Two bifunctional inositol pyrophosphate kinases/phosphatases control plant phosphate homeostasis
Figures
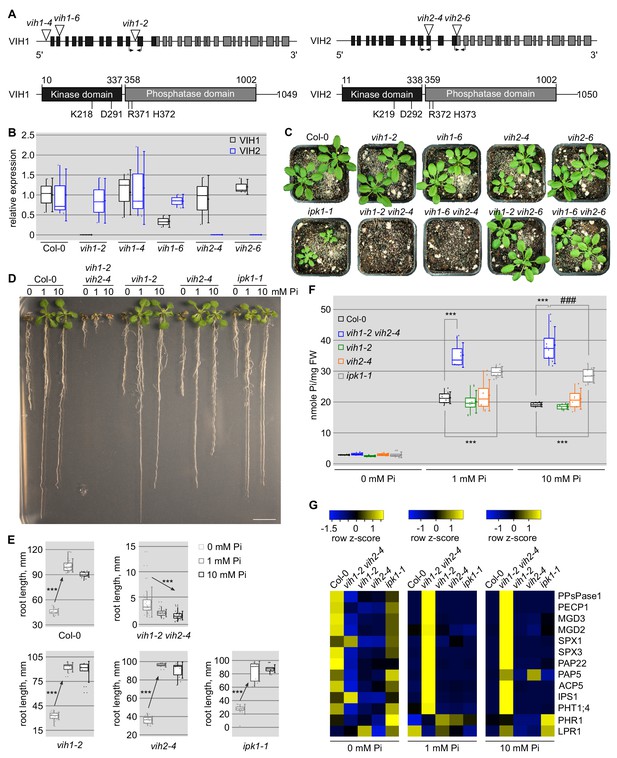
vih1 vih2 loss-of-function mutants show severe growth phenotypes and hyperaccumulate Pi.
(A) Schematic overview of VIH1 and VIH2: (upper panel) VIH1 and VIH2 genes with exons described as rectangles, introns as lines. T-DNA insertions are depicted as triangles, primer positions used in qRT-PCR analyses are indicated by arrows. (lower panel) VIH1 and VIH2 protein architecture, with kinase domains shown in black, phosphatase domains in gray and putative linkers/unstructured regions as lines. The point mutations used in this study are shown below the domain schemes. (B) qRT-PCR expression analysis of VIH1 and VIH2 transcripts in the T-DNA mutant allele backgrounds. Shown are 2-∆CT values relative to Col-0 wild-type. Quantifications were done using three biological replicates. (C) Growth phenotypes of vih1 and vih2 single mutant, and of vih1 vih2 double mutants. Shown are plants 20 DAG in soil compared to Col-0 wild-type. (D) Growth phenotypes of vih1-2 vih2-4, vih1-2, vih2-4, and of ipk1-1 compared to Col-0 wild-type seedlings. Plants were germinated in vertical 1/2MS plates for 8 d, transferred to Pi-deficient 1/2MS plates supplemented with either 0 mM, 1 mM or 10 mM Pi and grown for additional 6 d. Scale bars correspond to 2 cm. (E) Trend analysis of seedling root length vs. cellular Pi concentration for the seedlings described in (D). For each boxed position, root length measurements were performed for seedlings from three independent MS plates. (F) Pi contents of the seedlings shown in (D) 14 DAG. For each boxed position, four independent plants were measured with two technical replicates. (G) qRT-PCR quantification of PSI marker genes (PPsPase1, PECP1, MGD3, MGD2, SPX1, SPX3, PAP22, PAP5, ACP5, IPS1, PHT1;4) and of the non-PSI genes PHR1 and LPR1 in Col-0 wild-type, vih1-2 vih2-4, vih1-2, vih2-4 and ipk1-1 seedlings described in (D). Expression levels are represented as Z-scores. The original data are shown in Supplementary file 3a.
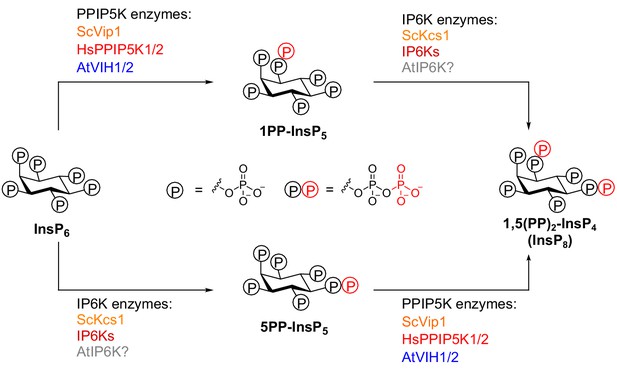
Overview of PP-InsP isoforms and kinases involved in inositol pyrophosphate metabolism.
Two classes of inositol pyrophosphate kinases, PPIP5K and IP6K, are responsible for the synthesis of inositol pyrophosphates. The scheme outlines PPIP5K enzymes from Saccharomyces cerevisiae ScVip1, Homo sapiens HsPPIP5K11 and 2, and Arabidopsis thaliana VIH1 and VIH2. These enzymes synthesize 1PP-InsP5 and 1,5(PP)2-InsP4 (InsP8) by phosphorylating InsP6and 5PP-InsP5 at the C1 position, respectively. IP6K enzymes from Saccharomyces cerevisiae KCS1 and Homo sapiens IP6Ks synthesize 5PP-InsP5 and 1,5(PP)2-InsP4 (InsP8) by phosphorylating InsP6and1PP-InsP5 at the C5 position, respectively. Plant IP6Ks have not been reported thus far.
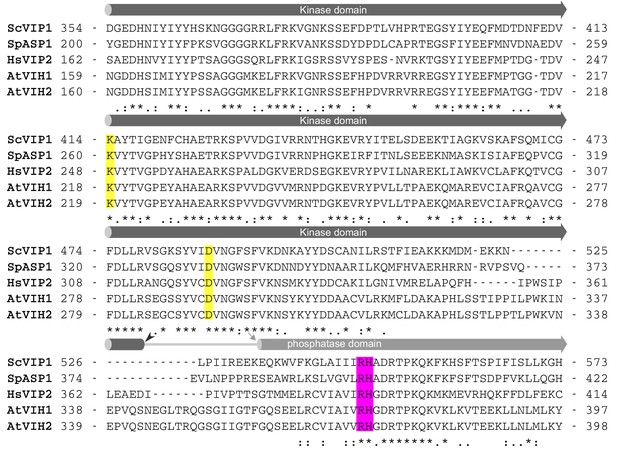
The kinase domain and phosphatase domain of VIHs/PPIP5Ks are conserved among different species.
Structure-based sequence alignment of the kinase - phosphatase cores of Saccharomyces cerevisiae Vip1 (Uniprot (https://www.uniprot.org/) identifier: Q06685), Schizosaccharomyces pombe ASP1 (Uniprot identifier: O74429), Homo sapiens PPIP5K22 (Uniprot identifier: O43314), Arabidopsis thaliana VIH1 (Uniprot identifier: Q84WW3), and VIH2 (Uniprot identifier: F4J8C6). Predicted kinase domain and the N-termini of the phosphatase domains in VIHs/PPIP5Ks are shown as dark and light gray cylinders, respectively. Linker regions are indicated by gray lines. Conserved residues that were mutated in this study are highlighted in yellow for the kinase domain and in magenta for the phosphatase domain.
