B cells extract antigens at Arp2/3-generated actin foci interspersed with linear filaments
Figures
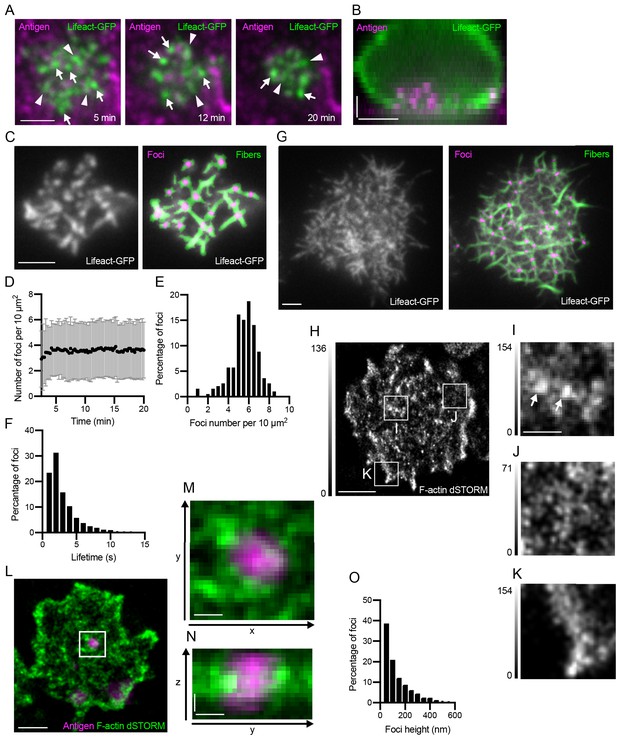
Actin structure of the B cell synapse.
(A) TIRF images of a Lifeact-GFP (green) expressing primary mouse B cell making a synapse with a PMS loaded with the surrogate antigen anti-Igκ F(ab’)2 (magenta), at indicated times from the initial contact. Examples of actin foci and filaments are marked by white arrows and arrowheads, respectively. Scalebar 2 μm. (B) Sideview of a Lifeact-GFP (green) expressing B cell after 20 min of contact with a PMS loaded with anti-Igκ F(ab’)2 (magenta). Scalebars 2 μm. (C) Example of actin foci and fiber segmentation in primary mouse B cells prepared as in (A). Left panel shows raw TIRF image of Lifeact-GFP, right panel shows segmented foci (magenta) and fibers (green) overlaid on the original image. Scalebar, 2 μm. (D) Number of actin foci quantified from primary B cells incubated as in (A) at the indicated times after synapse formation. Data points are mean ± SD from 310 cells in one representative experiment. (E) Distribution of the numbers of actin foci in 193 cells from a representative experiment at 20 min after synapse formation. (F) Distribution of the lifetime of actin foci in B cells from the experiment in (E). (G) Actin structure and segmentation of foci and fibers in a Ramos B cell interacting with a PMS loaded with anti-IgM. Scalebar, 2 μm. (H) dSTORM reconstruction of F-actin stained with phalloidin-AlexaFluor647 at the focal plane of a synapse using TIRF illumination of a primary mouse B cell interacting with a PMS loaded with anti-Igκ F(ab’)2. Scale bar, 2 μm. (I, J, K) Magnified areas show actin foci (arrows), filament meshwork, and lamellipodia, respectively. Gray intensity scales are different from H and are shown on the left of each image. Scale bar, 0.5 μm. (L) 3D dSTORM reconstruction of F-actin, stained with phalloidin-AlexaFluor647 (green) overlaid onto epifluorescence images of PMS loaded with anti-Igκ F(ab’)2 (antigen, magenta). Image shows a section containing actin 250–300 nm above the PMS and the corresponding epifluorescence section. Scale bar 1 μm. (M) Magnified area containing an antigen cluster lifted from the PMS, surrounded by actin foci. (N) Vertical section through this area shown as maximum intensity projection along x. Scale bars in (M), (N) 0.2 μm. (O). Histogram of foci size in the z-direction (height). Data are from 1369 foci in 58 cells from one representative experiment.
