A contractile injection system stimulates tubeworm metamorphosis by translocating a proteinaceous effector
Figures
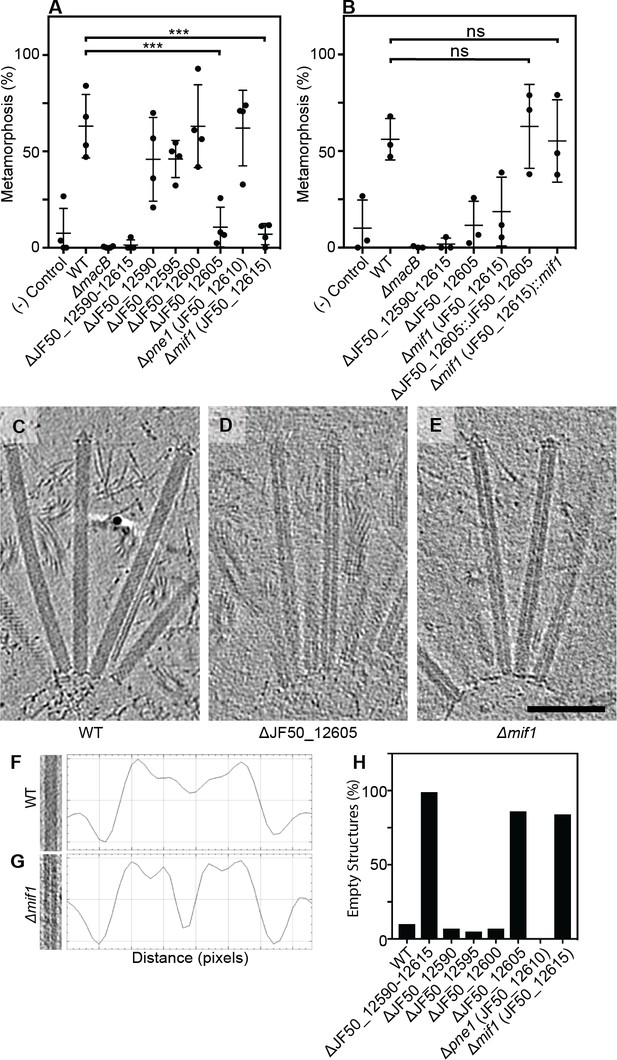
Two bacterial genes are important for inducing Hydroides metamorphosis and deletions of these genes produce MACs with an ‘empty’ phenotype.
(A) Metamorphosis (%) assay of Hydroides larvae in response to biofilms of P. luteoviolacea wildtype (WT) and different gene deletion strains. Deletion of JF50_12605 or mif1 (JF50_12615) showed a significant loss in the ability to induce metamorphosis when compared to wildtype. (B) Restoration of JF50_12605 and mif1 (JF50_12615) into their native chromosomal loci restored function. Graphs in (A/B) show an average of biological replicates, where each point represents one biological replicate. *p-value≤0.05, ns = not significant. (C–E) Representative cryotomographic images of the ‘filled’ phenotype from wildtype MACs (C), and ‘empty’ phenotype from ΔJF50_12605 (D), and Δmif1 (E) MACs. Scale bar, 100 nm. (F/G) Shown are representative MAC structures (on left; taken from C/E) and their density plots. The wildtype ‘filled’ phenotype shows a relatively homogeneous density profile across the diameter of the MAC. The Δmif1 ‘empty’ phenotype shows a low-density region in the center of the MAC. (H) Shown is the fraction of empty structures for different deletion mutants as observed by cryoET imaging. Note that the ‘empty’ phenotype correlates with the inability to induce metamorphosis (A).
