Bridging the gap between single-cell migration and collective dynamics
Figures
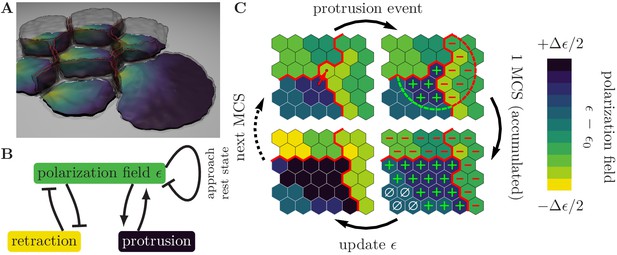
Illustration of the computational model with the pertinent simulation steps.
(A) Illustration of a small cell cohort that adheres to a surface (-plane). The polarization field, , is defined on the contact surface with the adhesion plane. The magnitude of the polarization field, which is indicated by the colorbar in Figure (C), encodes the local strength of cell-substrate adhesions and emulates the local mass of force-generating (pushing) cytoskeletal structures. Cell-cell adhesions are indicated in red. (B) Cytoskeletal structures respond to external mechanical stimuli through reaction networks involving different feedback loops. We greatly simplify these complex processes into two prototypic feedback loops, which break detailed balance and drive cell migration, as follows. The polarization field induces membrane protrusions and inhibits retractions. In turn, protrusions increase the polarization field (positive feedback) and therefore the likelihood of further protrusive activity, while retractions decrease the polarization field (negative feedback). In the absence of mechanochemical signals, the polarization field approaches its rest state. (C) Zoom-in to a common boundary shared between the substrate contact areas of three cells (bounded by the red lines), each represented by a contiguous set of occupied grid sites (hexagons). Top left: The upper right corner of the lower left cell (source cell) initiates a protrusion event against a neighboring element in the cell to its right (target cell), as indicated by the arrow, in an attempt to displace it. The success of each such attempted elementary event depends on the balance between contractile forces, cytoskeletal forces, and cell adhesion. Top right: If the protrusion event is successful, then the levels of regulatory factors are increased (decreased) in integer steps, at all lattice sites inside the source (target) cell that lie within a radius of the accepted protrusion event (as indicated by the plus and minus signs). Bottom right: During the course of one MCS, different levels of regulatory factors accumulate locally within each cell, with positive levels of regulatory factors (green plus signs) promoting a build-up of cytoskeletal structures, negative levels of regulatory factors (red minus signs) causing degradation of cytoskeletal structures, and neutral levels of regulatory factors (white zero signs) causing relaxation towards a resting state, as indicated in the lower left image. The color code indicates local levels of cytoskeletal structures, .
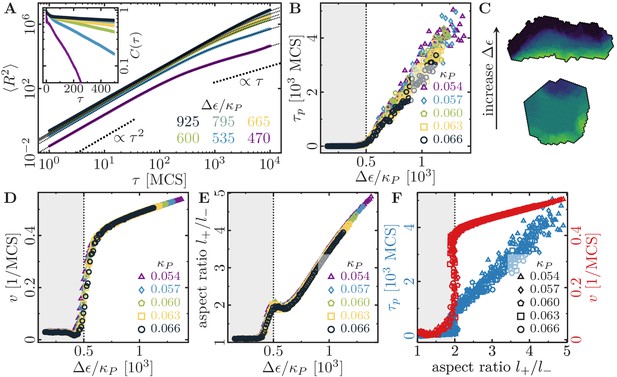
Cell shape and persistence of migration as a function of cell polarizability.
(A) Mean-squared displacement (MSD) for single-cell movements at different maximum cell polarity (stiffness parameters , ; average polarization field ; signaling radius ; cell-substrate dissipation ; cell-substrate adhesion penalty ; cytoskeletal update rate ; independent simulations for each set of parameters). Single cells perform a persistent random walk, i.e. they move ballistically () for , and diffusively () for . Inset: Normalized velocity auto-correlation function for the same parameters as in the main figure. (B) Persistence time of directed cell migration plotted as a function of maximum cell polarity , and perimeter stiffness (area stiffness ; average polarization field ; signaling radius ; cell-substrate dissipation ; cell-substrate adhesion penalty ; cytoskeletal update rate ; independent simulations for each set of parameters). The persistence time of the random walk increases with increasing cytoskeletal polarity and decreasing perimeter elasticity. (C) Cytoskeletal polarity also controls cell shapes, with crescent cell shapes (long persistence times) being observed at large cytoskeletal polartities, and round cell shapes (short persistence times) at small cytoskeletal polarities. Color code: cell polarization; cf. color bar in Figure 1C. (D) Single cell speed plotted as a function of maximum cell polarity , and perimeter stiffness . (E) Single cell aspect ratio plotted as a function of maximum cell polarity , and perimeter stiffness . (F) Speed and persistence time of single cells are correlated with the cell aspect ratio.
