Dynamic relocalization of replication origins by Fkh1 requires execution of DDK function and Cdc45 loading at origins
Figures
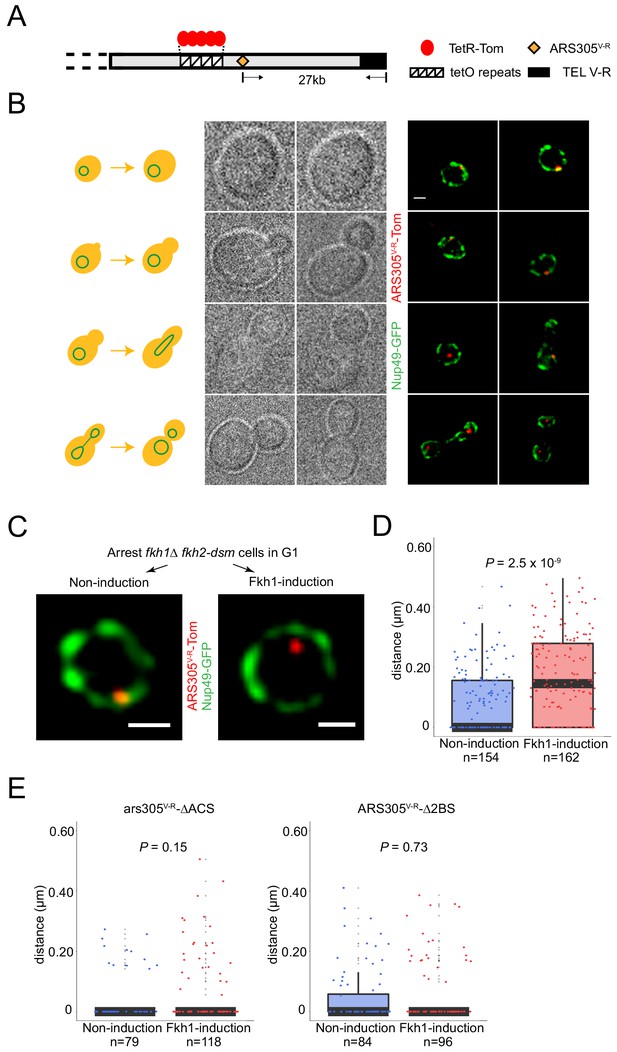
Fkh1-induced origin activation re-positions a subtelomeric origin in G1 phase.
(A) Schematic of chromosome V-R showing tetO repeats inserted adjacent to the ARS501 locus, which has been replaced with ARS305 (designated ARS305V-R); TetR-Tomato binds to and illuminates the locus as a single focus. (B) Images of cells from an unsynchronized culture of strain HYy132 (fkh1∆ fkh2-dsm ARS305V-R-Tomato NUP49-GFP GAL-FKH1) are shown sorted according to cell morphology; all images are at the same magnification: scale bar = 0.5 μm. (C) FKH1 induction scheme: HYy132 cells grown at 25°C in raffinose medium were arrested in G1 phase with 1x α-factor 2.5 hr, incubated an additional 2 hr in raffinose (Non-induction) or galactose (Fkh1-induction) with 1.7x α-factor, and images of live cells captured, examples of which are shown; scale bar = 0.5 μm. (D) The shortest distance from the ARS305V-R-Tomato focus to the nuclear periphery (Nup49-GFP) in each cell was measured and plotted as quartile boxplots (median shown as thick black segment) for non-induction and FKH1-induction; the result of a z-test comparing the two distributions is given as P. (E) Cells of fkh1∆ fkh2-dsm GAL-FKH1 NUP49-GFP strains HYy119 (ars305-∆ACSV-R-Tomato) and HYy120 (ars305-∆2BSV-R-Tomato) were treated and analyzed as above.