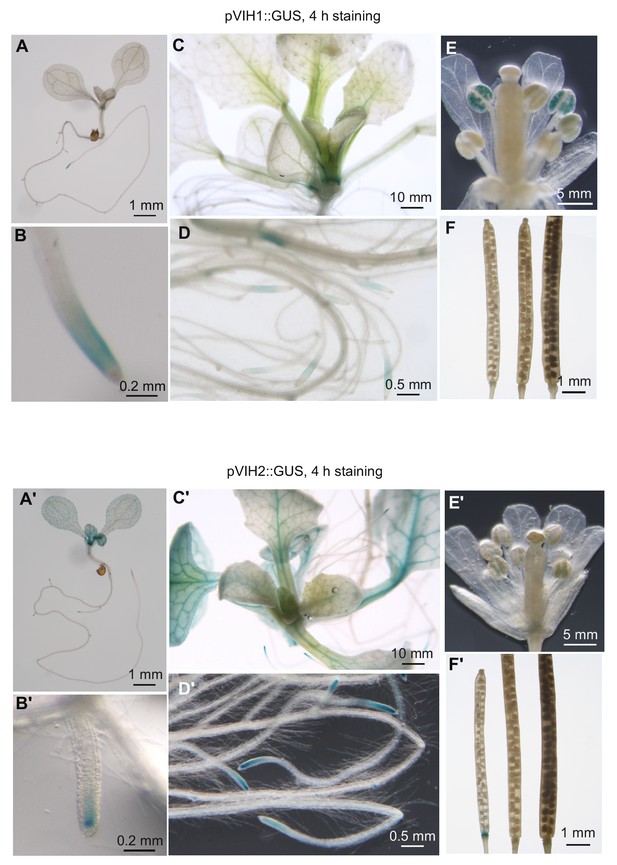
VIH1 and VIH2 show partly unique and partially overlapping expression patterns.
Transgenic Arabidopsis lines expressing pVIH1::GUS and pVIH2::GUS in the Col-0 wild-type background were stained for 4 hr and analyzed for β-glucuronidase activity. Images were acquired from seedlings 7 DAG (A,B and A’,B’) and 14 DAG (C,D and C’,D’), flowers (E and E’) and siliques (F and F’).
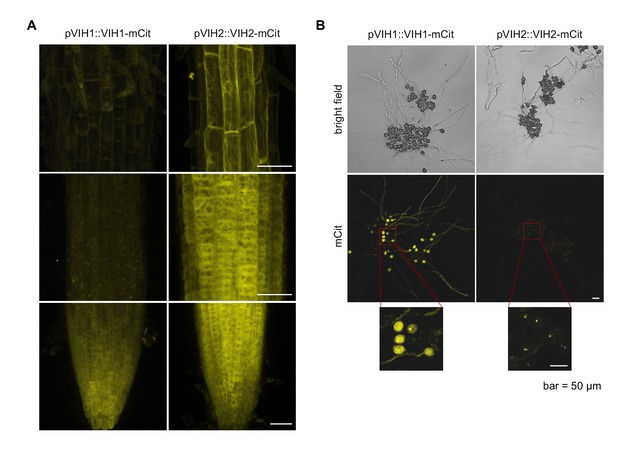
VIH1 and VIH2 are cytoplasmic enzymes with partially overlapping expression domains.
Stacked slices of (A) confocal scanning microscopy images showing the root tip of transgenic Arabidopsis plants expressing pVIH1::VIH1-mCit (left) and pVIH2::VIH2-mCit (right) in the vih1-2 and vih2-4 background, respectively. (B) Confocal scanning microscopy images of pollen and pollen tubes from the plants shown in (A), incubated for 4 hr on agarose plates containing 5 mM CaCl2, 5 mM KCl, 1 mM MgSO4, 0.01% (w/v) H3BO4 and 10% (w/v) sucrose, and 1.5% (w/v) agarose pH 7.5. Similar localization and expression patterns were observed in at least three independent transgenic lines. Scale bars = 50 μm.
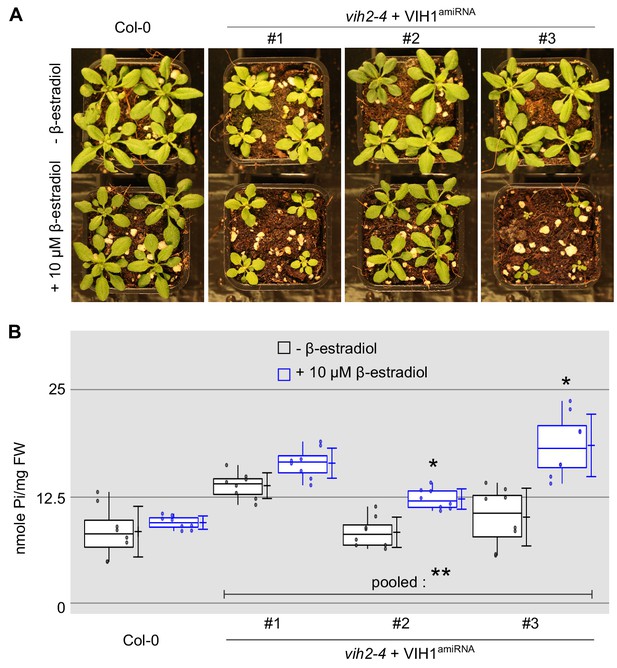
Inducible knock-down of VIH1 in the vih2-4 background impairs plant growth and leads to shoot Pi accumulation.
(A) Growth phenotypes of Col-0 wild-type and inducible mutant lines. Seedlings 7 DAG were transferred to soil and sprayed daily with tap water containing either 10 μM β-estradiol diluted from a 100 mM stock solution (100% [v/v] ethanol), or the corresponding concentration of ethanol only. (B) Shoot Pi content of plants 20 DAG treated or not with 10 μM β-estradiol as described in (A). For each boxed position, four independent plants were measured using two technical replicates.
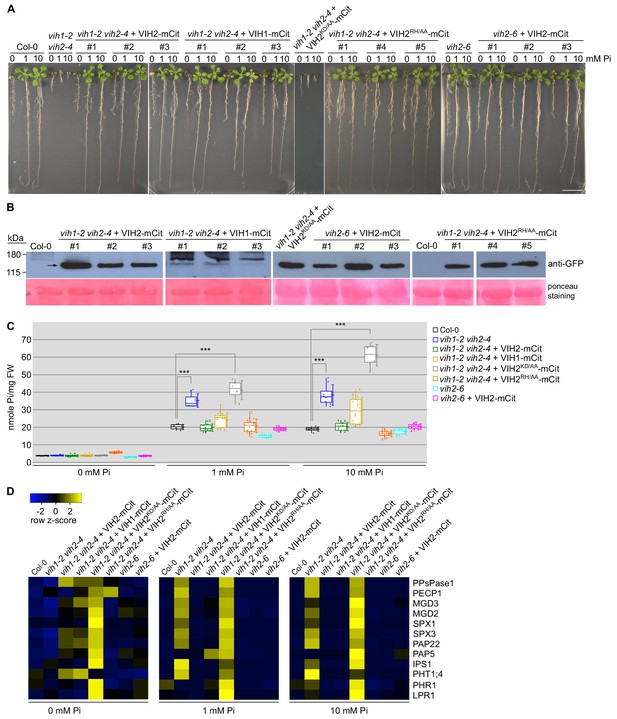
VIH kinase and phosphatase activities regulate plant Pi homeostasis.
