurog3 | Aal(4-16) | Ngn3 CreERT | Lineage tracing Transplant | (Nakagawa et al., 2010) |
| Pax7 | As | Pax7-CreERT2 | Lineage tracing | (Aloisio et al., 2014) |
| Pdx1 | As, Apr | Antibody staining | Lineage tracing | (La et al., 2018) |
Differential expression of SSC identity transcripts in Plzf +/+ and Plzf lu/lu mutant cells.
https://doi.org/10.7554/eLife.43352.017| Gene | Log2 FC | P value |
|---|---|---|
| Gfra1 | -0.779 | 1.31E-12 |
| Eomes | -0.766 | 1.41E-09 |
| Plzf | -1.27 | 4.09E-21 |
| Id4 | -0.046 | 0.73 |
| Bmi1 | -0.021 | 0.8 |
| Pax7 | 0.09 | 0.278 |
| T | -1.45 | 2.11E-19 |
| Lin28a | 0.046 | 0.71 |
| Nanos2 | 0.039 | 0.67 |
| Neurog3 | -0.177 | 0.08 |
| Pdx1 | -0.738 | 6.11E-09 |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Mus musculus) | B6/N | The Jackson Laboratory | JR # 05304 | |
| Strain, strain background (Mus musculus) | B6.129P2-Gt(ROSA) 26 Sortm1(CAG-Brainbow2.1)Cle/J | The Jackson Laboratory | JR# 017492 | |
| Strain, strain background (Mus musculus) | Zbtb16lu/J (Plzf lu) | The Jackson Laboratory | JR#000100 | |
| Strain, strain background (Mus musculus) | B6(Cg)-Tyr c-2J/J | The Jackson Laboratory | JR# 000058 | |
| Strain, strain background (Mus musculus) | B6.Cg-Gt(ROSA)26Sor tm6(CAG-ZsGreen)Hze/J | The Jackson Laboratory | JR# 007906 | |
| Strain, strain background (Mus musculus) | Tg(Ctsl-Gdnf) B6.C3H.F1 | Sharma & Braun 2018 | PMCID: 29440301 | |
| Strain, strain background (Mus musculus) | Id4 -/- | Received from Mathew Havrda and Mark Israel; Dartmouth Medical School | PMCID: 15469968 | |
| Cell line (M. musculus) | B6N-JM8 ES | The Jackson Laboratory | RRID:CVCL_J957 | |
| Antibody | Mouse mono-clonal anti-PLZF | Santa Cruz Biotech Inc, Santa Cruz, CA | Cat# sc-28319 | Dilution – 1:250 |
| Antibody | Goat poly-clonal anti-GFRA1 | R and D systems, Minneapolis, MN | Cat# AF560 | Dilution – 1:500 |
| Antibody | Rat poly-clonal anti-LIN28A | Gift from Dr. Eric G. Moss, Rowan University, School of Osteopathic Medicine, One Medical Center Drive, Stratford, NJ 08084 | Dilution – 1:500 | |
| Antibody | Rabbit poly-clonal anti-EOMES | Abcam, Cambridge, MA | Cat# ab23345 | Dilution – 1:500 |
| Antibody | Rabbit polyclonal anti-SOHLH1 | Abcam, Cambridge MA | Cat# ab49272 | Dilution – 1:400 |
| Antibody | Rabbit mono-clonal anti-ID4 | BioCheck Inc, Foster City, CA | Cat # BCH-9/ # 82–12 | Dilution – 1:500 |
| Chemical compound, drug | Tamoxifen diet | https://www.envigo.com/products-services/teklad/laboratory-animal-diets/ | TD. 130859 | |
| Chemical compound, drug | EdU | Invitrogen, https://www.thermofisher.com/us/en/home/brands/invitrogen.html | Cat# A10044 | |
| Chemical compound, drug | Busulfan | Sigma-Aldrich, St. Louis, MO | Cat# B2635 | |
| Commercial assay or kit | Click-IT EdU assay kit | Invitrogenhttps://www.thermofisher.com/us/en/home/brands/invitrogen.html | Cat # A20185 | |
| Commercial assay or kit | RNA isolation PureLink RNA Mini kit | Ambion https://www.thermofisher.com/us/en/home/brands/invitrogen/ambion.html | Cat# 12183018A | |
| Commercial assay or kit | Arcturus PicoPure RNA kit | Applied Biosystems https://www.thermofisher.com/us/en/home/brands/applied-biosystems.html | Cat # 12204–01 | |
| Commercial assay or kit | TruSeq RNA Sample Prep Kit v2 | Illuminahttps://www.illumina.com/ | ||
| Commercial assay or kit | Nextera Xt DNA sample kit | Illumina https://www.illumina.com/ | ||
| Sequence-based reagent | Forward primers for RT-PCR of Eomes | 5’- ctggttctgttctttgcacaggcgcatg-3’ | ||
| Sequence-based reagent | Reverse primers for RT-PCR of Eomes | 5’- atcagccacaccagacacagagatc-3’ | ||
| Sequence-based reagent | Forward primers for RT-PCR B-actin | 5’- ccagttcgccatggatgacgatat-3' | ||
| Sequence-based reagent | Reverse primers for RT-PCR B-actin | 5’- gtcaggatacctctctgctctga-3' | ||
| Software, algorithm | TopHat v 2.0.7 | Kim et al., 2013 | PMCID:PMC4053844 | |
| Software, algorithm | EdgeR v2.6.10 | Robinson et al., 2010 | PMCID:PMC2796818 | |
| Software, algorithm | BamTools v 1.0.2 | Barnett et al., 2011 | PMCID:PMC3106182 | |
| Software, algorithm | Cell View | Bolisetty et al., 2017 | http://doi.org/10.1101/123810 |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.43352.019