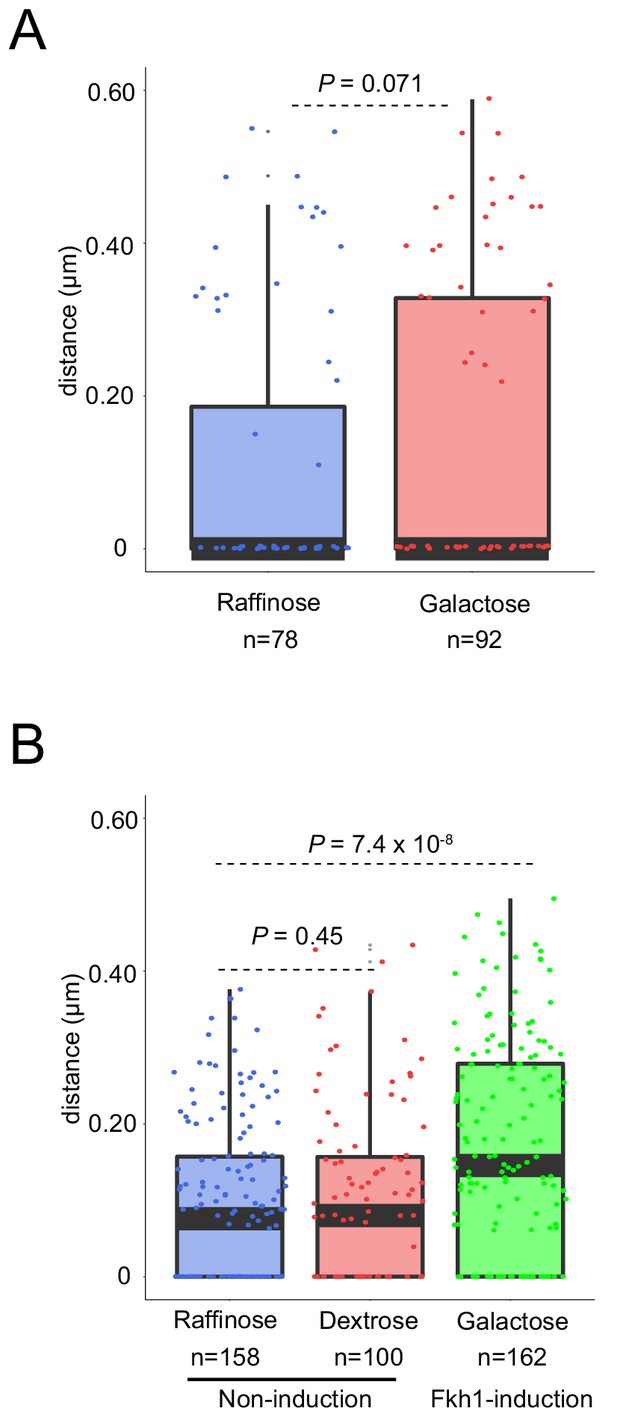
Fkh1-induction is required to re-position a subtelomeric origin in G1 phase.
(A) HYy80 (fkh1∆ fkh2-dsm ARS305V-R-Tomato NUP49-GFP) cells were treated and analyzed as in Figure 1C and D legends. (B) HYy132 (fkh1∆ fkh2-dsm GAL-FKH1 ARS305V-R-Tomato NUP49-GFP) cells were treated as in Figure 1C legend, except that dextrose was substituted for galactose, and analyzed as in Figure 1D legend. The galactose data are the same as in Figure 1D.
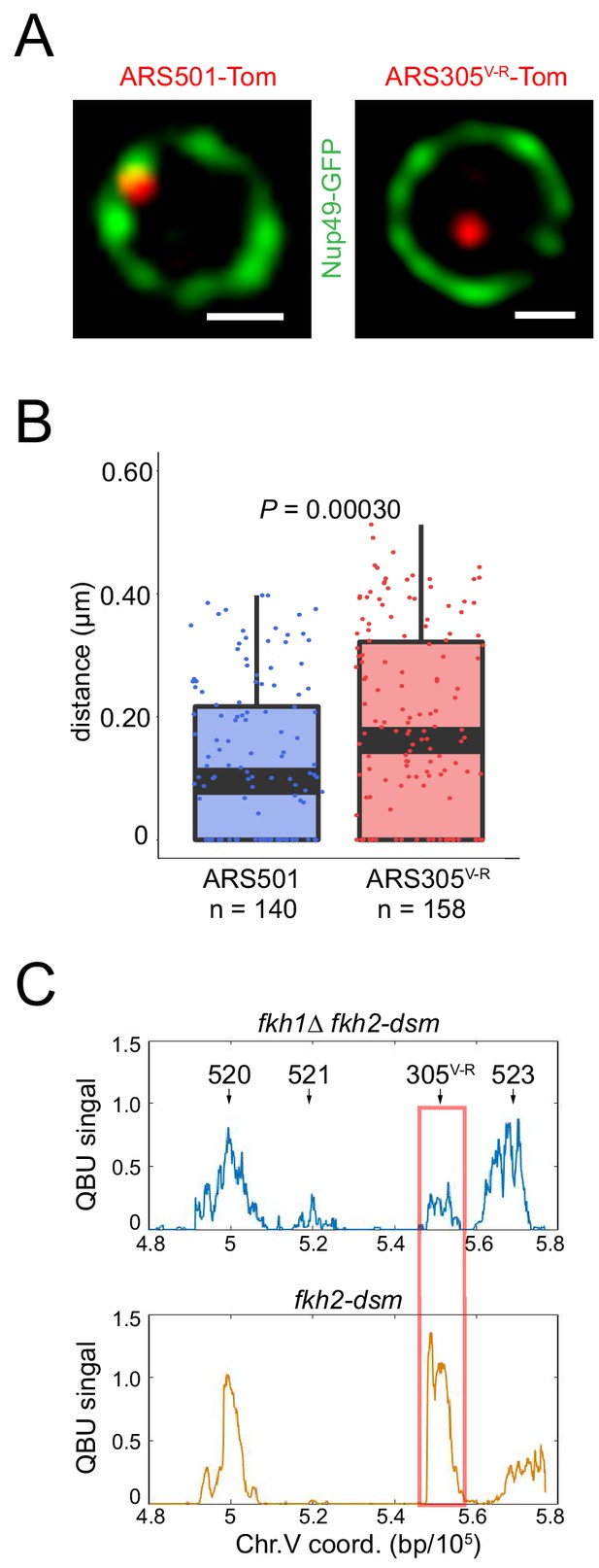
Normal dosage of Fkh1 is sufficient to relocalize ARS305V-R and advance its firing time.
