Thermosensitive alternative splicing senses and mediates temperature adaptation in Drosophila
Figures
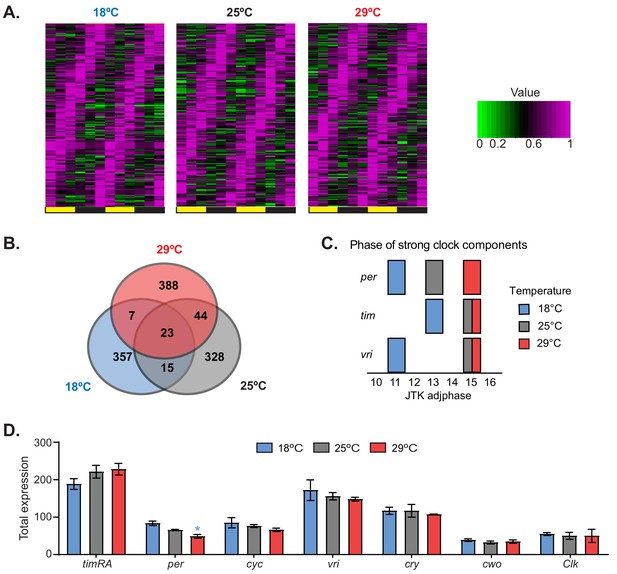
Temperature remodels the circadian transcriptome.
(A) Heat maps of normalized expression of genes that display 24 hr cycling expression at different temperatures. Flies were entrained at the indicated temperatures for 3 days in 12:12 LD conditions. Fly heads were collected every 4 hr. Cycling expression was assessed as described in the 'Materials and methods' section. (B) Venn diagrams of the numbers of oscillating genes at different temperatures. (C) Phases of the core clock genes tim, per, and vri that display strong cycling at 18°C, 25°C, and 29°C. The blue, gray, and red blocks indicate the phases of these genes at 18°C, 25°C, and 29°C, respectively. (D) Total expression of the indicated clock genes in the microarray for each temperature. Only per shows a significant difference between 18°C and 29°C. Plotted are means ± SEM.
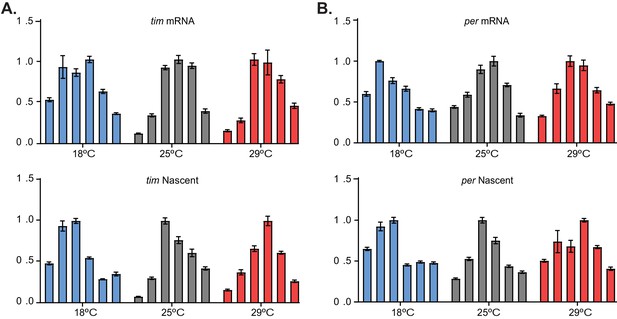
tim transcription is more efficient at 18°C than at 25°C or 29°C.
(A) qRT-PCR analyses of tim mRNA (top) and chromatin-bound tim (bottom) from the heads of wild-type flies entrained at 18°C (blue), 25°C (gray), or 29°C (red) isolated at timepoints ZT3, ZT7, ZT11, ZT15, ZT19, and ZT23 (consecutive bars). The values were normalized to the maximum, and each bar represents the mean ± SEM. (B) qRT-PCR analysis for per performed as described in panel (A).
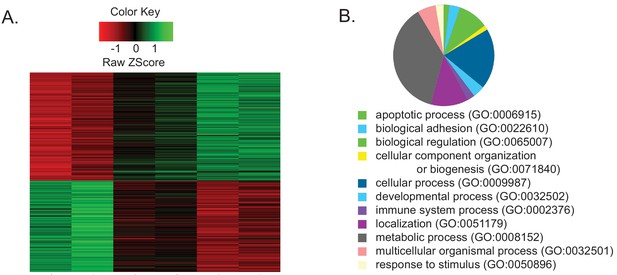
Temperature regulates gene expression.
(A) Heat map showing an unsupervised clustering of differentially expressed genes from a microarray experiment performed on RNA extracted from heads of flies entrained at three temperatures. Each lane represents a mix of all time points at a given temperature. (B) A pie chart demonstrating GO terms that are enriched among the differentially expressed genes.
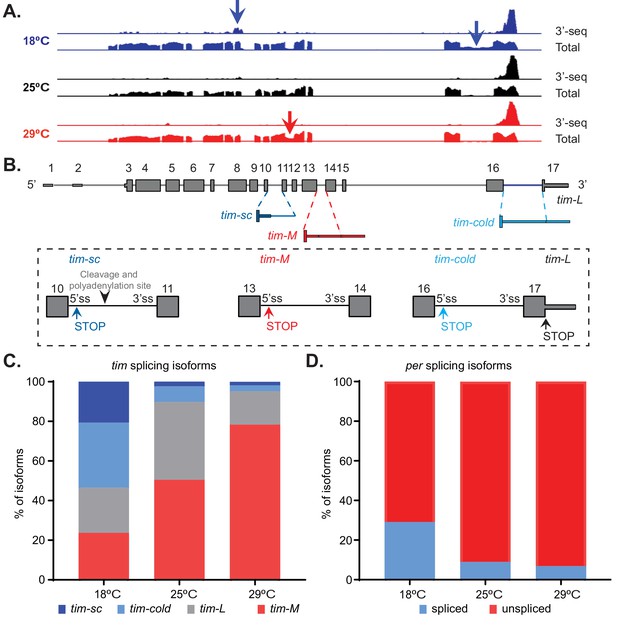
Temperature modulates tim alternative splicing.