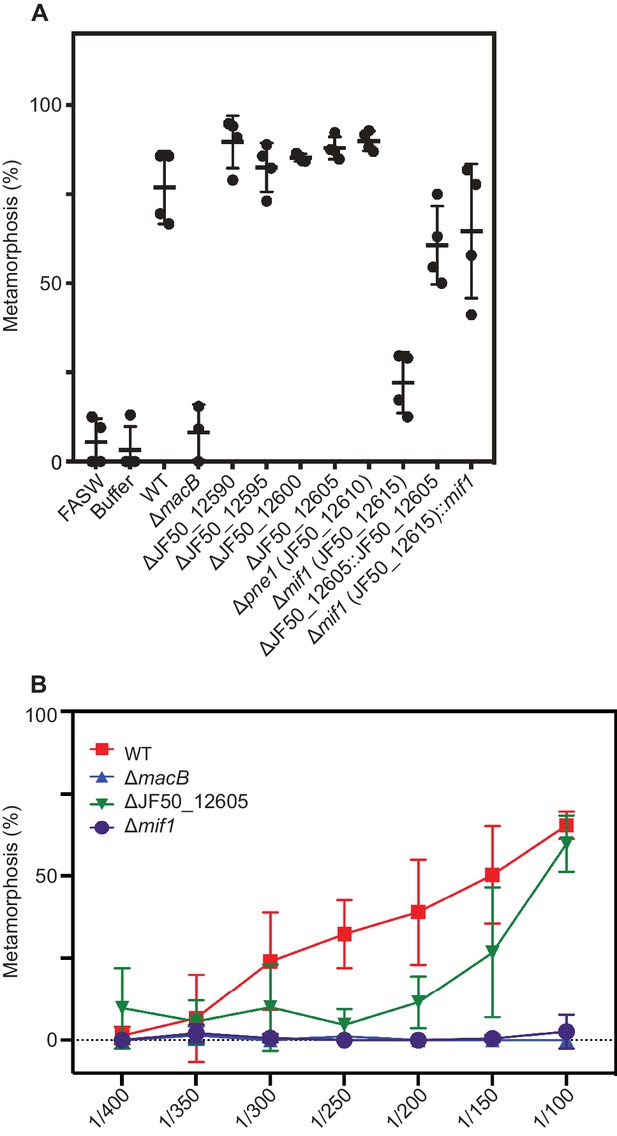
Metamorphic response of Hydroides larvae to cell-free MAC extracts from wild type P. luteoviolacea and individual gene mutants.
(A) Metamorphosis (%) of Hydroides larvae 24 h after exposure to extracted MACs from P. luteoviolacea wildtype (WT) and mutants. MAC extracts were diluted 1:100 before being mixed with larvae. (B) Dose-response curve of MAC extracts from WT (red), ∆macB (blue), ∆JF50_12605 (green), and ∆mif1 (purple) mutants.

Wildtype and ΔJF50_12590-12615 have structurally similar arrays.
(A-D) MAC arrays were present in (A/B) wildtype (WT) and (C/D) ΔJF50_12590-12615 MAC extracts. Both strains showed arrays that comprised individual contractile structures in extended and contracted conformations. Shown are cryotomographic slices. Scale bar, 100 nm.
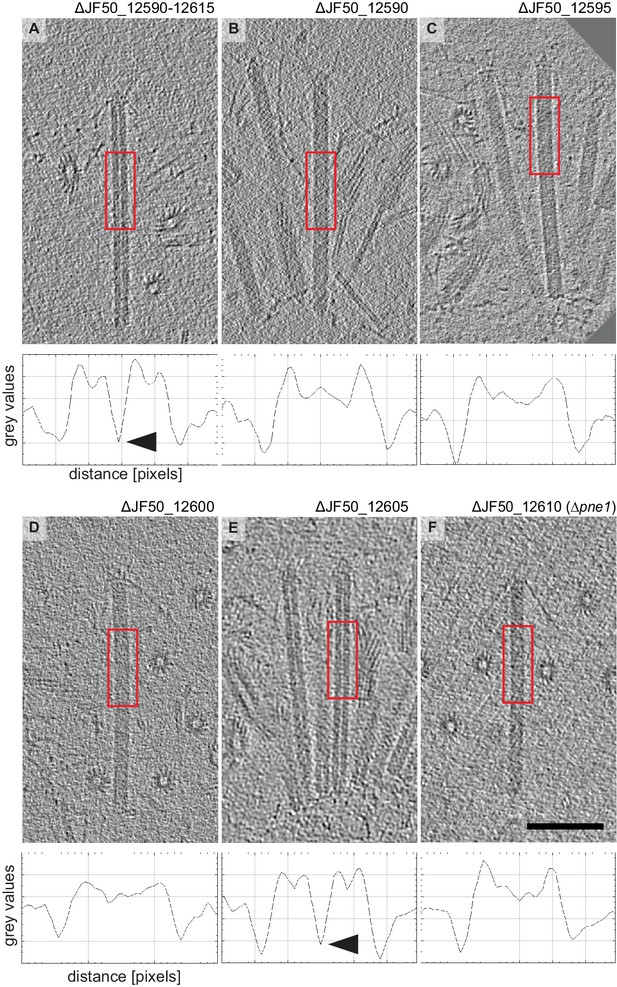
‘Filled’ and ‘empty’ phenotypes in all studied gene mutant strains.