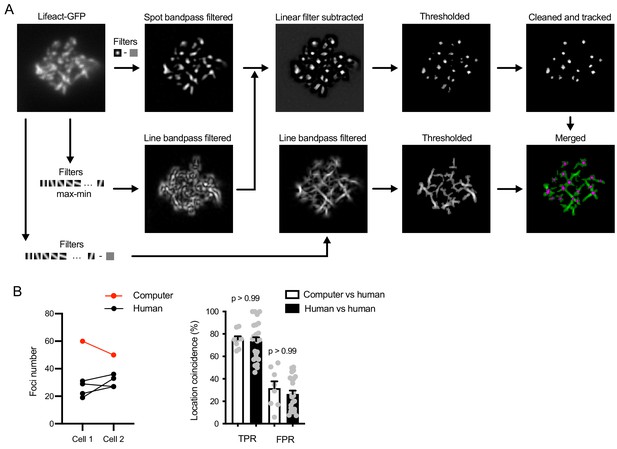
Actin foci and fiber segmentation.
(A) Workflow of foci and fiber segmentation. To segment foci, Lifeact-GFP image was first bandpass-filtered using a Gaussian spot matching the apparent size of the foci (about 300 nm in diameter). In parallel, the image is filtered with linear elements of the apparent width of the actin filaments (about 250 nm) and oriented in all directions using 18-degree rotation. The difference between maximum and minimum filter values is used to suppress linear structures in the spot-filtered image. Foci segmentation mask is further thresholded using a value derived from the mean Lifeact-GFP intensity and the resulting mask is cleaned up by removing small scattered spots, spots that are highly elongated and spots that do not persist for at least two consecutive frames. To segment fibers, Lifeact-GFP image is bandpass-filtered using the linear filaments and taking the maximum value. The filtered image is thresholded again using a value derived from the mean Lifeact-GFP intensity. The segmentation mask is zeroed in pixels already segmented as foci and combined with the foci mask for two-color display. (B) Comparison of the computer-produced automated foci segmentation to manual segmentation by humans. Foci in two sample cell timelapses were segmented by a computer using the above algorithm or by four experienced researchers. Left, comparison of foci numbers detected in each cell. Right, comparison of foci positions. Positions of the foci detected by the computer were compared to those segmented by humans with 5-pixel and 1-frame tolerance. True positive rates (TPR) of the computerized segmentation were calculated for each computer-human pair as the percent of the human-detected foci that were also detected by the computer. False positive rates (FPR) were calculated as the percent of foci detected by the computer and not detected by the human. As a comparison, segmentation data obtained by the human researchers were compared to each other using the same calculations. P, significance value in Kruskal-Wallis multiple comparison test comparing the indicated columns.
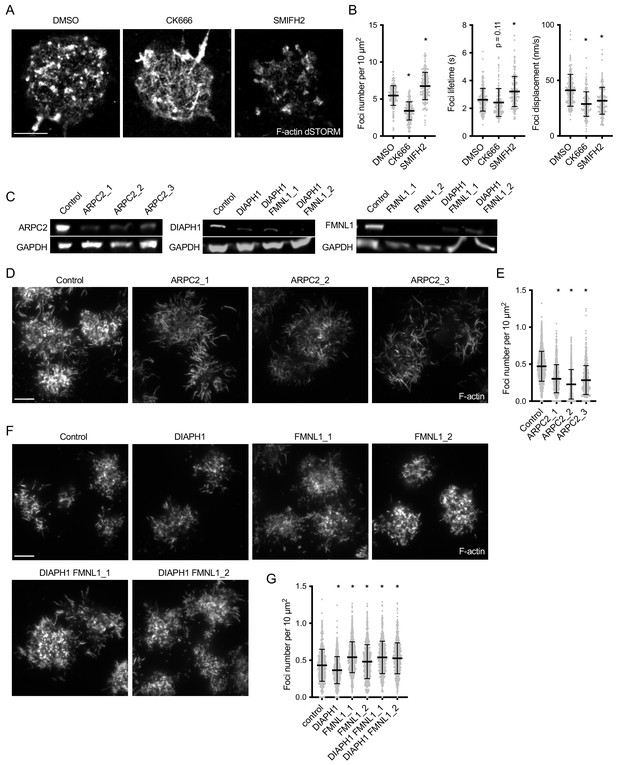
Arp2/3 is responsible for actin foci generation, while formins produce the interspersed actin filaments.
