Functional cross-talk between allosteric effects of activating and inhibiting ligands underlies PKM2 regulation
Figures
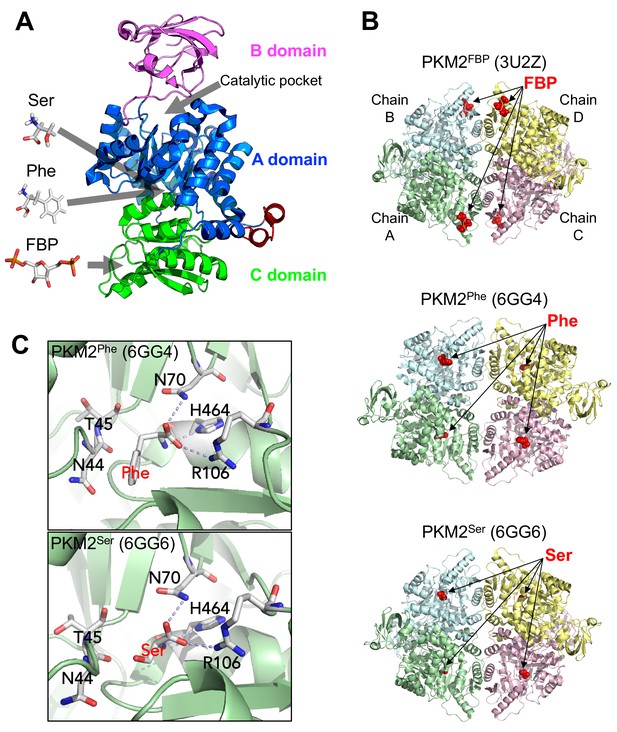
Overview of PKM2 structure and allosteric ligand binding sites.
(A) Domain structure of PKM2 monomer (PDBID: 3BJT) depicting orthosteric and allosteric sites discussed in this work. (B) Published crystal structures of PKM2 bound to FBP, Phe or Ser (PDBID shown in parentheses). (C) Phe and Ser bind to the same pocket in PKM2. Close-up view of the Phe (top) and Ser (bottom) binding pockets with key contact residues [Asn(N)44, Thr(T)45, Asn(N)70, Arg(R)106, His(H)464] shown in stick configuration. Dashed lines indicate hydrogen bonds between PKM2 residues and Phe or Ser.
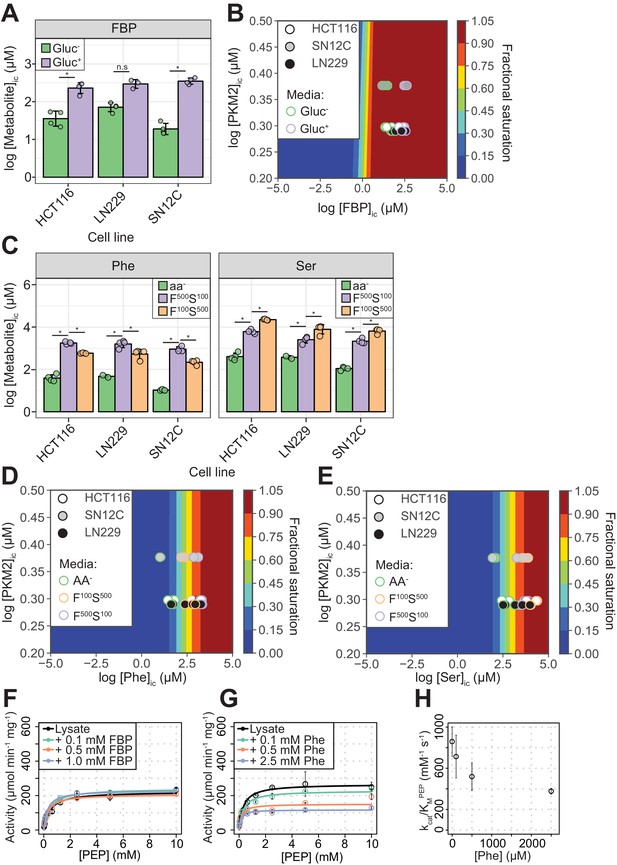
PKM2 allosteric effector concentrations in cells predict saturating binding of FBP and sub-saturating binding of Phe and Ser.
(A) Intracellular concentrations of FBP ([FBP]ic) measured using liquid-chromatography mass spectrometry (LC-MS), from HCT116 (colorectal carcinoma), LN229 (glioblastoma) and SN12C (renal cell carcinoma) cells cultured in RPMI media containing 11 mM glucose (Gluc+), or 0 mM (Gluc–) for 1 hr. Statistical significance was assessed using a Wilcoxon rank-sum test. Asterisk (*) marks significant changes (p-value<0.05). (B) Phase diagram for intracellular FBP binding to PKM2 computed for a range of [FBP] and [PKM2] values using 174 nM as the upper-limit estimate of the KDFBP, obtained as shown in Figure 2—figure supplement 2. Colour scale represents fractional saturation of PKM2 with ligand. A fractional saturation of 0 indicates no FBP bound to PKM2 and a fractional saturation equal to one indicates that each FBP binding site in the cellular pool of PKM2 would be occupied. Experimental fractional saturation values were estimated from [FBP]ic obtained from (A) and [PKM2]ic was determined using targeted proteomics (see Materials and methods and Supplementary file 1). The predicted fractional saturation for each of the three cell lines (four technical replicates) is shown as shaded open circles in the phase diagram. (C) Intracellular concentrations of Phe ([Phe]ic) and Ser ([Ser]ic) measured as in (A), in HCT116, LN229 and SN12C cells cultured in Hank’s Balanced Salt Solution (HBSS) without amino acids (aa-), HBSS containing 100 µM Phe and 500 µM Ser (F100 S500), or HBSS containing 500 µM Phe and 100 µM Ser (F500 S100). The low concentrations of Phe and Ser are similar to human serum concentrations (Tardito et al., 2015). [Phe]ic and [Ser]ic were not affected by extracellular glucose concentration (Figure 2—figure supplement 4A), neither did extracellular Phe and Ser concentrations influence [FBP]ic (Figure 2—figure supplement 4B). Statistical significance was assessed as in (A). (D) Phase diagram for Phe computed as in (B) using [Phe]ic from (C). (E) Phase diagram for Ser computed as in (B) using [Ser]ic from (C). (F) PKM2 activity in lysates of HCT116 cells cultured in RPMI (Gluc+). Measurements were repeated following the addition of either 0.1, 0.5 or 1.0 mM of exogenous FBP. Initial velocity curves were fitted using Michaelis-Menten kinetics and the absolute concentration of PKM2 in the lysates was estimated using quantitative Western blotting (Figure 2—figure supplement 5B), to calculate PKM2 specific activity. (G) PKM2 activity in HCT116 cell lysates as in (F), but with addition of exogenous Phe. (H) Plot of kcat/KM versus [Phe] from (G) revealing a dose-dependent inhibitory effect of Phe on the activity of PKM2 in HCT116 lysates.
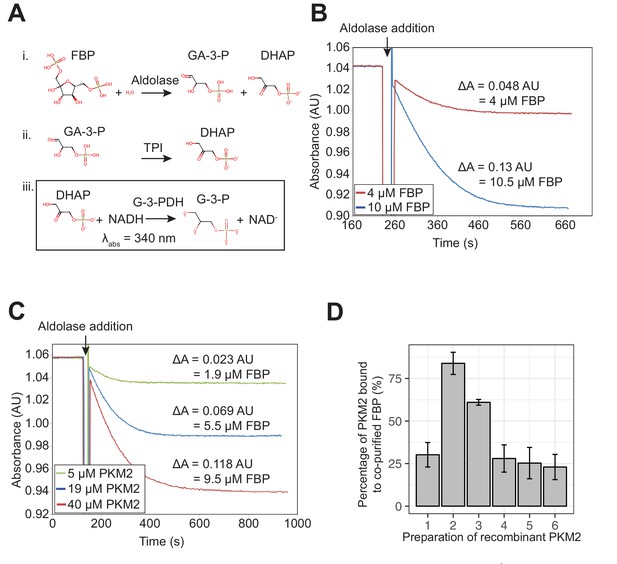
Detection of residual FBP in purified recombinant PKM2 preparations.
(A) FBP co-purified with recombinant PKM2 was quantified using a spectrophotometric coupled-enzyme assay that employs aldolase, triose phosphate isomerase (TPI) and glycerol-3-phosphate dehydrogenase (G-3-PDH). The final step of the three-enzyme reaction involves the oxidation of NADH, leading to a decrease in absorbance at 340 nm. (B) The aldolase coupled assay in (A) can reliably detect as low as 4 µM FBP. Sensitivity of the aldolase assay was tested by adding 4 or 10 µM purified FBP, and oxidation of NADH was initiated by adding the aldolase enzyme following an equilibration period. Amount of FBP was calculated from the measured decrease in the amount of NADH (one molecule of FBP consumed for every two molecules of NADH oxidised). (C) Levels of FBP detected in increasing amounts of recombinant PKM2. Indicated amounts of purified PKM2 were heat-precipitated at 95°C to release any co-purified FBP and the supernatant was used for FBP quantification with the aldolase assay as in (A). (D) Amounts of FBP quantified, as in (A), for six independent preparations of purified recombinant PKM2. The amounts of co-purified FBP are represented as a proportion of the amount of recombinant PKM2. Preparations of PKM2 determined to have < 25% co-purifying FBP were used throughout this study.
