Multiplex CRISPR/Cas screen in regenerating haploid limbs of chimeric Axolotls
Figures
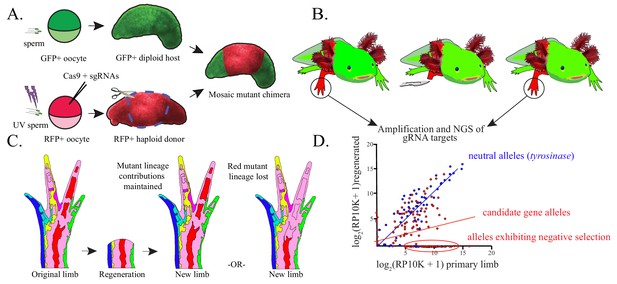
Haploid-diploid chimeric generation and lineage analysis.
(A) Schematic of haploid-diploid chimera generation. Gynogenetic haploids are generated by in vitro activation of unfertilized eggs with UV-enucleated sperm and subsequently mutagenized using CRISPR/Cas9. Chimeric embryos are generated by replacing the limb buds of GFP+ diploid embryos with corresponding tissue from mutagenized haploid donors. (B) DNA is extracted from original and fully regenerated haploid limbs of juvenile chimeric axolotls, target sequences are PCR amplified, and these products are subjected to NGS. (C) Schematic depicting the contribution of mutant cell lineages to the original and regenerated limb. Cell lineages mutant for non-essential candidate genes (light blue, dark blue, yellow, purple, red, green) may participate normally in regeneration and therefore contribute to the regenerated limb and original limb in a similar proportion. Cell lineages harboring deleterious mutant alleles deleterious (red, far right) are predicted to be reduced in regenerated limbs. (D) A hypothetical linear regression plot of the log2 of reads per ten thousand (RP10K+1) of mutant alleles before and after regeneration. Mutant alleles of a neutral gene, tyrosinase (blue), are faithfully preserved between original and regenerated limbs. Mutagenized genes essential for regeneration (red) will show a decrease in allele frequency or a complete loss of alleles in the regenerated limb.
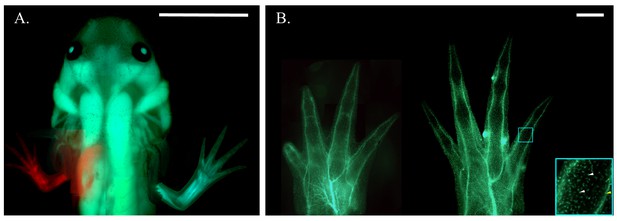
Haploid-diploid chimeric axolotl.
(A) Composite fluorescent image of a chimeric axolotl produced from a limb bud graft from an RFP+ haploid embryo to a GFP+ diploid host. Scale bar = 1 cm. (B) Composite fluorescent image of haploid (left) and diploid (right) limbs produced by embryonic limb bud grafting from a white donor embryo to a GFP+ diploid host. Both the GFP- haploid limb and GFP- diploid limb grafted to a GFP+ diploid host exhibit a GFP expression pattern that appears to be restricted to spinal nerves innervating the limb (yellow arrow) and individual sensory neurons and blood-derived cells (white arrows) stemming from the host body. Blue box is at 4x magnification (bottom right). Scale bars = 1 mm. Composite images were generated by manually compiling individual photos. Images have been adjusted with cropping, contrast, color correction, and gamma correction.
-
Figure 2—source data 1
The number of diploid white to diploid GFP+ grafts that were performed to determine the optimal embryonic stage for limb bud grafting.
- https://cdn.elifesciences.org/articles/48511/elife-48511-fig2-data1-v1.xlsx
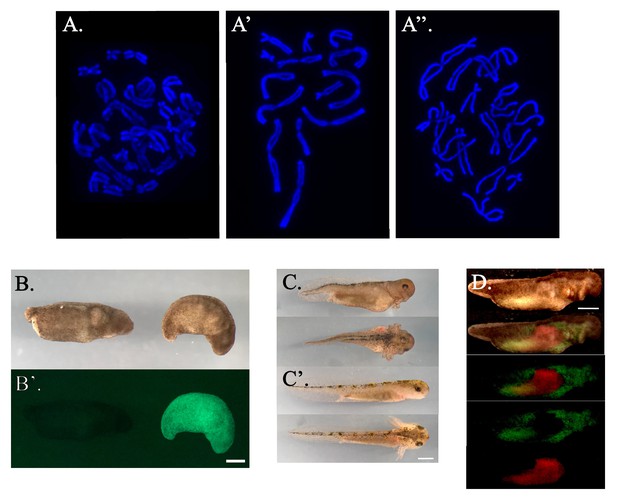
Characterization of haploid larvae.