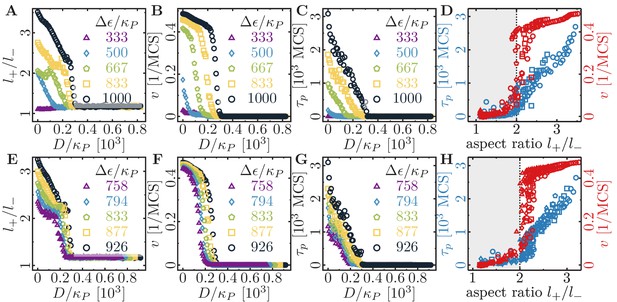
Role of substrate dissipation for cell shape and motility.
(A–D) Role of substrate dissipation for cells of varying maximum cell polarity . The aspect ratio (A), the speed (B), and the persistence time (C) as a function of substrate dissipation for a series of values for indicated in the graphs. (D) Cell persistence and cell speed are correlated with the aspect ratio of the cell. Only cells with an aspect ratio larger than 2 are motile. In the simulations, the substrate dissipation was varied in the interval , and the maximum cell polarity . Fixed parameters: average polarization field ; area elasticity ; membrane elasticity ; cytoskeletal update rate ; cell-substrate adhesion penalty ; independent simulations for each set of parameters. (E–F) Role of substrate dissipation for cells of varying membrane stiffness . The aspect ratio (E), the speed (F), and the persistence time (G) as a function of substrate dissipation for a series of values for indicated in the graphs. (H) Cell persistence and cell speed are correlated with the aspect ratio of the cell. Only cells with an aspect ratio larger than 2 are motile. In the simulations, the substrate dissipation was varied in the interval , and the membrane elasticity . Fixed parameters: average polarization field ; area elasticity ; maximum cell polarity ; cytoskeletal update rate ; cell-substrate adhesion penalty ; independent simulations for each set of parameters.
Single cell motility and shape for different maximum cell polarities (, ).
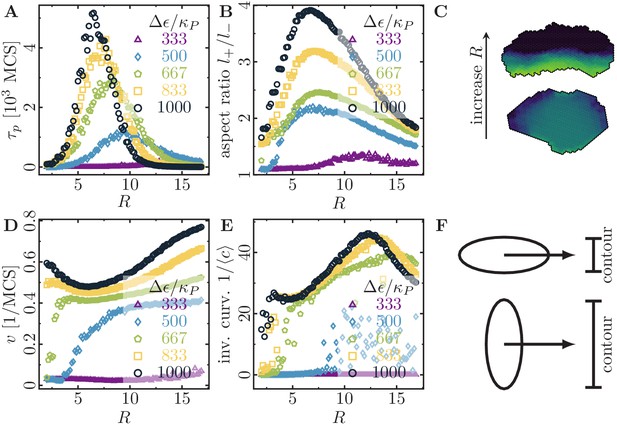
Migratory behavior of single cells as a function of the cell’s signaling radius at different values for the maximal cytoskeletal polarity .
(Stiffness parameters , ; average polarization field ; cell-substrate dissipation ; cell-substrate adhesion penalty ; cytoskeletal update rate ; independent simulations for each set of parameters). (A) The persistence times of directed migration of single cells exhibit a pronounced maximum at an optimal signaling radius, which depends on cell polarizability. (B) The shapes of single cells exhibit a pronounced maximal elongation at an optimal signaling radius, which depends on cell polarizability. (C) The signaling radius critically determines the synchronicity of internal cytoskeletal remodeling processes. Small signaling radii frequently lead to transient formation of mutually independent lamellipodia at different positions around the cell perimeter, thereby interrupting persistent motion (reducing persistence times of directed migration). Large signaling radii lead to structurally stable front-rear polarization profiles across the entire cell body (long persistence times of directed migration). Color code: cell polarization; cf. color bar in Figure 1C. (D) The speed of single cells does not drop to zero even when their persistence time of directed migration vanishes. This indicates single cell rotations. (E) The inverse curvature of the cell trajectories as a function of the signaling radius. (F) Depending on whether a cell migrates along its long axis (top) or short axis (bottom), it has to move a different projected contour length. If each protrusion takes roughly the same amount of time, then migration along the long axis (top; cell has to move a smaller projected contour length) allows for greater cell speeds than migration along the short axis (bottom; cell has to move a larger projected contour length).