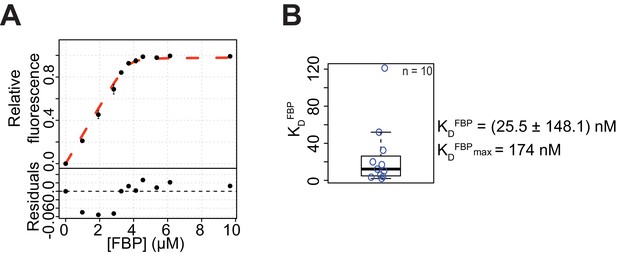
FBP binds to PKM2 with nM affinity.
(A) A representative curve of FBP binding to PKM2, derived from measuring fluorescence (λEX = 280 nm, λEM = 290–450 nm) of PKM2 tryptophan residues (two of which are proximal to the FBP binding pocket) at different FBP concentrations. The apparent binding constant (KDFBP) was estimated from a 1:1 binding model, fit to the experimental data (dashed red line) using a non-linear least-squares fitting procedure (see Materials and methods). The average measurements and standard deviations from four independent experiments are shown. (B) The average KDFBP = (25.5 ± 148.1) nM (KDFBP upper limit = 174 nM) calculated from ten replicate titration measurements as in (A). The large error in the affinity estimate was due to the high concentration of PKM2 (5 µM), relative to the apparent KDFBP, that was necessarily used to monitor intrinsic PKM2 fluorescence changes upon FBP addition (see Materials and methods for details).
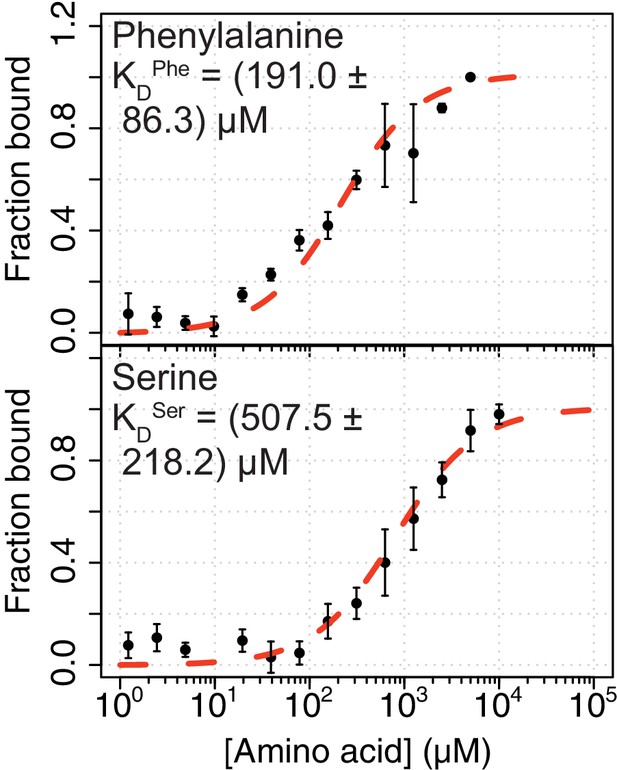
Affinities of Phe and Ser for PKM2.
Affinity of Phe and Ser for PKM2 determined using microscale thermophoresis (MST) measurements of fluorescein-labelled PKM2 (30 nM). Indicated binding affinities were estimated using a 1:1 binding model. The average measurements and standard deviations from four independent experiments are shown.
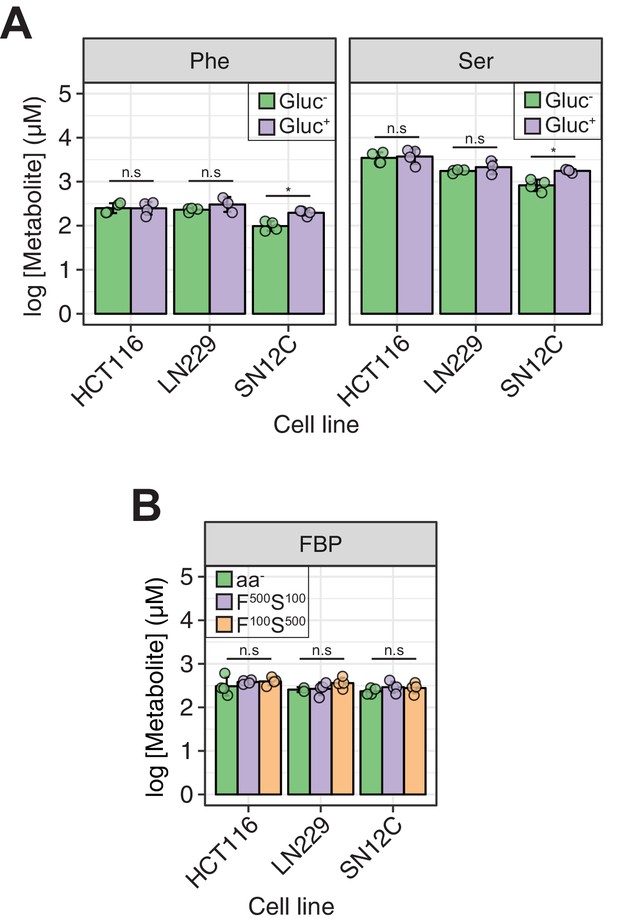
Acute (1 hr) modulation of glucose and Phe/Ser concentration in the media does not affect intracellular concentrations of Phe/Ser or FBP, respectively.
(A) [Phe]ic and [Ser]ic in cells as in Figure 2A showing no significant changes upon changes of glucose concentration in media. Statistical significance determined as in Figure 2A. (B) [FBP]ic in cells as in Figure 2C showing no significant changes depending on Ser or Phe concentration in HBSS. Statistical significance determined as in Figure 2A.
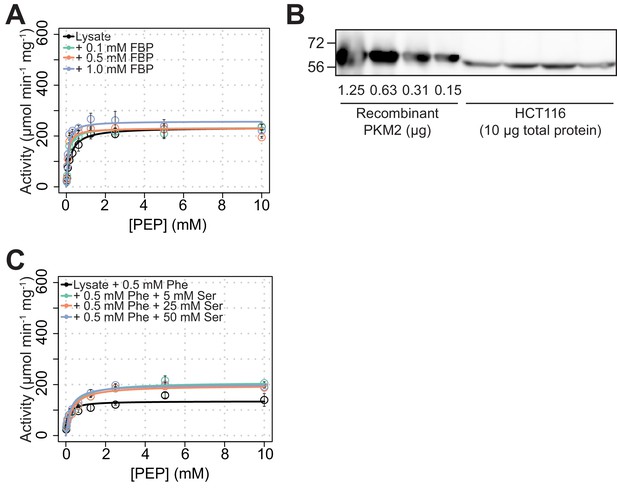
PKM2 activity in cell lysates can be significantly modulated by exogenous amino acids but not by FBP.
(A) PKM2 activity as in Figure 2F but in lysates of HCT116 cells cultured in RPMI without glucose (Gluc-). (B) Quantification of PKM2 in HCT116 lysates. Western blot of recombinant PKM2 or HCT116 cell lysates (Gluc+ media) was probed with a PKM2 antibody. An interpolation of the linear fit of the band intensities of four amounts of recombinant PKM2 was used to estimate the amounts of PKM2 protein per 1 µg of total protein in HCT116 cell lysates, in order to determine specific activity in panels (A) and (C), and in Figure 2F and G. (C) PKM2 activity in HCT116 cell lysates as in Figure 2G but with 0.5 mM Phe and increasing amounts of Ser, as indicated.
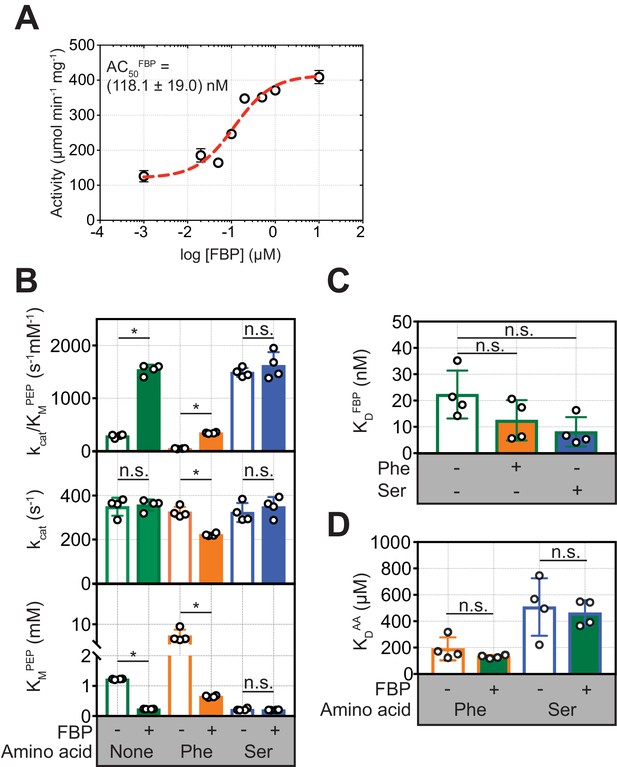
FBP influences the inhibition of PKM2 activity by Phe.