(A) Fluorescent image of a chromosome squash of a diploid cell (2n = 28). A’ and A’’ Fluorescent images of chromosome squashes from two haploid cells (1n = 14). Chromosomes were stained with Hoescht 33342. (B) Light image of stage 25 haploid (left) and diploid (right) embryos. B’ Green fluorescent image of GFP- haploid and GFP+ diploid embryos. (C) Lateral view (upper) and dorsal view (lower) of a haploid embryo 14 days post fertilization (dpf). C’ Lateral view (upper) and dorsal view(lower) of diploid embryo 14 dpf. (D) Bright field image of a stage 36 haploid-diploid chimera embryo with green and red channel fluorescent overlays in the two panels below. The two lowest panels are fluorescent images of the GFP+; RFP- diploid host body and GFP-; RFP+ haploid tissue graft. All scale bars = 1 mm. Images have been cropped and color corrected with brightness and contrast adjusted as necessary.
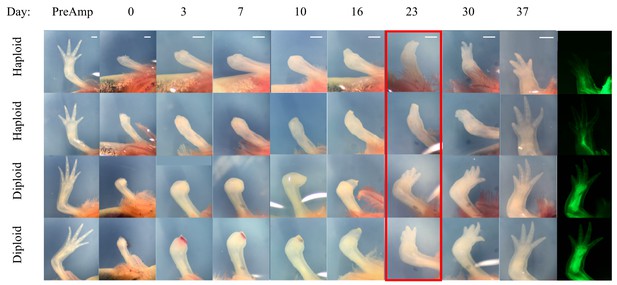
Time course of haploid and diploid limb regeneration.
Four diploid limbs and four haploid limbs of eight stage-matched animals were amputated and imaged on days 0, 3, 7, 10, 16, 23, 30, 37. Haploids (top two rows) regenerate complete limbs and retain the neural GFP expression pattern (last two panels), but show a slight delay around 23 days after amputation versus controls (red box). At this time point, most haploid limbs were still in the palette stage of regeneration (3/4) while diploid limbs were at the point of digital outgrowth (4/4). Haploid limbs are smaller and shorter than diploid limbs (PreAmp, first column). Scale bars = 1 cm. Images have been cropped and color corrected with brightness and contrast adjusted as necessary.
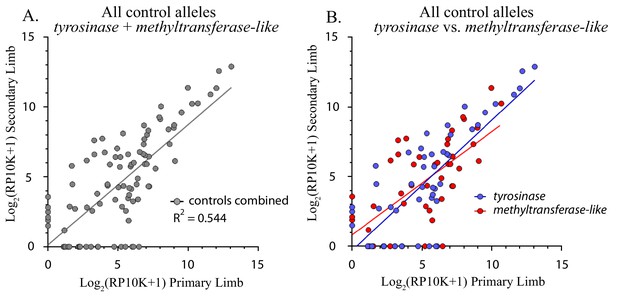
Control alleles.
(A) Comparison of all alleles generated in the controls (methyltransferase plus tyrosinase) in the original and regenerated haploid limbs of 12 animals. The log scores of the reads per ten thousand (RP10K) of every allele in the original limb are significantly correlated with those of the secondary limb (R2 = 0.544, p-value<0.0001). (B) Linear regression comparing the log scores of RP10K for alleles depicted in 3A, but separated by gene (methyltransferase-like in red and tyrosinase in blue). The slopes of the regression lines are not significantly different for the two genes (methyltransferase-like m = 0.740, tyrosinase m = 0.935, p-value=0.238, ANCOVA).