(A) Complementation of vih1 vih2 growth phenotypes. Shown are three independent lines for each construct grown in different Pi regimes. Plants were germinated in vertical 1/2MS plates for 8 d, transferred to Pi-free 1/2MS plates supplemented with either 0 mM, 1 mM or 10 mM Pi and grown for additional 6 d. Scale bars correspond to 2 cm. (B) Western blot showing the expression of VIH2-mCit, VIH1-mCit, VIH2KD/AA-mCit, VIH2RH/AA-mCit proteins (indicated by arrow heads) in the transgenic lines from (A) using an anti-GFP antibody. A Ponceau stain of the membrane is shown as loading control below. (C) Pooled Pi content of seedlings 14 DAG as shown in (A). For each position, four independent plants from each trangenic line were measured with two technical replicates. (D) qRT-PCR quantification of the PSI marker genes in seedlings shown in (A). Expression levels are represented as Z-scores. The original qRT-PCR data are shown in Supplementary file 3b.
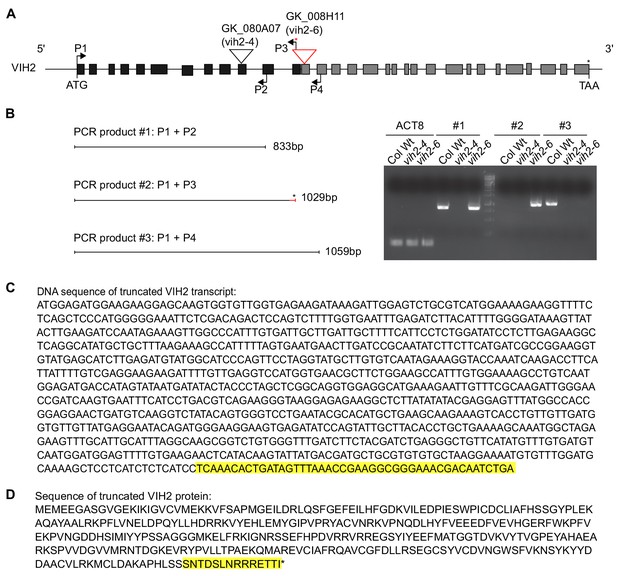
The vih2-6 transcript encodes a truncated VIH2 protein harboring the kinase domain only.
(A) Positions of T-DNA insertions in the vih2-4 and vih2-6 alleles and transcript detection by RT-PCR analysis at the VIH2 locus (At3g01310). A red asterisk indicates the putative stop codon residing in the T-DNA sequence in the vih2-6 mutant, which is included in the reverse primer P3 (vih2-6_TDNA_R). The primer set P1 (VIH2cDNA_F) + P4 (VIH2-c-R) encompasses the transcript region from the ATG start codon to sequences adjacent to the T-DNA insertion position in vih2-6. (B) RT - PCR products #1, 2 and 3 are amplified by primer set P1 + P2 (VIH2-b-R), P1 + P3 and P1 + P4 using cDNA reverse-transcribed from 10 DAG Col-0 wild-type, vih2-4 and vih2-6 mutant seedlings. Transcripts of ACTIN8 (ACT8) were used as a cDNA input control. (C) DNA sequence of the truncated VIH2 transcript in vih2-6. The highlighted (yellow) sequence is from the left border of the inserted T-DNA. (D) Putative protein sequence of the truncated VIH2 in vih2-6. The additional residues (highlighted in yellow) originate from the left border of the T-DNA insertion.
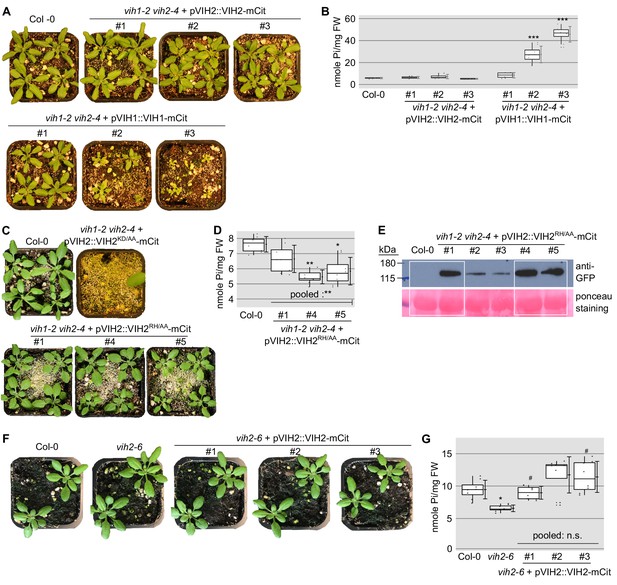
Growth phenotypes of soil-grown vih1-2 vih2-4 double mutant lines complemented with pVIH1::VIH1-mCit, pVIH2::VIH2-mCit, pVIH2::VIH2KD/AA-mCit or pVIH2::VIH2RH/AA-mCit.
(A) Expression of VIH1-mCit or VIH2-mCit under the control of their endogenous promoter can complement the vih1-2 vih2-4 phenotype. Seedlings 7 DAG were transferred to soil and grown for 23 d. Note that in the case of the VIH1-mCit construct, we also observed partially complementing lines. (B) Shoot Pi content of plants 22 DAG shown in (A). For each position, four independent plants were measured with two technical replicates each. (C) Growth phenotypes of independent vih1-2 vih2-4 lines, complemented with pVIH2::VIH2KD/AA-mCit or pVIH2::VIH2RH/AA-mCit. All plants were transferred to soil 7 DAG and grown for 21 d, Col-0 wild-type plants are shown alongside. (D) Shoot Pi contents of plants 20 DAG described in (C) except the lethal vih1-2 vih2-4 complemented by pVIH2::VIH2KD/AA-mCit. For each boxed position, four independent plants were measured with two technical replicates each. (E) Western blot showing the VIH2RH/AA-mCit protein levels in the complementation lines described in (C), detected using an anti-GFP antibody. The white box highlights the lines shown in Figure 2B. (F) Growth phenotypes of the vih2-6 mutant expressing the kinase domain-only, and three independent lines of vih2-6 complemented with pVIH2::VIH2-mCit. Plants 7 DAG were transferred to soil and grown for 21 d. (G) Shoot Pi content of plants 20 DAG as described in (F). For each position, four independent plants were measured with two technical replicates each.
