An elusive electron shuttle from a facultative anaerobe
Figures
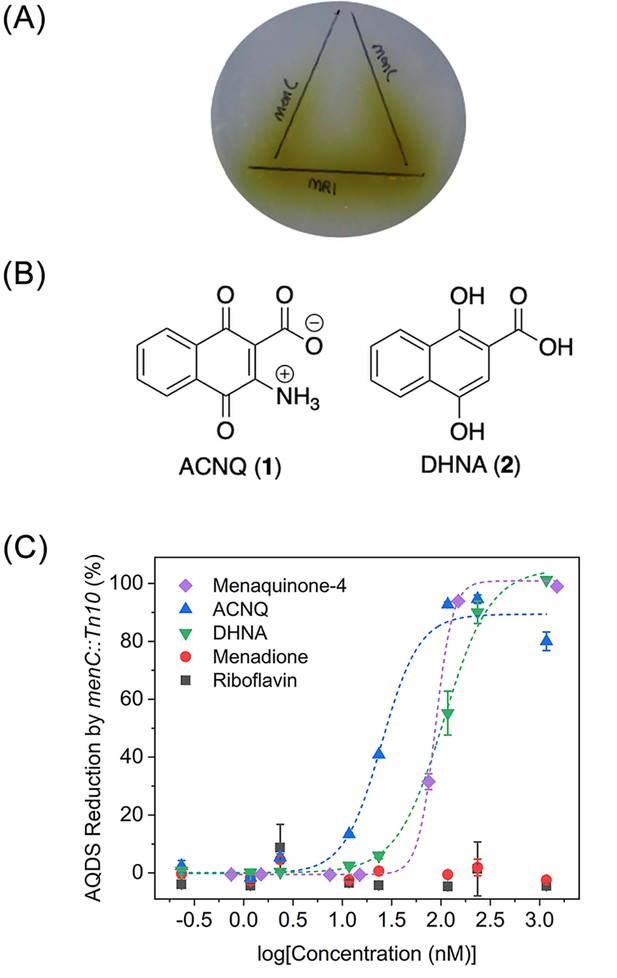
Functional analysis and structures of electron shuttles.
(A) Wild type S. oneidensis MR-1 was inoculated on the horizontal line and menC mutant was inoculated on the vertical lines (x2). Reduction of AQDS (colorless) to AHQDS (yellow) is only observed by menC mutant where it was growing in close proximity to MR-1. (B) Structures of ACNQ and DHNA. (C) Dose-response of ACNQ, DHNA, menaquinone-4 (MK4), menadione, riboflavin to recover reduction of AQDS by menC mutant. Fitting with ‘DoseResp’ function by OriginPro 2019 (OriginLab Corporation).
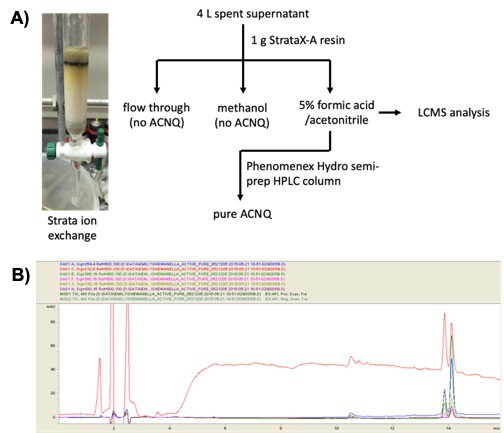
Purification scheme of ACNQ from spent supernatant.
(A) Optimized workflow for the purification of ACNQ. Cultures were grown for 72 hr under aerobic conditions at 30°C, 0 rpm. The cells were pelleted by centrifugation (7,000 rpm, 30 min) and the supernatant passed through a 0.22 μm filter. The pH of the sterile supernatant was adjusted to 6.5 and then was passed through 1 g of StrataX-A resin. ACNQ was eluted from the column with 15 mL of 5% formic acid/acetonitrile. This material was then processed on Phenomenex Hydro semi-prep HPLC column where pure ACNQ eluted around 33 min. (B) HPLC UV/Vis chromatogram of the reduced complex fraction used for analysis HR-q-TOF-MS. Three main metabolites can be visualized with UV/Vis - ~ 10.5, 13.9, and 14.1 min. ACNQ was the third metabolite.
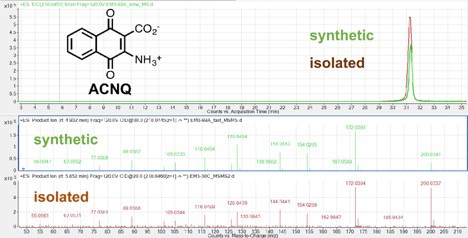
Comparison of positive mode fragmentation pattern of ACNQ isolated from MR-1 spent supernatant (bottom) and synthesized material (top).