Single cell motility and shape for different signaling radii (, ).
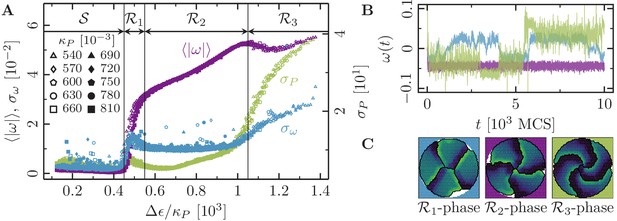
Phases of collective motion.
(-cell systems; confinement radius ; area stiffness ; average polarization field ; signaling radius ; cytoskeletal update rate ; cell-cell adhesion ; cell-cell dissipation ; cell-substrate dissipation ; cell-substrate adhesion penalty (), (); 100 independent simulations for each set of parameters). (A) Characteristic observables of collective cell rotation at different values of the cell perimeter stiffness parameter : mean () and standard deviation () of the magnitude of the cell cluster's angular velocity, and the standard deviation of the cell perimeter (). The statistics of collective cell motion depends only on the ratio of maximum cell polarity, , to cell contractility, (specific polarizability). (B) Representative angular trajectories and (C) cell shapes (color code represents cell polarization; cf. Figure 1C) for the different parameter regimes as described in the main text. The cellular dynamics in the different parameter regimes are shown in Figure 4—video 1, Figure 4—video 2 and Figure 4—video 3.
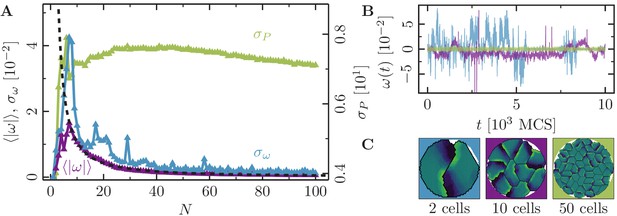
Collective motion for varying number of cells at low polarizability.
(-cell systems; confinement radius ; stiffness parameters , ; average polarization field ; maximum cell polarity ; signaling radius ; cytoskeletal update rate ; cell-cell adhesion ; cell-cell dissipation ; cell-substrate dissipation ; cell-substrate adhesion penalty (), (); independent simulations for each set of parameters). For this choice of parameters -cell populations rotate in the -phase. We observe a similar behavior here: the cell clusters rotate slowly and reorient frequently. (A) Characteristic observables of collective cell rotation at different values of the cell perimeter stiffness parameter : mean () and standard deviation () of the angular velocity magnitude of cell motion, and the standard deviation of the cell perimeter (). The black line corresponds to a power-law fit of the form with the fitted exponent . (B) Representative angular trajectories and (C) cell shapes (color code represents cell polarization; cf. Figure 1) for the different parameter regimes as described in the main text. .
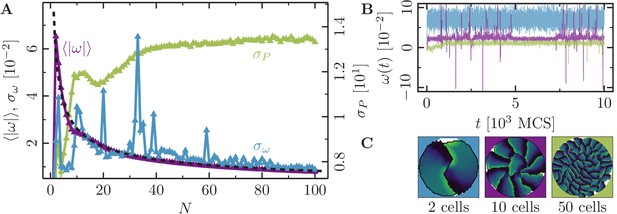
Collective motion for varying number of cells at intermediate polarizability.