(A-F) Shown are cryotomographic slices of representative extended MACs for different strains and the corresponding density plots (plots were calculated from the boxed image region). ‘Empty’ and ‘filled’ phenotypes are characterized by differences in densities in the MAC center (arrowheads indicate low-density region in empty structures). Scale bar, 100 nm.

Replacing JF50_12605 and mif1 into their native chromosomal loci generates MACs with filled tubes.
(A–B) When complemented into the native chromosomal loci, (A) JF50_12605 and (B) mif1 assemble wildtype-like MACs with a filled phenotype. Scale bar, 100 nm.
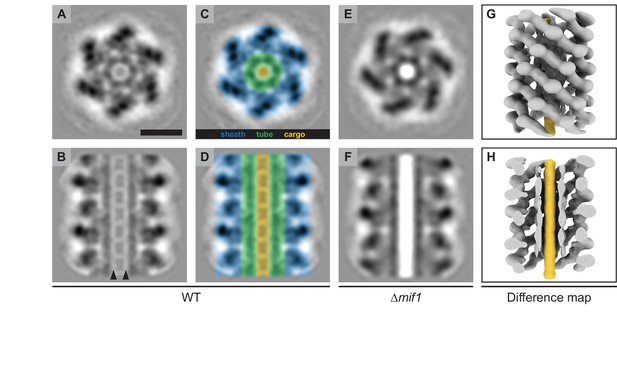
MACs from a ∆mif1 mutant lack electron density in the tube lumen.
(A–F) Cross sectional (A/C/E) and longitudinal (B/D/F) slices through subtomogram averages of the MAC sheath-tube complex from wildtype (WT; A–D) and ∆mif1 (E–F). The hexameric sheath and tube modules could be clearly discerned (indicated in C/D). The inner tube lumen displayed clear differences in density between WT and ∆mif1. The wildtype tube lumen was filled with densities that likely represent cargo (A-D, indicated in yellow), which was not present in the ∆mif1 lumen (E/F). Note the low-density region that separates the tube and cargo (indicated by arrowheads in B). (G/H) Shown are isosurfaces of the ∆mif1 structure (gray) and of a difference map (yellow; calculated from the wildtype and ∆mif1 structure), highlighting the additional density in the wildtype tube lumen. Scale bar, 10 nm.
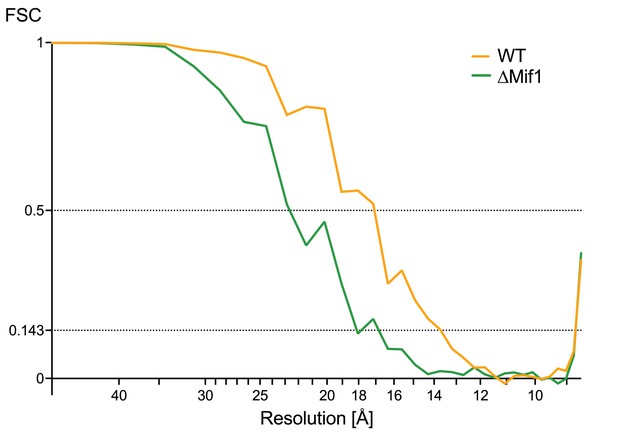
Fourier shell correlations.
Fourier shell correlation (FSC) between the two independently aligned and averaged half-datasets for the wildtype (yellow graph, WT) and the ∆mif1 (green graph) MAC subtomogram averages. Resolution estimates are ~17 Å and ~23 Å at the 0.5 threshold for wildtype and ∆mif1, respectively, and ~14 Å and ~18 Å at the 0.143 threshold.

Triggered MAC tubes show ‘empty’ phenotype.
Shown is a representative cryotomographic slices of WT MACs. Arrowheads indicate expelled tubes with ‘empty’ phenotype.