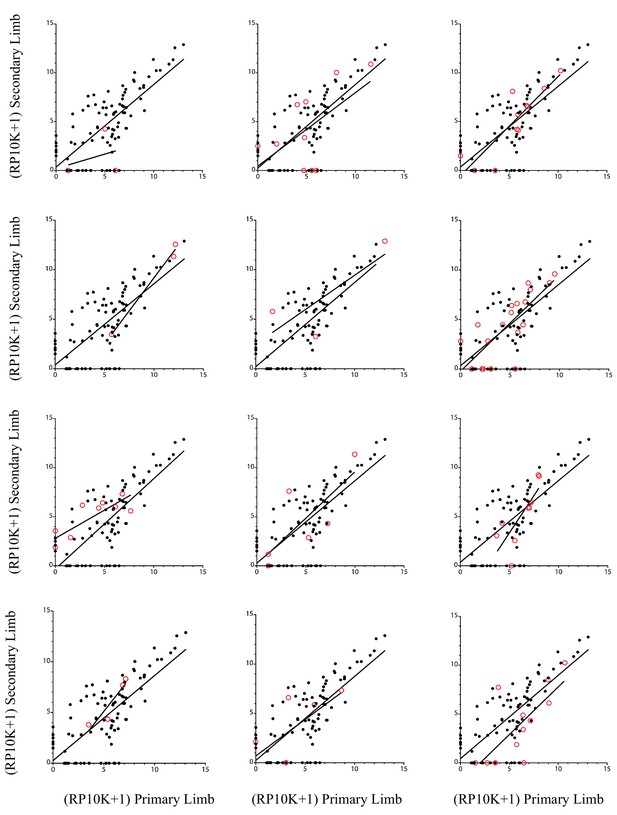
Comparison of all alleles generated in the controls (methyltransferase and tyrosinase) in the original and regenerated haploid limbs of 12 animals shown individually and compared to the entire remaining set of control alleles.
In each graph, the open red circles depicts the log scores of the RP10K of every allele in the original limb compared to that of the same allele in the regenerated limb of an individual animal, while the black circles depict those of the alleles for all other control animals. The best-fit line for the linear regression of the alleles of each individual animal and that of those of all other control animals are depicted. There are no significant differences between the slopes of any best-fit lines with those of the remaining controls (from top-left to bottom-right, p=0.408, 0.699, 0.490, 0.273, 0.614, 0.534, 0.597, 0.767, 0.203, 0.211, 0.737, 0.652).
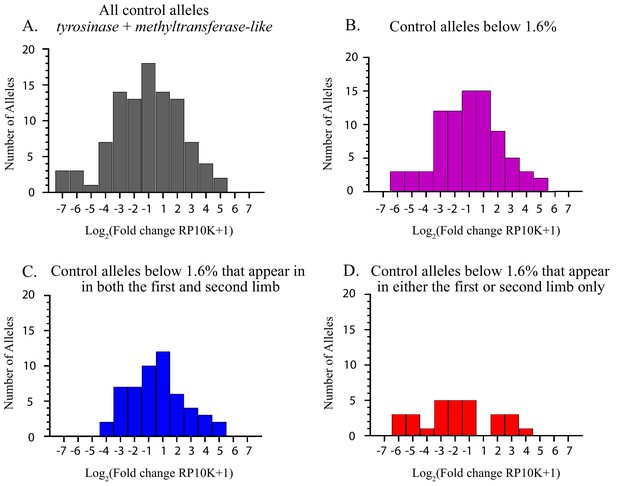
Histograms depicting the log of fold change after regeneration for alleles detected in the controls (methyltransferase-like and tyrosinase).
(A) All alleles detected in the controls. (B) All alleles that occur with a mutation frequency of 1.60% or less. The majority of these alleles undergo less than a two-fold change after regeneration. (C) All alleles that occur with a mutation frequency of 1.60% or less and are preserved between the first and second limb. (D) All alleles with a mutation frequency less than 1.60% that occur in either the first limb or the second limb only.