(-cell systems; confinement radius ; stiffness parameters , ; average polarization field ; maximum cell polarity ; signaling radius ; cytoskeletal update rate ; cell-cell adhesion ; cell-cell dissipation ; cell-substrate dissipation ; cell-substrate adhesion penalty (), (); independent simulations for each set of parameters). For this choice of parameters -cell populations rotate in the -phase. We observe a similar behavior here: highly correlated rotations with no changes in rotational direction. (A) Characteristic observables of collective cell rotation at different values of the cell perimeter stiffness parameter : mean () and standard deviation () of the angular velocity magnitude of cell motion, and the standard deviation of the cell perimeter (). The black line corresponds to a power-law fit of the form . (B) Representative angular trajectories and (C) cell shapes (color code represents cell polarization; cf. Figure 1) for the different parameter regimes as described in the main text. .
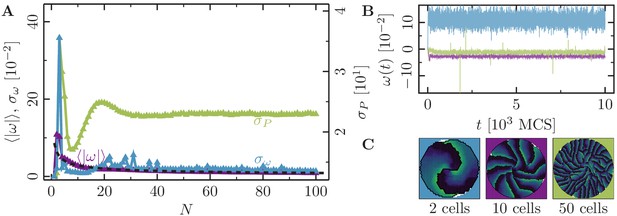
Collective motion for varying number of cells at high polarizability.
(-cell systems; confinement radius ; stiffness parameters , ; average polarization field ; maximum cell polarity ; signaling radius ; cytoskeletal update rate ; cell-cell adhesion ; cell-cell dissipation ; cell-substrate dissipation ; cell-substrate adhesion penalty (), (); independent simulations for each set of parameters). For this choice of parameters -cell populations rotate in the -phase. We observe a similar behavior here: highly correlated rotations. (A) Characteristic observables of collective cell rotation at different values of the cell perimeter stiffness parameter : mean () and standard deviation () of the angular velocity magnitude of cell motion, and the standard deviation of the cell perimeter (). The black line corresponds to a power-law fit of the form . (B) Representative angular trajectories and (C) cell shapes (color code represents cell polarization; cf. Figure 1) for the different parameter regimes as described in the main text.
Collective rotations of cells in the -phase (; ).
Collective rotations of cells in the -phase (; ).
Collective rotations of cells in the -phase (; ).
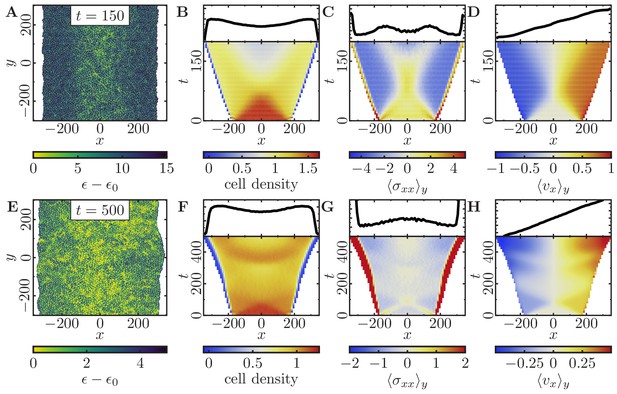
Expansion of a confluent epithelial cell sheet after removal of boundaries positioned at for two different parameter settings.
(Stiffness parameters , ; average polarization field ; signaling radius ; cytoskeletal update rate ; cell-cell adhesion ; cell-cell dissipation ; cell-substrate dissipation ; cell-substrate adhesion penalty ; independent simulations for each set of parameters). (A–D) Tissue expansion for a migration-dominated setup without explicit cell growth and mitosis. (-cell system; maximum cell polarity ). (E–H) Tissue expansion at low density and cell polarizability for a cell sheet comprised of dividing cells. (Initially a -cell system; maximum cell polarity ; growth time ; division time ; size threshold for cell growth , where is the size of a solitary cell in equilibrium). (A, E) Snapshots of the polarization field ; cf. Figure 5—video 1 and Figure 5—video 2. (B, F) Kymographs showing the cell density averaged over the -direction and (top) final snapshots of the cell density profiles. (C, G) Kymographs showing the component of the stress tensor averaged over the -direction and (top) final snapshots of the stress profiles. (D, H) Kymographs showing the component of the cell velocities averaged over the -direction and (top) final snapshot of the velocity profiles.