(A) dSTORM reconstruction of F-actin stained with phalloidin-AlexaFluor647 in synapses of primary mouse B cells interacting for 20 min with PMS loaded with anti-Igκ F(ab’)2. The cells were treated with the indicated inhibitors from 5 min after initial spreading. Scale bar, 2 μm. (B) Quantification of foci characteristics obtained from TIRF timelapses of live cells treated with inhibitors as in (A). Plots show foci numbers, mean lifetimes, and mean displacements per cell. (C) Immunoblot of lysates of Cas9 Ramos cells transfected with the indicated gRNAs, developed with anti-ARPC2 and anti-GAPDH antibodies. (D, F) TIRF images of F-actin stained with phalloidin-AlexaFluor647 in synapses of a Cas9-expressing Ramos B cells transduced with the indicated gRNAs. Cells were imaged after interacting for 20 min with PMS loaded with anti-IgM F(ab’)2. Scale bars, 5 μm. (E, G) Quantification of Ramos actin foci numbers per cell area. Data in B, E, G show values from individual cells pooled from two (B, G) or three (E) experiments. Bars indicate means and SDs. P, significance in one-way ANOVA compared to DMSO or control, *, p<0.0001.
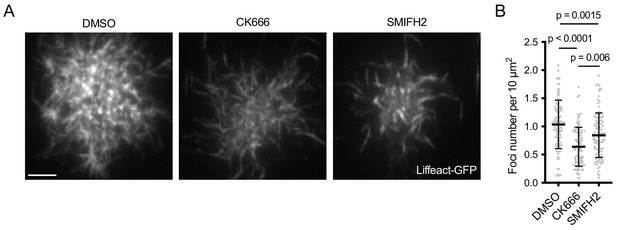
Actin architecture of Ramos B cell synapses after Arp2/3 and formin inhibition.
(A) F-actin imaged by TIRF microscopy in Ramos B cells expressing Lifeact-GFP 20 min after forming synapses with PMSs loaded with anti-IgM F(ab’)2. Cells were treated with either DMSO or the indicated inhibitors 5 min after their initial spreading on the PMS. See also Video 5. (B) Quantification of foci numbers in each cell normalized per cell area. Data show values from individual cells pooled from three experiments. Bars indicate means and SDs. P, significance in one-way ANOVA compared to DMSO.
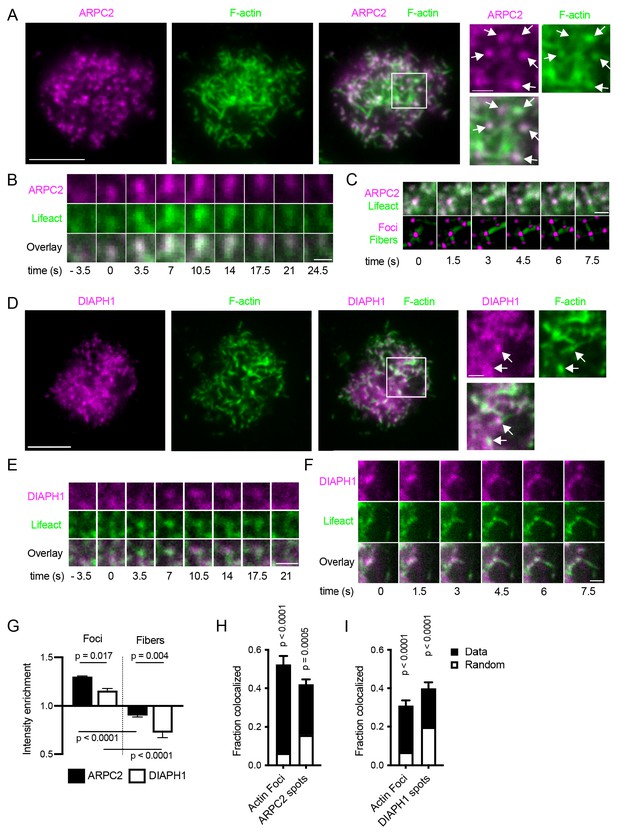
Localization and dynamics of ARPC2 and DIAPH1 in synapses of Ramos cells.
