Structure of the Helicobacter pylori Cag type IV secretion system
Figures
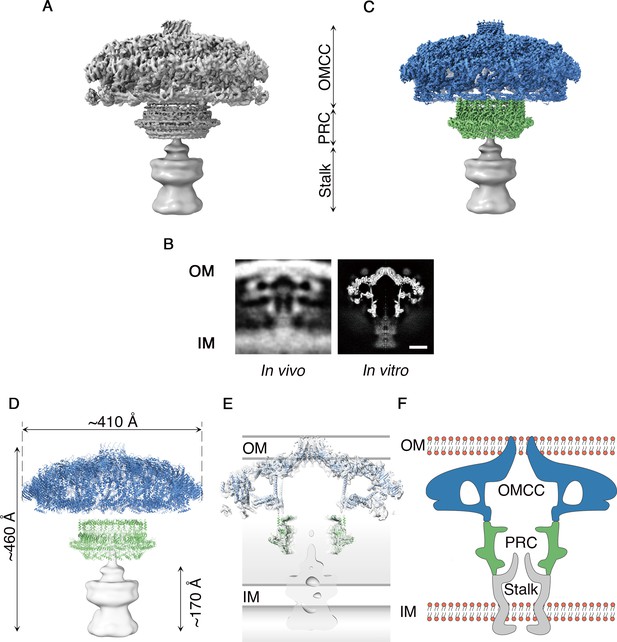
Cryo-EM structure of the H. pylori Cag T4SS.
(A) Reconstruction of the H. pylori Cag T4SS particles with no applied symmetry at 5.4 Å, showing three parts: the outer membrane core complex (OMCC), periplasmic ring complex (PRC), and Stalk density. The Stalk was not well defined in the structure and has been gaussian filtered to a resolution of 15 Å. (B) Comparison of central sections through the longitudinal plane of the T4SS 3D density determined by cryo-ET of intact H. pylori (left panel, EMD: 7474) (Chang et al., 2018) or by cryo-EM of purified particles (this study) (right panel). OM, Outer membrane, IM, inner membrane. Scale bar, 10 nm C. Combined high resolution structure of the H. pylori Cag T4SS that includes the 3.8 Å OMCC (blue) with 14-fold symmetry, the 3.5 Å PRC (green) with 17-fold symmetry, and the gaussian filtered Stalk (gray). (D) Secondary structure model of the refined OMCC (blue) and PRC (green) with gaussian filtered Stalk (gray). (E) Central axial slice view showing how refined structures of the OMCC (blue) and PRC (green) fit into the 3D map of the Cag T4SS (light gray). (F) Cartoon schematic showing the organization of the H. pylori Cag T4SS in relation to the OM and IM. OMCC (blue), PRC (Green) and Stalk (Gray).
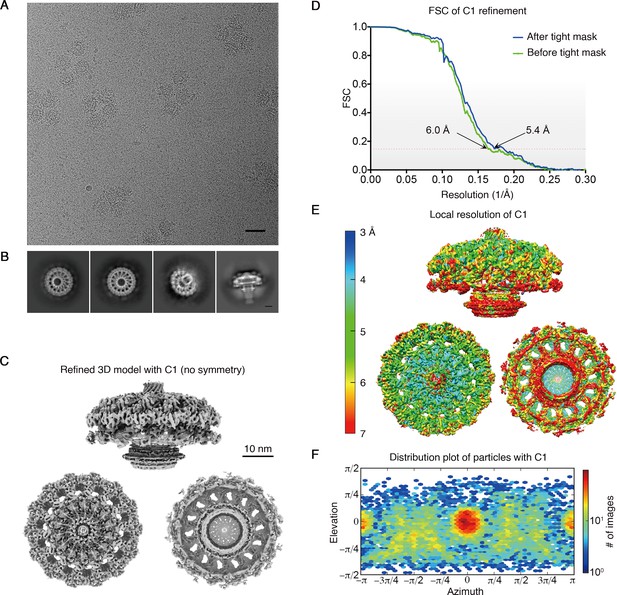
Single particle cryo-EM analysis of H. pylori Cag T4SS.
