Modeling the dynamics of Plasmodium falciparum gametocytes in humans during malaria infection
Figures
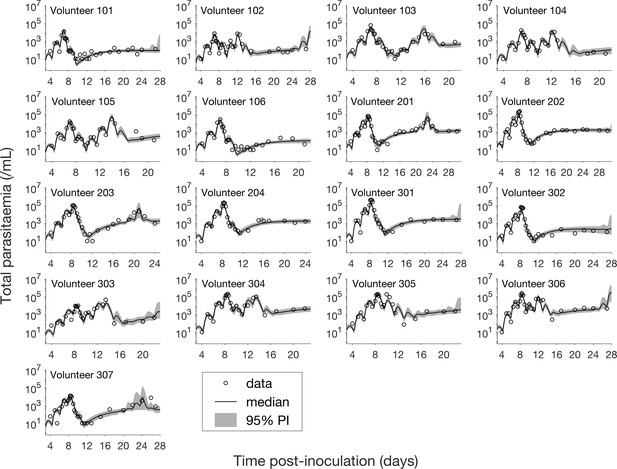
Results of model fitting for all 17 volunteers.
Data are presented by circles. The median of posterior predictions (solid line) and 95% prediction interval (PI, shaded area) are generated by 5000 model simulations based on 5000 samples from the posterior parameter distribution as described in the Materials and methods. The histograms showing the posterior distributions of population mean and standard deviation hyperparameters are given in Figure 1—figure supplements 1 and 2. The posterior distribution of each model parameter (see the Materials and methods) for individual volunteers are given in Figure 1—figure supplements 3–14 Posterior distributions for some biological parameters are given in Figure 1—figure supplement 15, which are generated based on the posterior samples of population mean parameters (see the Materials and methods) and will be used to support the results in Table 1 shown later. The source data and computer code with instructions of implementation to generate Figure 1 and Figure 1—figure supplements 1–15 are fully publicly available at https://doi.org/10.26188/5cde4c26c8201.
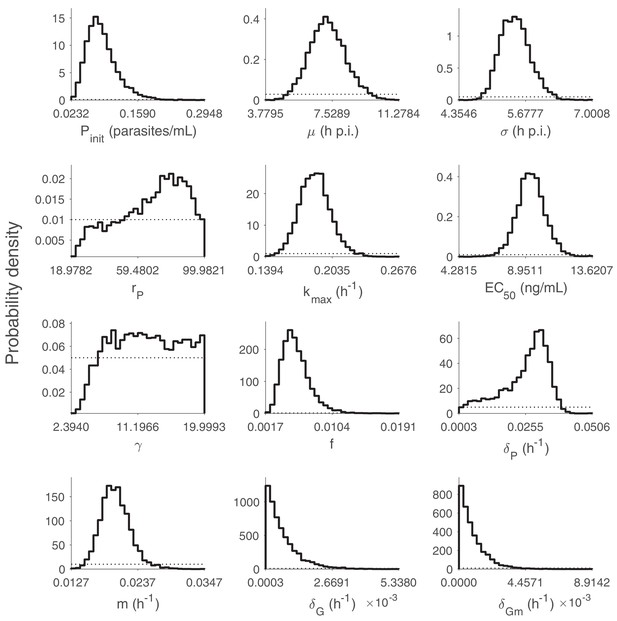
Marginal posterior distributions for the 12 population mean parameters (hyperparameters).
5000 samples are used to generate the distributions. The dashed curves indicate the uniform prior distributions. p.i.: post-inoculation. Note that the y-axis is probability density instead of number of samples. Relevant details of the hyperparameters are provided in the Materials and methods in the main text.
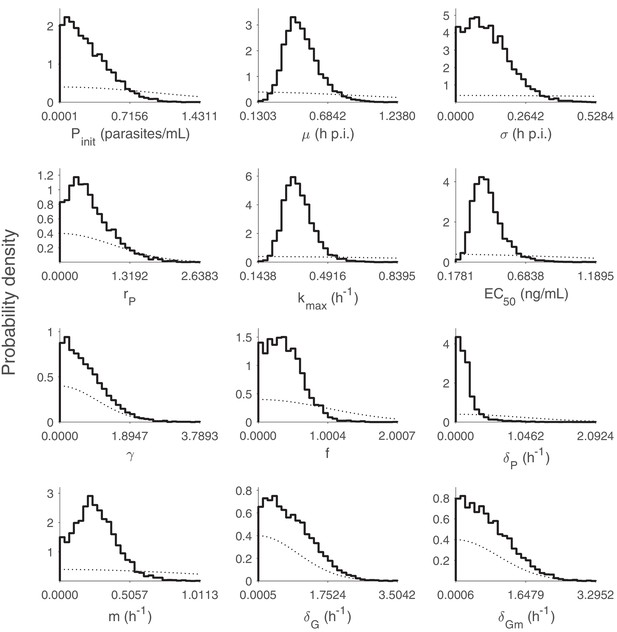
Marginal posterior distributions for the 12 population SD parameters (hyperparameters).