(A) HYy160 (ARS501-Tomato NUP49-GFP) and HYy157 (ARS305V-R-Tomato NUP49-GFP) cells were arrested in G1 phase at 25°C with 1x α-factor 2 hr and images were collected; scale bar = 0.5 μm. (B) Distances from origin foci to nuclear periphery were determined, plotted as quartile boxplots, and analyzed by a z-test. (C) Quantitative BrdU-IP-Seq (QBU) analysis was performed with ARS305V-R-bearing strains HYy113 (fkh2-dsm) and HYy38 (fkh1∆ fkh2-dsm) after G1 block-and-release into hydroxyurea in the presence of BrdU; averaged data from three experimental replicates was plotted for the V-R region with the positions of several replication origins indicated; ARS305V-R resides at the ARS522 (aka: ARS501) locus.
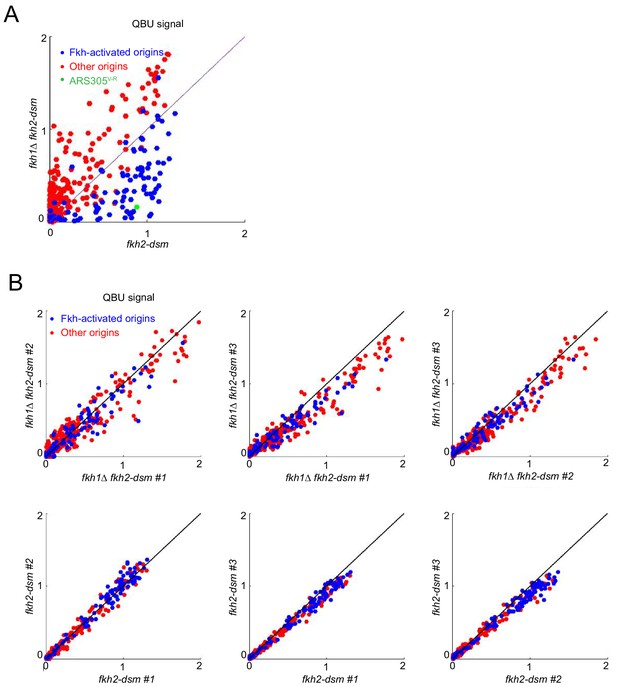
Normal dosage of Fkh1 is sufficient to advance firing time of ARS305V-R.