(A) Integrative Genomics Viewer (IGV) snapshot of the tim locus, indicating the expression of this gene at 18°C (blue), 25°C (black), and 29°C (red). The presented data include the aggregated data from the 3′ RNA-seq datasets presented in Figure 1 (upper traces) as well as full transcript polyA+ RNA-seq datasets (lower traces). The latter include two time points at 25°C and three time points at 18°C and 29°C. The arrows indicate the alternative splicing events that are regulated by temperature. (B) A schematic of the alternatively spliced tim isoforms. Constitutive exons are shown in gray, sequences found mainly at high temperatures in red, and sequences found mainly at low temperatures in blue. Close-ups of the exons surrounding each non-canonical isoform are shown in rectangles. Alternative stop codons (STOP) and cleavage and polyadenylation sites in these isoforms are also indicated. (C) Quantification of the relative amounts of tim-sc (dark blue), tim-cold (light blue), tim-M (red), and tim-L (gray) isoforms based on the data shown in panel (A). To quantify the different isoforms, we counted the relative number of spliced junctions that are specific for each isoform. (D) Quantification of the relative amount of spliced per (blue) and unspliced per (red) RNAs at the three assayed temperatures.
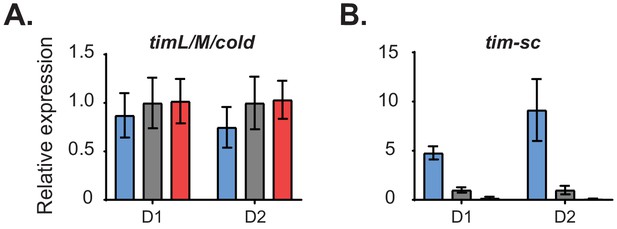
Relative expression of (A) tim-L/M/cold and (B) tim-sc in the two gene expression sets described in Figure 1.
Expression levels of tim-L, tim-cold, and tim-M are reported together as they share the same 3′ end. For each digital gene expression dataset, we averaged the expression of the different tim isoforms over the day at each temperature. Results are shown normalized to 25°C. Each bar represents the mean ± SEM. D1 = Dataset1; D2 = Dataset2.
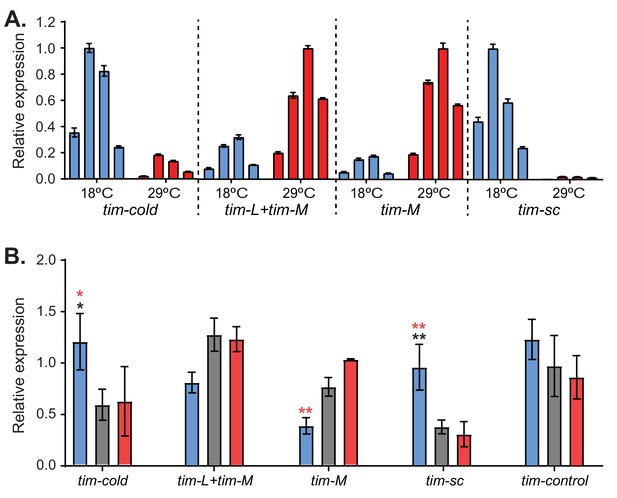
tim expression is regulated by temperature co- and post-transcriptionally.
(A) Isoform-specific qRT-PCR of tim in the heads of flies entrained at 18°C (blue) and 29°C (red). Each bar represents the mean at a different time point (ZT3, ZT9, ZT15, and ZT21) ± SEM. Values were normalized to the maximum of both temperatures. (B) Quantification of the different tim isoforms in the adult brain at ZT11 after entraining the flies at 18°C (blue), 25°C (gray), and 29°C (red). Each bar represents the mean levels ± SEM (n = 3). Black stars indicate statistically significant difference relative to flies kept at 25°C; red stars indicate significant difference relative to flies entrained at 29°C. Significance was determined by two-way ANOVA using Sidak's multiple comparisons test. *, p<0.05; **, p<0.01.
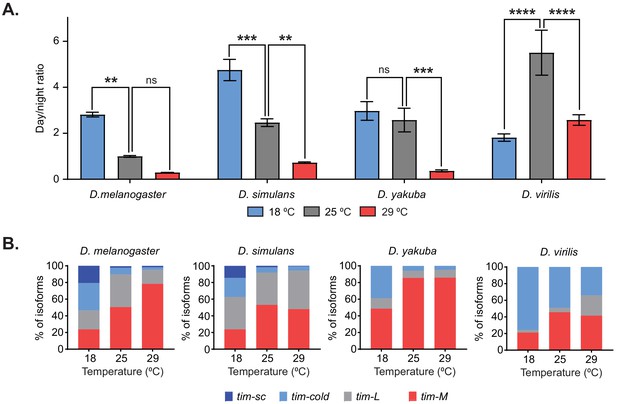
Temperature-specific alternative splicing is conserved across different Drosophila species.