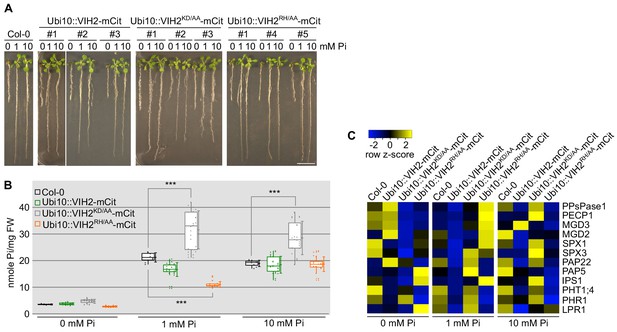
The kinase and phosphatase activities of VIH1 and VIH2 together control plant Pi homeostasis at seedling stage.
(A) Growth phenotypes of seedlings over-expressing VIH2-mCit, VIH2KD/AA-mCit or VIH2RH/AA-mCit with a Ubi10 promoter in Col-0 wild-type background. Col-0 wild-type and three independent transgenic lines for each construct are shown. Seedlings 7 DAG were transplanted from 1/2MS plates to Pi-free 1/2MS liquid supplemented with 0 mM, 1 mM or 10 mM Pi, and grown for additional 6 d. (B) Pooled Pi content of seedlings 14 DAG described in (A). Four independent plants from each transgenic line were measured with two technical replicates each. (C) qRT-PCR quantification of the PSI marker genes in seedlings shown in (A). Expression levels are represented as Z-scores. The original data of qRT-PCR are shown in Supplementary file 3d.
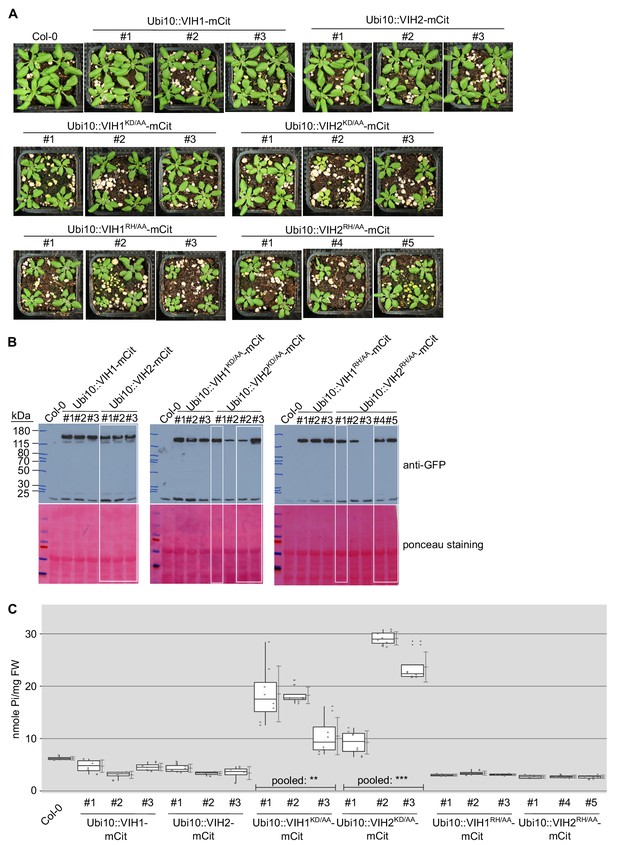
The kinase and phosphatase activities of VIH1 and VIH2 together control Pi homeostasis in soil-grown adult plants.
(A) Growth phenotype of plants expressing Ubi10::VIH1-mCit, Ubi10::VIH2-mCit, Ubi10::VIH1KD/AA-mCit, Ubi10::VIH2KD/AA-mCit, Ubi10::VIH1RH/AA-mCit and Ubi10::VIH2RH/AA-mCit. Seedings 7 DAG were transferred to soil and grown for 21 d. (B) Western blot of VIH1-mCit, VIH2-mCit, VIH1KD/AA-mCit, VIH2KD/AA-mCit, VIH1RH/AA-mCit and VIH2RH/AA-mCit proteins, detected by an anti-GFP antibody. White boxes correspond to the lines shown in panels A and C and in Figure 2—figure supplement 3. (C) Shoot Pi contents of plants 20 DAG from selected lines described in (A). Four independent plants were measured with two technical replicates each.
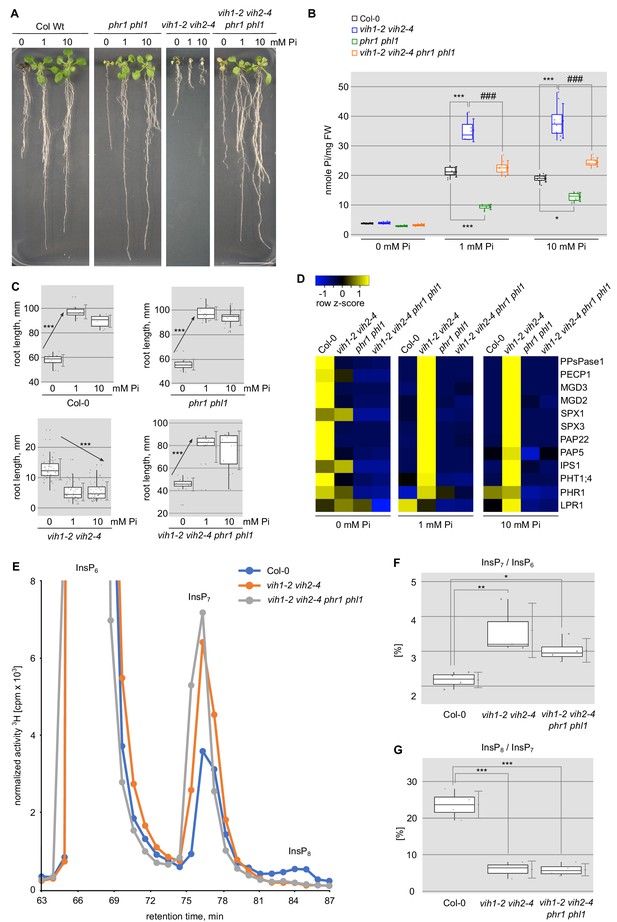
Deletion of PHR1 and PHL1 rescues vih1-2 vih2-4 growth phenotypes.
(A) Growth phenotype of Col-0, phr1 phl1, vih1-2 vih2-4, and vih1-2 vih2-4 phr1 phl1 seedlings grown in different Pi conditions. Seedlings 7 DAG were transplanted from 1/2MS plates to Pi-deficient 1/2MS liquid supplemented with 0 mM, 1 mM or 10 mM Pi, and grown for additional 6 d. (B) Cellular Pi content of seedlings 14 DAG described in (A). For each Pi concentration, at least four independent plants were measured with two technical replicates each. (C) Root length of seedlings 14 DAG described in (A). For each Pi concentration, seedlings from at least three independent plates were analyzed. (D) qRT-PCR quantification of the PSI marker genes in seedlings shown in (A). Expression levels are represented as Z-scores. The original data of qRT-PCR are shown in Supplementary file 3c. (E) Normalized HPLC profiles of [3H] inositol-labeled Col-0 (blue line), vih1-2 vih2-4 double mutant (orange line) and vih1-2 vih2-4 phr1 phl1 quadruple mutant (gray line). The InsP6, InsP7 and InsP8 regions are shown (termed as such, as specific PP-InsP5/(PP)2-InsP4 regioisomers cannot be resolved using this method). Fractions were collected each minute (solid dots) for radioactivity determination. The experiment was repeated at least three times with similar results, and representative profiles from one experiment are shown. The InsP7 / InsP6 ratio is plotted in (F) and the InsP8 / InsP7 ratio in (G).