(A) QBU analysis was performed as described in Figure 3C legend and presented as a scatter plot showing average of three replicates. The results show expected global effects on Fkh-activated origins while highlighting ARS305V-R. (B) Scatter plots show comparison of individual experimental replicates for the above experiment.
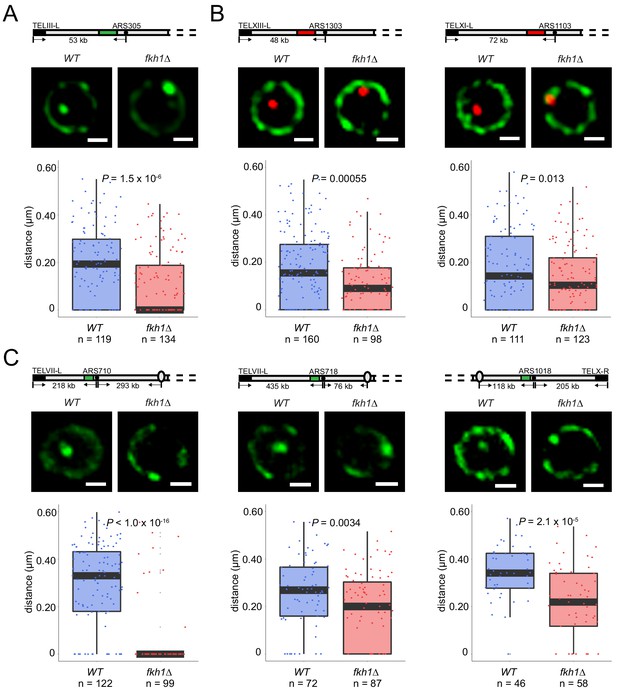
Fkh1 determines early origin positioning globally.
Diagrams of chromosomes with replication origins labeled with lacO/LacI-GFP (green-filled segment) or tetO/TetR-Tomato (red-filled segment) are shown above the corresponding images. Distances between origins (black-filled spheres) and telomeres, and in some cases centromeres (ovals), are indicated and include ~14 kb or 16 kb added by lacO or tetO repeats, respectively; elements are not drawn to scale. Cells of WT and fkh1∆ strains with ARS305-GFP (HYy151, HYy147) in (A), ARS1303-Tomato (HYy166, HYy173) and ARS1103-Tomato (HYy165, HYy172) in (B), ARS710-GFP (MPy6, MPy10), ARS718-GFP (MPy20, MPy21), and ARS1018-GFP (MPy19, MPy22) in (C), all expressing NUP49-GFP, were arrested in G1 phase at 25°C with 1x α-factor 2 hr and live images were captured; scale bar = 0.5 μm. Distances from origin foci to nuclear periphery were determined, plotted as quartile boxplots, and analyzed by a z-test.
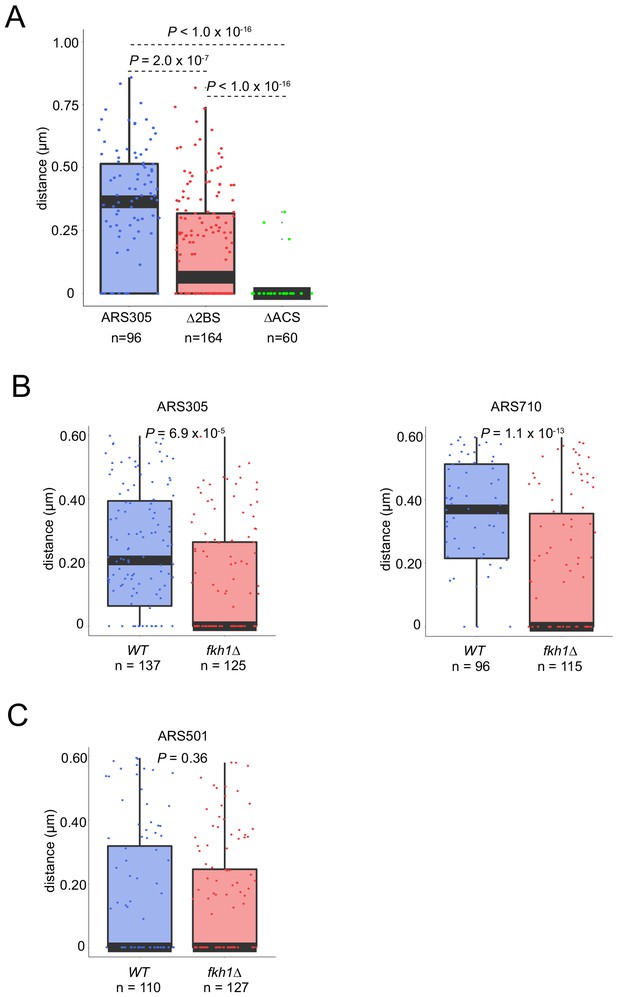
Fkh1 determines early origin positioning globally.