(A) Quantification of the ratio between day and night counts of locomotor activity for D. melanogaster, D. simulans, D. yakuba, and D. virilis at 18°C (blue), 25°C (gray), and 29°C (red). Each bar represents the ratio between the mean locomotor activity during the day and during the night over four days (n = 4) ± SEM. Significance was calculated by performing two-way ANOVA analysis (**, p<0.01; ***, p<0.001; ****, p<0.0001). (B) Quantification of the relative amounts of tim-sc (dark blue), tim-cold (light blue), tim-M (red), and tim-L (gray) from whole transcriptome polyA+ mRNA-seq at the indicated temperatures. Each different isoform was quantified as in Figure 2C.
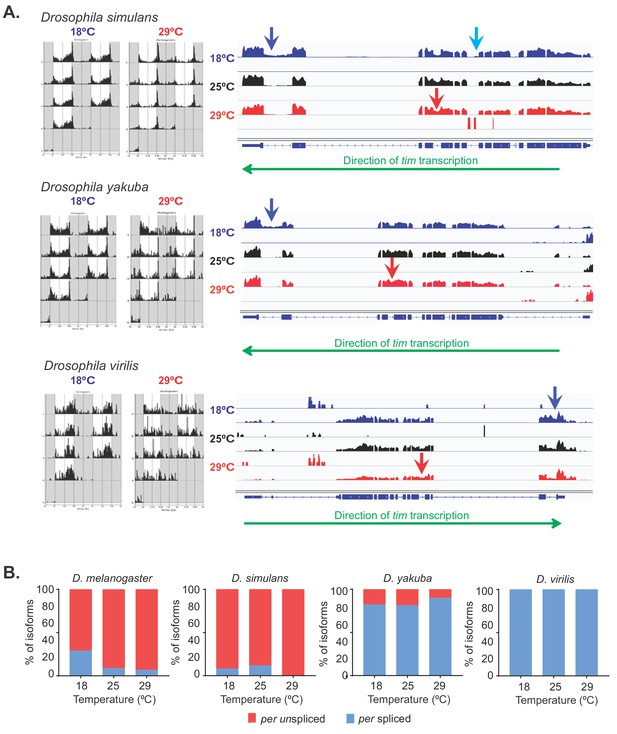
Temperature-specific alternative splicing is conserved across Drosophila species.
(A) IGV snapshot of whole-transcript RNA-seq showing the tim locus in D. simulans, D. yakuba, and D. virilis (top to bottom) at different temperatures. The upper panel (blue) corresponds to 18°C, the middle one (black) to 25°C and the lower one (red) to 29°C. Arrows indicate the sequences that are unique to the different isoforms: tim-sc (light blue), tim-M (red), and tim-cold (blue). Green arrows below the IGV snapshots indicate the direction of tim transcription. (B) Quantification of the relative amounts of spliced per (light blue) and unspliced per (red) from whole transcriptome polyA+ mRNA-seq in all species at the indicated temperatures. To quantify the different isoforms, we counted the relative number of spliced junctions that were specific for each isoform.
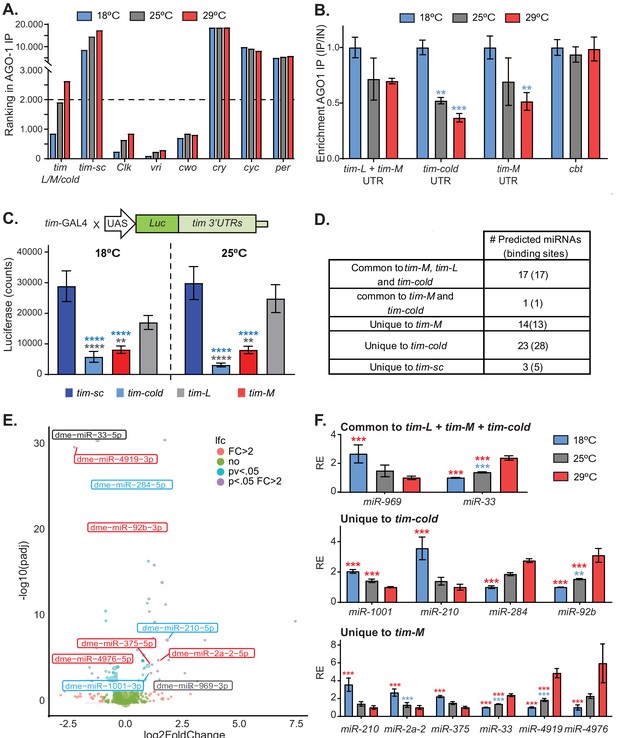
Alternatively spliced isoforms of tim are post-transcriptionally regulated.