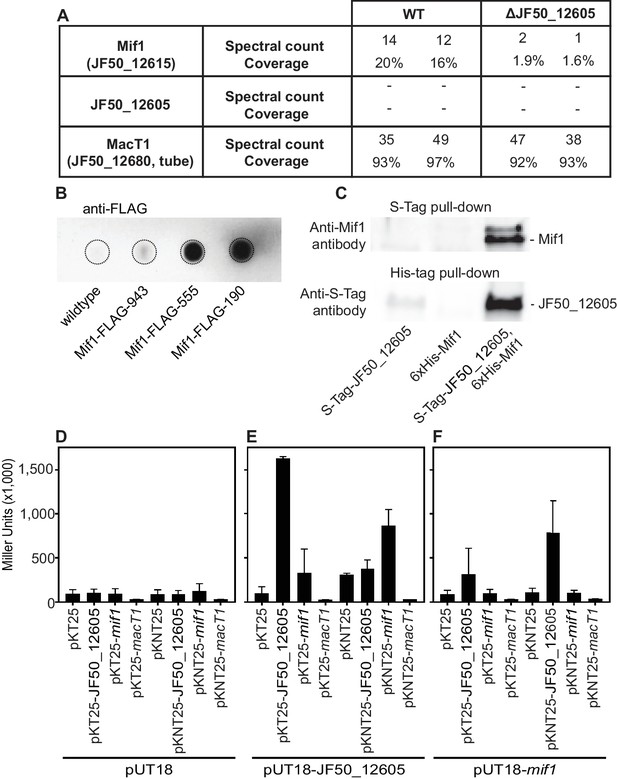
Mif1 is present in MAC complexes and JF50_12605 is required for Mif1’s association with the MAC complex.
(A) Mass spectrometry of wildtype MAC arrays detected Mif1 but not JF50_12605. Spectral counts for Mif1 were low for MACs purified from the ∆JF50_12605 mutant, indicating a possible chaperone-like function for JF50_12605. (B) Dot blot of purified MACs from wildtype or strains with Mif1 tagged at amino acid positions 943 [C-terminus] (Mif1-FLAG-943), 555 (Mif1-FLAG-555), and 190 (Mif1-FLAG-190) were probed with anti-FLAG antibody. The signal indicates association of Mif1 with MAC arrays. (C) Co-expression, reciprocal pull-down and western blotting of S-tagged JF50_12605 and 6xHis-tagged Mif1 indicate an interaction between both proteins. Strains in which only one component was tagged were used as controls. (D–F) Quantification of bacterial two-hybrid experiments were used to analyze possible interactions between JF50_12605, Mif1 and tube (MacT1) proteins. Briefly, the two fragments of CyaA (T18/T25) were fused to the respective target proteins with the CyaA activity only being restored by interaction between target proteins. JF50_12605 showed a strong interaction with itself and with Mif1.
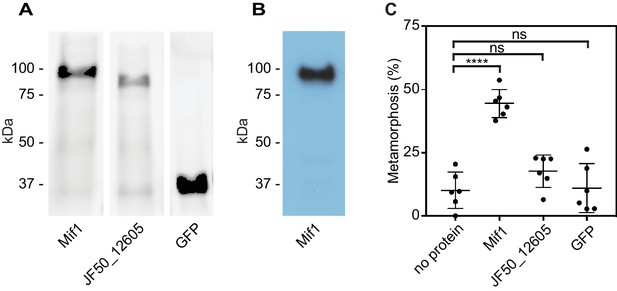
Mif1 is sufficient for stimulating metamorphosis when delivered by electroporation.
(A) Shown is an SDS page gel of purified Mif1, JF50_12605 and GFP. (B) Western blot of purified Mif1 protein probed with a C-terminal anti-Mif1 peptide antibody confirms Mif1 identity. (C) Metamorphosis (%) of Hydroides larvae 24 hr after electroporation with purified Mif1, JF50_12605 or GFP protein, shows induction of metamorphosis by electroporated Mif1. Graph shows an average of biological replicates, where each point represents one biological replicate. ****p-value≤0.0001 by t-test, ns = not significant.

Purified Mif1 is unable to induce metamorphosis when added exogenously.
Metamorphosis (%) of Hydroides larvae after being soaked in 250 ng/μl of purified GFP, JF50_12605, and JF50_12615 protein for 24 h. Wildtype (WT) MACs diluted 1:100 were used as a positive control for larval competence.