(A) PKM2 (5 nM) activity measured over a range of added FBP concentrations (0.01–10 µM) with constant substrate concentrations (PEP = 1.5 mM and ADP = 5 mM). The apparent activation constant (AC50FBP) was estimated by fitting the resulting data to a binding curve (red line) assuming a 1:1 stoichiometry. Means and standard deviations of four separate experiments are plotted. (B) Steady-state kinetic parameters of purified recombinant human PKM2, in the absence of added ligands; in the presence of 2 µM FBP, 400 µM Phe and 200 mM Ser alone; and after addition of either 400 µM Phe or 200 mM Ser to PKM2 pre-incubated with 2 µM FBP. Initial velocity curves were fit to Michaelis-Menten kinetic models. Each titration was repeated four times. Statistical significance was assessed using a Wilcoxon rank-sum test. Asterisk (*) marks significant changes (p-value<0.05). (C) Binding constant of FBP to PKM2 obtained from fluorescence emission spectroscopy measurements in the absence and in the presence of either 400 µM Phe or 200 mM Ser. (D) Binding constants for Phe and Ser to PKM2, in the absence or presence of 2 µM FBP, obtained from microscale thermophoresis (MST) measurements. MST was used, rather than intrinsic fluorescence spectroscopy as for FBP, due to the absence of tryptophan residues proximal to the amino acid binding pocket on PKM2. Significance was assessed as in (A).
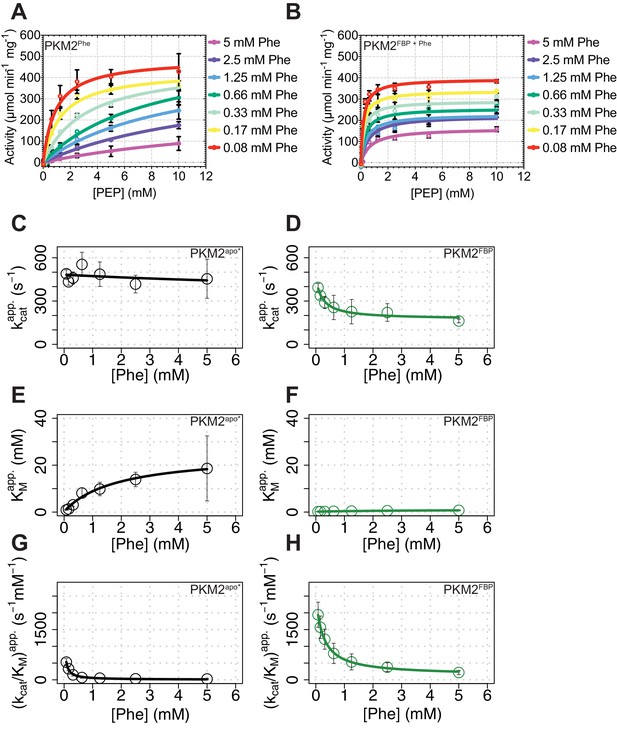
FBP influences the kinetics of PKM2 inhibition by Phe - experimental data.
(A) Purified recombinant PKM2 activity was assayed over a range of phosphoenolpyruvate (PEP) concentrations and with the indicated concentrations of phenylalanine (Phe). The averages and standard deviations of four separate experiments are shown. Initial velocity curves were fitted using the Michaelis-Menten model. (B) PKM2 activity was assayed in the presence of increasing concentrations of the inhibitor Phe, as in (A), in the presence of a constant concentration of 2 µM FBP. Rate curves were fitted to Michaelis-Menten kinetic models, from which the parameters kcat, KMPEP and kcat/KMPEP were computed for Phe inhibition of PKM2apo* (black; C, E and G) and PKM2FBP (green; D, F and H). The dependence of the three steady-state kinetic parameters kcat, KMPEP and kcat/KMPEP were fit to the steady-state modifier-rate equations (see Materials and methods).
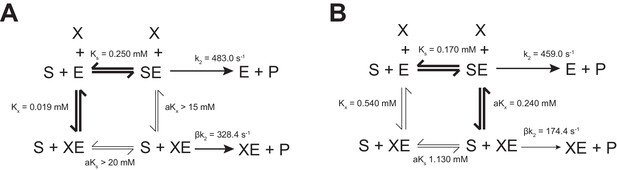
FBP influences the kinetics of PKM2 inhibition by Phe - kinetic mechanism models.
(A) The steady-state mechanism of allosteric inhibition is shown from a single-substrate-single-modifier representation of PKM2apo catalysis; where E is the enzyme, S is the substrate PEP and X is the allosteric effector Phe. Equilibrium constants were assigned from fittings of the dependence of the kinetic parameters on the concentration of Phe, determined above. The mechanism of PKM2 inhibition by Phe in the absence of FBP is found to be hyperbolic-specific (Baici, 2015). (B) The mechanism of Phe inhibition of PKM2FBP was determined as in (A), and was found to be of the variety of hyperbolic-mixed inhibition (Baici, 2015).
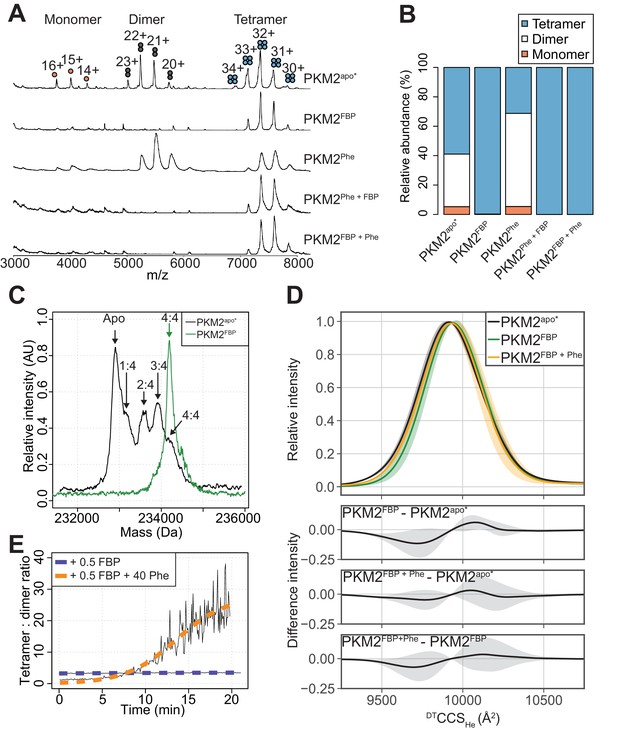
FBP modulates the effects of Phe on PKM2 oligomerisation.
(A) Native mass spectra of 10 µM PKM2 in 200 mM ammonium acetate at pH 6.8, in the absence of allosteric ligands (PKM2apo*), or in the presence of: 10 µM FBP (PKM2FBP), 300 µM Phe (PKM2Phe), 300 µM Phe followed by addition of 10 µM FBP (PKM2Phe + FBP) or 10 µM FBP followed by addition of 300 µM Phe (PKM2FBP + Phe). (B) Relative abundance of PKM2 monomers, dimers and tetramers obtained from the spectra shown in (A) by computing the area of the peaks corresponding to each of the three oligomeric states. Relative peak areas were calculated as a percentage of the total area given by all charge-state species in a single mass spectrum. (C) Deconvolved mass spectra of PKM2 tetrameric species in the absence of any added ligands (PKM2apo*) or presence of FBP (PKM2FBP). PKM2apo* has five distinct mass peaks, separated by approximately 340 Da (equivalent to the weight of FBP), corresponding to tetrameric PKM2apo*, and tetrameric PKM2 bound to 1, 2, 3 and 4 molecules of FBP, respectively. See Table 4 for the theoretical, and Table 5 for the experimentally measured masses of PKM2, FBP and their complexes. The spectrum of PKM2FBP contains a single peak, corresponding to tetrameric PKM2 bound to four molecules of FBP. (D) DTCCSHe distribution of PKM2apo*, PKM2FBP and PKM2FBP+Phe tetramers calculated from analyses of arrival time distribution measurements of PKM2 tetramer peaks (see Materials and methods). The plots at the bottom show the mean difference of the DTCCSHe distributions between the indicated liganded states. Grey shaded regions show the standard deviations of the distribution differences. (E) Change, over time, in the oligomeric state of PKM2 upon addition of sub-stoichiometric FBP and saturating Phe concentrations. Oligomerisation is reported as the ratio of the tetramer 32 + charge state peak relative to the dimer 22 + charge state peak, obtained from mass spectra of 10 µM PKM2 following addition of either 5 µM FBP, or 5 µM FBP and 400 µM Phe over the course of 20 min. The kinetics of tetramerisation were estimated from a two-state sigmoidal model (orange and blue dashed lines, see Materials and methods). In the legend, 0.5 FBP and 40 Phe indicate molar ratio of these ligands compared to PKM2.