(A) Graph of the ranking of enrichment of RNAs in AGO-1 immunoprecipitates isolated from flies entrained at 18°C (blue), 25°C (gray), and 29°C (red). Briefly, we ranked RNAs according to their binding enrichment (lower ranking value corresponds to higher binding to AGO-1). To assess the ranking, we averaged the residuals in a linear model: signal in AGO1-immunoprecipitate/signal in input (n = 2 for each temperature). (B) tim isoform-specific qPCR of AGO1-bound mRNAs at 18°C (blue), 25°C (gray), and 29°C (red). Each bar represents an average of all time points normalized to a negative control (cyc). Blue stars indicate statistically significant levels relative to those at 18°C. Significance was determined by two-way ANOVA analysis using Tukey's multiple comparisons test (**, p<0.01; ***, p<0.001). Four different time points from each temperature were used as biological replica. Bars display the means ± SEM. (C) Luciferase assay of whole flies expressing luciferase fused to different tim 3′ UTRs at 18°C (left) and 25°C (right). Blue and gray stars represent statistically significant difference relative to tim-sc or tim-L, respectively, determined by performing two-way ANOVA analysis using Tukey's multiple comparisons test (*, p<0.05; **, p<0.01). Means ± SEM are plotted (n = 8). (D) Table summarizing the number of putative miRNA-binding sites identified in the different tim isoforms using TargetScan. (E) Volcano plot showing miRNAs that have differential expression at 18°C and 29°C (n = 4 for each temperature). The color code represents the significance for each miRNA differentiating all the combinations, for fold change >2 and adjusted p value <0.05. miRNAs with predicted binding sites present in any of the tim isoform 3′ UTRs were identified using TargetScanFly.6.2. Blue, gray, and red indicate putative targets for tim-cold, common sequence, or tim-M 3′ UTRs, respectively. (F) Relative abundances of miRNAs that are differentially expressed (fold change >2 and adjusted p value < 0.05) at different temperatures and that are predicted to target the 3′ UTR of the different isoforms. For each miRNA, the values were normalized to the minimum and their relative expression (RE) is plotted ± SEM. Blue and red stars represent statistical significance relative to expression at 18° or 29°C, respectively, determined using a negative binomial model employing DeSeq2. See details in the 'Materials and methods' (**, p<0.01; ***, p<0.001).
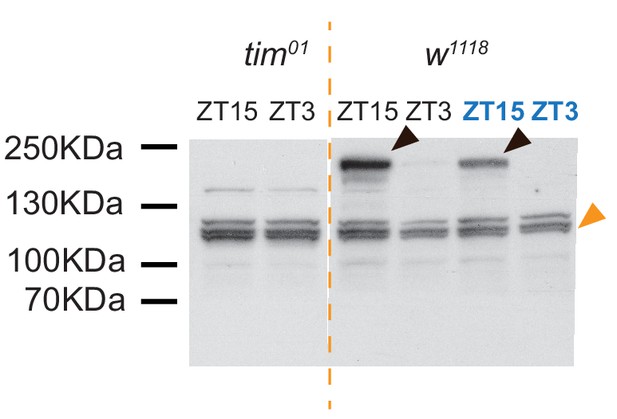
Western blot of TIM at 25°C (black) and 18°C (blue) using heads from tim01 and w1118 flies at ZT3 and ZT15.
Black arrowheads mark the canonical TIM-L isoform; orange arrowheads mark the ~60 kDa smaller degradation product.
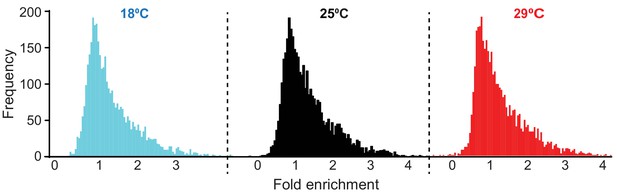
Density plot representing 4000 genes selected as ‘AGO1-associated’.
The distributions are the frequencies of each fold enrichment for a particular temperature.
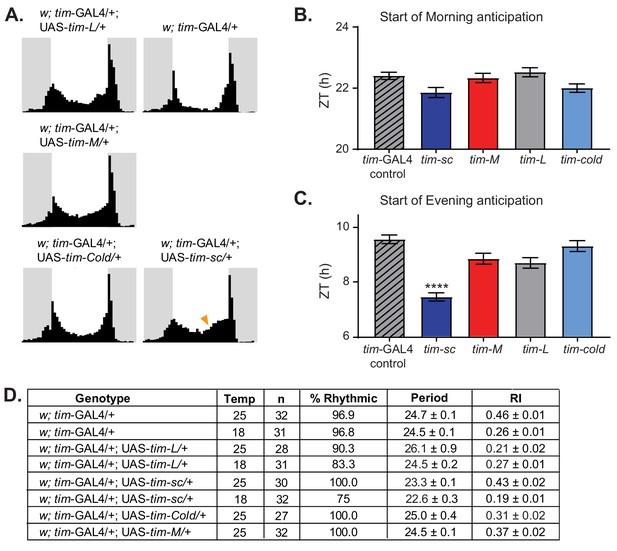
Overexpression of tim isoforms at 25°C results in various behavioral defects.