(A) Sub-area of a motion corrected image of vitrified H. pylori Cag T4SS particles. Scale bar, 50 nm. (B) Selected 2D class averages. Scale bar, 10 nm. (C) Rotated views of the 3D reconstructed model with no applied symmetry (C1, 5.4 Å resolution). Scale bar, 10 nm. (D) FSC of C1 refinement before (green line) and after (blue line) applying a tight mask in cryoSPARC. Dotted red line, Fourier Shell Correlation (FSC) = 0.143. (E) Heat map showing local resolution of C1 3D reconstruction. The heat map scales are exponential. (F) Orientation direction distribution plot of Euler angles of particles in the T4SS complex with C1 symmetry.
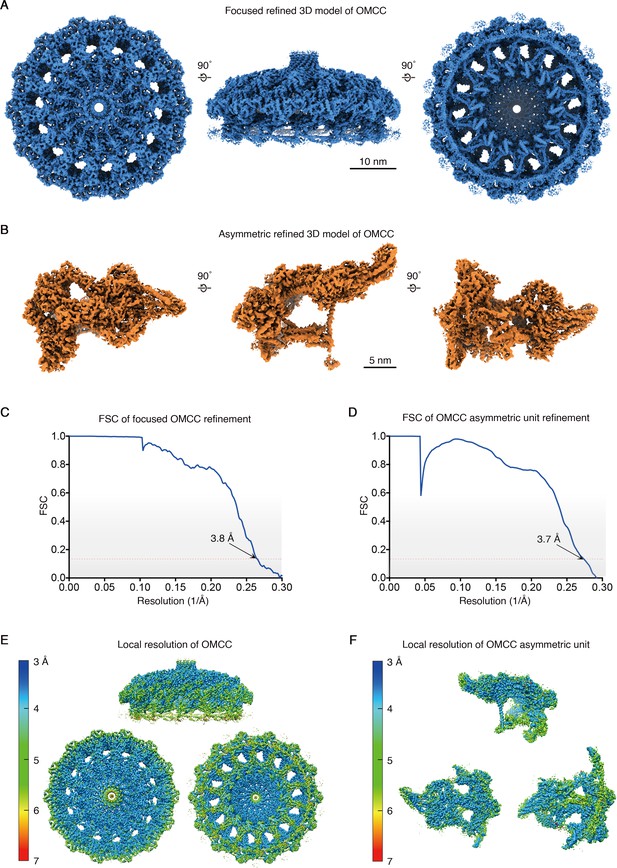
Focused and asymmetric refinement of the H. pylori Cag T4SS OMCC.
(A) Top (left), side (middle), and bottom (right) view of the focused refined 3D model of C14 OMCC (3.8 Å resolution). Scale bar, 10 nm. (B) Top (left), side (middle), and bottom (right) view of an OMCC asymmetric unit (3.7 Å resolution). Scale bar, 5 nm. (C–D) FSC of focused C14 OMCC refinement (C) and of OMCC asymmetric unit refinement (D) Dotted red line, FSC = 0.143. (E) Map of local resolution of focused OMCC refinement. (F) Map of local resolution of OMCC asymmetric unit refinement.
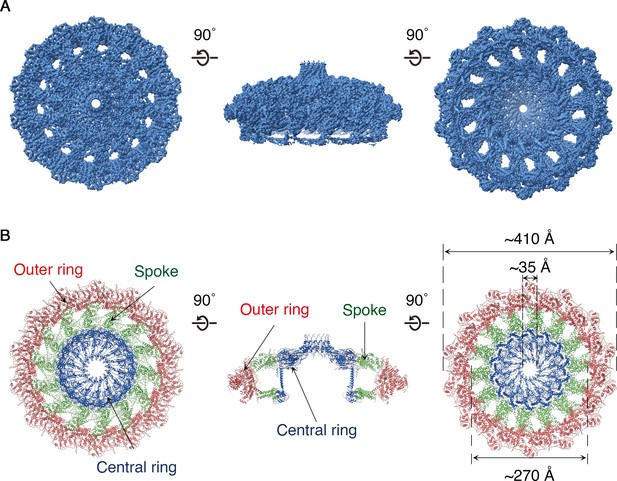
Structure of the H. pylori Cag T4SS OMCC.
(A) 3.8 Å resolution cryo-EM density map of the Cag T4SS OMCC rotated 90° around the X-axis. The structure has 14-fold symmetry. Left panel represents view of the OMCC looking in from outer membrane towards the inner membrane, and right panel represents the view looking out from the inner membrane towards the outer membrane. (B) Secondary structure model of the OMCC. In the en face view on the left (i.e. view looking in from the outer membrane), the structure has a central ring (blue) and outer ring (red) connected by 14 spokes (green). The middle panel represents a central slice of the OMCC. The right panel shows the view looking out from the inner membrane.