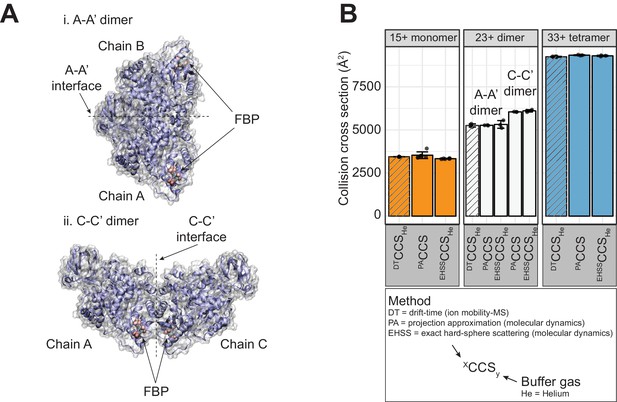
Evidence from IM-MS and MD simulations that PKM2 dimers predominantly adopt the A-A’ configuration.
(A) Representation of the two possible dimer assemblies based on PKM2 symmetry: i. the A-A’ dimer (protomers associated along the A-A’ interface) and ii. the C-C’ dimer (protomers associated along the C-C’ interface). (B) Evidence from IM-MS that the observed PKM2 dimeric species adopt the A-A’ configuration. The collisional cross section (DTCCSHe) of 15 + monomer species (orange), 23 + dimer species (white) and 33 + tetramer species (blue) PKM2apo* were experimentally determined using ion mobility-mass spectrometry (IM-MS; striped bars). The exact hard sphere collision cross section (EHSSCCSHe) and projection approximation collision cross section (PACCS) were calculated from short in vacuo molecular dynamics simulations of 15 + monomeric (orange), 23 + dimer (white) and 33 + tetrameric (blue) PKM2apo models. The DTCCSHe of the dimeric species is very similar to that of the A-A’ dimer calculated by both EHSSCCSHe and PACCS.
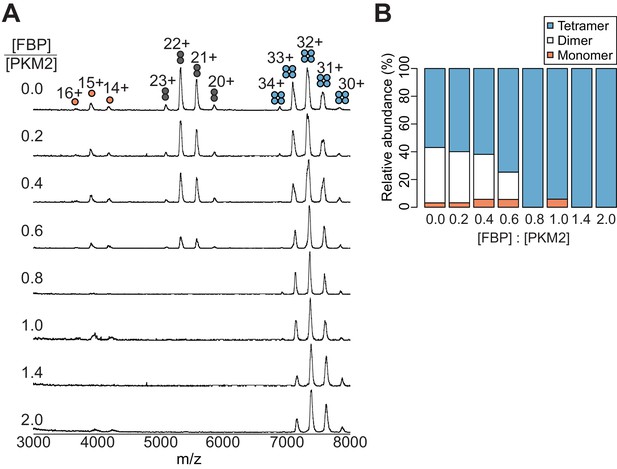
FBP promotes a dose-dependent conversion of PKM2 monomers and A-A’ dimers into the tetrameric species.
(A) FBP-dose–dependent conversion of PKM2 monomers and dimers into tetramers. Mass spectra of PKM2 obtained following pre-incubation with increasing amounts of FBP resulting in the indicated FBP/PKM2 ratios. (B) Relative abundances of PKM2 monomers, dimers and tetramers determined from (A).
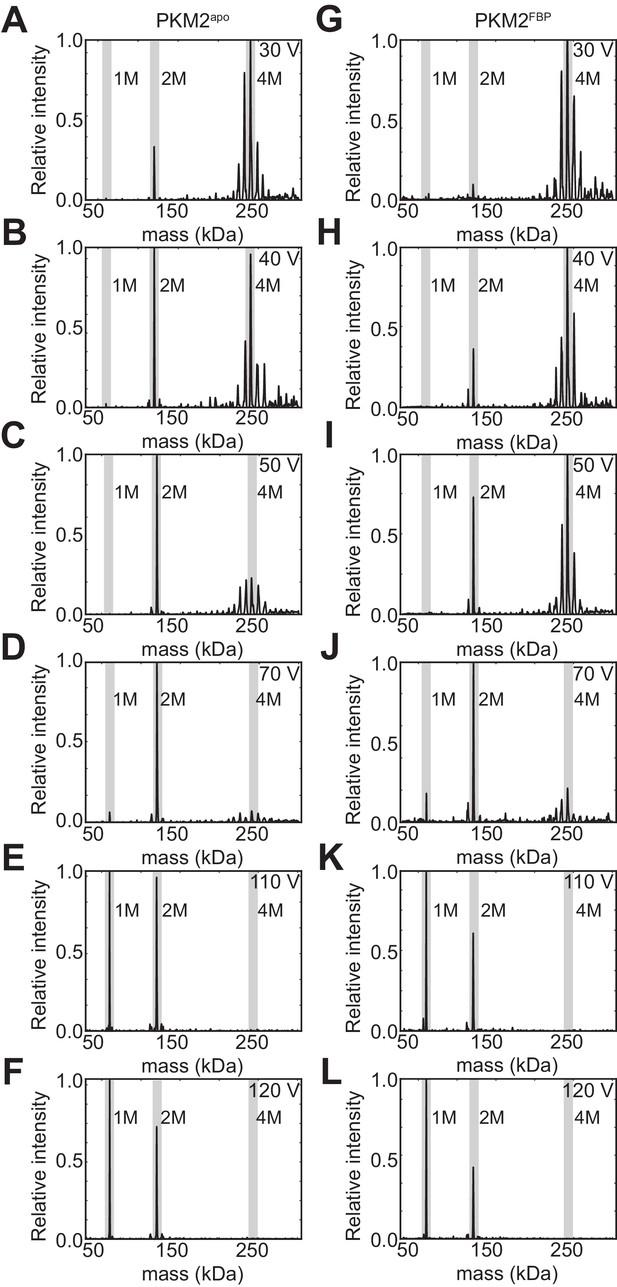
FBP binding increases the stability of PKM2 tetramers.
Mass deconvolved spectra showing the relative abundance of oligomeric species produced from the 32 + charge state precursor of tetrameric PKM2 at increasing collision voltages (40 V – 120 V; lab-frame), obtained using surface-induced dissociation (SID) (see Materials and methods) for PKM2apo* (A-F) and PKM2FBP (G-L). The positions corresponding to the tetramer, dimer and monomer species are highlighted in grey.
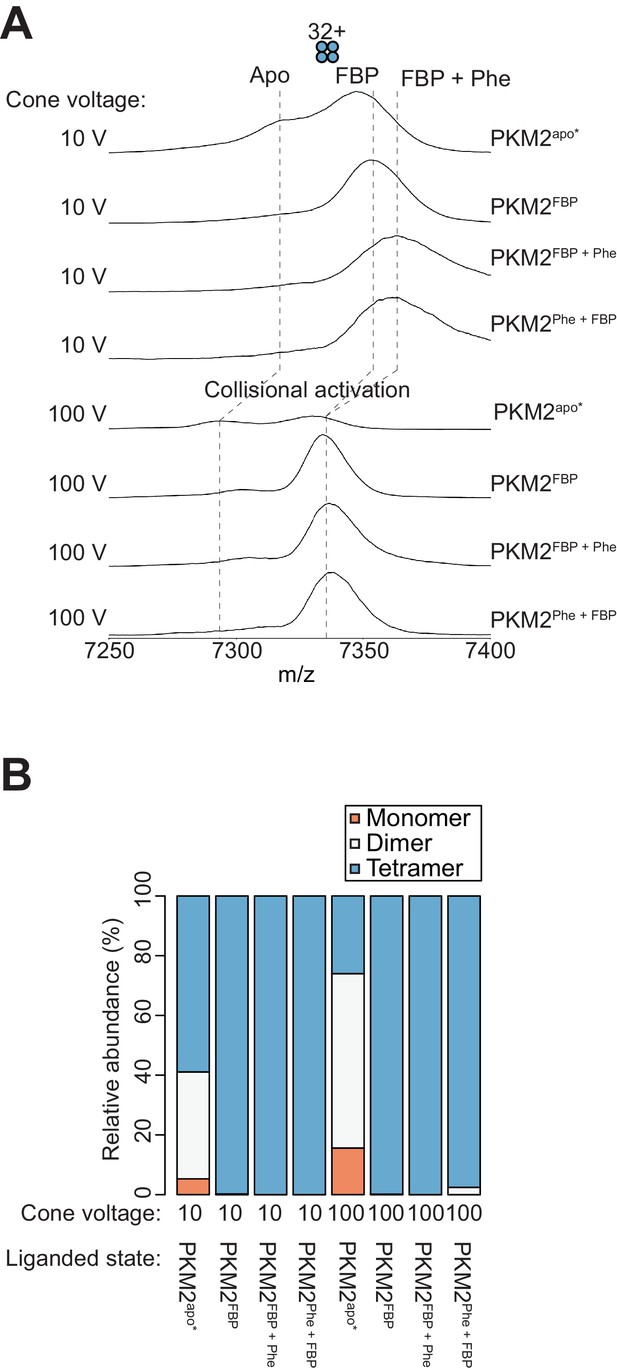
Phe and FBP can simultaneously bind to PKM2.