(A) Average activity over three days in 12:12 LD at 25°C of flies of the indicated genotypes. Light phase is represented in white and dark-phase in gray. (B) Quantification of the start of the morning activity component in the last three days of LD (n = 27–31) ± SEM. (C) Quantification of the start of the evening activity component in the last three days of LD (n = 27–31) ± SEM. Stars represent statistical significance relative to tim-Gal4 control calculated by one-way ANOVA using Tukey's multiple comparisons test (****, p<0.0001). (D) Table summarizing the behavior of the indicated genotypes in DD at 18°C or 25°C, indicating percent rhythmic, period length, rhythmicity index (RI), and respective SEMs.
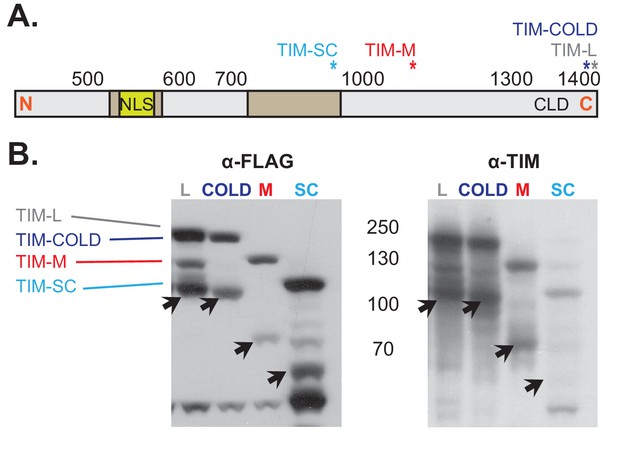
Overexpression of different tim isoforms with a FLAG-tagged C-terminus.
(A) Schematic representation of the TIM protein. The stars mark the locations of the different FLAG-tags for each isoform. NLS, Nuclear Localization Signal; CLD, Cytoplasmic Localization Domain. (B) Western blot membranes showing anti-FLAG (left) and anti-TIM (right) staining of the overexpression of the four 3′ FLAG-tagged UAS-tim isoforms in S2 cells. Arrows mark a degradation product that is ~60 kDa smaller than the original TIM isoform.
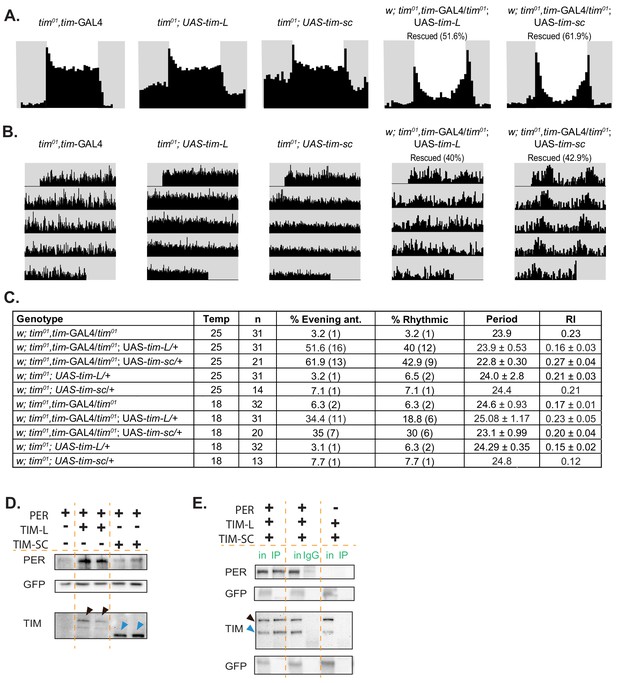
Overexpression of tim-sc at 25°C partially rescues tim01 flies in LD and DD, and TIM-SC binds to PER.
(A) Average activity over three days in 12:12 LD at 25°C of flies of the indicated genotypes. The light phase is represented in white and the dark-phase in gray. The percentage of rescued flies (displaying evening anticipation) is reported in brackets. (B) Actograms of each genotype in DD at 25°C. (C) Summary of the percentage of rescue in LD (evening anticipation component) and in DD (percent rhythmic, period length, rhythmicity index, and their respective SEMs). (D) Representative western blot showing staining for PER (top), GFP (middle), and TIM (bottom) of protein lysates from S2 cells transfected with pAcPerV5 (PER), pAcTimL-HA (TIM-L), and pAcTimSC-HA (TIM-SC); pAcGFP was used as a transfection control (n = 3). Arrows in the TIM staining point to TIM-L (black) and TIM-SC (blue). (E) Representative western blot for PER, GFP, and TIM of PER immunoprecipitates in S2 cells co-transfected with pAcPerV5 (PER), pAcTimL-HA (TIM-L), or pAcTimSC-HA (TIM-SC) (n = 2). IgG and no PER controls were loaded in the middle and right of the membrane. pAcGFP was used as a transfection control. Arrows indicate TIM-L (black) and TIM-SC (blue). in, input; IP, immunoprecipitation.