5000 samples are used to generate the distributions. The dashed curves indicate the half-normal prior distributions. p.i.: post-inoculation. Note that the y-axis is probability density instead of number of samples. Relevant details of the hyperparameters are provided in the Materials and methods in the main text.
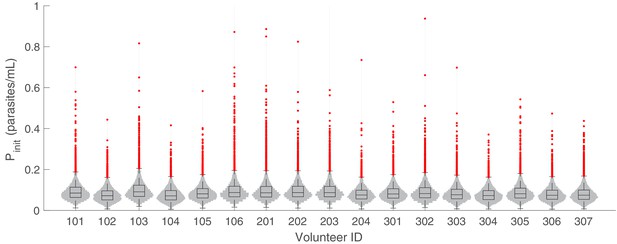
The marginal posterior distributions of the individual parameter of (inoculation size) for all 17 volunteers.
The violin plots (gray area) show the distributions of 5000 posterior samples. Box plots show the 25%, 50% (median) and 75% quantiles with outliers indicated by red dots. Relevant details of the individual parameter are provided in the Materials and methods in the main text.
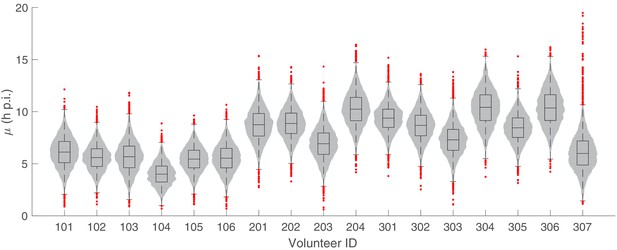
The marginal posterior distributions of the individual parameter of µ (mean of the initial parasite age distribution) for all 17 volunteers.
The violin plots (gray area) show the distributions of 5000 posterior samples. Box plots show the 25%, 50% (median) and 75% quantiles with outliers indicated by red dots. Relevant details of the individual parameter are provided in the Materials and methods in the main text.
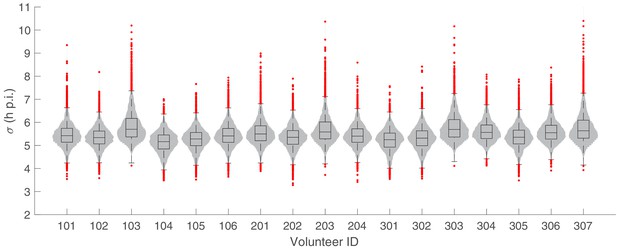
The marginal posterior distributions of the individual parameter of (Standard deviation of the initial parasite age distribution) for all 17 volunteers.
The violin plots (gray area) show the distributions of 5000 posterior samples. Box plots show the 25%, 50% (median) and 75% quantiles with outliers indicated by red dots. Relevant details of the individual parameter are provided in the Materials and methods in the main text.
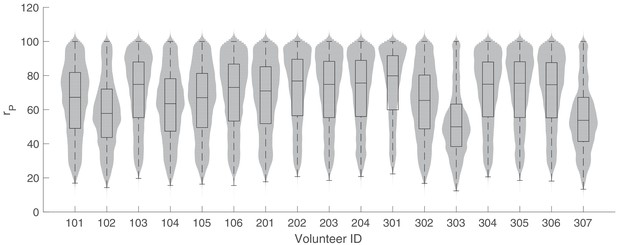
The marginal posterior distributions of the individual parameter of (parasite replication rate) for all 17 volunteers.
The violin plots (gray area) show the distributions of 5000 posterior samples. Box plots show the 25%, 50% (median) and 75% quantiles with outliers indicated by red dots. Relevant details of the individual parameter are provided in the Materials and methods in the main text.
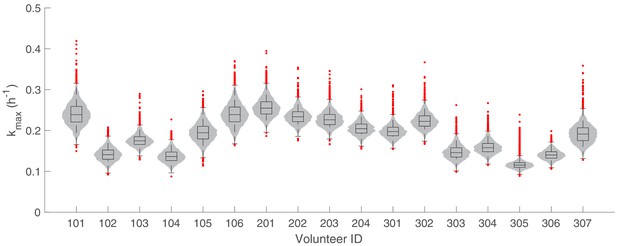
The marginal posterior distributions of the individual parameter of (maximum rate of parasite killing by PQP) for all 17 volunteers.