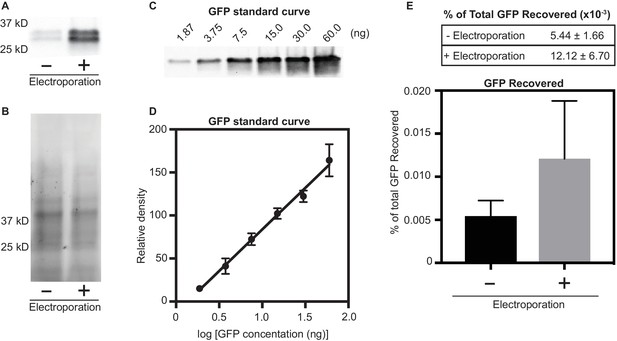
Quantification of GFP protein associated with larvae after electroporation.
We tested if electroporation results in a higher abundance of protein within tubeworm larvae. To this end, tubeworm larvae were mixed with purified GFP protein (0.625 µg/µl, 50 µg total) and electroporated at 30 V (150 V/cm) at 10 ohms and 3000 µF. As a control, a second treatment of tubeworm larvae mixed with GFP without electroporation was performed. After electroporation, larvae were washed five times to remove unassociated GFP protein. (A) Western blot analysis of larval lysate with and without electroporation resulted in higher recovery of GFP in electroporated larvae (+) when compared to un-electroporated control (-). (B) To normalize protein loading, we visualized the stain-free SDS-PAGE gel. (C/D) To quantify the amount of protein recovered after electroporation, a GFP standard curve was created by performing a Western blot on known GFP concentrations (n = 3). (E) To determine the amount of GFP associated with tubeworm larvae, we quantified GFP with (+) and without (-) electroporation. Values are a mean percent of total GFP electroporated (n = 4 biological replicates, ± SD).
Tables
Strains and plasmids used in this work.
https://doi.org/10.7554/eLife.46845.015| Strain no. | Genotype | Source or reference | |
|---|---|---|---|
| Strain | |||
| NJS5 | HI1 StrR | P. luteoviolacea HI1, StrR | (Huang et al., 2012) |
| NJS23 | ∆macB | P. luteoviolacea HI1, StrR∆macB | (Shikuma et al., 2014) |
| NJS235 | ∆JF50_12590-F50_12615 | P. luteoviolacea HI1, StrR ∆R4 | (Shikuma et al., 2016) |
| NJS289 | ∆JF50_12590 | P. luteoviolacea HI1, StrR∆JF50_12590 | This Study |
| NJS287 | ∆JF50_12595 | P. luteoviolacea HI1, StrR∆JF50_12595 | This Study |
| NJS285 | ∆JF50_12600 | P. luteoviolacea HI1, StrR∆JF50_12600 | This Study |
| NJS283 | ∆JF50_12605 | P. luteoviolacea HI1, StrR∆JF50_12605 | This Study |
| NJS281 | ∆JF50_12610 | P. luteoviolacea HI1, StrR∆JF50_12610 | This Study |
| NJS279 | ∆JF50_12615 | P. luteoviolacea HI1, StrR∆JF50_12615 | This Study |
| NJS294 | ∆JF50_12605::12605 | P. luteoviolacea HI1, StrR∆JF50_12605::JF50_12605 | This Study |
| NJS295 | ∆JF50_12615::12615 | P. luteoviolacea HI1, StrR∆JF50_12615::JF50_12615 | This Study |
| Plasmid | |||
| pNJS007 | pCVD443 | AmpR, KmR, sacB, pGP704 derivative | (Huang et al., 2012) |
| pNJS266 | pCVD443_∆12590 | pCVD443::∆12590 AmpR, KmR | This Study |