(A) Mass spectra of PKM2 were acquired in the absence of any added ligands (PKM2apo*), and following addition of stoichiometric amounts of FBP (PKM2FBP); addition of 400 µM Phe to 10 µM PKM2 pre-incubated with 10 µM FBP (PKM2FBP + Phe); or addition of 10 µM FBP to 10 µM PKM2 pre-incubated with 400 µM Phe (PKM2Phe + FBP). Spectra were acquired with the cone-voltage of the electrospray-ionisation (ESI) source set at 10 V. An m/z shift is observed upon the addition of FBP, and a further shift is observed upon addition of Phe after FBP binding. Increasing the cone voltage of the ESI source to 100 V led to the collisional removal of phenylalanine, resulting in the conversion of PKM2FBP+Phe and PKM2Phe+FBP peaks to an PKM2FBP-like state. The m/z shift of PKM2apo* upon collisional activation (cf. 10 V vs. 100 V) is likely due to a desalting effect. Positions of the m/z peaks for PKM2apo*, PKM2FBP, PKM2FBP + Phe and PKM2Phe + FBP are shown as dashed lines, for reference, at each of the two cone voltage conditions. (B) The relative abundances of monomer (orange) dimer (white) and tetramer (blue) species quantified from the spectra in (A).
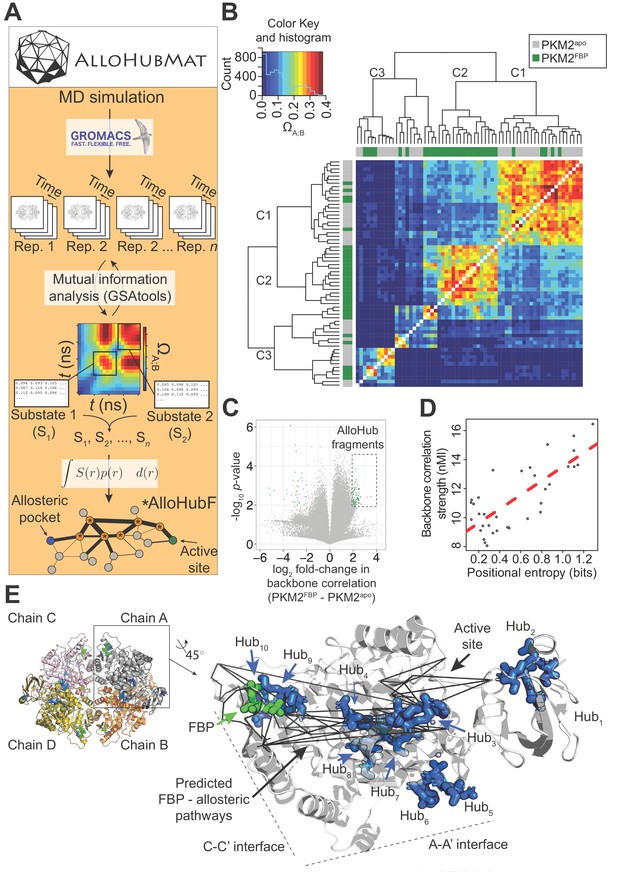
AlloHubMat predicts candidate residues that mediate the allosteric effect of FBP on PKM2, from molecular dynamics (MD) simulations.
(A) Schematic of the AlloHubMat computational pipeline, developed to identify residues that are involved in the transmission of allostery between an allosteric ligand pocket and the active site. Multiple replicate molecular dynamics (MD) simulations are seeded from the 3D protein structure using the GROMACS molecular dynamics engine. All MD simulations are encoded with the M32K25 structural alphabet (Pandini et al., 2010), and the protein backbone correlations over the MD trajectory are computed with GSAtools (Pandini et al., 2013) using information theory mutual information statistics. The backbone correlations are explicitly used to identify and extract configurational sub-states from the MD trajectories. A global allosteric network is then constructed by integrating over the correlation matrices, and their respective probabilities, from which allosteric hub fragments (AlloHubFs) are extracted. Each AlloHubF comprises four consecutive amino acid residues. (B) Correlation matrices cluster according to the liganded state of PKM2 in the MD simulations. AlloHubMat, described in (A), was used to identify correlation matrices of the conformational substates from five separate 400 ns MD simulations of PKM2apo (grey) and PKM2FBP (green). In total, we identified eight sub-states for all simulations of PKM2apo and eight for PKM2FBP. For every sub-state, the network of correlations from each of the four protomers is presented individually. To investigate whether the correlated motions for each sub-state could be attributed to the liganded state of PKM2, the correlation matrices were compared with a complete-linkage hierarchical clustering (see Materials and methods). The matrix covariance overlap (ΩA;B) was used as a distance metric, represented by the colour scale. A high ΩA;B score indicates high similarity between two correlation matrices, and a low ΩA;B score indicates that the correlation matrices are dissimilar. The clustering analysis revealed three clusters, denoted C1-C3. Cluster C1 was dominated by correlation matrices extracted from PKM2apo simulations, and cluster C2 was exclusively occupied by PKM2FBP correlation matrices. C3 consisted of correlation matrices from PKM2apo and PKM2FBP simulations. (C) A volcano plot showing difference in protein backbone correlations – derived from the AlloHubMat analysis – between PKM2apo and PKM2FBP. Each point corresponds to a correlation between two distal protein fragments; points with a positive log2 fold-change represent correlations that are predicted to increase in strength upon FBP binding. Correlations with a log2fold-change ≥ 2 and a false discovery rate ≤ 0.05% (determined from a Wilcoxon ranked-sum test) between PKM2apo and PKM2FBP were designated as AlloHubFs, highlighted in green. A total of 72 AlloHubFs were predicted from this analysis. (D) The positional entropy of the PKM2 fragment-encoded structure correlates linearly with the correlation strength of the fragment. The total mutual information content was computed by summing over the correlations for each of the top AlloHubFs. nMI: normalised mutual information. (E) Left: PKM2 structure depicting the spatial distribution of the top ten predicted AlloHubFs. Right: zoom into a single protomeric chain shown in cartoon representation. AlloHubFs (blue) and FBP (green) are shown as stick models. Black lines indicate minimal distance pathways between the FBP binding pocket and the active site, predicted using Dijkstra’s algorithm (see Materials and methods) with the complete set of correlation values as input.
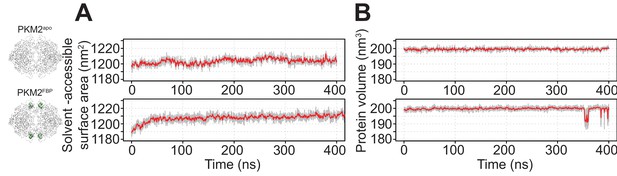
Solvent accessibility and volume analyses of MD simulations of PKM2.
(A) Five separate molecular dynamics (MD) simulations of PKM2apo and PKM2FBP analysed for solvent-accessibility. The grey trace shows the variance of the solvent-accessibility over the replicate simulations and the red trace is the running average of the variance. (B) MD simulations as in (A) but analysed for volume.
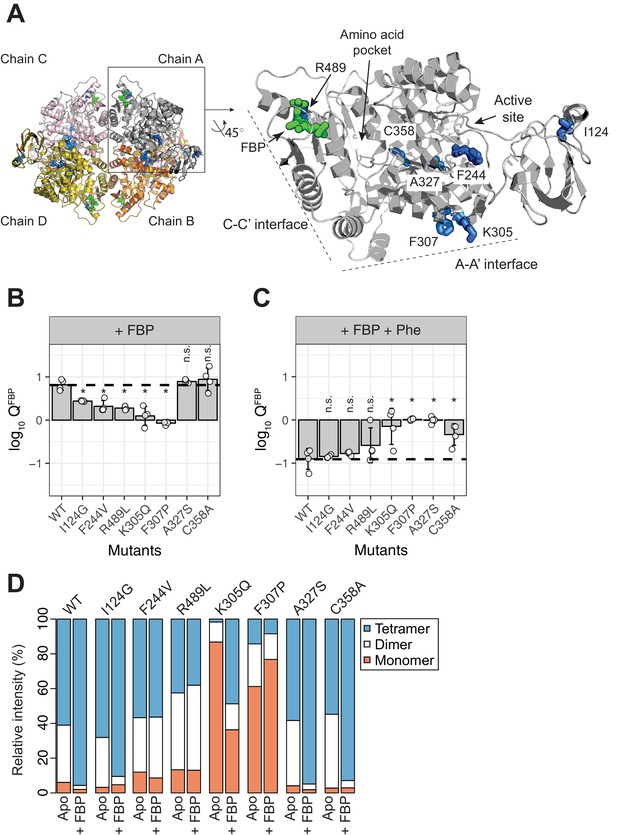
AlloHubF mutants (AlloHubMs) either interfere with FBP-induced PKM2 activation or mediate its disruption by Phe.