(A) Ramos cells expressing ARPC2-mRuby (magenta) were imaged by TIRF microscopy on PMSs loaded with anti-IgM F(ab’)2. F-actin was stained with phalloidin-AlexaFluor647 (green). Scale bar, 5 μm. Panels on the right show magnified area in the white box. Arrows show ARPC2 clusters colocalized with actin foci. Scale bar 1 μm. (B) Example of dynamics of ARPC2-mRuby in a single actin focus visualized with Lifeact-GFP. Time zero corresponds to initial focus formation. Scalebar 1 μm. (C) Example of a dynamic filament growth from ARPC2-positive actin foci in Ramos cells co-expressing ARPC2-mRuby and Lifeact-GFP. Bottom panel shows results of actin and fiber segmentation. Scalebar 1 μm. (D) Ramos cells expressing DIAPH1-mRuby (magenta) were imaged as in (A). Scale bar, 5 μm. Panels on the right show magnified area in white box. Arrows show clusters of DIAPH1 colocalized with actin foci. Scale bars 1 μm. (E) Example of dynamics of DIAPH1-mRuby in a single actin focus visualized with Lifeact-GFP. Time zero corresponds to initial focus formation. Scalebar 1 μm. (F) Example of a fiber outgrow from a DIAPH1 cluster in Video 7. Scalebar 1 μm. (G) Quantification of relative enrichment or depletion of ARPC2-mRuby and DIAPH1-mRuby fluorescence in actin foci and filaments. Data are mean ± SEM from n = 4 experiments each containing 12–213 cells. P, significance from one-way ANOVA. (H, I) Fraction of actin foci colocalized with ARPC2 (G) or DIAPH1 (H) spots and vice versa. Total bar heights show rates of colocalization, white bars indicate colocalization obtained after randomization of locations of the corresponding structures. Values are mean ± SEM of n = 4 of the same experiments as in F. P, significance in two-way repeated measures ANOVA comparing the measured with the randomized data.
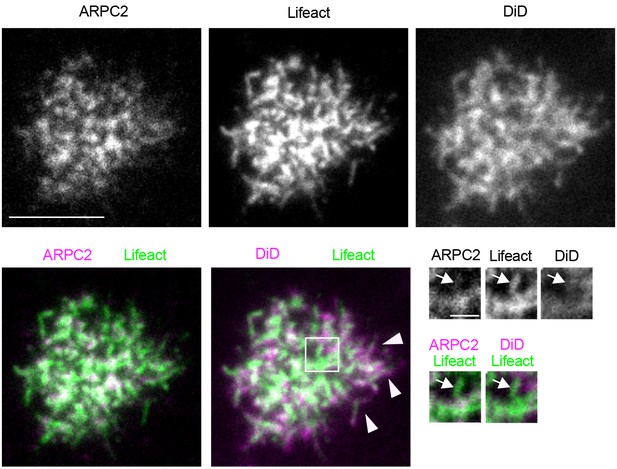
Actin fibers within the B cell synapse are not associated with membrane structures.
TIRF images of a Ramos cell expressing ARPC2-mRuby and Lifeact-GFP, and labeled with the plasma membrane dye DiD on PMS loaded with anti-IgM F(ab’)2. Magnified inset shows an ARPC2-negative actin fiber associated with an ARPC2-positive actin focus in the absence of DiD membrane signal (arrow), in contrast to DiD-labeled filopodia at the cell periphery (arrowheads). Scalebar 5 μm (1 μm in inset). Contrast was enhanced in inset images.
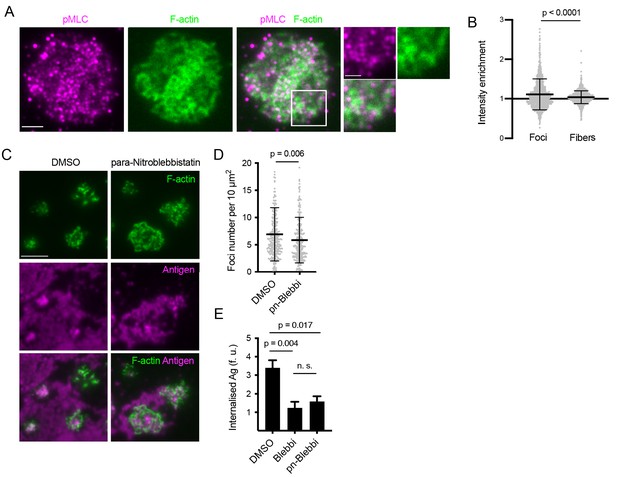
Myosin IIa resides outside of actin foci and its activity is not required for foci formation.