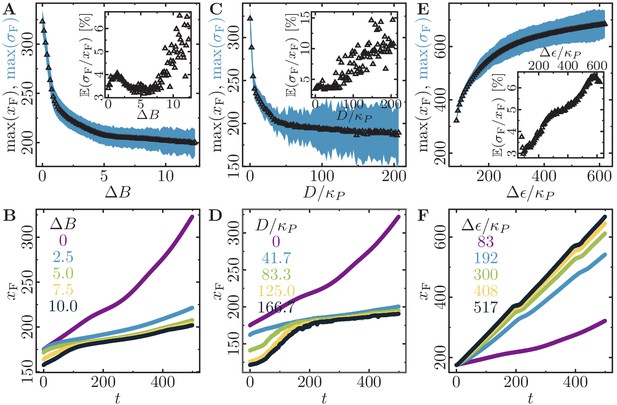
Monolayer expansion depends on dissipation and cell polarizability.
Cell monolayer expansion depends on the cell-cell dissipation, cell-substrate dissipation, and maximum cell polarity. (Initially a -cell system; stiffness parameters , ; average polarization field ; maximum cell polarity ; signalling radius ; cytoskeletal update rate ; cell-cell adhesion ; cell-substrate adhesion penalty ; growth time ; division time ; size threshold for cell growth , where is the size of a solitary cell in equilibrium; independent simulations for each set of parameters). (A,B) Cell monolayer expansion depends on the cell-cell dissipation (maximum cell polarity ; cell-substrate dissipation ). (A) Maximal monolayer extension and roughness (we exclude an initial time interval of 200 MCS because it takes at least that long for first daughter cells to appear). Inset: Relative roughness of the spreading monolayer relative to its size. (B) Time traces for selected values of . (C,D) Cell monolayer expansion depends on the cell-substrate dissipation (maximum cell polarity ; cell-cell dissipation ). (C) Maximal monolayer extension and roughness (we exclude an initial time interval of 200 MCS because it takes at least that long for first daughter cells to appear). (D) Time traces for selected values of . (E,F) Cell monolayer expansion depends on the maximum cell polarity (cell-cell dissipation ; cell-substrate dissipation ). (E) Maximal monolayer extension and roughness (we exclude an initial time interval of 200 MCS because it takes at least that long for first daughter cells to appear). (F) Time traces for selected values of . .
Motility-dominated tissue dynamics.
Proliferation-dominated tissue dynamics.
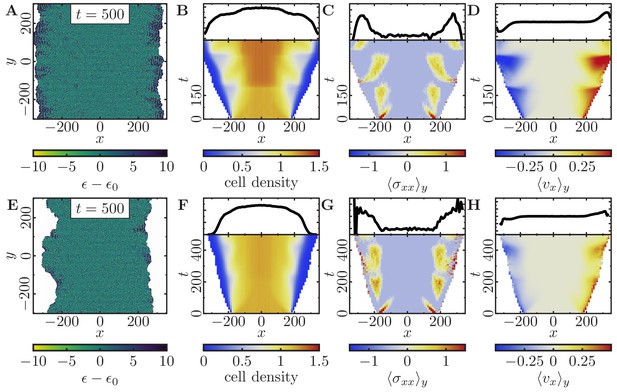
Expansion of a confluent epithelial cell sheet after removal of boundaries positioned at for two different parameter settings that produce rough tissue fronts.
(Initially a -cell system; stiffness parameters , ; average polarization field ; maximum cell polarity ; signaling radius ; cytoskeletal update rate ; cell-cell adhesion ; cell-cell dissipation ; cell-substrate dissipation ; cell-substrate adhesion penalty ; growth time ; division time ; independent simulations for each set of parameters). (A–D) Tissue expansion at low density and cell polarizability for a cell sheet comprised of quickly dividing cells. (Size threshold for cell growth , where is the size of a solitary cell in equilibrium). (E–H) Tissue expansion at low density and cell polarizability for a cell sheet comprised of slowly dividing cells. (Size threshold for cell growth , where is the size of a solitary cell in equilibrium). (A, E) Snapshots of the polarization field ; cf. Figure 6—video 1 and Figure 6—video 2. (B, F) Kymographs showing the cell density averaged over the -direction and (top) final snapshots of the cell density profiles. (C, G) Kymographs showing the component of the stress tensor averaged over the -direction and (top) final snapshots of the stress profiles. (D, H) Kymographs showing the component of the cell velocities averaged over the -direction and (top) final snapshot of the velocity profiles.
Weak monolayer roughening (fingering) in motility-dominated tissue with quick proliferation.
Strong monolayer roughening in motility-dominated tissue with slow proliferation.