(A) AlloHubF mutants characterised in this study, shown on the PKM2 protomer structure. (B) The allosteric response of PKM2(WT) and AlloHubF mutant enzymatic activities to FBP, quantified by the allosteric coefficient Q, which denotes the change of the KMPEP in the absence and in the presence of saturating concentrations of FBP (see Materials and methods). A Q-coefficient > 0, indicates allosteric activation; and Q-coefficient < 0 indicates allosteric inhibition. The Q-coefficient for PKM2(WT) is shown as a dotted line for comparison. Each of the Q-coefficients of the AlloHubF mutants were statistically compared to PKM2(WT) using a Wilcoxon ranked-sum test (n = 4); a p-value<0.05 was deemed significant (denoted by an asterisk); n.s.: not significant. (C) The magnitude of allosteric inhibition by Phe, in the presence of FBP, determined for PKM2(WT) and AlloHubF mutants, quantified by the allosteric co-efficient Q as in (B). (D) Relative abundance of monomers, dimers and tetramers for PKM2 (WT) and PKM2 AlluHubF mutants in the absence or presence of saturating FBP.
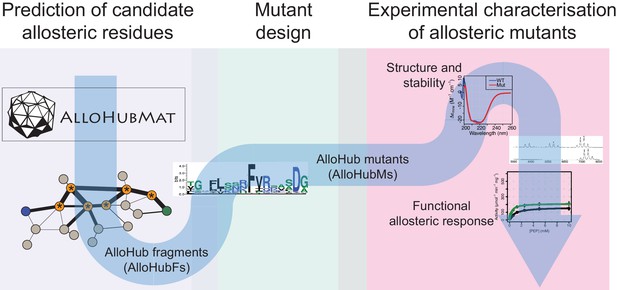
Schematic depicting the integrated computational and experimental strategy used to identify protein residues involved in allosteric regulation.
Molecular dynamics (MD) simulations were analysed using the AlloHubMat analysis software (Figure 5A) to generate a network of correlated protein dynamics that was used to identify allosteric hub fragments (AlloHubFs). The design of mutant variants (AlloHubMs) was guided by protein sequence conservation analysis of residues comprising AlloHubFs. AlloHubMs were experimentally characterised for their structural integrity using CD spectroscopy, for their oligomerisation state using ESI-MS, and enzymatic properties to assess their allosteric response to FBP and the ability of Phe to interfere with it. See text and methods for details.
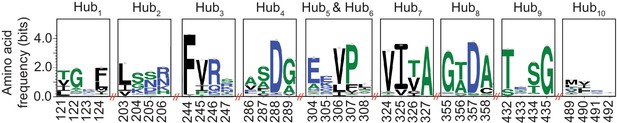

Sequence conservation analysis of AlloHubFs.
A multiple sequence alignment of PKM2 homologues was generated through a BLAST search of the NCBI non-redundant protein database (Altschul et al., 1990). Sequences with a pairwise identity of greater than 0.8 were removed, leaving a total of 5381 homologues, which were aligned using the HMMER suite (Eddy, 1998). Sequence conservation logo plots were generated from the alignment for each of the top-ten predicted AlloHubF residues.
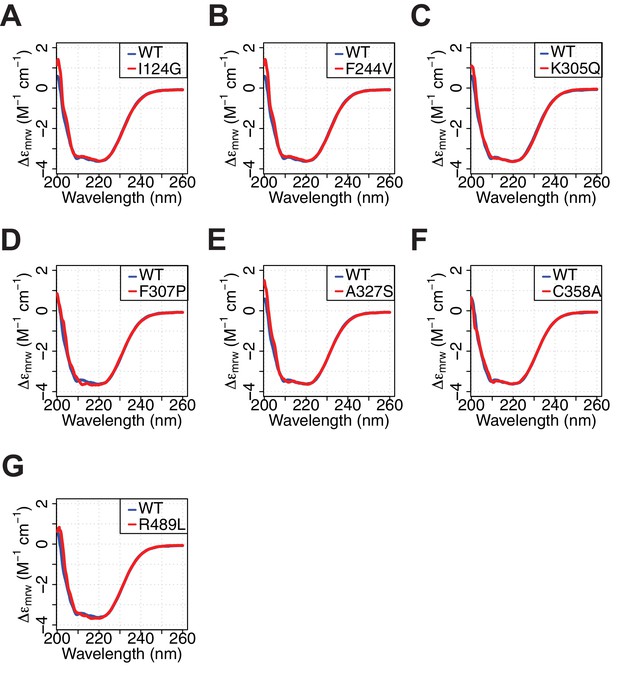
Purified AlloHubF mutants have similar secondary structure content to PKM2(WT).
(A–G) Smoothened far-UV circular dichroism spectra (200–260 nm) of each of the AlloHubMs (red) superimposed to that of PKM2(WT) (blue).
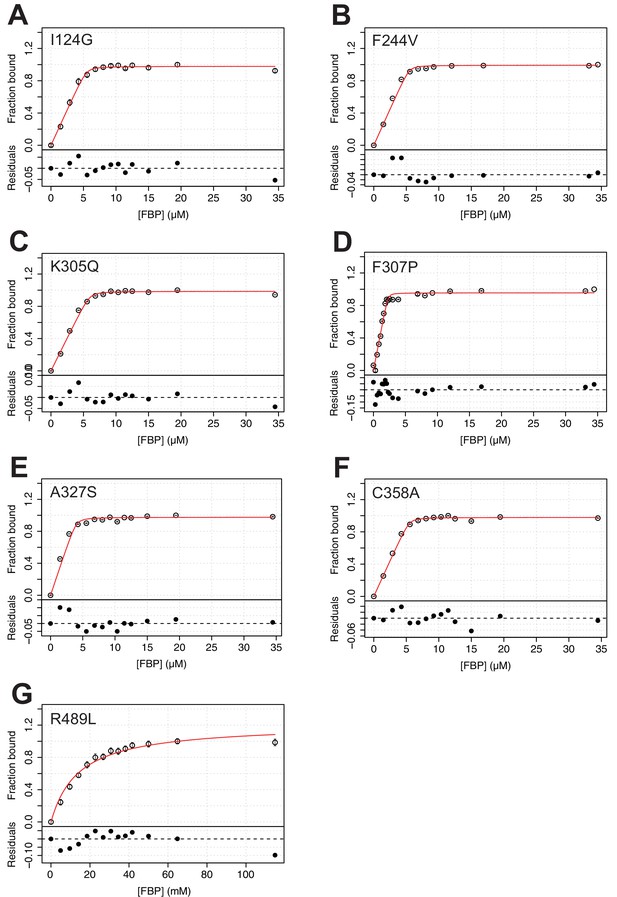
FBP has a similar affinity for AlloHubMs to that for PKM2(WT), with the exception of PKM2(R489L).
(A–G) Fluorescence emission spectroscopy measurements were used to monitor binding of FBP to PKM2 for each of the AlloHubF mutants. Each binding curve was fit to a 1:1 binding model (see Materials and methods), shown as a red line.
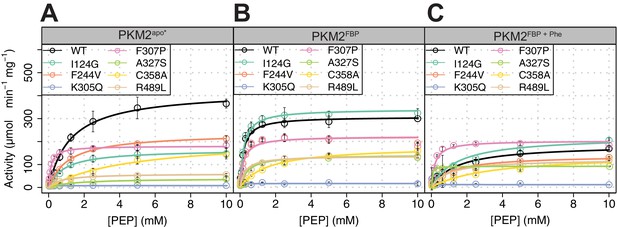
Steady-state enzyme kinetics of the AlloHubMs.
(A) Specific enzyme activity for PKM2(WT) and for each of the AlloHubMs was determined over a range of PEP concentrations between 0 mM and 10 mM, in the absence of any added allosteric ligands. Enzyme activity was assayed with a constant concentration of ADP (5 mM) at 37°C. Means and standard deviations of four separate repeats are shown. Rate curves were fit using Michaelis-Menten kinetics. (B) As in (A) but in the presence of FBP. (C) As in (A) but in the presence of FBP and Phe.
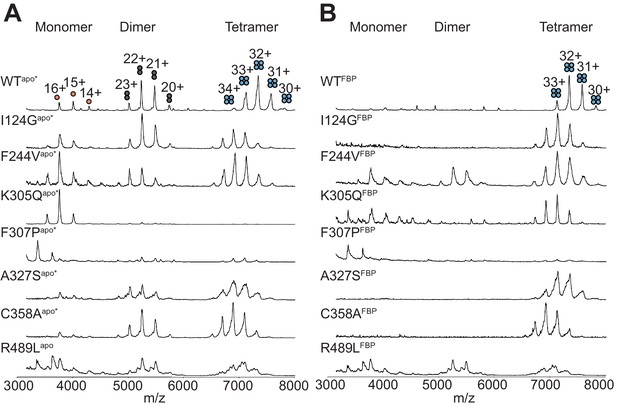
Native mass spectra of the AlloHubMs.
(A) nESI-MS mass spectra of the AlloHubMs in the absence of any added ligands. (B) As in (A) but in the presence of FBP at a 10:1 molar ratio of FBP:PKM2.