(A) Images of a primary mouse B cell interacting with anti-Igκ F(ab’)2- loaded PMS, stained for phosphorylated myosin light chain (pMLC, magenta) and for F-actin (green). Scalebar 2 μm. Panels on the right show magnified region. Scalebar 1 μm. (B) Quantification of pMLC fluorescence intensity shows little enrichment in foci or fibers detected in F-actin images. Data points show values from 272 and 278 individual cells in one representative experiment. Bars indicate means and SDs. P, significance in t-test. (C) Images of Lifeact-GFP (green) expressing B cells interacting with PMSs loaded with anti-Igκ F(ab’)2 (magenta) and treated after spreading with para-nitroblebbistatin. Scalebar 5 μm. (D) Quantification of foci numbers in each cell. Data show values from individual cells pooled from two experiments. Bars indicate means and SDs. P, significance in t-test. (E) Control shows that para-nitroblebbistatin has a similar inhibitory activity on extraction of anti-Igκ F(ab’)2from PMSs as blebbistatin. Data show means ± SEM of n = 120–360 cells in a representative experiment. P, significance in non-parametric Kruskal-Wallis test with Dunn’s correction.
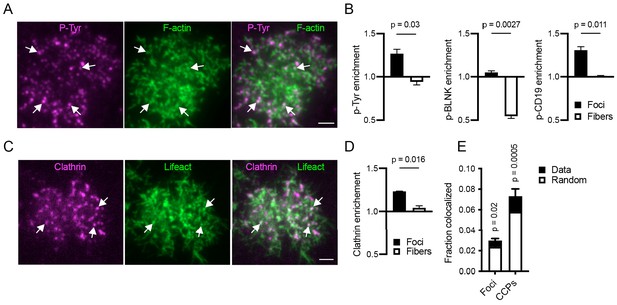
Relationship of actin structures to BCR signaling and endocytic complexes.
(A) TIRF image of a Ramos cell interacting with anti-IgM F(ab’)2-loaded PMS and stained for F-actin (green) and phosphotyrosine (magenta). Arrows show colocalization of phosphotyrosine signal with actin foci. Scale bar, 2 μm. (B) Quantification of enrichment of the indicated stains in actin foci or fibers. Data are mean ± SEM from n = 4 (phosphotyrosine) or n = 3 (phospho-BLNK and phospho-CD19) experiments. P, significance in paired t-tests. (C) TIRF image of a Ramos cell interacting with anti-IgM F(ab’)2-loaded PMS and expressing Lifeact-GFP (green) and CLTA-mCherry (magenta). Arrows show colocalization of CCPs with actin foci. Scale bar, 2 μm. (D) Quantification of enrichment of clathrin in actin foci and fibers. Mean ± SEM from n = 4 experiments. P, significance in paired t-test. (E) Fraction of actin foci colocalized with CCPs and vice versa. Total bar heights show rates of colocalization, the white bars colocalization obtained after randomization of locations of the corresponding structures. Values are mean ± SEM of n = 4 of the same experiments as in D. P, significance in two-way repeated measures ANOVA comparing the measured with the randomized data.
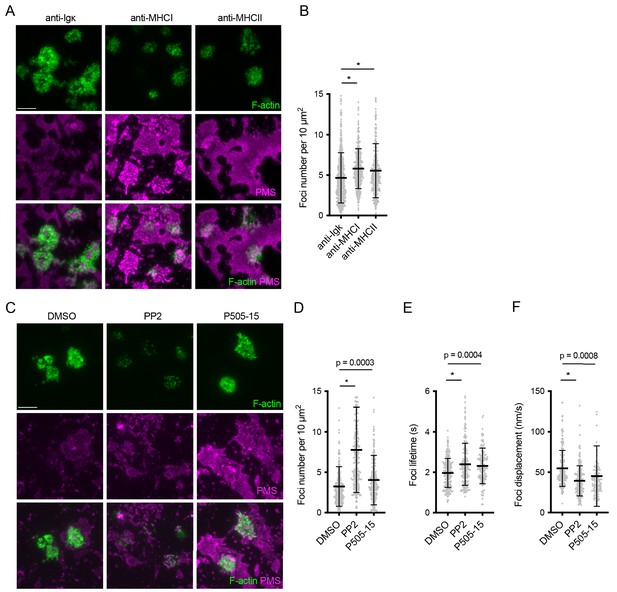
Regulation of actin architecture in primary mouse B cells by BCR signaling.
(A) Images of Lifeact-GFP (green) expressing B cells interacting with PMSs (magenta) loaded with the indicated antibodies. (B) Quantification of foci numbers in each cell normalized per cell area. (C) Images of Lifeact-GFP (green) expressing B cells interacting with PMSs loaded with anti-Igκ F(ab’)2 (magenta) and treated after spreading with the indicated inhibitors. (D, E, F) Quantification of the number of actin foci, their lifetime and their mean displacement as a measure of mobility. Data in (B, D, E, F) show values from individual cells pooled from two experiments. Bars indicate means and SDs. P, significance in one-way ANOVA compared to DMSO. *, p<0.0001 compared to anti-Igκ F(ab’)2 or DMSO.