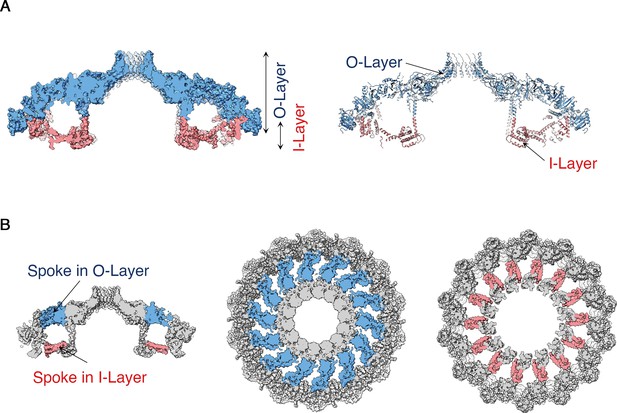
Structural features of the H. pylori Cag T4SS OMCC I-Layer and O-layer.
(A) Central axial slice view of the OMCC highlighting the position of the outer-layer (O-layer) and inner-layer (I-layer) colored in blue and red, respectively. Left panel, Surface representation; Right panel, Ribbon representation of secondary structure model. (B) Central axial slice view (left) and two cross sections of the OMCC highlighting the position of the spokes in O-layer (middle panel, blue, looking in from the outer membrane) and I-layer (right panel, red, looking out from the inner membrane).
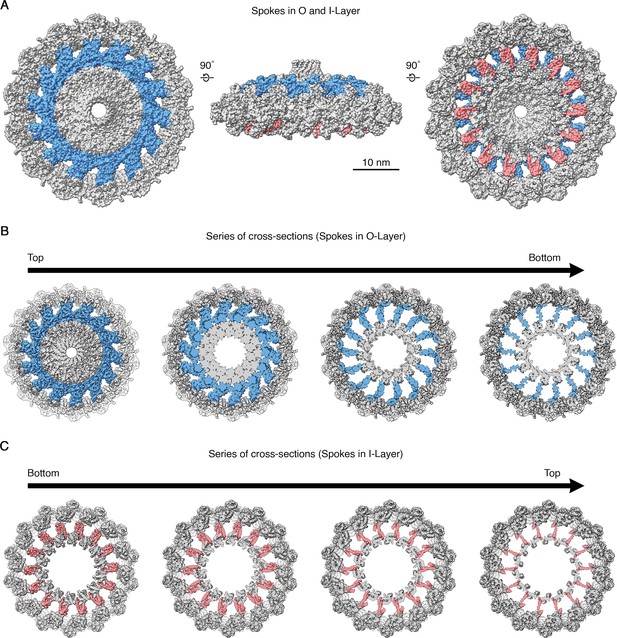
Spokes in O-layer and I-layer of OMCC.
(A) Top (left), side (middle), and bottom (right) view of the OMCC highlighting the position of spokes in O-layer (blue) and I-layer (red). (B–C) Cross-section series of OMCC in different planes highlighting the position of spokes in O-layer. (B) View from the O-layer to I-layer (i.e. view looking in from the outer membrane towards the inner membrane) and C) View from the I-layer to O-layer (i.e. view looking out from inner membrane towards the outer membrane).
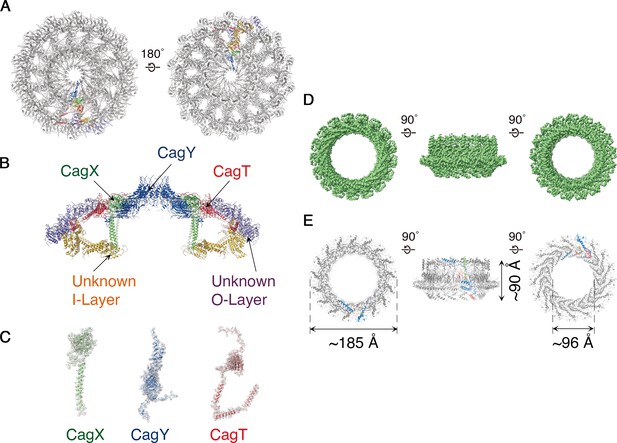
Molecular organization of the H. pylori Cag T4SS.