The violin plots (gray area) show the distributions of 5000 posterior samples. Box plots show the 25%, 50% (median) and 75% quantiles with outliers indicated by red dots. Relevant details of the individual parameter are provided in the Materials and methods in the main text.
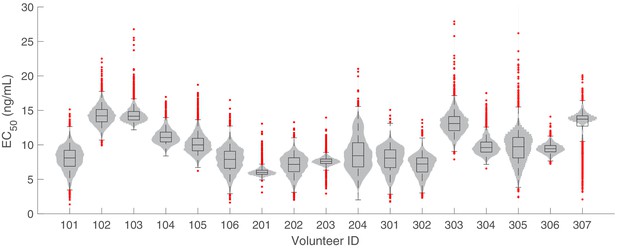
The marginal posterior distributions of the individual parameter of (half-maximum effective PQP concentration) for all 17 volunteers.
The violin plots (gray area) show the distributions of 5000 posterior samples. Box plots show the 25%, 50% (median) and 75% quantiles with outliers indicated by red dots. Relevant details of the individual parameter are provided in the Materials and methods in the main text.
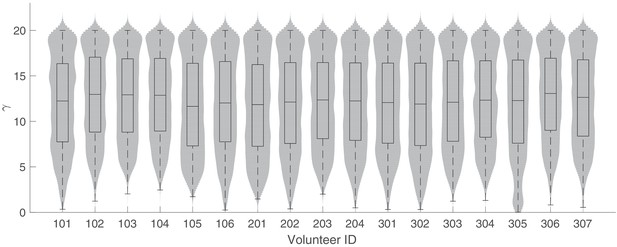
The marginal posterior distributions of the individual parameter of (Hill coefficient for PQP) for all 17 volunteers.
The violin plots (gray area) show the distributions of 5000 posterior samples. Box plots show the 25%, 50% (median) and 75% quantiles with outliers indicated by red dots. Relevant details of the individual parameter are provided in the Materials and methods in the main text.
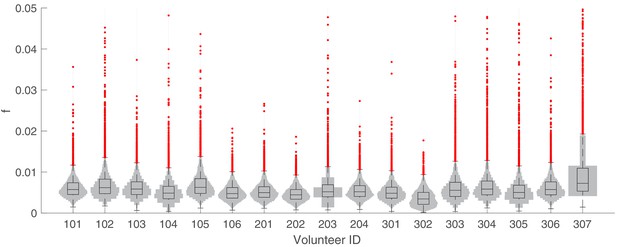
The marginal posterior distributions of the individual parameter of (sexual commitment rate; not converted to percentage) for all 17 volunteers.
The violin plots (gray area) show the distributions of 5000 posterior samples. Box plots show the 25%, 50% (median) and 75% quantiles with outliers indicated by red dots. Relevant details of the individual parameter are provided in the Materials and methods in the main text.
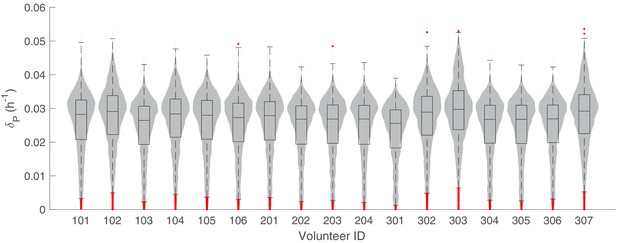
The marginal posterior distributions of the individual parameter of (death rate of asexual and sexual parasites) for all 17 volunteers.
The violin plots (gray area) show the distributions of 5000 posterior samples. Box plots show the 25%, 50% (median) and 75% quantiles with outliers indicated by red dots. Relevant details of the individual parameter are provided in the Materials and methods in the main text.
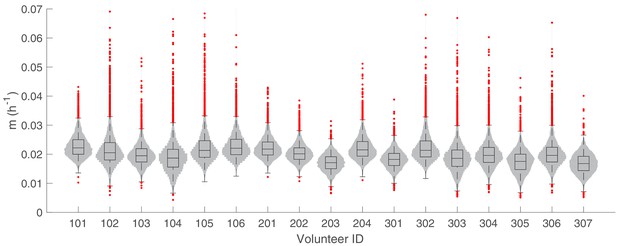
The marginal posterior distributions of the individual parameter of (maturation rate of gametocytes) for all 17 volunteers.
The violin plots (gray area) show the distributions of 5000 posterior samples. Box plots show the 25%, 50% (median) and 75% quantiles with outliers indicated by red dots. Relevant details of the individual parameter are provided in the Materials and methods in the main text.