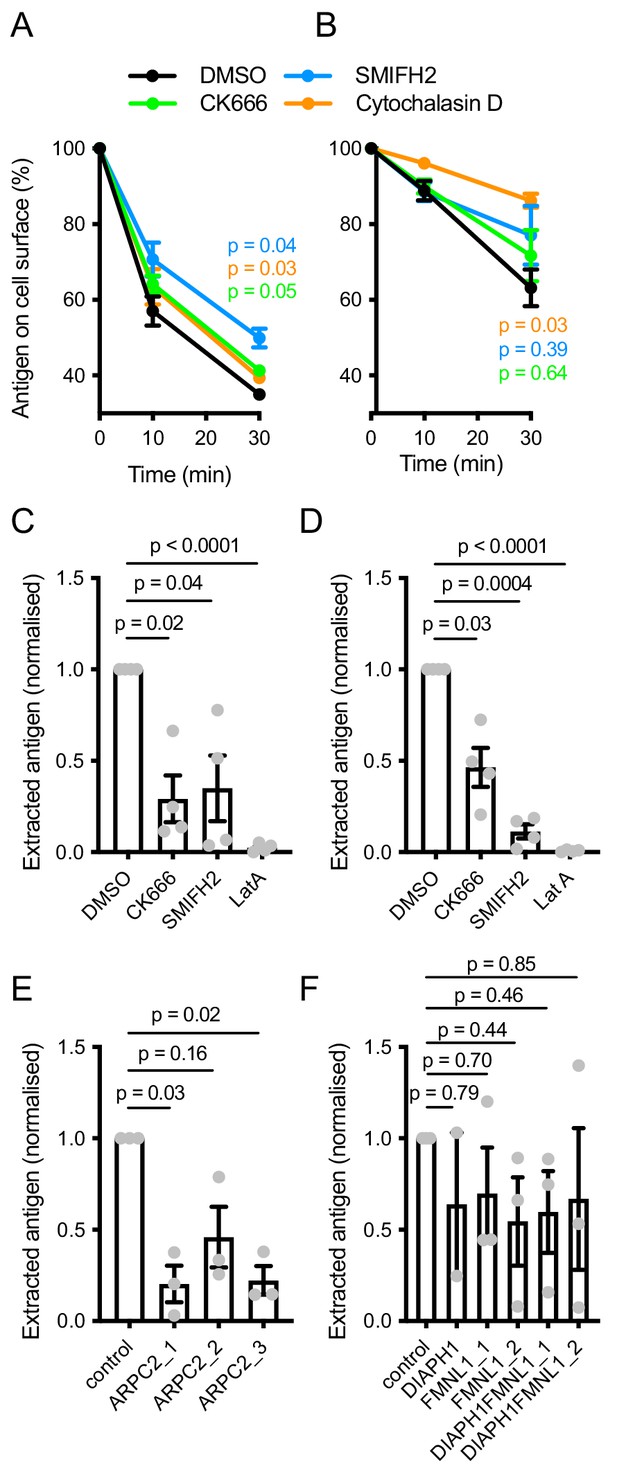
Importance of actin polymerization pathways in membrane antigen extraction.
(A, B) Internalization of soluble anti-Igκ F(ab’)2 by naive mouse B cells (A) or soluble anti-IgM F(ab’)2 by Ramos cells treated with the indicated inhibitors. Data are means ± SEM of n = 3 (A) or 4 (B) experiments. P, significance of the 30 min timepoint in two-way ANOVA comparing treated cells to DMSO. (C, D) Extraction of PMS-bound anti-Igκ F(ab’)2 by naive mouse B cells (C) or anti-IgM F(ab’)2 by Ramos cells (D) treated with inhibitors. Data points represent mean cell values from individual experiments normalized to DMSO controls. Bars show means ± SEM n = 4 experiments. (E, F) Extraction of PMS-bound anti-IgM F(ab’)2 by Ramos cells targeted with the indicated gRNAs. Data points represent mean cell values from individual experiments normalized to controls. Bars show means ± SEM n = 2–3 experiments. In C-F, p values from one-way repeated measures ANOVA are shown.