Growth phenotypes of soil-grown phr1 phl1, vih1-2 vih2-4 and vih1-2 vih2-4 phr1 phl1 mutant adult plants.
(A) Growth phenotype of phr1 phl1, vih1-2 vih2-4, and vih1-2 vih2-4 phr1 phl1 mutant plants in comparison to the Col-0 control. Seedlings 7 DAG were transferred to soil and grown for 21 d. (B) Shoot Pi content of plants 20 DAG as described in (A). Four independent plants were measured with two technical replicates each.
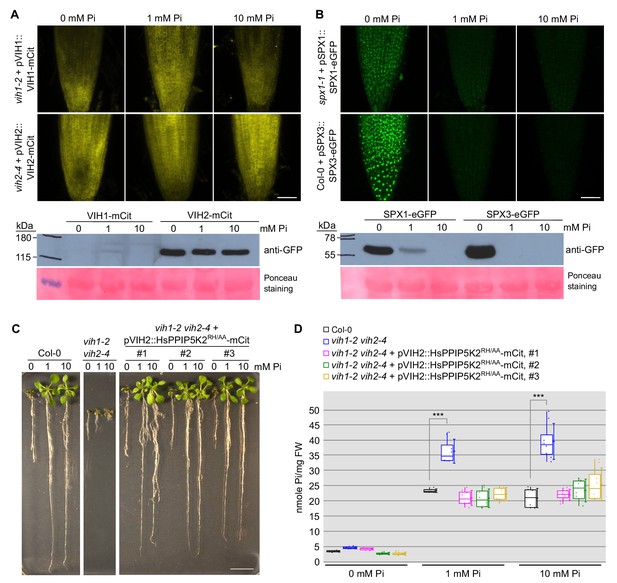
VIH1/VIH2 protein levels remain stable in different Pi growth regimes.
(A) Representative confocal scanning microscopy images showing a mCit fluoresecent signal in the root tip of vih1-2 and vih2-4 seedlings transformed with pVIH1::VIH1-mCit or pVIH2::VIH2-mCit respectively. Seedlings 7 DAG were transferred from 1/2MS plates to Pi-deficient 1/2MS liquid medium supplemented with 0 mM, 1 mM or 10 mM Pi, and grown for 3 d. Scale bar = 50 μm. Protein levels of VIH1-mCit and VIH2-mCit in plants detected by an anti-GFP antibody are shown below, the ponceau stained membrane serves as loading control. (B) Representative confocal scanning microscopy images showing a GFP fluorescent signal in the root tip of spx1-1 and Col-0 wild-type seedlings transformed with pSPX1::SPX1-eGFP and pSPX3::SPX3-eGFP respectively. Seedlings were treated as described in (E). Scale bar = 50 μm. Protein levels of SPX1-eGFP and SPX3-eGFP in plants detected by an anti-GFP antibody are shown below, the ponceau stained membrane is included as loading control. (C) Growth phenotypes of Col-0, vih1-2 vih2-4 and 3 independent lines of vih1-2 vih2-4 seedlings complemented with pVIH2::HsPPIP5K2RH/AA-mCit. Seedlings 7 DAG were transplanted from 1/2MS plates to Pi-deficient 1/2MS liquid supplemented with 0 mM, 1 mM or 10 mM Pi, and grown for additional 6 d. Scale bars correspond to 2 cm. (D) Pi content of seedlings 14 DAG as described in (D). Four independent plants from each transgenic line were measured with two technical replicates each.
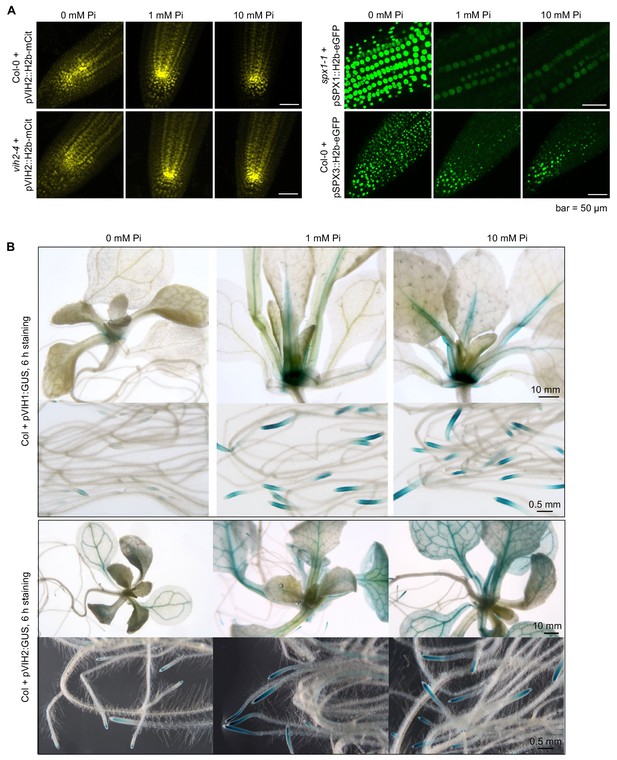
VIH1 and VIH2 promoter activities are not induced in low or high Pi conditions.
(A) Representative confocal images showing the expression of mCit-tagged histone H2b under the control of VIH2 promoter in root tip of Col-0 wild-type and vih2-4 seedlings (in yellow). Shown in green is the expression of eGFP-tagged H2b under the control of SPX1 promoter and SPX3 promoter in the root tip of spx1-1 and Col-0 wild-type seedlings, respectively. Seedlings 7 DAG were transferred from 1/2MS plates to Pi-deficient 1/2MS liquid medium supplemented with 0 mM, 1 mM or 10 mM Pi, and grown for 3 d. Scale bar = 50 μm. (B) Representative staining (6 hr) of GUS reporter lines expressing pVIH1::GUS and pVIH2::GUS in a Col-0 wild-type background. Seedlings 7 DAG were transferred from 1/2MS plates to Pi-deficient 1/2MS liquid medium supplemented with 0 mM, 1 mM or 10 mM Pi, and grown for 6 d.