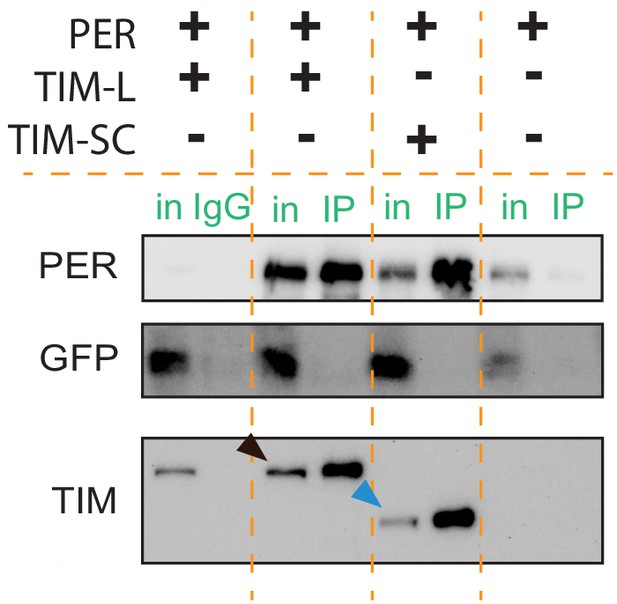
TIM-SC binds to PER.
Representative western blot against PER (top), GFP (middle), and TIM (bottom) of HA immunoprecipitate in S2 cells co-transfected with pAcPerV5 (P) and pAcTimL-HA (L) or pAcTimSC-HA (SC). IgG and no TIM controls are loaded onto each side of the membrane. pAcGFP was used as a transfection control. Arrows in the TIM staining point to TIM-L (black) and TIM-SC (blue). in, input; IP, immunoprecipitation. n = 3.
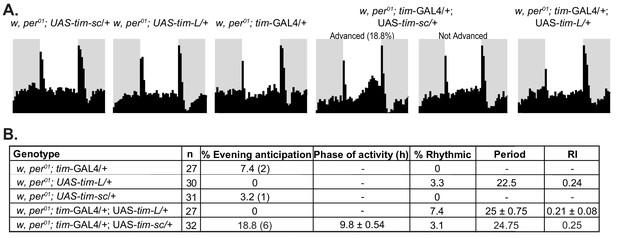
Overexpression of TIM-SC in per01 results in a peak of activity resembling the evening activity component in a small fraction of the flies.
(A) Average activity of three days in 12:12 LD at 25°C of flies of the indicated genotypes. The light phase is represented in white and the dark phase in gray. The percentages of advanced flies are reported in brackets. (B) Summary of the percentages of evening anticipation in LD.
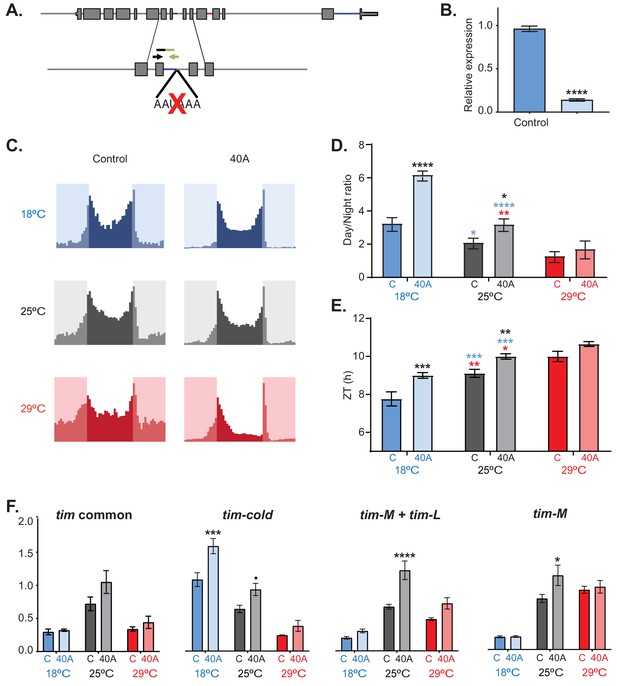
tim-sc null flies have behavioral and molecular defects.