(A) Cells with ARS305-GFP (HYy151), ARS305-∆2BS-GFP (MPy46), and ars305-∆ACS-GFP (MPy43) all expressing NUP49-GFP, were treated and analyzed as in Figure 3 legend; scale bar = 0.5 μm. (B) Cycling cultures of WT and fkh1∆ strains with ARS305-GFP (HYy151, HYy147) and ARS710-GFP (MPy6, MPy10) were imaged and G1 phase (unbudded) cells were analyzed as described in Figure 3 legend. (C) Cells of WT and fkh1∆ strains with ARS501-Tom (HYy198, HYy201) were arrested in G1 phase and analyzed as described in Figure 3 legend.
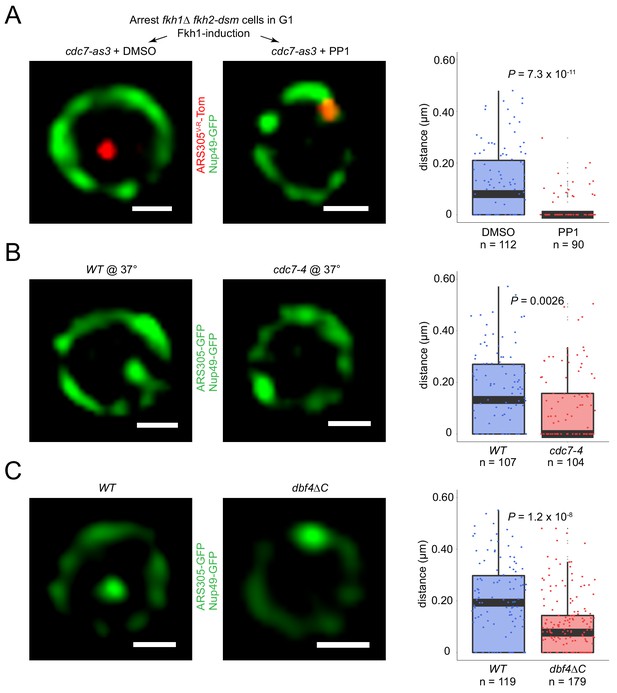
Origin localization in G1 is DDK regulated.
(A) HYy186 (fkh1∆ fkh2-dsm GAL-FKH1 ARS305V-R-Tomato NUP49-GFP cdc7-as3) cells were subjected to FKH1-induction scheme as described in Figure 1C legend except that PP1 or DMSO (vehicle) was included with α-factor, and images were captured. (B) Cells of ARS305-GFP NUP49-GFP strains HYy151 (WT) and HYy191 (cdc7-4) were arrested in G1 with 1x α-factor 2 hr at 25°C followed by 1 hr incubation at 37°C with 2x α-factor, and images were captured. (C) HYy181 (ARS305-GFP NUP49-GFP dbf4∆C) cells were arrested in G1 phase with 1x α-factor 2 hr at 25°C and live images were captured. The control experiment with WT cells (HYy151) is shown in Figure 3A. (A–C) Scale bar = 0.5 μm. Distances from origin foci to nuclear periphery were determined, plotted as quartile boxplots, and analyzed by a z-test.