All fragmentation matched and both metabolites eluted at the same time thus confirming the structure of the natural product.

Complete deuterium exchange on the LCMS, using mixtures of D2O/acetonitrile as the mobile phase, revealed that the active component (ACNQ) has three exchangeable protons.
(A) EIC of ACNQ in D2O; (A1) Observed ion for ACNQ at [M + D]+ 222.0702; (B) EIC of ACNQ in H2O;( B1) Observed ion for ACNQ at [M + H]+ 218.0450 indicated a mass difference of 3 Da, thus three exchangeable protons.
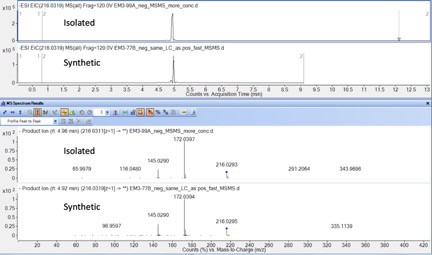
Comparison of negative mode fragmentation pattern of ACNQ isolated from MR-1 spent supernatant (top) and synthesized material (bottom).
All fragmentation matched and both metabolites eluted at the same time thus confirming the structure of the natural product.
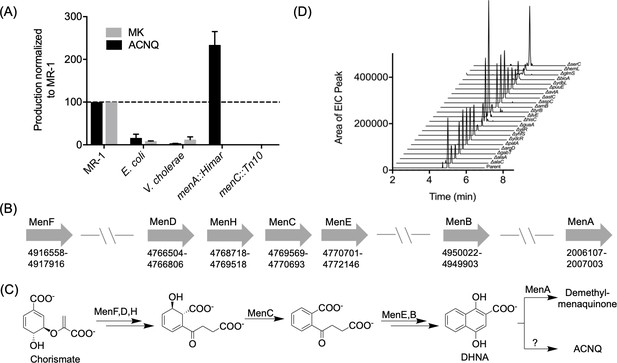
Production of and biosynthetic pathway for ACNQ.
(A) Relative production levels of ACNQ and menaquinones by S. oneidensis MR-1 (MR1), E. coli, V. cholerae, S. oneidensis menA, and menC mutants. The ability of E. coli to produce ACNQ is in contrast to the original report by Newman and Kolter, however E. coli produces significantly lower quantities of ACNQ than S. oneidensis, which is likely below the detection limit of the technique used in the previous report. (B) Genes responsible for the biosynthesis of menaquinone and their location on the chromosome. (C) Biosynthesis scheme of dimethyl-menaquinone and ACNQ. (D) Production of ACNQ by all E. coli aminotransferase/transaminases knockout mutants.
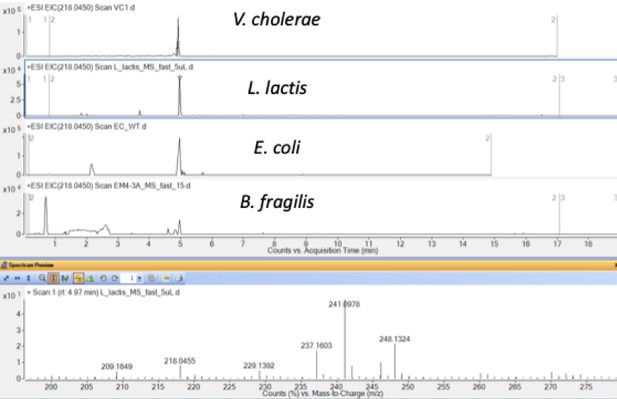
Production of ACNQ by V. cholerae V52, Lactococcus lactis lactis (ATCC 25285), E. coli K12, and Bacteroides fragilis (DSM 20481).
https://doi.org/10.7554/eLife.48054.009
Various bacterial strains that can recover reduction of AQDS by S. oneidensis menC::Tn10.
Each of the different bacterial strains was inoculated on the horizontal line and menC::Tn10 was inoculated on the vertical lines (x2). Reduction of AQDS (colorless) to AHQDS (yellow) is only observed by menC::Tn10 where it was growing in close proximity to any of the other bacterial strains. This suggests ACNQ is being produced by each of the bacterial strains except menC::Tn10.