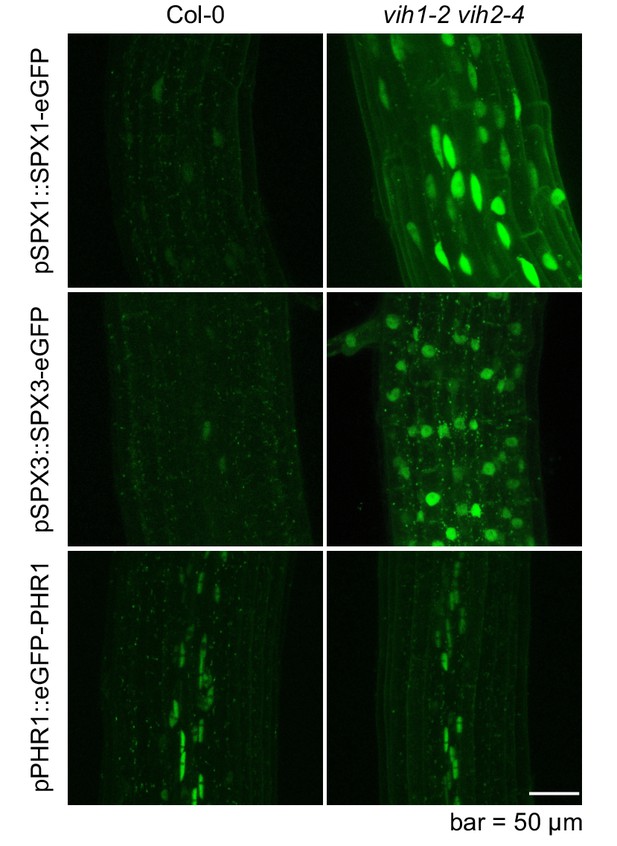
PSI marker proteins SPX1 and SPX3 accumulate in the vih1-2 vih2-4 mutant seedlings grown in Pi replete condition.
Representative confocal scanning microscopy images showing a GFP fluorescent signal in the root tip of Col-0 wild-type and vih1-2 vih2-4 mutant seedlings expressing pSPX1::SPX1-eGFP and pSPX3::SPX3-eGFP, respectively, and grown in Pi replete conditions. pPHR1-eGFP-PHR1 expressing control lines expressed in Col-0 and ih1-2 vih2-4 mutant background are shown for comparison. Scale bar = 50 μm.
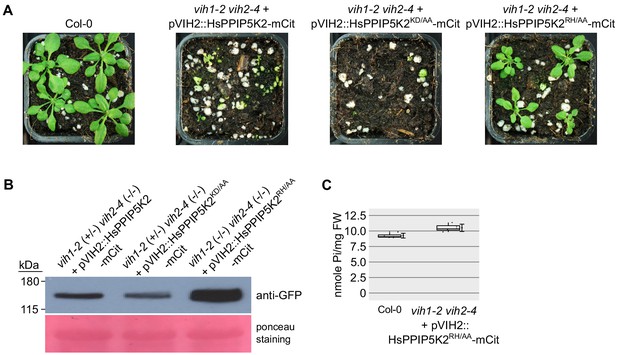
Growth phenotypes of soil-grown vih1-2 vih2-4 double mutant lines complemented with pVIH2::HsPPIP5K2-mCit, pVIH2::HsPP5PIK2KD/AA-mCit or pVIH2::HsPPIP5K2RH/AA-mCit.
(A) Growth phenotype of vih1-2 vih2-4 double mutants complemented with HsPPIP5K2-mCit, HsPPIP5K2KD/AA-mCit and HsPPIP5K2RH/AA-Cit under the control of the AtVIH2 promoter. Seedlings 7 DAG are transferred to soil and grown for 21 d. (B) Expression of HsPPIP5K2-mCit, HsPPIP5K2KD/AA-mCit and HsPPIP5K2RH/AA-Cit proteins, detected by an anti-GFP antibody. (C) Shoot Pi content of plants 20 DAG described in (A). Four independent plants were analysed with two technical replicates each.
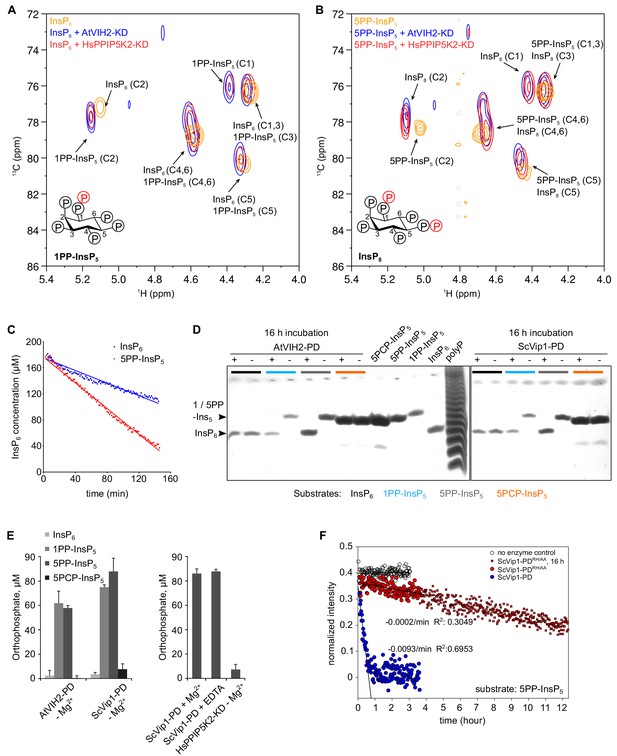
The AtVIH2 kinase domain has 1-kinase activity and produces 1PP-InsP5 and InsP8, the phosphatase domain is a 1 - and 5 - pyrophosphatase.
(A, B) 2D 1H-13C-HMBC spectra of the products produced by plant AtVIH2-KD (blue trace) and human HsPPIP5K2-KD (red trace) in the presence of InsP6 (A) or 5PP-InsP5 (B). Substrate standards are colored in yellow. (C) Decay of the InsP6 or 5-PP-InsP5 substrate during the NMR-time course experiment shown in (A) and (B), respectively. A fit of the initial decay indicates a turnover number of ~0.4/min with InsP6 as a substrate and ~1/min using 5-PP-InsP5 as a substrate. (D) Qualitative native PAGE phosphatase activity assay. Reactions containing recombinant phosphatase domain of AtVIH2 (AtVIH2-PD,~5 µg) or ScVip1 (ScVip1-PD,~27 µg) were incubated with 175 μM of either InsP6 (black), 1PP-InsP5 (blue), 5PP-InsP5 (gray) or a non-hydrolyzable 5PCP-InsP5 analog (orange) for 16 hr at 37°C. 40 μl of the reaction were separated in a 35% acrylamide gel. The bands corresponding to InsP6 and 1 or 1/5PP-InsP5 are indicated by an arrowhead. (E) Malachite green-based phosphatase activity assay. Reactions containing recombinant AtVIH2-PD (~5 µg) or ScVip1-PD (~27 µg) were incubated with 175 μM InsP6, 1PP-InsP5, 5PP-InsP5 or 5PCP-InsP5 for 16 hr at 37°C (left). 1 mM Mg2+ or 5 mM EDTA were supplemented as indicated (right; 5PP-InsP5 only). Recombinant HsPPIP5K2-KD (~17 µg) was used as a negative control and tested only with 5PP-InsP5 (right). Reactions were performed in quadruplicates and released orthophosphate was quantified using a malachite green assay (Baykov et al., 1988). (F) NMR time course experiment comparing the phosphatase activities of 2 µM ScVip1-PD and ScVip1-PDRH/AA using 40 µM [13C6]5PP-InsP5 as substrate. Samples were measured in a pseudo-2D spin-echo difference experiment and the relative intensities of the C2 peaks of InsP6 and 5PP-InsP5 were quantified.