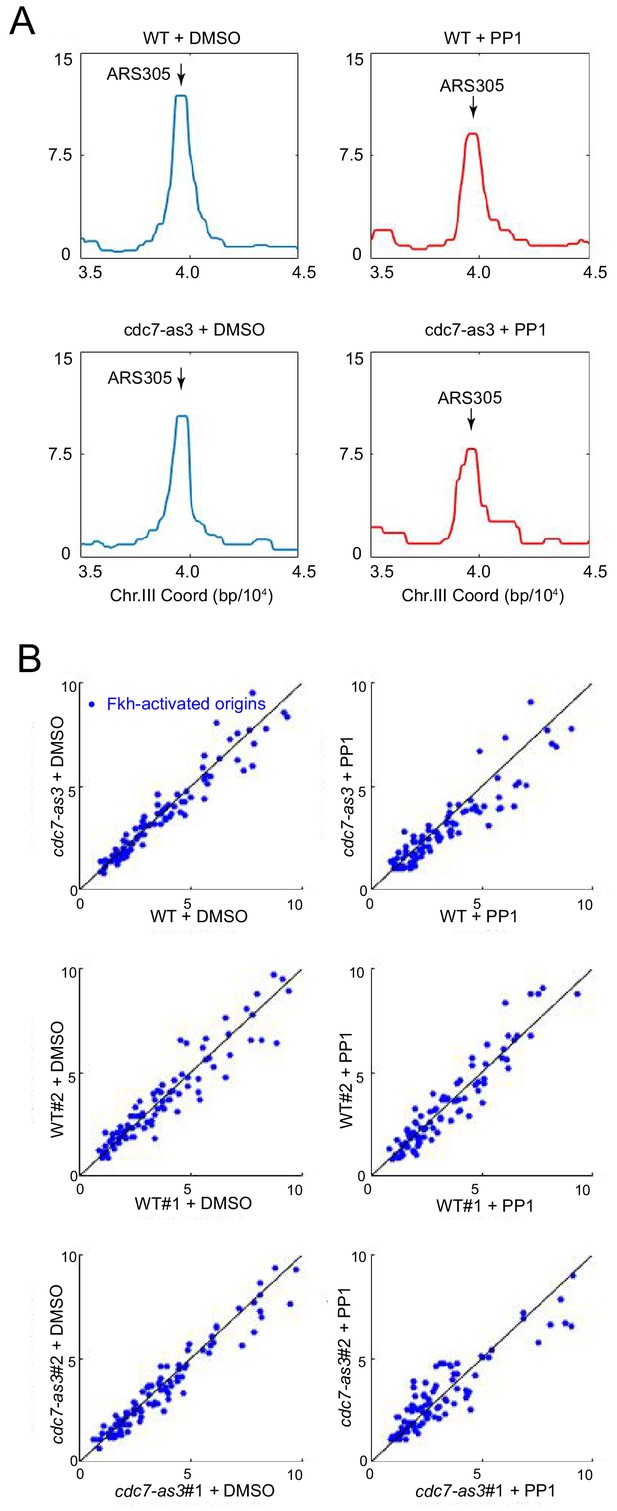
Fkh1 binds origins independently of Cdc7 function.
FKH1-FLAG-tagged strains OAy1102 (WT) and HYy123 (cdc7-as3) were arrested in G1 at 23°C and treated with PP1 or DMSO (vehicle), and subjected to ChIP-seq analysis. (A) Averaged data from duplicate experiments is plotted for the ARS305 chromosomal region. (B) Data for Fkh-activated origins are plotted. The upper panels present averaged data from the individual replicates presented in the middle and lower panels.
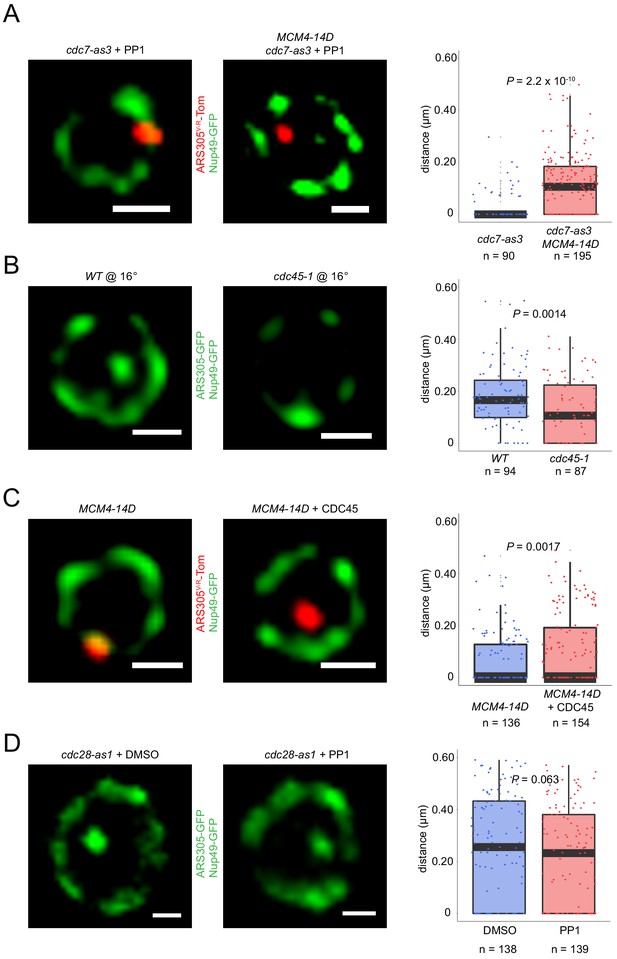
DDK regulation of origin localization reflects its phosphorylation of Mcm4 and consequent Cdc45 loading.