(A) Schematic of the mutation performed by CRISPR used to generate tim-sc null flies (40A flies). The black and green arrows indicate the locations of the forward and reverse primers used for the verification of tim-sc depletion. (B) Levels of tim-sc in 40A flies. The displayed qPCR result was obtained using RNA obtained from the heads of flies entrained at 18°C (n = 5). (C) Average activity during three days of control (left) and 40A (right) flies at 18°C (blue), 25°C (gray), and 29°C (red) in 12:12 LD cycles (n = 23–32). (D) Ratio between total light/dark activity at 18°C (blue), 25°C (gray), and 29°C (red) in 12:12 LD. Blue and red stars represent statistically significant difference between flies kept at 18°C or 29°C and those at 25°C; black stars indicate differences relative to the control flies. Significance was determined by two-way ANOVA using Sidak's multiple comparisons test (n = 16–31). (E) Quantification of the start of the evening activity component on day 3 of 12:12 LD. Blue and red stars represent statistically significant differences between flies kept at 18°C or 29°C and those kept at 25°C, whereas black stars represent differences relative to the control flies, as determined by two-way ANOVA using Sidak's multiple comparisons test (n = 16–31). (F) Assessment by qRT-PCR of exons 5 and 6 (common to all tim isoforms, left), tim-cold (middle left), tim-L + tim-M (middle right), and tim-M (right) at 18°C (blue), 25°C (gray), and 29°C (red) at ZT15 in 12:12 LD. Significance was determined by two-way ANOVA using Sidak's multiple comparisons test for each isoform (n = 5). Each bar represents the mean ± SEM. *, p<0.05; **, p<0.01; ***, p<0.001; ****, p<0.0001.
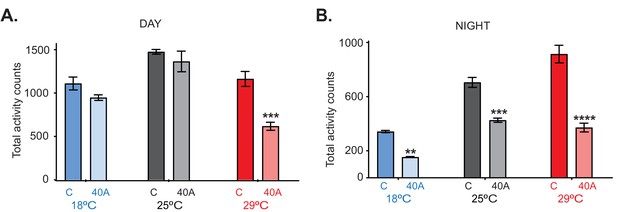
Mean total activity counts in control and 40A mutant flies at 18°C (blue), 25°C (gray), and 29°C (red) in 12:12 LD during the day (A) and night (B) ± SEM (n = 23–32).
Stars indicate significant differences between the control and mutants from the same temperature, as determined by two-way ANOVA using Sidak's multiple comparisons test. **, p<0.01; ***, p<0.001; ****, p<0.0001.
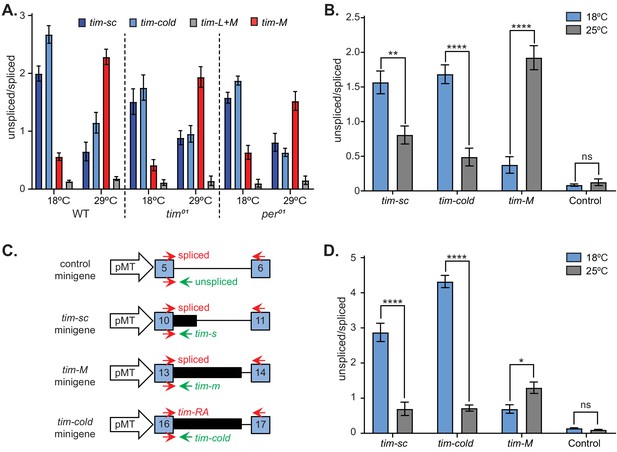
Splicing of timeless is temperature dependent and clock independent.
(A) qRT-PCR results from wild-type, tim01, and per01 clock mutant flies entrained at 18°C or at 29°C. Each bar represents the mean ratio (± SEM) between the unspliced tim and tim-sc (dark blue), tim-cold (light blue), tim-M (red), and tim control (gray). (B) Drosophila S2 cells transfected with pMT-Clk display the same tim alternative splicing patterns observed in living flies at 18°C (blue) and at 25°C (gray). Each bar represents the mean ratio (± SEM) between the unspliced and spliced variant of tim-sc, tim-cold, tim-M and tim control. Significance was determined by two-way ANOVA using Sidak's multiple comparisons test (n = 3). (C) Schematic of the tim isoform minigenes and locations of the primers used to measure the spliced (top) and unspliced (bottom) isoforms of each intron. (D) Mean unspliced/spliced ratios (± SEM) for the indicated minigenes at 18°C (blue) and at 25°C (gray). Significance was determined by two-way ANOVA using Sidak's multiple comparisons test (n = 3). *, p<0.05; **, p<0.01; ***, p<0.001.
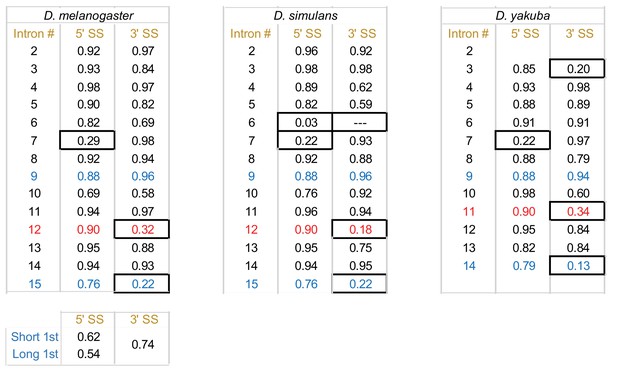
Table summarizing the 3′ and 5′ splicing strengths of each tim intron for D. melanogaster, D. simulans, and D. yakuba.