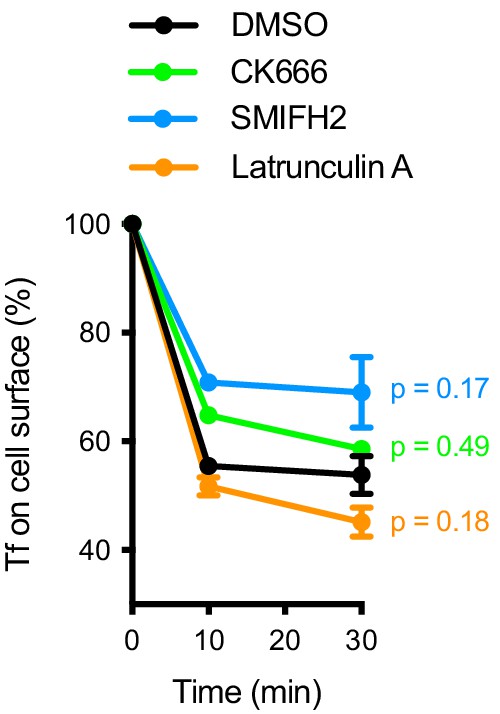
Internalization of soluble transferrin (Tf) by naive mouse B cells in the presence of the indicated inhibitors.
Data show means ± SEM from n = 6 experiments. P, significance of the 30 min timepoint in two-way ANOVA comparing treated cells to DMSO.
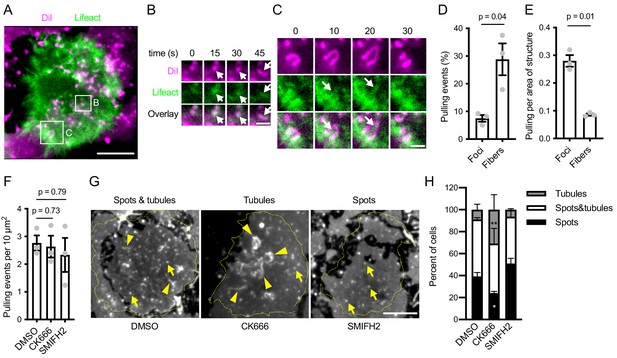
Effect of actin structures on mechanical activity during antigen extraction.
(A) TIRF image of a Lifeact-GFP-expressing Ramos cell (green) interacting with anti-IgM F(ab’)2-loaded PMS stained with DiI (magenta). Spots and tubules in the DiI image indicate deformations of the PMS by the cell. A region containing a DiI spot associated with an actin focus (arrow), later elongating into a fiber is magnified in (B), another region in which a fiber (arrow) elongates a tubule is shown in (C). In B, C the channel intensities were rescaled to the local ranges. Time is shown as relative duration from the initial frame shown. (D) Quantification of the percent of pulling events associated with actin foci or fibers. (E) The rate of pulling events normalized to the area of the cells covered by the two actin structures in the same experiments as in E. Data points in D, E are means of 28–40 cell values from individual experiments, bars show means ± SEM of n = 3 experiments. P, significance in paired t-tests. (F) Quantification of the total number of pulling events in Ramos cells treated with DMSO or the indicated inhibitors. Data points are means of 28–40 cell values from individual experiments, bars show means ± SEM of n = 3 experiments. P, significance in one-way ANOVA. (G) Examples of the shape of membrane deformations in Ramos synapses used to quantify the effect of the inhibitors. Yellow lines show cell outlines. Arrows indicate examples of DiI spots, arrowheads examples of tubules. (H) Quantification of the frequency of synapses containing the indicated membrane shapes in Ramos cells treated with DMSO or the inhibitors. Data are means ± SEM from n = 3 experiments. *, p=0.03, **, p=0.003 in two-way ANOVA against the DMSO control. Scale bars, 5 μm (A, G), 1 μm (B, C).
Relationship of mechanical activity and F-actin in the B cell synapse.
Forces exerted by a Lifeact-expressing Ramos cell on an anti-IgM F(ab’)2-loaded PMS labeled with DiI are visualized as dynamically appearing and moving spots, tubules and ridges in the DiI signal. Related to Figure 6A. Elapsed time is in minutes and seconds.
Mechanical activity in a DMSO-treated Lifeact-expressing Ramos cell on an anti-IgM F(ab’)2-loaded PMS labeled with DiI.
Dynamic spots, tubules and ridges in the DiI signal indicate mechanical activity of the cell. Related to Figure 6G. Elapsed time is in minutes and seconds.
Mechanical activity in a CK666-treated Lifeact-expressing Ramos cell on an anti-IgM F(ab’)2-loaded PMS labeled with DiI.