(A) Fkh1-induction scheme with PP1 as described in Figure 4A legend was carried out with fkh1∆ fkh2-dsm GAL-FKH1 ARS305V-R-Tomato NUP49-GFP strains HYy186 (cdc7-as3) and HYy177 (cdc7-as3 MCM4-14D), and images captured. (B) ARS305-GFP NUP49-GFP strains HYy151 (WT) and HYy184 (cdc45-1) were arrested in G1 with 1x α-factor 1 hr at 30°C followed by 2 hr incubation at 16°C with 1x α-factor, and images were captured. (C) Cells of strain HYy177 harboring no plasmid or high-copy plasmid expressing CDC45 were arrested in G1 with 1x α-factor 2 hr at 25°C and images captured. (D) ARS305-GFP NUP49-GFP strain HYy197 (cdc28-as1) cells were arrested in G1 phase with 0.5x α-factor 2 hr at 25°C, PP1 or DMSO was added and incubated one additional hour with 0.5x α-factor, and images were captured. (A–D) Scale bar = 0.5 μm. Distances from origin foci to nuclear periphery were determined, plotted as quartile boxplots, and analyzed by a z-test.
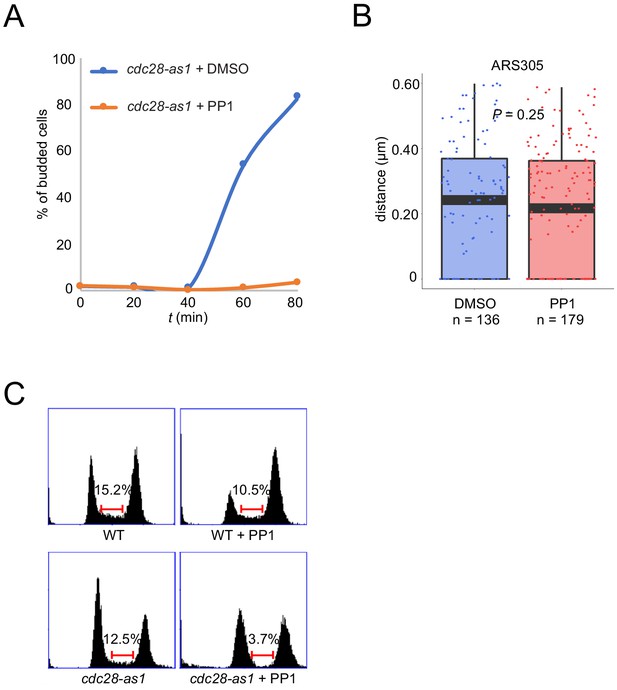
CDK activity is dispensable for origin localization in G1.
(A) Cells treated as described in Figure 5D legend were released from α-factor block with PP1 or DMSO and budding morphology was quantified and plotted. (B) ARS305-GFP cdc28-as1 cells (HYy197) were treated with PP1 or DMSO for two hours at 25°C and origin localization was analyzed in G1 phase (unbudded) cells as described in Figure 5 legend. (C) Cells treated as in (B) were subjected to DNA content analysis; the brackets estimate the proportion of cells in S phase based on DNA content between 1C and 2C.