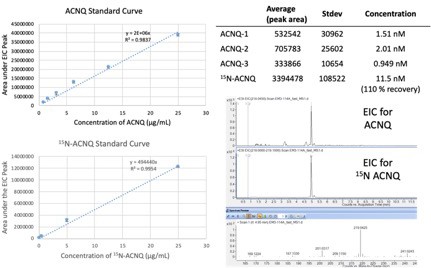
Absolute quantification of ACNQ in MR-1 culture using 15N-ACNQ as an internal standard to determine recovery yields.
Standard curves were generated using 4-point calibration curves with biological triplicates.
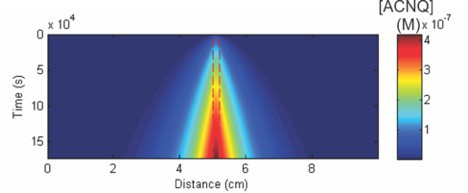
Simulation of ACNQ production and diffusion in an agarose gel.
Calculated local concentration of ACNQ (shown in colormap) as a function of time and distance from a population of Shewanella oneidensis MR-1 that produces ACNQ. Since the diffusion of ACNQ is quite slow relative its production, the local concentration of ACNQ becomes high, approaching ~0.4 μM in 2 days. The details of this calculation are described in the Materialand method Section.
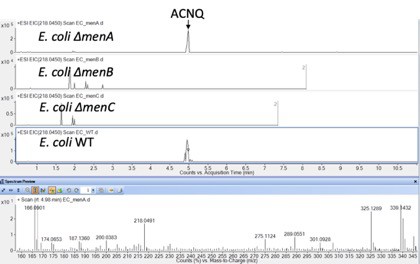
Production of ACNQ by various E. coli K12 men mutants.
Production was only observed in the WT and ΔmenA mutant, consistent with the MR-1 data.
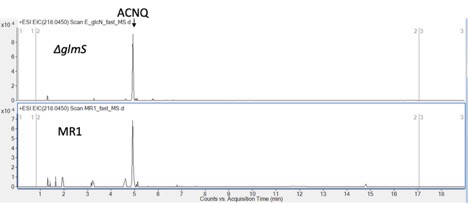
Production of ACNQ by S. oneidensis ΔglmS.
Production level in ΔglmS mutant is consistent with WT when grown under the same condition (GlcNAc supplement).
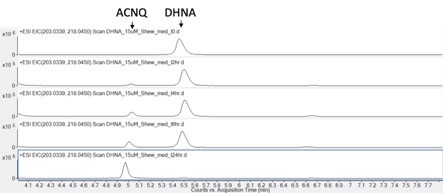
DHNA conversion to ACNQ in the same minimal media that was used to culture MR-1.
No DHNA was observed after 24 hr of incubation at room temperature under aerobic conditions.

DHNA conversion to ACNQ in ammonium phosphate, pH 8.
No DHNA was observed after 24 hr of incubation at room temperature under aerobic conditions.

DHNA conversion to ACNQ in minimal media with amino acids as the only nitrogen source.
No DHNA was observed after 6 hr of incubation.
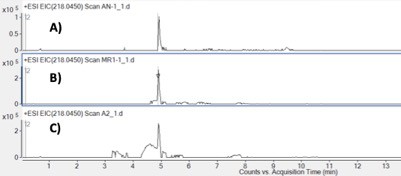
Production of ACNQ when grown with (A) NH4SO4, (B) NH4SO4 and amino acid mixture and (C) amino acid mixture as the sole nitrogen source.
ACNQ production was observed under each growth condition. [Amino acid mixture (10x mixture): L-arginine: 1.26 g/L, L-cystine: 0.24 g/L, L-histidine HCl: 0.42 g/L, L-isoleucine: 0.52 g/L, L-leucine: 0.52 g/L, L-lysine HCl: 0.73 g/L, L-methionine: 0.15 g/L, L-phenylalanine: 0.33 g/L, L-threonine: 0.48 g/L, L-tryptophan: 0.1 g/L, L-tyrosine: 0.36 g/L, L-valine: 0.467 g/L].
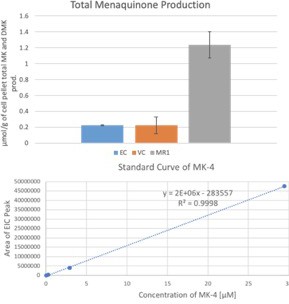
Absolution quantification of menaquinone (MK) in MR-1, V. cholerae V52, and E. coli K12 cultures.
Menaquinone-4 was used to generate the standard curve (4-point). The error bars in the top graph represent variance in biological replicates. (EC – E. coli K12, VC – V. cholerae V52, MR1 – S. oneidensis MR-1).