Dynamic spots, tubules and ridges in the DiI signal indicate mechanical activity of the cell. Related to Figure 6G. Elapsed time is in minutes and seconds.
Mechanical activity in a SMIFH2-treated Lifeact-expressing Ramos cell on an anti-IgM F(ab’)2-loaded PMS labeled with DiI.
Dynamic spots, tubules and ridges in the DiI signal indicate mechanical activity of the cell. Related to Figure 6G. Elapsed time is in minutes and seconds.
Videos
Actin dynamics at the B cell synapse.
Left panel shows raw TIRF image stream with 100 ms time resolution of a primary mouse B cell expressing Lifeact-GFP spread on anti-Igκ F(ab’)2. Right panel shows the cell image overlaid with segmented actin foci and fibers.
Actin foci transiently colocalise with antigen clusters.
Video shows TIRF timelapse of lifeact-GFP (lifeact) in a primary mouse B cell interacting with anti-Igκ F(ab’)2-AlexaFluor647-loaded PMS (antigen). Elapsed time is shown in minutes and seconds. Circles highlight actin foci colocalizing with an antigen cluster.
3D super-resolution image of actin (green) overlaid with epifluorescence images of antigen (magenta).
The video steps through 50 nm-thick sections of the cell shown in Figure 1L.
Actin dynamics after Arp2/3 and formin inhibition.
The panels show TIRF timelapses with 100 ms time resolution of Lifeact-GFP in primary mouse B cells spread on anti-Igκ F(ab’)2-loaded PMSs and treated with DMSO, CK666 and SMIFH2.
Actin dynamics in Ramos cells after Arp2/3 and formin inhibition.
The panels show TIRF timelapses with sub-second time resolution of Lifeact-GFP in Ramos cells spread on anti-IgM F(ab’)2-loaded PMSs and treated with DMSO, CK666 and SMIFH2. Related to Figure 2—figure supplement 1A.
Localization of ARPC2 in Ramos B cell synapse.
TIRF timelapse of Ramos cells co-expressing ARPC2-mRuby and Lifeact-GFP spread on PMSs loaded with anti-IgM F(ab’)2. Panels show ARPC2-mRuby (left), Lifeact-GFP (middle), and their overlay (right). Elapsed time is in seconds.
Localization of DIAPH1 in Ramos B cell synapse.
TIRF timelapse of Ramos cells co-expressing DIAPH1-mRuby and Lifeact-GFP spread on PMSs loaded with anti-IgM F(ab’)2. Panels show DIAPH1-mRuby (left), Lifeact-GFP (middle), and their overlay (right). Elapsed time is in seconds.
Actin dynamics after myosin IIa inhibition.
The panels show TIRF timelapses with 100 ms time resolution of Lifeact-GFP in primary mouse B cells spread on anti-Igκ F(ab’)2-loaded PMSs and treated with DMSO (left) or pn-Blebbistatin (right).
Actin dynamics in B cells interacting with non-stimulatory substrates.
The panels show TIRF timelapses with 100 ms time resolution of Lifeact-GFP in primary mouse B cells spread on anti-Igκ F(ab’)2 (left), anti-MHC I (middle), or anti-MHC II (right)-loaded PMSs.
Actin dynamics after Src-family and Syk kinase inhibition.
The panels show TIRF timelapses with 100 ms time resolution of Lifeact-GFP in primary mouse B cells spread on anti-Igκ F(ab’)2-loaded PMSs and treated with DMSO (left), PP2 (middle), or P505-15 (right).
F-actin and ARPC2 dynamics during uptake of an antigen cluster.
The panels show a TIRF microscopy timelapse of a Ramos cells expressing Lifeact-GFP (green) and ARPC2-mRuby (magenta) interacting with a PMS loaded with anti-IgM F(ab’)2(white). Arrow indicates antigen cluster being taken up by the cell.
F-actin and DIAPH1 dynamics during uptake of an antigen cluster.
The panels show a TIRF microscopy timelapse of a Ramos cells expressing Lifeact-GFP (green) and DIAPH1-mRuby (magenta) interacting with a PMS loaded with anti-IgM F(ab’)2(white). Arrow indicates antigen cluster being taken up by the cell.
Additional files
-
Source code 1
Image Segmentation.
- https://cdn.elifesciences.org/articles/48093/elife-48093-code1-v2.zip
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/48093/elife-48093-transrepform-v2.docx