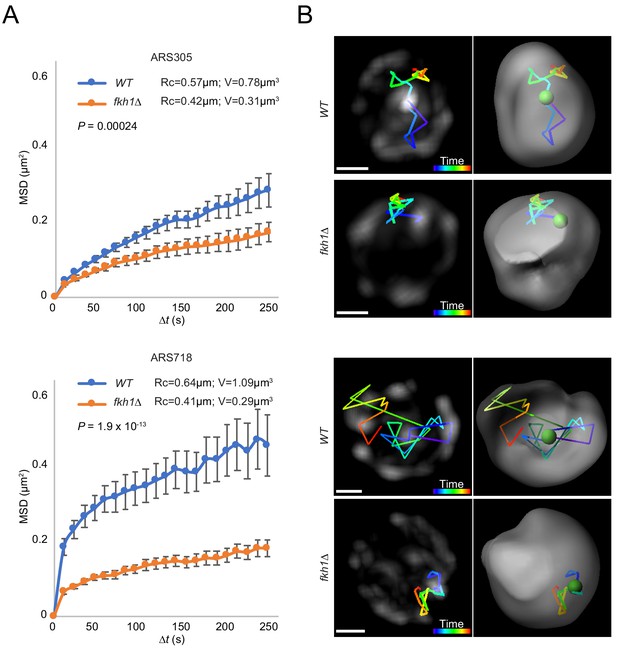
Origin mobility increases with origin relocalization.
(A) Mean-squared-displacement (MSD) analysis of tracking data for ARS305-GFP strains HYy151 (WT) and HYy147 (fkh1∆) and ARS718-GFP strains MPy20 (WT) and MPy21 (fkh1∆). Radius of constraint (Rc) and volume searched (V) are given, and statistical significance comparing WT and fkh1∆ was estimated by two-tailed Mann-Whitney test. (B) Images (left) and 3D reconstructions with Imaris (right) showing examples of tracks of origin focus over time (color corresponding to time progression); scale bar = 0.4 μm. Movies of the individual ARS305 time-courses are available as Figure 6—video 1 and Figure 6—video 2.
Origin tracking in WT.
https://doi.org/10.7554/eLife.45512.013Origin tracking in fkh1∆.
https://doi.org/10.7554/eLife.45512.014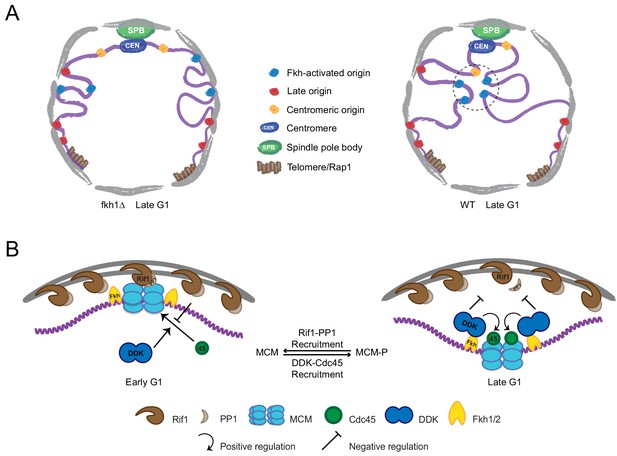
Model of origin localization linked to initiation.
(A) Absent Fkh1, most replication origins are enriched at nuclear periphery, however, Fkh1 binding to a subset of origins allows execution of the DDK-dependent step of initiation, resulting in release from the nuclear periphery and/or capture in the nuclear interior to form early origin clusters. (B) Hypothetical mechanism for origin tethering to the nuclear periphery regulated by Rif1-PP1 versus Fkh1-DDK-Cdc45 activities. Rif1 associates with inner nuclear membrane and with licensed replication origins, while associated PP1 antagonizes execution of the DDK-dependent step. Fkh1-dependent recruitment of DDK results in phosphorylation of MCM, Cdc45 loading and local release from Rif1 and PP1. See text for further discussion.
Additional files
-
Supplementary file 1
Plasmid list.
- https://doi.org/10.7554/eLife.45512.016
-
Supplementary file 2
Primer list.
- https://doi.org/10.7554/eLife.45512.017
-
Supplementary file 3
Strain list.
- https://doi.org/10.7554/eLife.45512.018
-
Supplementary file 4
Reagent list.
- https://doi.org/10.7554/eLife.45512.019
-
Transparent reporting form
- https://doi.org/10.7554/eLife.45512.020