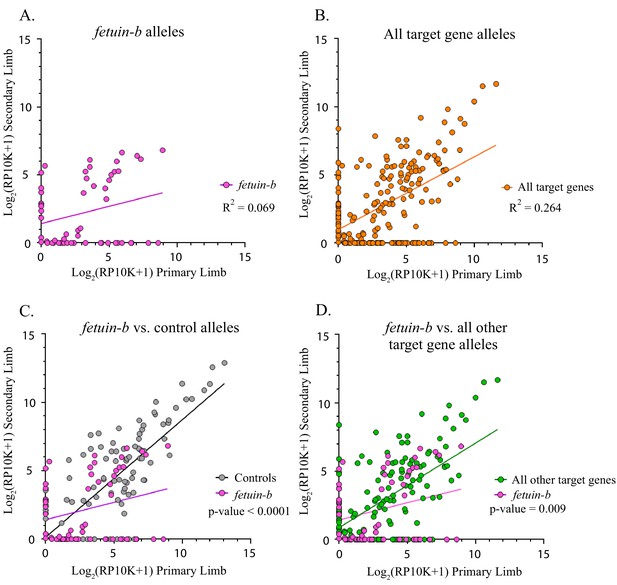
Fetuin-b alleles compared to all other target gene and control alleles.
(A) Linear regression plot of the log2(RP10K) score for all alleles of fetuin-b detected in the first and regenerated haploid limbs of 11 animals. The log scores of alleles in the primary limb poorly predict the log scores of alleles in the secondary limb. (R2 = 0.069, p-value=0.046). (B) Linear regression plot of the log2(RP10K) score for all alleles of all targets detected in the primary and regenerated limb (R2 = 0.264, p<0.0001). (C) Comparison of linear regression plots of fetuin-b (pink) with controls (gray). The slopes of the regression lines are significantly different (fetuin-b m = 0.254, controls m = 0.861, p-value<0.0001, ANCOVA). (D) Comparison of linear regression plots of fetuin-b (pink) with all other targets (green). The slopes of the regression lines are significantly different (fetuin-b m = 0.254, all other targets m = 0.619, p-value=0.009, ANCOVA).
-
Figure 4—source data 1
Raw number of reads, normalized reads, and log2(RP10K) score for all mutant alleles of every targeted gene in each mutant limb in this study.
- https://cdn.elifesciences.org/articles/48511/elife-48511-fig4-data1-v1.xlsx
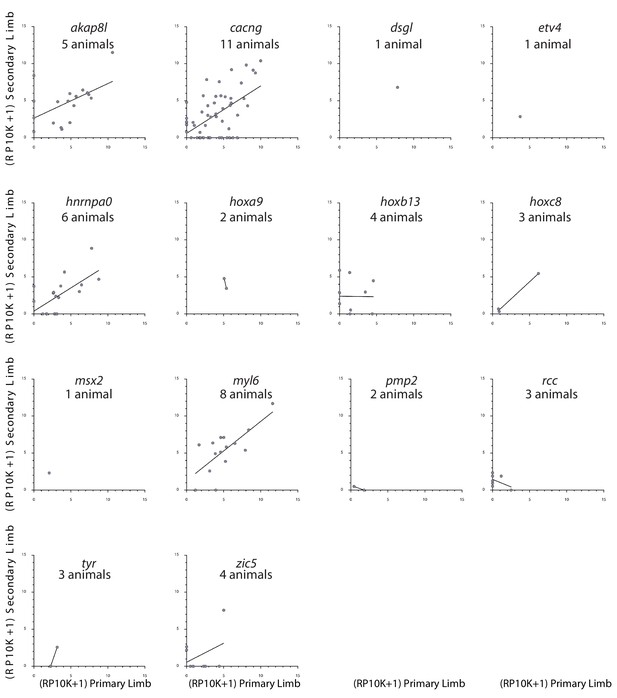
Linear regression plot of the log2(RP10K) score for all alleles in primary and secondary of each targeted gene for which no significant deviation was detected from that of control alleles (akap8l, p=0.069; cacng, p=0.166; hnrnpa0, p=0.371; hoxa9, p=0.637; hoxb13, p=0.053; myl6, p=0.850; pmp2, p=0.624; rcc, p=0.176; tyr, p=0.532,; zic5, p=0.480; ANCOVA).