(A) Secondary structure model of the OMCC. The left panel represents view looking in from the outer membrane and the right panel represents the view looking out from the inner membrane. (A–C) Blue, CagY; Green, CagX; Red, CagT; Gold, I-layer; and Purple, O-layer. (B) Central axial slice view of the secondary structural model of the OMCC highlighting the position of CagY, CagX, CagT, I-layer, and O-layer. (C) Cryo-EM densities with built models of CagX, CagY, and CagT. Cryo-EM density map is in gray mesh and model is shown as ribbon diagram. (D) 3.5 Å resolution cryo-EM density map of the Cag T4SS PRC rotated 90° around the X-axis. The structure has 17-fold symmetry. (E) Secondary structure model of the PRC. Chain 1 (67 residues, green), Chain 2 (96 residues, blue) and Chain 3 (136 residues, red). In both D and E the left panel represents view looking in from outer membrane and right panel represents the view looking out from the inner membrane.
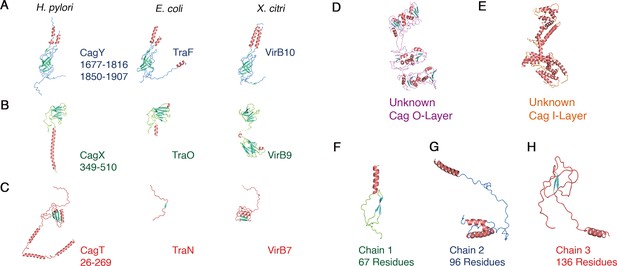
Structures of T4SS components.
(A-C) Structures of H. pylori Cag T4SS components determined based on the cryo-EM data (left panels) compared to structural homologs in E. coli (middle panels) and X. citri (right panels). (A) Ribbon representation of CagY (residues 1677–1816 and 1850–1907), E. coli TraF (PDB 3JQO), and X. citri VirB10 (PDB 6GYB). (B) Ribbon representation of CagX (residues 349–510), E. coli TraO (PDB 3JQO), and X. citri VirB9 (PDB 6GYB). (C) Ribbon representation of CagT (residues 26–269), E. coli TraN (PDB 3JQO), and X. citri VirB7 (PDB 6GYB). (D–E) Ribbon representation of regions of the O-layer and I-layer that have traceable secondary structure but could not be assigned to specific proteins. (F–H) Ribbon representation of the poly-alanine model of the three traceable protein chains in the PRC.
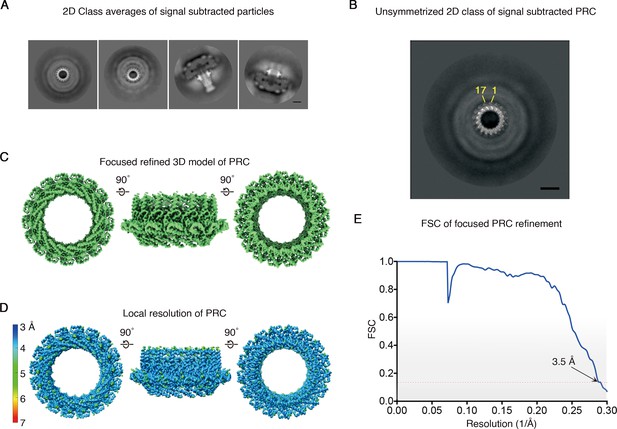
Focused refinement of the H.pylori Cag T4SS PRC.
(A) Selected 2D class average images without applied symmetry of signal subtracted particles leaving only the PRC density (i.e. OMCC density is subtracted). Scale bar, 10 nm. (B) Enlarged 2D class average shown in A (left panel, no applied symmetry) with apparent symmetry labeled. Scale bar, 10 nm. (C) Top (left), side (middle), and bottom (right) view of the focused refined 3D map of C17 PRC (3.5 Å resolution). (D) Heat map of the local resolution of the PRC with C17 applied symmetry. (E) FSC of focused refinement with C17 PRC. Dotted red line, FSC = 0.143.
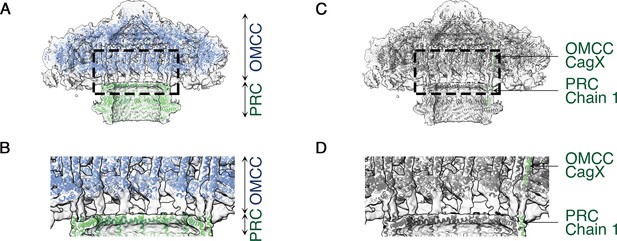
C1 symmetry density map showing connection of the OMCC to the PRC.