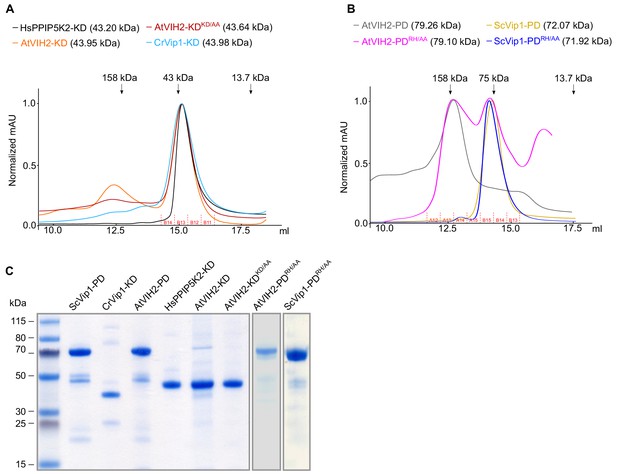
The purification of recombinant proteins.
(A) Size-exclusion chromatography traces of purified recombinant HsPPIP5K2 kinase domain (HsPPIP5K2-KD, residues 41–366), AtVIH2 kinase domain (AtVIH2-KD, residues 11–338), AtVIH2 kinase domain mutant (AtVIH2-KDKD/AA, residues 11–338), and CrVip1 kinase domain from Chlamydomonas (CrVip1-KD, residues 20–350). Fractions isolated for enzymatic assays are indicated below. (B) Size-exclusion chromatography traces of purified recombinant AtVIH2 phosphatase domain (AtVIH2-PD, Gly359-Arg1002), AtVIH2 phosphatase domain mutant (AtVIH2-PDRH/AA, residues 359–1002), ScVip1 phosphatase domain (ScVip1-PD, residues 515–1088), and ScVip1 phosphatase domain mutant (ScVip1-PDRH/AA, residues 515–1088). For enzyme activity assays, the indicated fractions containing the purified recombinant protein were pooled (fractions A12-A15 for AtVIH2-PD; A12-B13 for AtVIH2-PDRH/AA; and A15-B13 for both ScVip1-PD and ScVip1-PDRH/AA). (C) Coomassie-stained SDS-PAGE gel showing the recombinant proteins used in this study. For each lane, around 3 μg of protein from the respective concentrated fractions were loaded.
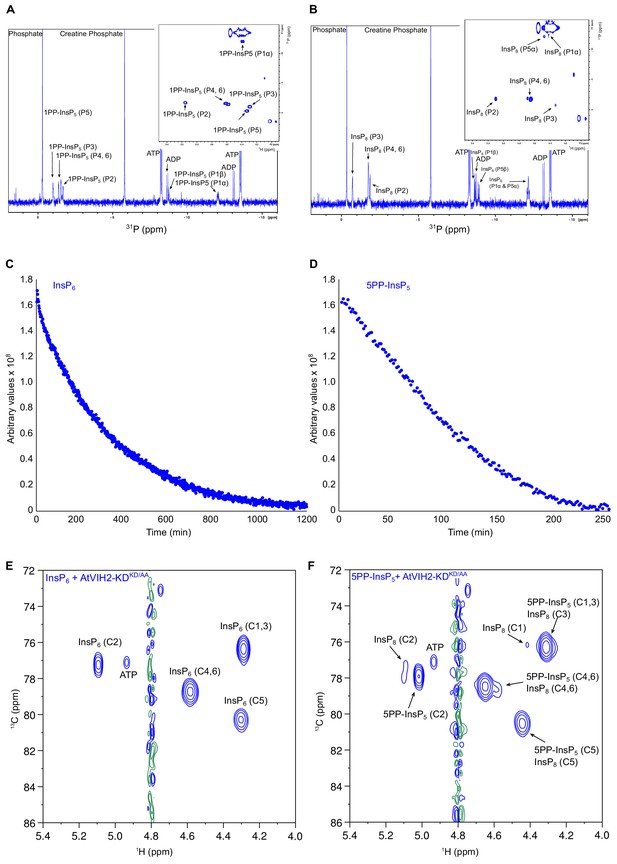
1D 31P and 2D 1H-31P-HMBC spectra of the products produced by AtVIH2-KD, and 2D 1H-13C-HMBC spectra of the products produced by plant AtVIH2-KDKD/AA.
(A,B) 1D 31P spectra of the products produced by the plant AtVIH2-KD in the presence of InsP6 (A) and 5PP-InsP5 (B). Inset panels shows the 2D 1H-31P-HMBC spectra. 1,5(PP)-InsP4 refers to InsP8. (C, D) Decay of the InsP6 (C) or 5-PP-InsP5 (D) substrate during the NMR-time course experiment. (E, F) 2D 1H-13C-HMBC spectra of the products produced by plant AtVIH2-KDKD/AA (blue trace) in the presence of InsP6 (E) or 5PP-InsP5 (F).
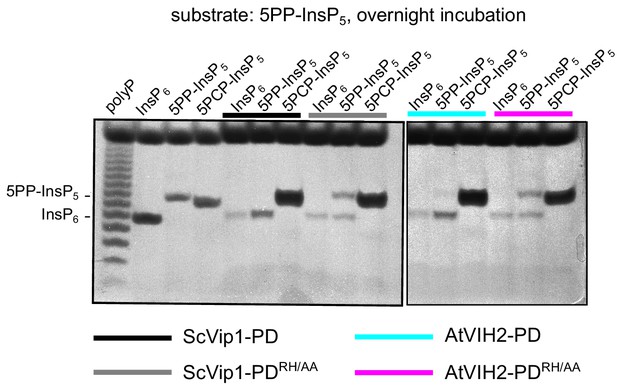
The ScVip1-PDRH/AA and AtVIH2-PDRH/AA recombinant enzymes show reduced phosphatase activity.
Qualitative native PAGE phosphatase activity assay. Reactions containing 2 µM ScVip1-PD (black), ScVip1-PDRH/AA(gray), AtVIH2-PD (cyan) and AtVIH2-PDRH/AA (magenta) were incubated with 80 μM 5PP-InsP5 for 16 hr at 37°C. The reactions were then separated in a 35% acrylamide gel. The bands corresponding to InsP6 and 1 or 5PP-InsP5 are indicated by an arrow head.
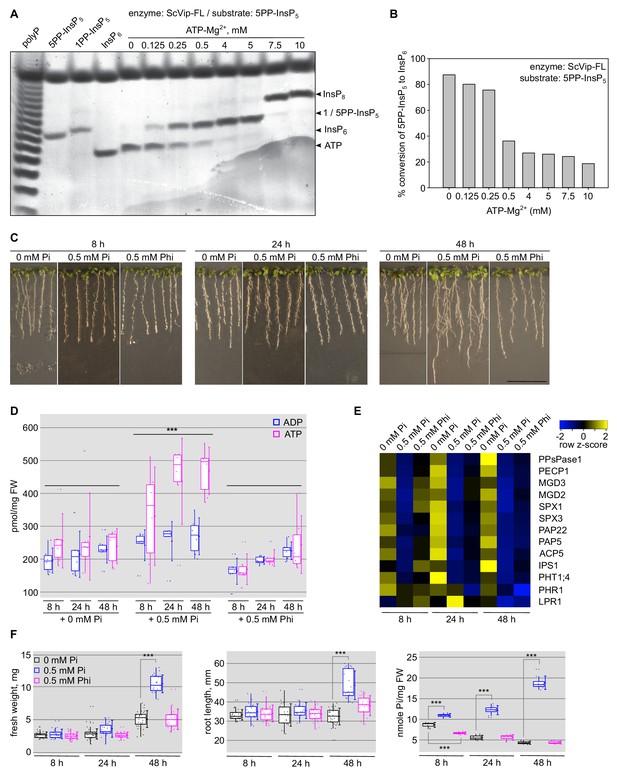
Changes in cellular ATP levels affect the relative PPIP5K kinase and phosphatase activities.