Best-fit lines are added for genes for which more than one allele was detected. Numbers of animals in which mutations were detected are listed under each gene name.
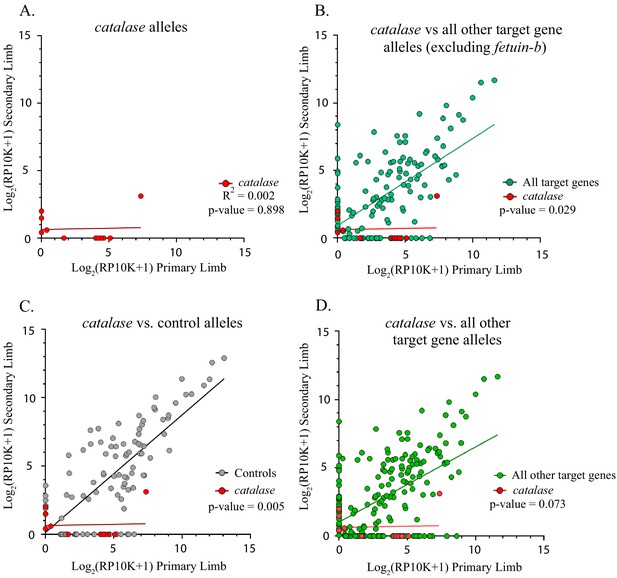
Catalase alleles compared to all other target gene and control alleles.
(A) Linear regression plot of the log2(RP10K) score for all alleles of catalase detected in the first and regenerated haploid limbs of three animals. The log scores of alleles in the primary limbs do not predict the log scores of alleles in the secondary limbs. (R2 = 0.002, p-value=0.898). (B) Comparison of linear regression plots of catalase (red) with all other targets excluding fetuin-b (teal). The slopes of the regression lines are significantly different (catalase m = 0.018, all other targets excluding fetuin-b m = 0.645, p-value=0.029, ANCOVA). (C) Comparison of linear regression plots of catalase (red) with controls (gray). The slopes of the regression lines are significantly different (catalase m = 0.018, controls m = 0.861, p-value=0.005). (D) Comparison of linear regression plots of catalase (red) with all other targets (green). The slopes of the regression lines are not significantly different (catalase m = 0.018, all other targets m = 0.550, p-value=0.073, ANCOVA).
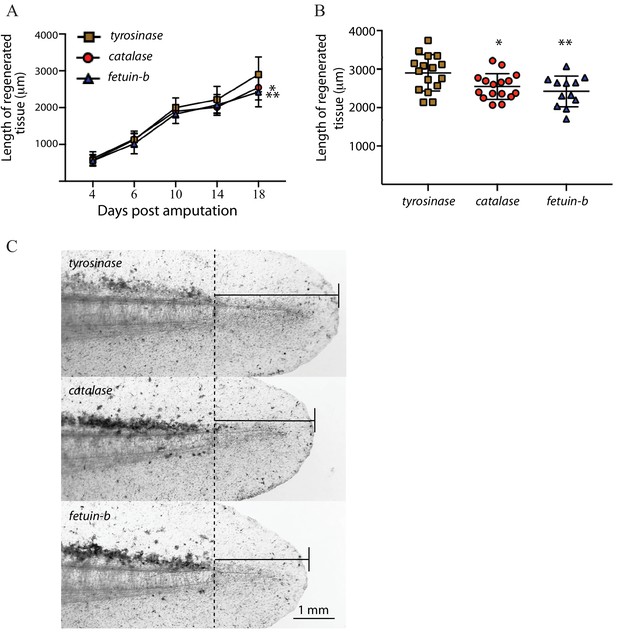
Larval tail regeneration in tyrosinase, catalase, and fetuin-b mutants.