(A) Central axial slice view showing refined structures of the OMCC (blue) and PRC (green) fit into the C1 3D map of the Cag T4SS (light gray). The map is contoured at a low level to show less-ordered features. (B) Zoomed-in view of the region delimited by the dashed line in (a) showing the connection region of OMCC and PRC. (C) Central axial slice view showing refined structures of OMCC CagX (top, green) and PRC Chain 1 (bottom, green) fit into the C1 3D map of Cag T4SS complex (light gray). (D) Zoomed-in view of the region delimited by the dashed line in (c) showing candidates for the connection of OMCC and PRC.
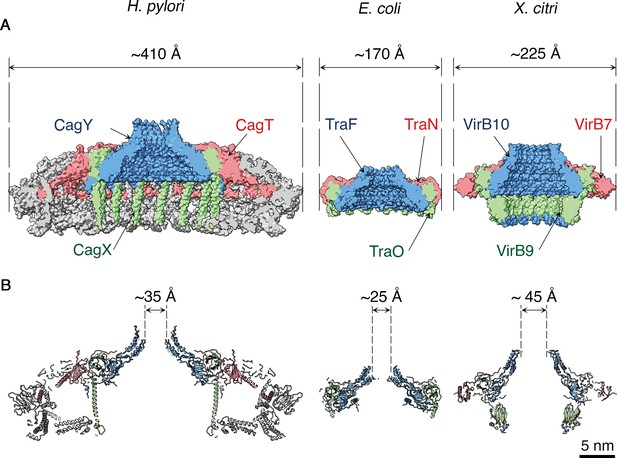
Structural comparison of H. pylori Cag T4SS OMCC components with E. coli and X. citri OMCC components.
Side views (A) and central axial slice views (B) of H. pylori Cag T4SS (left), E. coli pKM101 T4SS (middle; PDB 3JQO [Chandran et al., 2009]) and X. citri T4SS (right; PDB 6GYB [Sgro et al., 2018]). Structural homologs CagY, TraF, and VirB10 are labeled blue. Structural homologs CagX, TraO, and VirB9 are labeled green. Structural homologs CagT, TraN, and VirB7 are labeled red. Scale bar, 5 nm.
Videos
Movie showing the cryo-EM density maps of the H. pylori T4SS.
https://doi.org/10.7554/eLife.47644.006Movie showing the cryo-EM density of the H. pylori T4SS and the molecular map of the OMCC sub-complex.
https://doi.org/10.7554/eLife.47644.007Movie showing the cryo-EM density of the H. pylori T4SS and the molecular map of the PRC sub-complex.
https://doi.org/10.7554/eLife.47644.008Additional files
-
Supplementary file 1
Flow chart of cryo-EM processing steps.
~25,000 Cag T4SS particles were manually picked in RELION and analyzed by two different image processing software packages (CryoSPARC and RELION). The processing steps done in CryoSPARC are on a gray background and the processing steps done using RELION are on a tan background. The particles were exported into cryoSPARC for 2D alignment and ab initio 3D classification. The best class was chosen for further refinement, without symmetry (C1). The 5.4 Å 3D model with no symmetry applied was filtered to 60 Å resolution and used as an initial model for 3D structure determination in RELION using 3D refinement without applied symmetry. Focused 3D classification (without alignment) was used to determine higher resolution maps of the OMCC (with 14-fold symmetry) and the PRC (with 17-fold symmetry). The maps of the OMCC and PRC were further refined using focused 3D refinement (with local refinement), resulting in 3D maps of the OMCC (14-fold symmetry) and the PRC (17-fold symmetry) at 3.8 Å and 3.5 Å, respectively. To improve the resolution of the OMCC, symmetry expansion was applied, resulting in the 3D reconstruction at 3.7 Å resolution.
- https://doi.org/10.7554/eLife.47644.017
-
Supplementary file 2
Summary of Data collection, model refinement, and model validation.
(A) Information about the cryo-EM data collection and data sets. (B) Information about model and model validation.
- https://doi.org/10.7554/eLife.47644.018
-
Supplementary file 3
FSCs of the half maps against the refined models agree with each other, suggesting that the models are not over-refined.
A-F. FSC of the half map against the refined model of CagT (A), CagX (B), CagY (C), O-layer (D), I-layer (E), and PRC (F).
- https://doi.org/10.7554/eLife.47644.019
-
Transparent reporting form
- https://doi.org/10.7554/eLife.47644.020