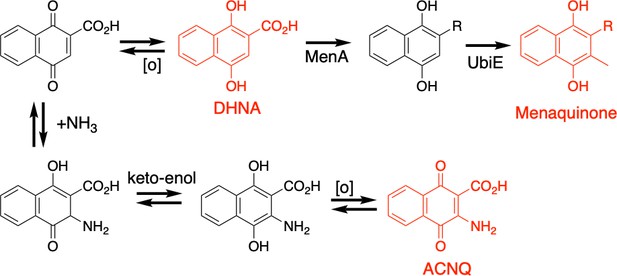
Proposed mechanism for the non-enzymatic conversion of DHNA to ACNQ.
Initiated by oxidation of DHNA then conjugate addition of ammonium at the C3 position followed by enol formation and subsequent oxidation of the hydroquinone.
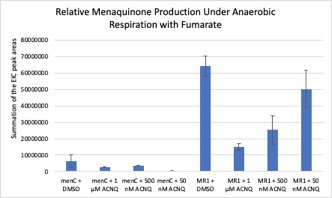
Menaquinone analysis of S. oneidensis menC::Tn10 (menC) and S. oneidensis MR-1 (MR-1) cell pellets when grown with 0.05, 0.5, and 1 μM of ACNQ supplemented to the media under anaerobic conditions with fumarate as the sole TEA.
No evidence of ACNQ being converted to menaquinone or being prenylated to form an amino-modified menaquinone was observed.
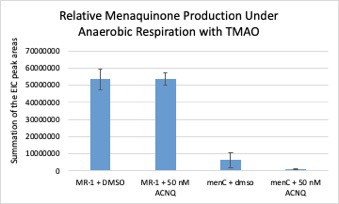
Menaquinone analysis of S. oneidensis menC::Tn10 (menC) and S. oneidensis MR-1 (MR-1) cell pellets when grown with 0.05 μM of ACNQ supplemented to the media under anaerobic conditions with TMAO as the sole TEA.
No evidence of ACNQ being converted to menaquinone or being prenylated to form an amino-modified menaquinone was observed.
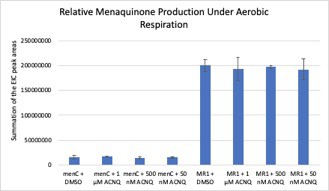
Menaquinone analysis of S. oneidensis menC::Tn10 (menC) and S. oneidensis MR-1 (MR-1) cell pellets when grown with 0.05, 0.5, and 1 μM of ACNQ supplemented to the media under aerobic conditions.
No evidence of ACNQ being converted to menaquinone or being prenylated to form an amino-modified menaquinone was observed.
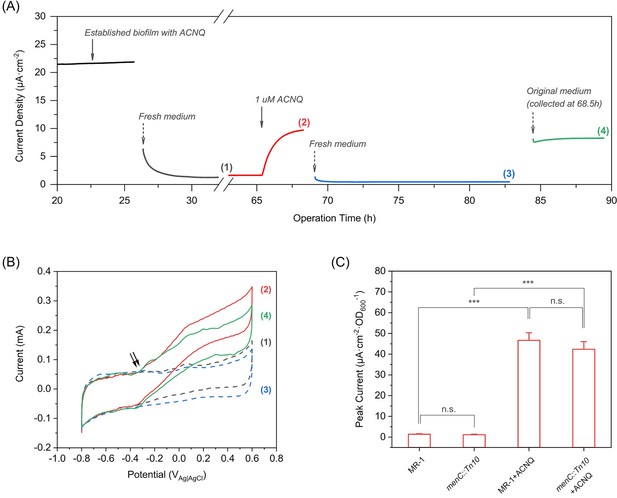
ACNQ acts as an extracellular electron transfer mediator for MR-1 in a bioelectrochemical system.
(A) The current production of an established MR-1 biofilm over time in the presence of different media. Oxidation current drops after the introduction of fresh medium (gray trace), but recovers immediately after the addition of 1 μM ACNQ (red trace). The second replacement of fresh media (blue trace) again causes the current to drop, and the introduction of original ACNQ-containing media after filtration (green trace) leads to high oxidation current. Numbered gaps indicate CV analysis. (B) Turnover CV scan of the MR-1 biofilm with different media where the color and number indicate when the analysis was performed in(A). The double arrow highlights the initiation of the catalytic wave. (C) Peak current (steady state current, collected after at least 12 hr operation) from chronoamperometry experiments (electrode bias at −0.1 VAg/AgCl) with wild type S. oneidensis (MR-1) and menC mutant (menC::Tn10), with or without 1 μM ACNQ. ***p<0.001, ANOVA with Tukey Test by OriginPro 2019 (OriginLab Corporation), n = 3.
-
Figure 3—source data 1
Source Data for Figure 3C.