(A) Regenerative outgrowth of tail in high-level tyrosinase, catalase, and fetuin-b F0 mutants. While no significant difference is detected at early time points, both fetuin-b and catalase mutants display tail reduced tail regeneration compared to tyrosinase mutants at later time points (catalase vs tyrosinase, Day 4, p=0.205, Day 6, p=0.400, Day 10, p=0.111. Day 14, p=0.026, Day 18, p=0.011; fetuin-b vs tyrosinase, Day 4, p=0.450, Day 6, p=0.129, Day 10, p=0.047, Day 14, p=0.109, Day 18, 0 = 0.002, Welch’s t-test). Bars indicate standard deviation. (B) Plots of lengths of regenerate in individual tyrosinase, catalase, and fetuin-b F0 mutants at 18 days post-amputation; **=fetuin b, p=0.002, *=catalase, p=0.011. (C) Brightfield images of individual tyrosinase, catalase, and fetuin-b F0 mutants at 18 days post-amputation (dpa) showing median amount of tail regeneration at 18 dpa. Dotted line indicates the amputation plane.
-
Figure 6—source data 1
Regenerative outgrowth measurements and genotyping data for tyrosinase, catalase, and fetuin-b F0 mutants.
- https://cdn.elifesciences.org/articles/48511/elife-48511-fig6-data1-v1.xlsx
Tables
The numbers of all alleles in the first limbs of controls, all targets, fetuin-b, all targets excluding fetuin-b, catalase, and all targets excluding catalase that are sorted by mutation frequency and log of fold change.
| Controls | All targets | |||||||
|---|---|---|---|---|---|---|---|---|
| Allele Frequency | Log of fold change | Allele Frequency | Log of fold change | |||||
| (Low) Frequency < 1.6% | < 2 | > 2 | (Low) Frequency < 1.6% | < 2 | > 2 | |||
| Alleles Lost | 22 | 5 | 17 | Alleles Lost | 60 | 24 | 36 | |
| Alleles Preserved | 53 | 35 | 18 | Alleles Preserved | 94 | 71 | 23 | |
| Sum | 75 | 40 | 35 | Sum | 154 | 95 | 59 | |
| Allele Frequency | Log of fold change | Allele Frequency | Log of fold change | |||||
| Frequency > 1.6% | < 2 | > 2 | Frequency > 1.6% | < 2 | > 2 | |||
| Alleles Lost | 0 | 0 | 0 | Alleles Lost | 2 | 0 | 2 | |
| Alleles Preserved | 17 | 15 | 2 | Alleles Preserved | 20 | 13 | 7 | |
| Sum | 17 | 15 | 2 | Sum | 22 | 13 | 9 | |
| Total alleles: 92 | Total alleles: 176 | |||||||
| fetuin-b | All targets except fetuin-b | |||||||
| Allele Frequency | Log of fold change | Allele Frequency | Log of fold change | |||||
| (Low) Frequency < 1.6% | < 2 | > 2 | (Low) Frequency < 1.6% | < 2 | > 2 | |||
| Alleles Lost | 20 | 9 | 11 | Alleles Lost | 40 | 15 | 25 | |
| Alleles Preserved | 25 | 21 | 4 | Alleles Preserved | 69 | 50 | 19 | |
| Sum | 45 | 30 | 15 | Sum | 109 | 65 | 44 | |
| Allele Frequency | Log of fold change | Allele Frequency | Log of fold change | |||||
| Frequency > 1.6% | < 2 | > 2 | Frequency > 1.6% | < 2 | > 2 | |||
| Alleles Lost | 2 | 0 | 2 | Alleles Lost | 0 | 0 | 0 | |
| Alleles Preserved | 1 | 0 | 1 | Alleles Preserved | 19 | 13 | 6 | |
| Sum | 3 | 0 | 3 | Sum | 19 | 13 | 6 | |
| Total alleles: 48 | Total alleles: 128 | |||||||
| catalase | All other targets except catalase | |||||||
| Allele Frequency | Log of fold change | Allele Frequency | Log of fold change | |||||
| (Low) Frequency < 1.6% | < 2 | > 2 | (Low) Frequency < 1.6% | < 2 | > 2 | |||
| Alleles Lost | 6 | 1 | 5 | Alleles Lost | 54 | 23 | 31 | |
| Alleles Preserved | 1 | 1 | 0 | Alleles Preserved | 93 | 70 | 23 | |
| Sum | 7 | 2 | 5 | Sum | 147 | 93 | 54 | |
| Allele Frequency | Log of fold change | Allele Frequency | Log of fold change | |||||
| Frequency > 1.6% | < 2 | > 2 | Frequency > 1.6% | < 2 | > 2 | |||
| Alleles Lost | 0 | 0 | 0 | Alleles Lost | 2 | 0 | 2 | |
| Alleles Preserved | 1 | 0 | 1 | Alleles Preserved | 19 | 13 | 6 | |
| Sum | 1 | 0 | 1 | Sum | 21 | 13 | 8 | |
| Total alleles: 8 | Total alleles: 168 | |||||||