(A) Bi-functional PP-InsP activity assay of ScVip1. Reactions containing 2 μM protein, 40 μM 5PP-InsP5 and various ATP-Mg2+ concentrations were incubated at 37°C for 45 min. Product PP-InsPs were separated on a native PAGE gel and stained with toluidine blue. The bands corresponding to InsP6, 1/5PP-InsP5, InsP8 and ATP are indicated by arrow heads. (B) Quantification of conversion of the 5-PP-InsP5 (substrate) to InsP6 (product) by full-length ScVip1 (FL) enzyme in reactions containing different concentrations of ATP-Mg2+, recorded by NMR specroscopy. (C) Col-0 seedlings 6 DAG were transplanted from 1/2MS plates to Pi-deficient 1/2MS plates and incubate 3 days more. Then, the seedlings were transplanted again to Pi-deficient 1/2MS plates supplemented with 0 mM Pi, 0.5 mM Pi or 0.5 mM Phi, and grown for additional 8 hr, 24 hr or 48 hr. (D) Determination of the cellular ATP and ADP concentrations of the seedlings shown in (C). (E) qRT-PCR quantification of PSI marker genes in the seedlings shown in (C). Expression levels are represented as Z-scores. The original qRT-PCR data are shown in Supplementary file 3e. (F) Quantification of seedling fresh weight, primary root length and cellular Pi concentrations of the plants shown in (C).
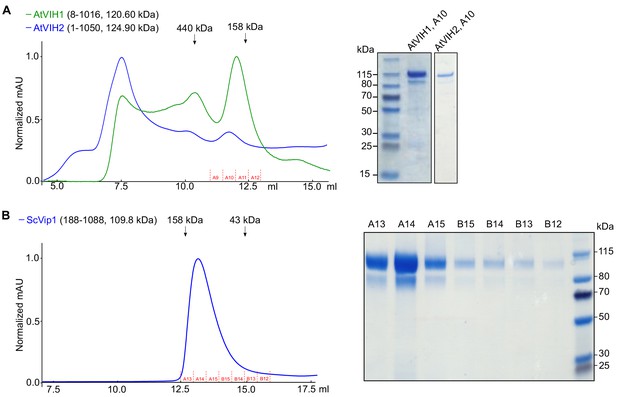
Purification of full-length AtVIH1, AtVIH2 and ScVip1 proteins from insect cells.
(A) Size-exclusion chromatography traces of purified full-length AtVIH1 (residues 8–1016) and AtVIH2 (residues 1–1050). A coomassie-stained SDS-PAGE gel showing the content of the peak fraction (A10) is shown alongside. (B) Size-exclusion chromatography trace of purified recombinant ScVip1 (residues 188–1088) and fractions harvested for the enzymatic assay shown in C (fractions A13-B12). A coomassie-stained SDA-PAGE gel showing the respective fractions of ScVip1 is shown alongside.
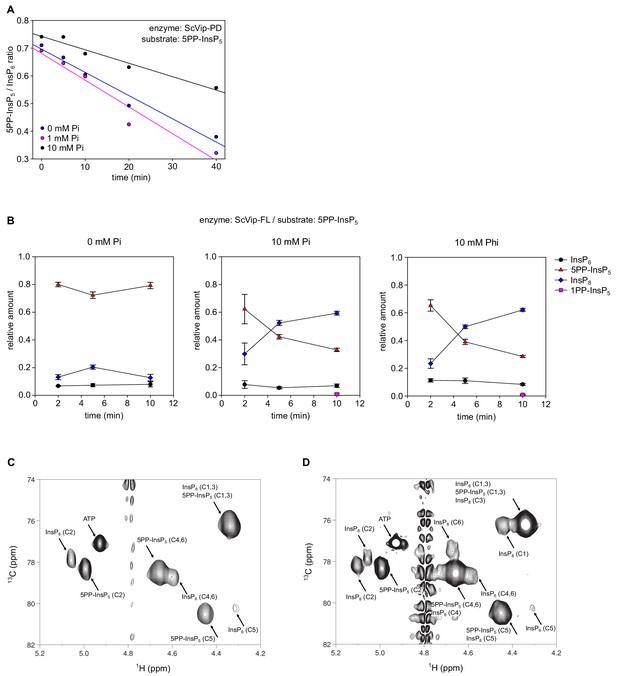
Pi inhibits the ScVip1-PD phosphatase activity, but promotes synthesis of InsP8 catalyzed by full-length ScVip1.
(A) Decay of the [13C6]5PP-InsP5 substrate during the NMR-time course experiment. 2 µM ScVip1-FL was incubated with [13C6]5-PP-InsP5 in the presence of 0 mM Pi (blue), 1 mM Pi (magenta) or 10 mM Pi (black). (B) Relative amount of products recorded by NMR in time-course kinase assays, incubating 1 µM full-length ScVip1 (FL) with 80 µM [13C6]5PP-InsP5 in the presence of 0 mM Pi, 10 mM Pi or 10 mM Pi, respectively. (C, D) 2D 1H-13C-HMBC spectra of the products produced by ScVip1-FL incubated with [13C6]5PP-InsP5 in the presence of 0 mM Pi (C) or 10 mM Pi (D) at 10 min.
Additional files
-
Supplementary file 1
Transgenic lines and genetic crosses used in the study.
(a) Crossing of transgenic Arabidopsis thaliana VIH T-DNA lines. ♀: used as mother for crossing; ♂: used as father for crossing. D: double mutant that shows a characteristic vih1-2 vih2-4 growth phenotype; W: double mutant that displaying a wild-type like phenotype. (b) Segregation ratio analysis in the progeny of independent lines of heterozygous plants shown in (a). (c) Overview of all transgenic lines used in this study.
- https://doi.org/10.7554/eLife.43582.026
-
Supplementary file 2
Primers used in this study.
(a) Primers for characterizing T-DNA mutants. (b) Primers for the cloning of binary vectors. (c) Primers for cloning constructs for recombinant protein expression in insect cells. (d) Primers for q/RT-PCR experiments.
- https://doi.org/10.7554/eLife.43582.027
-
Supplementary file 3
Original qPCR data (expressed as 2-ΔCT values) used to generate expression heat maps.
(a) Data corresponding to heat maps shown in Figure 1G. (b) Data corresponding to heat maps shown in Figure 2D. (c) Data corresponding to heat maps shown in Figure 3D. (d) Data corresponding to heat maps shown in Figure 2—figure supplement 3C. (e) Data corresponding to heat maps shown in Figure 6E.
- https://doi.org/10.7554/eLife.43582.028
-
Transparent reporting form
- https://doi.org/10.7554/eLife.43582.029