| pNJS265 | pCVD443_∆12595 | pCVD443::∆12595 AmpR, KmR | This Study |
| pNJS264 | pCVD443_∆12600 | pCVD443::∆12600 AmpR, KmR | This Study |
| pNJS263 | pCVD443_∆12605 | pCVD443::∆12605 AmpR, KmR | This Study |
| pNJS262 | pCVD443_∆12610 | pCVD443::∆12610 AmpR, KmR | This Study |
| pNJS261 | pCVD443_∆12615 | pCVD443::∆12615 AmpR, KmR | This Study |
| pNJS256 | pCVD443_12590–615 complement | AmpR, KmR | (Shikuma et al., 2016) |
| pNJS282 | pCVD443_12605 complement | AmpR, KmR | This Study |
| pNJS074 | pCVD443_∆12585 | AmpR, KmR | This Study |
| pNJS267 | pUT18 | AmpR | (Karimova et al., 2000) |
| pNJS268 | pUT18C | AmpR | (Karimova et al., 2000) |
| pNJS269 | pKT25 | KmR | (Karimova et al., 2000) |
| pNJS270 | pKNT25 | KmR | (Karimova et al., 2000) |
| pNJS283 | pUT18_12605 | AmpR | This Study |
| pNJS299 | pUT18_12615 | AmpR | This Study |
| pNJS527 | pUT18_12680 | AmpR | This Study |
| pNJS284 | pKT25_12605 | KmR | This Study |
| pNJS285 | pKT25_12615 | KmR | This Study |
| pNJS529 | pKT25_12680 | KmR | This Study |
| pNJS286 | pKNT25_12605 | KmR | This Study |
| pNJS300 | pKNT25_12615 | KmR | This Study |
| pNJS530 | pKNT25_12680 | KmR | This Study |
| pNJS393 | pET15b_12605 | AmpR | This Study |
| pNJS395 | pET15b_12615 | AmpR | This Study |
| pNJS397 | pET15b_GFP | AmpR | This Study |
Primers used in this work.
https://doi.org/10.7554/eLife.46845.016| Primer | Sequence |
|---|---|
| 1556_dA | TGATGGGTTAAAAAGGATCGATCCTCTAGATTGGAGCAATAAACGGGTTC |
| 1556_dB | GTTCATAATTAAACTGCGATCGCAGCCATAAGGCCTCCTTGATA |
| 1556_dC | TATCAAGGAGGCCTTATGGCTGCGATCGCAGTTTAATTATGAAC |
| 1556_dD | TTTTGAGACACAACGTGAATTCAAAGGGAGAGCTCCGCTTTGGGTACTGGCTTTA |
| 1556_intF | CCGAGCAAACGTTATCACAA |
| 1556_intR | TCAGCGCTCTCATTATGTGC |
| 1555_dA | TGATGGGTTAAAAAGGATCGATCCTCTAGACCGAGCAAACGTTATCACAA |
| 1555_dB | CCTTGCATGAGGTTAAGAAAGTTTGACGTACCCTTCAGCCATATT |
| 1555_dC | AATATGGCTGAAGGGTACGTCAAACTTTCTTAACCTCATGCAAGG |
| 1555_dD | TTTTGAGACACAACGTGAATTCAAAGGGAGAGCTCGATGCGGTAACGGTTGTTCT |
| 1555_intF | AGCGATTGATGCTGAACAAA |
| 1555_intR | ACCATCGCATAACCCGTAAC |
| 1554_dA | TGATGGGTTAAAAAGGATCGATCCTCTAGATACGCCGTCCAGTTAGGACT |
| 1554_dB | GTTTGTTAACGTCACGGCAGCTGCATTGCCATTTAAACTCC |
| 1554_dC | GGAGTTTAAATGGCAATGCAGCTGCCGTGACGTTAACAAAC |
| 1554_dD | TTTTGAGACACAACGTGAATTCAAAGGGAGAGCTCATTGATTGGAAGCGCGATAG |
| 1554_intF | TTTATGAGGCACCAACGACA |
| 1554_intR | GCCTGTGCCGTTTTATCTGT |
| 1553_dA | TGATGGGTTAAAAAGGATCGATCCTCTAGAGGCGATCAGTGGAGTGAAGT |
| 1553_dB | AATACTTCTTGCTCAGCCCCGCGTGCTTCTTCTGTCATGT |