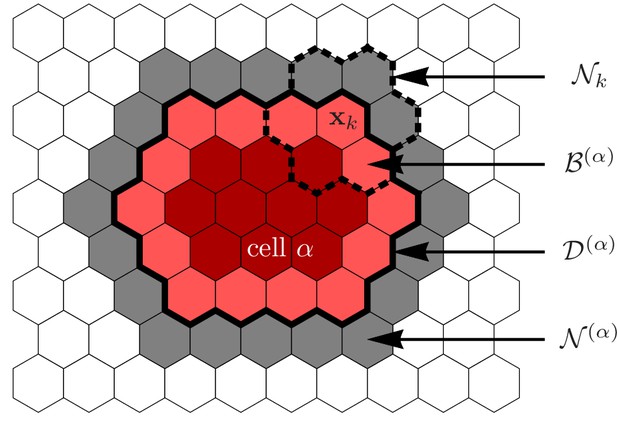
Illustration of the various sets defining a cell and its environment.
Grid sites occupied by cell , i.e. its domain , are indicated in red colors. The cell’s membrane sites, , are indicated by the lighter red color, the cell’s immediate neighborhood, , is indicated in gray. Elementary events involving cell always involve one grid site in and one grid site in . For the hexagonal lattices used in this work, each grid site is surrounded by nearest neighbors which we collectively denote by .
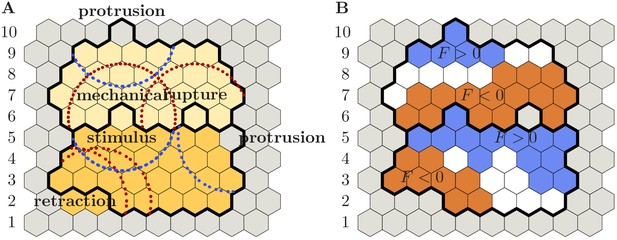
Distribution of regulatory factors on the basis of accepted elementary events.
For ease of reference, grid rows have been numbered from to . Left (A): Solid black lines indicate cells’ membrane positions after acceptance of the respective elementary event; colors indicate cellular domains before the respective elementary event has been accepted (gray: substrate; shades of yellow: cells). Blue and red circular arcs (of radius ) delineate areas of local increase or decrease in the level of regulatory factors, respectively. The following elementary events are depicted: (i) lower cell retracts (two grid sites in row ); (ii) lower cell protrudes (row ); (iii) upper cell protrudes (row ). In addition, the following elementary events occur across the cell-cell boundary: (iv) retraction of upper cell leads to rupture of cell-cell contacts (row , right event); (v) either the lower cell protrudes and pushes the upper cell or the upper cell retracts and pulls on the lower cell (row , left event). Specifically, event (v) entails mechanical signaling between the upper and lower cell and, therefore, affects the distribution of regulatory factors in both cells. Right (B): Identical copy of the left image (A). Colors indicate local levels of regulatory factors (blue: is positive; white: is zero; red: is negative; gray: substrate site). Note, in particular, that a substrate grid site has been inserted where cell rupture occurred (row , right grid site). The following cases can be distinguished: (i) Grid site lies in the zone of influence of only positive (blue circles) or negative (red circles) chemical feedback, in which case the level of regulatory factors is positive or negative, respectively (e.g. red grid sites in row , or blue grid sites in row ). (ii) Grid site lies outside of any zone of influence, in which case the level regulatory factors is zero (e.g. white grid sites in row ). (iii) Grid site lies in the zone of influence of equally many positive and negative feedbacks, in which case the level of regulatory factors remains zero (e.g. fourth grid site in row ). (iv) Grid site lies in a zone of predominantly positive or negative feedback, in which case the level of regulatory factors is positive or negative, respectively (e.g. third grid site in row ). Recall that only the sign of is of significance to update the cells’ polarization field; cf. Equation S13.
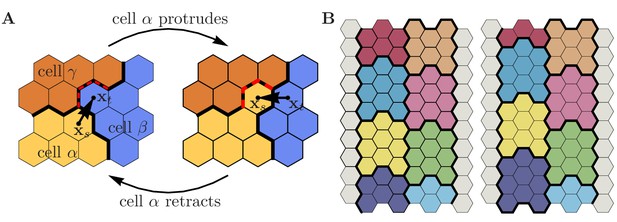
Cell-cell adhesion.