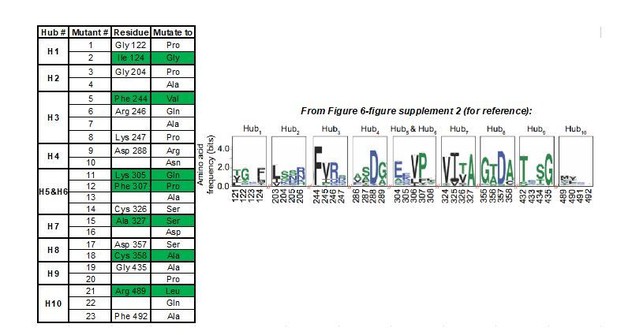
Summary of AlloHub mutants generated for this study.
https://doi.org/10.7554/eLife.45068.038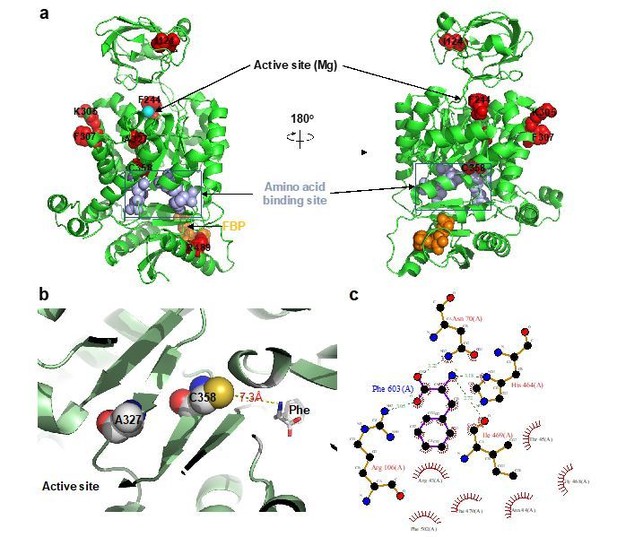
AlloHub mutant locations relative to catalytic and ligand binding pockets.
(a) PDBID:3u2z (PKM2+FBP), (b) PDBID:6gg4, (c) Phe binding pocket in 6gg4 plotted using LigPlot [Wallace et al. (1996). Protein Eng., 8:127-134].
Tables
Apparent steady-state binding constants of PKM2 for FBP, Phe and Ser.
https://doi.org/10.7554/eLife.45068.009| Titrated ligand | Constant ligand | KDapparent |
|---|---|---|
| FBP | – | (21.4 ± 9.0) nM |
| Ser (5 mM) | (12.5 ± 7.7) nM | |
| Phe (1.5 mM) | (8.1 ± 5.6) nM | |
| Phe | – | (191.0 ± 86.3) µM |
| FBP (50 μM) | (132.0 ± 11.7) µM | |
| Ser | – | (507.5 ± 218.2) µM |
| FBP (50 μM) | (462.0 ± 97.1) µM |
Intracellular concentrations of FBP, Phe and Ser in cancer cell lines.
https://doi.org/10.7554/eLife.45068.010| Metabolite | Media condition | Cell line | [Metabolite]ic (µM) | Fractional saturation |
|---|---|---|---|---|
| FBP | Gluc- (RPMI) | HCT116 | 38.4 ± 16.6 | 0.99 ± 0.00 |
| LN229 | 73.4 ± 18.4 | 0.99 ± 0.00 | ||
| SN12C | 19.9 ± 7.5 | 0.98 ± 0.00 | ||
| Gluc+ (RPMI) | HCT116 | 238.4 ± 67.4 | 0.99 ± 0.00 | |
| LN229 | 302.0 ± 75.3 | 0.99 ± 0.00 | ||
| SN12C | 355.1 ± 66.2 | 0.99 ± 0.00 | ||
| Gluc+ aa- (HBSS) | HCT116 | 337.2 ± 185.5 | 0.99 ± 0.00 | |
| LN229 | 257.0 ± 33.9 | 0.99 ± 0.00 | ||
| SN12C | 236.8 ± 41.1 | 0.99 ± 0.00 | ||
| Gluc+ Phe500 Ser100 (HBSS) | HCT116 | 384.5 ± 41.8 | 0.99 ± 0.00 | |
| LN229 | 277.3 ± 83.4 | 0.99 ± 0.00 | ||
| SN12C | 300.3 ± 89.1 | 0.99 ± 0.00 | ||
| Gluc+ Phe100 Ser500 (HBSS) | HCT116 | 400.9 ± 82.2 | 0.99 ± 0.00 | |
| LN229 | 372.2 ± 106.0 | 0.99 ± 0.00 | ||
| SN12C | 286.4 ± 73.6 | 0.99 ± 0.00 | ||
| Phe | Gluc- (RPMI) | HCT116 | 256.3 ± 65.8 | 0.56 ± 0.06 |
| LN229 | 231.6 ± 22.6 | 0.55 ± 0.03 | ||
| SN12C | 100.4 ± 24.6 | 0.34 ± 0.06 | ||
| Gluc+ (RPMI) | HCT116 | 258.2 ± 79.6 | 0.56 ± 0.08 | |
| LN229 | 318.7 ± 115.8 | 0.61 ± 0.09 | ||
| SN12C | 197.5 ± 26.6 | 0.51 ± 0.04 | ||
| Gluc+ aa- (HBSS) | HCT116 | 41.1 ± 15.8 | 0.17 ± 0.05 | |
| LN229 | 31.7 ± 28.0 | 0.13 ± 0.12 | ||
| SN12C | 10.6 ± 0.6 | 0.05 ± 0.02 | ||
| Gluc+ Phe500 Ser100 (HBSS) | HCT116 | 1776.2 ± 225.1 | 0.90 ± 0.01 | |
| LN229 | 1666.5 ± 543.0 | 0.89 ± 0.04 | ||
| SN12C | 926.1 ± 252.7 | 0.82 ± 0.04 | ||
| Gluc+ Phe100 Ser500 (HBSS) | HCT116 | 582.4 ± 23.8 | 0.75 ± 0.01 | |
| LN229 | 575.7 ± 224.4 | 0.73 ± 0.11 | ||
| SN12C | 221.6 ± 49.1 | 0.53 ± 0.06 | ||
| Ser | Gluc- (RPMI) | HCT116 | 3580.9 ± 1016.9 | 0.87 ± 0.03 |
| LN229 | 1755.0 ± 159.5 | 0.77 ± 0.02 | ||
| SN12C | 854.3 ± 240.7 | 0.62 ± 0.07 | ||
| Gluc+ (RPMI) | HCT116 | 3943.5 ± 1363.1 | 0.88 ± 0.05 | |
| LN229 | 2226.7 ± 757.7 | 0.80 ± 0.06 | ||
| SN12C | 1766.0 ± 142.9 | 0.78 ± 0.01 | ||
| Gluc+ aa- (HBSS) | HCT116 | 426.9 ± 179.2 | 0.44 ± 0.09 | |
| LN229 | 249.6 ± 221.8 | 0.28 ± 0.25 | ||
| SN12C | 111.0 ± 18.4 | 0.18 ± 0.02 | ||
| Gluc+ Phe500 Ser100 (HBSS) | HCT116 | 6157.3 ± 1334.4 | 0.92 ± 0.01 | |
| LN229 | 2641.1 ± 825.1 | 0.83 ± 0.06 | ||
| SN12C | 2197.7 ± 605.8 | 0.81 ± 0.04 | ||
| Gluc+ Phe100 Ser500 (HBSS) | HCT116 | 22360.4 ± 1554.2 | 0.98 ± 0.00 | |
| LN229 | 8464.7 ± 3214.5 | 0.93 ± 0.04 | ||
| SN12C | 6603.9 ± 1500.4 | 0.93 ± 0.02 |
Steady-state Michaelis-Menten kinetic parameters for PKM2apo, and following the addition of FBP, Phe and Ser.
https://doi.org/10.7554/eLife.45068.011| PKM2ligand | KMPEP (mM) | kcat (s−1) | kcat/KMPEP (s−1 mM−1) |
|---|---|---|---|
| PKM2apo | 1.22 ± 0.02 | 349.3 ± 40.9 | 285.6 ± 34.1 |
| PKM2Phe | 7.08 ± 1.58 | 324.7 ± 23.9 | 46.8 ± 6.0 |
| PKM2Ser | 0.22 ± 0.04 | 323.1 ± 43.2 | 1489.8 ± 84.7 |
| PKM2FBP | 0.23 ± 0.04 | 356.7 ± 25.7 | 1540.4 ± 96.9 |
| PKM2 FBP + Phe | 0.65 ± 0.03 | 222.3 ± 6.3 | 342.1 ± 11.1 |
| PKM2 FBP + Ser | 0.20 ± 0.04 | 348.7 ± 44.3 | 1620.0 ± 253.6 |
Theoretical masses of PKM2 (see Supplementary file 2 for PKM2 sequence), FBP and PKM2 + FBP.
https://doi.org/10.7554/eLife.45068.020| Molecule | Mass (Da) |
|---|---|
| PKM2 monomer | 58218.2 |
| PKM2 tetramer | 232872.6 |
| FBP | 340.0 |
| PKM2 tetramer + 1 FBP | 233212.6 |
| PKM2 tetramer + 2 FBP | 233552.6 |
| PKM2 tetramer + 3 FBP | 233892.6 |
| PKM2 tetramer + 4 FBP | 234232.6 |
Calculated masses from maximum-entropy deconvolution of PKM2 mass spectra.