| 1553_dC | ACATGACAGAAGAAGCACGCGGGGCTGAGCAAGAAGTATT |
| 1553_dD | TTTTGAGACACAACGTGAATTCAAAGGGAGAGCTCTCAGAACCAGCAGTCTCACG |
| 1553_intF | CGGGCCTAGAAATCACTCAA |
| 1553_intR | TCGACGTCAAATCAGTCGAG |
| 1552_dA | TGATGGGTTAAAAAGGATCGATCCTCTAGAGAGAGCAAGAAGTGGCGAGT |
| 1552_dB | TAGCCTTTTAGTGCCGCTTTTGAGGCGTCCATATCTGACA |
| 1552_dC | TGTCAGATATGGACGCCTCAAAAGCGGCACTAAAAGGCTA |
| 1552_dD | TTTTGAGACACAACGTGAATTCAAAGGGAGAGCTCTGCTGACCAAGCAGATTGAC |
| 1552_intF | GGGCAATTGTTGTGGATTTT |
| 1552_intR | TGATCCCAAACCACTTGTGA |
| 1551_dA | TGATGGGTTAAAAAGGATCGATCCTCTAGAGACTGCTGGTTCTGATTCGAT |
| 1551_dB | AACAGATCATTACATTAAAATGAGCCTCTGTTCTTGTTGTTGCATTTCA |
| 1551_dC | TGAAATGCAACAACAAGAACAGAGGCTCATTTTAATGTAATGATCTGTT |
| 1551_dD | TTTTGAGACACAACGTGAATTCAAAGGGAGAGCTCCTTCTCCATTTTCGCCTTTG |
| 1551_intF | CGTTTTCAGTGACCATCACG |
| 1551_intR | CGGTGGGCAAAAAGGTATAA |
| pUT18_605_F1 | CAGCTATGACCATGATTACGCCAAGCTTGCATGCCATGACAGAAGAAGCACGCGAAAAAA |
| pUT18_605_R1 | CTGGCGGCTGAATTCGAGCTCGGTACCCGGGGATCATTCACAAGTGCTAATTGATAAAAT |
| pUT18_615_F1 | CAGCTATGACCATGATTACGCCAAGCTTGCATGCCATGCAACAACAAGAACAGGAGCAAG |
| pUT18_615_R1 | CTGGCGGCTGAATTCGAGCTCGGTACCCGGGGATCCATTAAAATGAGCCTTTCTTTTTCA |
| pUT18_680_F | CATGATTACGCCAAGCTTGCATGCCATGGCTACTACTAAAGCAGATATCG |
| pUT18_680_R | AATTCGAGCTCGGTACCCGGGGATCATGGAACTCAATCTTGATGTCATCT |
| pKT_605_F1 | CCGATTACCTGGCGCGCACGCGGCGGGCTGCAGGGATGACAGAAGAAGCACGCGAAAAAA |
| pKT_605_R1 | AACGACGGCCGAATTCTTAGTTACTTAGGTACCCGCTAATTCACAAGTGCTAATTGATAA |
| pKT_615_F1 | CCGATTACCTGGCGCGCACGCGGCGGGCTGCAGGGATGCAACAACAAGAACAGGAGCAAG |
| pKT_615_R1 | AACGACGGCCGAATTCTTAGTTACTTAGGTACCCGTTACATTAAAATGAGCCTTTCTTTT |
| pKT_680_F1 | CCTGGCGCGCACGCGGCGGGCTGCAATGGCTACTACTAAAGCAGATATCG |
| pKT_680_R1 | GCCGAATTCTTAGTTACTTAGGTACTTAATGGAACTCAATCTTGATGTCA |
| pKNT_605_F1 | CAGCTATGACCATGATTACGCCAAGCTTGCATGCCATGACAGAAGAAGCACGCGAAAAAA |
| pKNT_605_R1 | TGATGCGATTGCTGCATGGTCATTGAATTCGAGCTATTCACAAGTGCTAATTGATAAAAT |
| pKNT_615_F1 | CAGCTATGACCATGATTACGCCAAGCTTGCATGCCATGCAACAACAAGAACAGGAGCAAG |
| pKNT_615_R1 | TGATGCGATTGCTGCATGGTCATTGAATTCGAGCTCATTAAAATGAGCCTTTCTTTTTCA |
| pKNT_615_F2 | CATGATTACGCCAAGCTTGCATGCCATGCAACAACAAGAACAGGAGCAAG |
| pKNT25-680_F | CATGATTACGCCAAGCTTGCATGCCATGGCTACTACTAAAGCAGATATCG |
| pKNT25-680_R | GCTGCATGGTCATTGAATTCGAGCTATGGAACTCAATCTTGATGTCATCT |
| pET15b_605_F1 | TGCCGCGCGGCAGCCATATGATGACAGAAGAAGCACGCG |
| pET15b_605_R1 | GCTTTGTTAGCAGCCGGATCCCTAATTCACAAGTGCTAATT |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.46845.017