(A) Adhesive energy contribution in a cyclic process, where a protrusion of source cell against target cell is followed by the inverse retraction event. Both events involve a third party cell , leading to net energy dissipation after the cyclic process has been completed. Protrusion: (i) Three pre-existing cell-cell contacts between and are torn apart (red dashed contacts); (ii) three new contacts between and are formed; (iii) the contact length between source cell and target cell increases by one unit of length. This implies . Retraction: (i) Three pre-existing cell-cell contacts between and are torn apart (red dashed contacts); (ii) three new contacts between and are formed; (iii) the contact length between source cell and target cell decreases by one unit of length. This implies . Altogether, this leads to , i.e. a (non-negative) dissipative contribution, whose magnitude depends on the dissipation matrix . (B) Shear viscosity due to cell-cell adhesion. Consider two rows of adhesive cells sliding past each other as indicated in the figure (left row of cells moves up by one grid site; colors indicate different cells). The associated adhesion energy change (per cell) reads , where denotes the number of cells sliding past each other, and where we assumed cells of like type, i.e. and (). The condition , Equation S15e, thus implies positive friction associated with cellular shear flows, whose magnitude is proportional to the number of cells sliding past each other. Note that this shear viscosity vanishes for , i.e. for zero dissipation matrix.
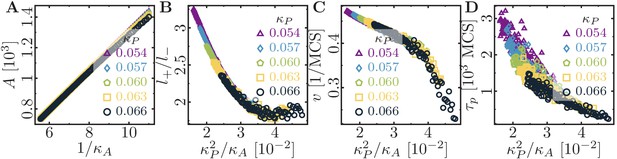
Role of area stiffness for cell size and motility.
(A) The cell area increases linearly with . The aspect ratio (B), speed (C) and persistence (D) of the cell decrease with increasing cell size. In the simulations, the area elasticity was varied in the interval , and the membrane elasticity was chosen from . Fixed parameters: average cell polarization field ; maximum cell polarity ; signaling radius ; cytoskeletal update rate ; cell-substrate dissipation ; cell-substrate adhesion penalty .
Tables
Source and parameter files used for each figure.
All source and parameter files are found in Source data 1.
| Figure | Simulation code | Processing code | Parameters |
|---|---|---|---|
| Figure 2 | CPM_NoDivision | TrajectoryAnalysisSingle | single_Q |
| Figure 2—figure supplement 1 (A-D) | CPM_NoDivision | TrajectoryAnalysisSingle | single_DQ |
| Figure 2—figure supplement 1 (E-H) | CPM_NoDivision | TrajectoryAnalysisSingle | single_DM |
| Figure 3 | CPM_NoDivision | TrajectoryAnalysisSingle | single_R |
| Figure 4 | CPM_NoDivision | TrajectoryAnalysisCircularPattern | rotation_Q |
| Figure 4—figure supplement 1 | CPM_NoDivision | TrajectoryAnalysisCircularPattern | rotation_N_R1 |
| Figure 4—figure supplement 2 | CPM_NoDivision | TrajectoryAnalysisCircularPattern | rotation_N_R2 |
| Figure 4—figure supplement 3 | CPM_NoDivision | TrajectoryAnalysisCircularPattern | rotation_N_R3 |
| Figure 5 (A-D) | CPM_Division | wound_nodiv | |
| Figure 5 (E-H) | CPM_Division | wound_div | |
| Figure 5—figure supplement 1 (A-B) | CPM_Division_Supplement | FrontAnalysis | wound_div_A |
| Figure 5—figure supplement 1 (C-D) | CPM_Division_Supplement | FrontAnalysis | wound_div_D |
| Figure 5—figure supplement 1 (E, F) | CPM_Division_Supplement | FrontAnalysis | wound_div_Q |
| Figure 6 (A-D) | CPM_Division | wound_div_fing_1.0 | |
| Figure 6 (E-H) | CPM_Division | wound_div_fing_1.1 | |
| Appendix 2—figure 1 | CPM_NoDivision | TrajectoryAnalysisSingle | single_A |
Additional files
-
Source code 1
Simulation code, processing code and parameter files.
- https://cdn.elifesciences.org/articles/46842/elife-46842-code1-v2.tar
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/46842/elife-46842-transrepform-v2.pdf