https://doi.org/10.7554/eLife.45068.021| Protein/ligand mixture | Cone voltage (V) | Number of tetramer peaks | Mass species (Da) |
|---|---|---|---|
| 10 µM PKM2 | 100 | 5 | 232880, 233220, 233580, 233930, 234290 |
| 10 µM PKM2 + 10 µM FBP | 10 | 1 | 235030 |
| 10 µM PKM2 + 10 µM FBP | 100 | 1 | 234190 |
Summary of molecular dynamics simulations.
https://doi.org/10.7554/eLife.45068.022| Liganded state simulated | PDB ID of starting structure | Number of protein atoms | Number of water atoms | Simulation time | Number of replicas |
|---|---|---|---|---|---|
| PKM2apo | 3BJT | 20104 | 263384 | 400 ns | 5 |
| PKM2FBP | 3U2Z | 20122 | 261594 | 420 ns | 5 |
Apparent steady-state binding constants of FBP to the PKM2 allosteric hub mutants.
https://doi.org/10.7554/eLife.45068.025| AlloHubM | KDFBP |
|---|---|
| I124G | (39.5 ± 33.5) nM |
| F244V | (30.7 ± 33.1) nM |
| K305Q | (39.4 ± 34.1) nM |
| F307P | (4.0 ± 12.8) nM |
| A327S | (43.2 ± 48.7) nM |
| C358A | (35.3 ± 23.6) nM |
| R489L | (14.0 ± 2.7) mM |
Steady-state Michaelis-Menten kinetic parameters for AlloHubF mutants.
https://doi.org/10.7554/eLife.45068.033| AlloHubM | Ligand | KMPEP (mM) | kcat (s−1) | kcat/KMPEP (s−1 mM−1) |
|---|---|---|---|---|
| I124G | Apo | 1.07 ± 0.13 | 190.27 ± 7.31 | 177.82 ± 56.23 |
| FBP | 0.29 ± 0.04 | 307.10 ± 6.11 | 1058.97 ± 152.75 | |
| FBP + Phe | 1.44 ± 0.34 | 221.92 ± 17.26 | 153.98 ± 37.88 | |
| F244V | Apo | 1.14 ± 0.11 | 237.44 ± 7.36 | 210.44 ± 27.15 |
| FBP | 0.55 ± 0.07 | 279.62 ± 9.93 | 522.42 ± 87.51 | |
| FBP + Phe | 1.15 ± 0.30 | 222.51 ± 18.29 | 195.87 ± 55.46 | |
| K305Q | Apo | 0.01 ± 0.01 | 8.06 ± 0.57 | 790.01 ± 605.56 |
| FBP | 0.01 ± 0.04 | 8.40 ± 0.43 | 1038.01 ± 535.50 | |
| FBP + Phe | 0.04 ± 0.01 | 12.21 ± 1.40 | 408.37 ± 136.10 | |
| F307P | Apo | 0.13 ± 0.01 | 180.47 ± 3.05 | 1413.87 ± 133.12 |
| FBP | 0.15 ± 0.02 | 227.20 ± 5.02 | 1508.53 ± 190.71 | |
| FBP + Phe | 0.21 ± 0.03 | 328.71 ± 12.37 | 1568.32 ± 268.58 | |
| A327S | Apo | 1.37 ± 0.42 | 31.8 ± 2.43 | 23.21 ± 5.79 |
| FBP | 0.17 ± 0.02 | 119.01 ± 6.86 | 700.06 ± 343.00 | |
| FBP + Phe | 0.15 ± 0.02 | 100.19 ± 5.32 | 667.93 ± 266.00 | |
| C358A | Apo | 4.71 ± 0.99 | 214.73 ± 21.40 | 45.59 ± 21.62 |
| FBP | 0.58 ± 0.18 | 191.13 ± 16.22 | 329.53 ± 90.11 | |
| FBP + Phe | 1.25 ± 0.07 | 199.01 ± 21.87 | 159.20 ± 31.24 | |
| R489L | Apo | 0.69 ± 0.19 | 60.05 ± 4.91 | 89.38 ± 36.93 |
| FBP | 0.36 ± 0.10 | 112.25 ± 7.60 | 317.20 ± 125.71 | |
| FBP + Phe | 1.44 ± 0.40 | 132.99 ± 15.74 | 158.19 ± 122.56 |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Protein (Homo sapiens) | PKM2 | N/A | Uniprot ID: P14618-1 | |
| Cell line (Homo sapiens) | HCT116 | ATCC, Manassas, VA, USA | ATCC Cat# CCL-247, RRID:CVCL_0291 | |
| Cell line (Homo sapiens) | LN229 | ATCC, Manassas, VA, USA | ATCC Cat# CRL-2611, RRID:CVCL_0393 | |
| Cell line (Homo sapiens) | SN12C | Kaelin Lab, DFCI, Boston, MA, USA | RRID:CVCL_1705 | |
| Antibody | Rabbit Anti-PKM2, Clone D78A4 | Cell Signaling Technology | Cat# 4053; RRID:AB_1904096 | 1:1000 in 5% BSA/T-BST |
| Recombinant DNA reagent | pET28a-His-PKM2(WT) | Anastasiou et al., 2011 | RRID:Addgene_42515 | Available from AddGene (Cambridge MA, USA). See Supplementary file 2 for PKM2 sequence. |
| Recombinant DNA reagent | pET28a-His-PKM2(I124G) | This study | 1-step mutagenesis using pET28a-His-PKM2(WT) as template | |
| Recombinant DNA reagent | pET28a-His-PKM2(F244V) | This study | 1-step mutagenesis using pET28a-His-PKM2(WT) as template | |
| Recombinant DNA reagent | pET28a-His-PKM2(R489L) | This study | 1-step mutagenesis using pET28a-His-PKM2(WT) as template | |
| Recombinant DNA reagent | pET28a-His-PKM2(K305Q) | This study | 1-step mutagenesis using pET28a-His-PKM2(WT) as template | |
| Recombinant DNA reagent | pET28a-His-PKM2(F307P) | This study | 1-step mutagenesis using pET28a-His-PKM2(WT) as template | |
| Recombinant DNA reagent | pET28a-His-PKM2(A327S) | This study | 1-step mutagenesis using pET28a-His-PKM2(WT) as template | |
| Recombinant DNA reagent | pET28a-His-PKM2(C358A) | This study | 1-step mutagenesis using pET28a-His-PKM2(WT) as template | |
| Peptide | ITLDNAYMEK[13C615N2] | This study | Synthesised by the Crick Peptide Synthesis STP | |
| Peptide | GDLGIEIPAEK[13C615N2] | This study | Synthesised by the Crick Peptide Synthesis STP | |
| Peptide | APIIAVTR[13C615N4] | This study | Synthesised by the Crick Peptide Synthesis STP | |
| Peptide | LFEELVR[13C615N4] | This study | Synthesised by the Crick Peptide Synthesis STP | |
| Peptide | LAPITSDPTEATAVGAVEASFK[13C615N2] | This study | Synthesised by the Crick Peptide Synthesis STP | |
| Chemical compound | Potassium phosphoenolpyruvate (2,3-13C2) | Cambridge Isotope Laboratories | CLM-3398-PK | |
| Chemical compound | D-fructose 1,6-bisphosphate sodium salt hydrate (U-13C6) | Cambridge Isotope Laboratories | CLM-8962-PK | |
| Chemical compound | L-Phe (ring-13C6) | Cambridge Isotope Laboratories | CLM-1055-PK | |
| Chemical compound | L-Ser (13C6) | Cambridge Isotope Laboratories | CLM-1574-H | |
| Software | sa_encode.R | This study | Software to encode trajectory into stacked alignment of structural alphabet strings (Kleinjung, 2019; copy archived at https://github.com/elifesciences-publications/ALLOHUBMAT) | |
| Software | kabsch.R | This study | Kabsch superpositioning routine (Kleinjung, 2019; copy archived at https://github.com/elifesciences-publications/ALLOHUBMAT) | |
| Software | MI.R | This study | Mutual Information and other entropy metrics between two character vectors, here intended for two alignment columns (Kleinjung, 2019; copy archived at https://github.com/elifesciences-publications/ALLOHUBMAT) | |
| Software | Xcalibur QualBrowser | Thermo Fisher Scientific (Waltham MA, USA) | N/A | |
| Software | Tracefinder v4.1 | Thermo Fisher Scientific (Waltham MA, USA) | N/A |
Comparison of intra- and inter-subunit Mutual Information signal strength for allosteric hubs identified in this study.
https://doi.org/10.7554/eLife.45068.040| Hub | Intra | Inter |
|---|---|---|
| H1 | 42.9 | - |
| H2 | 35.4 | - |
| H3 | 22.3 | - |
| H4 | 88.8 | 18.2 |
| H5_6 | 18.8 | 21.5 |
| H7 | 20.5 | 19.7 |
| H8 | 21.4 | 18.7 |
| H9 | 22.3 | - |
| H10 | 29.9 | - |
Additional files
-
Supplementary file 1
Targeted proteomics data used to quantify intracellular PKM2 concentrations.
- https://doi.org/10.7554/eLife.45068.034
-
Supplementary file 2
Human PKM2 as cloned in pET28a vector (His-tagged).
- https://doi.org/10.7554/eLife.45068.035
-
Transparent reporting form
- https://doi.org/10.7554/eLife.45068.036