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (Ambystoma mexicanum) | cagg:egfp | Ambystoma Genetic Stock Center (Sobkow et al., 2006) | AGSC Cat# 110A, RRID:AGSC_110A | |
| Genetic reagent (Ambystoma mexicanum) | cagg:nls-mcherry | Ambystoma Genetic Stock Center (Kragl et al., 2009) | AGSC Cat# 112A, RRID:AGSC_112A | |
| Chemical compound, drug | MS-222 | Western Chemical | ANADA #200–226 | |
| Chemical compound, drug | Human chorionic gonadotropin (Chorulon,) | Merck Animal Health | NADA 140–927 | |
| Gene (Ambystoma mexicanum) | msx2 | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000067092 | |
| Gene (Ambystoma mexicanum) | prmt1 | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000062704 | |
| Gene (Ambystoma mexicanum) | myl6 | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000067862 | |
| Gene (Ambystoma mexicanum) | fetub | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000227254 | |
| Gene (Ambystoma mexicanum) | hoxc8 | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000065333 | |
| Gene (Ambystoma mexicanum) | akap8l | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000192860 | |
| Gene (Ambystoma mexicanum) | hrnrpa0 | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000081837 | |
| Gene (Ambystoma mexicanum) | hsd17b10 | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000257015 | |
| Gene (Ambystoma mexicanum) | hoxb9 | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000035333 | |
| Gene (Ambystoma mexicanum) | tyrosinase | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000179254 | |
| Gene (Ambystoma mexicanum) | etv4 | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000233035 | |
| Gene (Ambystoma mexicanum) | cacng1 | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000081988 | |
| Gene (Ambystoma mexicanum) | catalase | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000186723 | |
| Gene (Ambystoma mexicanum) | hoxb13 | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000007929 | |
| Gene (Ambystoma mexicanum) | zic5 | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000057641 | |
| Gene (Ambystoma mexicanum) | ecm1 | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000123229 | |
| Gene (Ambystoma mexicanum) | cornifelin | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000173184 | |
| Gene (Ambystoma mexicanum) | dsg-like | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000056512 | |
| Gene (Ambystoma mexicanum) | enpp2 | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000217071 | |
| Gene (Ambystoma mexicanum) | fabp2 | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000084459 | |
| Gene (Ambystoma mexicanum) | pmp2 | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000238807 | |
| Gene (Ambystoma mexicanum) | kcne1 | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000121776 | |
| Gene (Ambystoma mexicanum) | krt6a | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000060835 | |
| Gene (Ambystoma mexicanum) | rcc1 | Axolotl transcriptome assembly 3.4 | AMEXTC_0340000210022 | |
| Recombinant DNA reagent | MLM3613 | (Hwang et al., 2013) | RRID: Addgene plasmid 42251 | Cas9 expression vector |
| Peptide, recombinant protein | Cas9 | PNABio | Cat. #: CP04-500 | |
| Commercial assay or kit | mMessage mMachine Kit | ThermoFisher | Cat. #: Am1345 | |
| Commercial assay or kit | MAXIscript SP6/T7 Transcription Kit | ThermoFisher | Cat. #: Am1322 | |
| Chemical compound, drug | MS-222 | Sigma Aldrich | SML1656 | |
| Software, algorithm | Geneious Software | Biomatters | RRID:SCR_010519 |
Additional files
-
Supplementary file 1
Primer and gRNA sequences used in study.
- https://cdn.elifesciences.org/articles/48511/elife-48511-supp1-v1.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/48511/elife-48511-transrepform-v1.docx