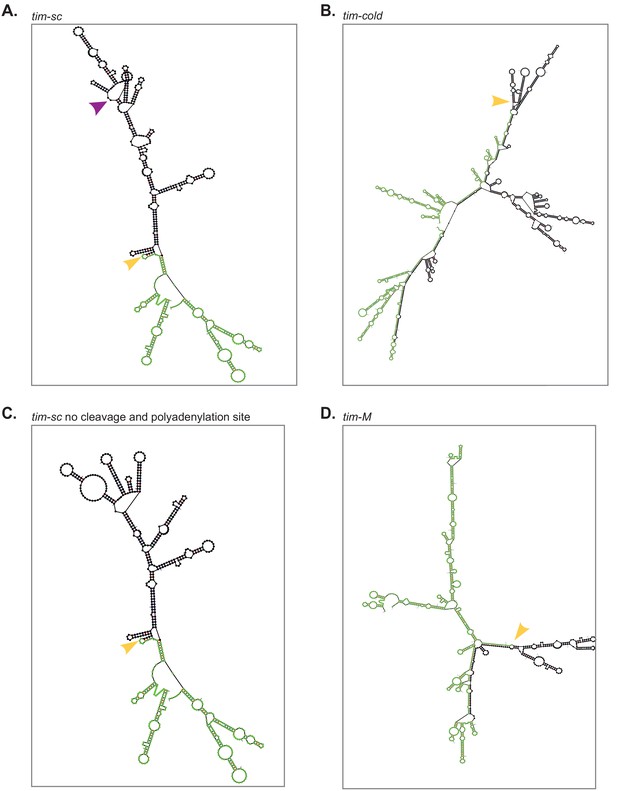
Predicted RNA secondary structures for (A) tim-sc, (B) tim-cold, (C) the tim-sc mutant without the cleavage and polyadenylation site, and (D) tim-M at 25°C for each exon-intron-exon region.
Exons are represented in green; introns are black. Orange arrowheads mark the 5′ splice sites and purple arrowheads indicate the cleavage and polyadenylation site present in tim-sc. Only 350 bp from exons surrounding tim-cold intron were included in panel (C).
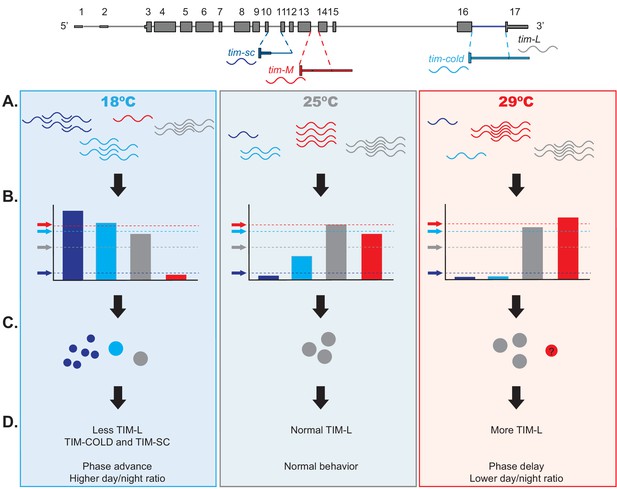
Model of the effects of thermosensitive alternative splicing of timeless in sensing and mediating temperature adaptation.
(A) Temperature affects each one of the four tim isoforms (tim-sc, tim-cold, tim-M, and tim-L, represented in dark blue, light blue, red, and gray, respectively) differently. tim-sc and tim-cold are more abundant at colder temperatures, whereas tim-M is expressed at higher temperatures. Abundance of the canonical isoform, tim-L, does not change with temperature. (B) In addition, miRNAs set the threshold for the amount of mRNA that is translated from each isoform. tim-sc has the lowest and tim-M the highest threshold at all temperatures. (C–D) Differences between the transcript levels and their thresholds result in changes in TIM expression (C) and behavioral changes in the phase and day/night ratios (D) that are linked to temperature adaptation.
Additional files
-
Supplementary file 1
Circadian oscillation analysis.
The circadian analysis was performed using the MetaCycle algorithm. In the table, the p-value, phase, and amplitude are reported for each gene along with its normalized expression at each circadian time point.
- https://cdn.elifesciences.org/articles/44642/elife-44642-supp1-v2.xlsx
-
Supplementary file 2
Gene ontology analysis for the genes with circadian oscillation at each temperature.
Genes are ordered by ranking in a Fisher exact test. Three different ontologies were analyzed: biological process, molecular function, and cellular component. The top 100 genes are reported.
- https://cdn.elifesciences.org/articles/44642/elife-44642-supp2-v2.xlsx
-
Supplementary file 3
miRNA binding sites predicted using TargetScan for each tim RNA isoform 3′ UTR.
Putative miRNA binding sites are reported as chromosome coordinates. Conservation in different insect species is also reported.
- https://cdn.elifesciences.org/articles/44642/elife-44642-supp3-v2.xlsx
-
Supplementary file 4
Differential miRNA expression at each temperature.
Log2(fold change) and adjusted p-value from DeSeq2 are reported for each miRNA.
- https://cdn.elifesciences.org/articles/44642/elife-44642-supp4-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/44642/elife-44642-transrepform-v2.pdf