- https://doi.org/10.7554/eLife.48054.028
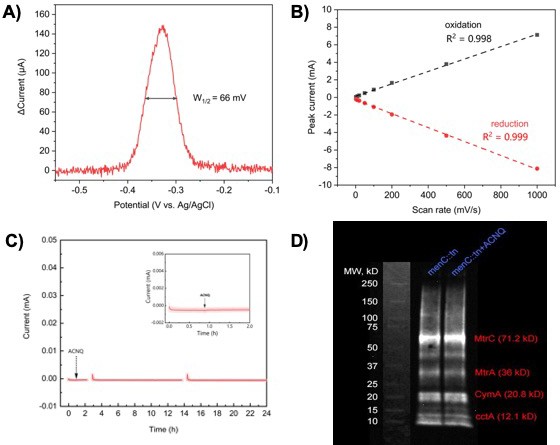
ACNQ acts as an electron shuttle and does not affect cytochrome c protein expression.
(A) Differential pulse voltammetry (DPV) of ACNQ in M9 media. (B) The relationship between ACNQ redox peak current and CV scan rate from non-turnover MR-1 biofilm. (C) Chronoamperometry (−0.1 VAg/AgCl) of clean carbon felt electrode in M9 media, 1 μM ACNQ was added after the oxidation current is stable. (D) Enhanced chemiluminescence (ECL) analysis of menC::Tn10 and menC::Tn10 with ACNQ, key cytochrome c proteins involved in extracellular electron transfer are noted in red.
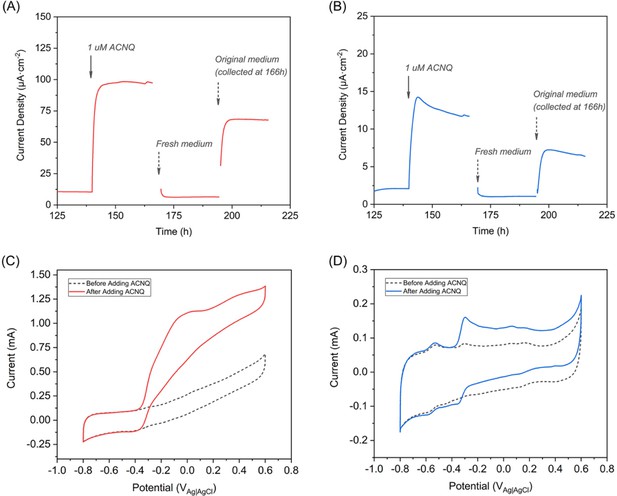
ACNQ acts as an extracellular electron transfer mediator for S. oneidensis Δbfe and Δmtr.
(A) Chronoamperometry of S. oneidensis Δbfe. (B) Chronoamperometry of S. oneidensis Δmtr. (C) CV scan of the S. oneidensis Δbfe. (D) CV scan of the S. oneidensis Δmtr.
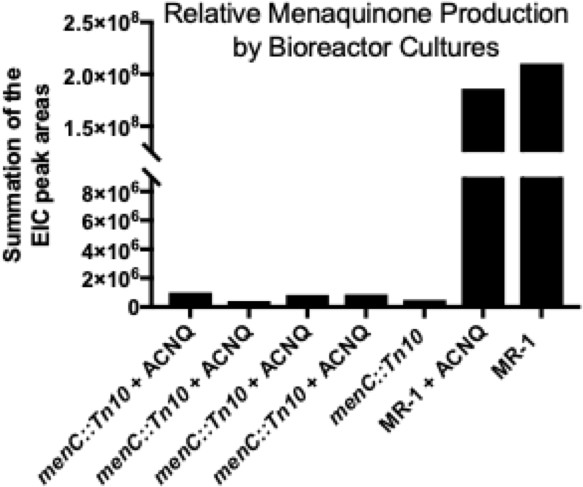
Menaquinone analysis of S. oneidensis menC::Tn10 (menC) and S. oneidensis MR-1 (MR-1) cell pellets when grown with 0.05, 0.5, and 1 μM of ACNQ supplemented to the media under aerobic conditions.
No evidence of ACNQ being converted to menaquinone or being prenylated to form an amino-modified menaquinone was observed.
Additional files
-
Supplementary file 1
(A) Acquisition of non-E. coli CGCS bacterial strains. (B) E. coli strains purchased from Coli Genetic Stock Center.
- https://doi.org/10.7554/eLife.48054.029
-
Supplementary file 2
Statistical report for Figure 3C.
- https://doi.org/10.7554/eLife.48054.030
-
Transparent reporting form
- https://doi.org/10.7554/eLife.48054.031