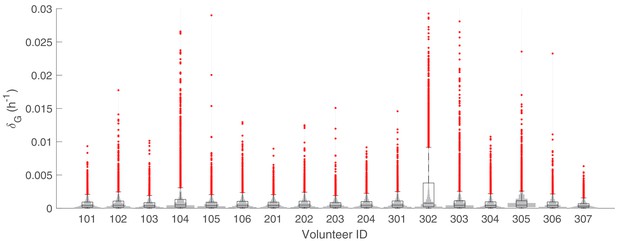
The marginal posterior distributions of the individual parameter of (death rate of sequestered gametocytes) for all 17 volunteers.
The violin plots (gray area) show the distributions of 5000 posterior samples. Box plots show the 25%, 50% (median) and 75% quantiles with outliers indicated by red dots. Relevant details of the individual parameter are provided in the Materials and methods in the main text.
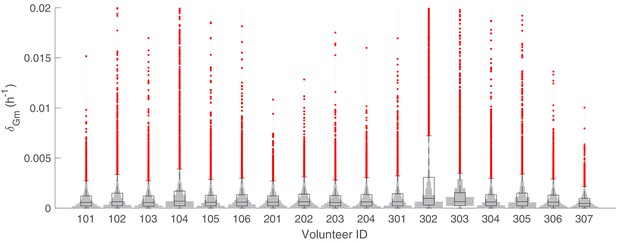
The marginal posterior distributions of the individual parameter of (death rate of circulating gametocytes) for all 17 volunteers.
The violin plots (gray area) show the distributions of 5000 posterior samples. Box plots show the 25%, 50% (median) and 75% quantiles with outliers indicated by red dots. Relevant details of the individual parameter are provided in the Materials and methods in the main text.
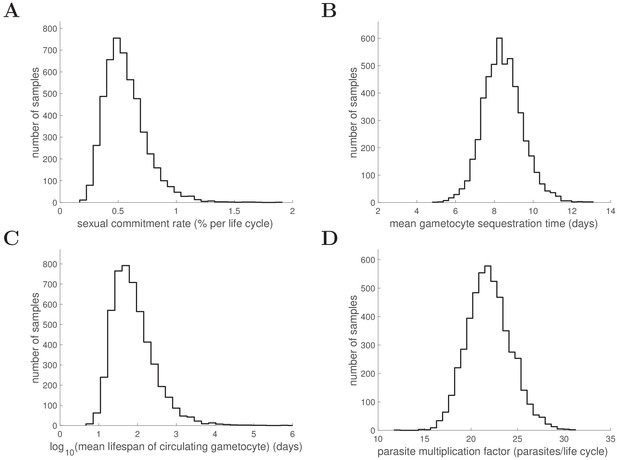
Marginal posterior distributions of some key biological parameters.
5000 samples from the posterior parameter distribution are used to generate the figures. Full details about the definitions and expressions of those biological parameters are provided in the Materials and methods in the main text. Note that a logarithm is taken for the mean lifespan of circulating gametocyte for a better visualisation of the distribution.
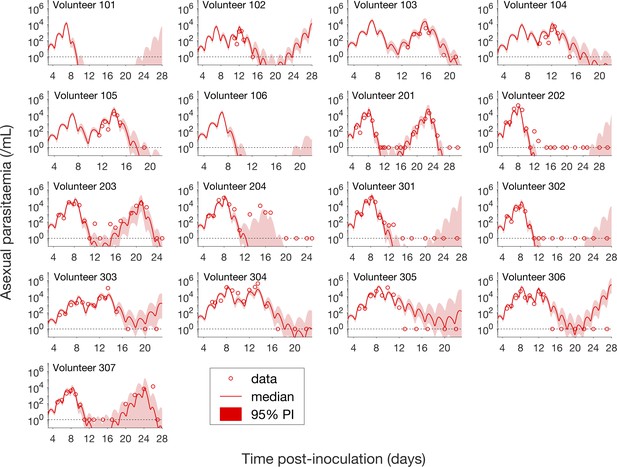
Comparison of model predictions and clinical data for the asexual parasitemia for all 17 volunteers.
Data are presented by circles. The median of posterior predictions (solid curve) and 95% PI (shaded area) are generated by 5000 model simulations based on 5000 samples from the posterior parameter distribution as described in the Materials and methods. The data points with one parasite/mL (i.e., those points which lie on the dotted line) indicate measurements for which no parasites were detected. No data are available for Volunteer 101 and 106 to validate the model predictions. The source data and computer code with instructions of implementation to generate Figure 2 are fully publicly available at https://doi.org/10.26188/5cde4c26c8201.
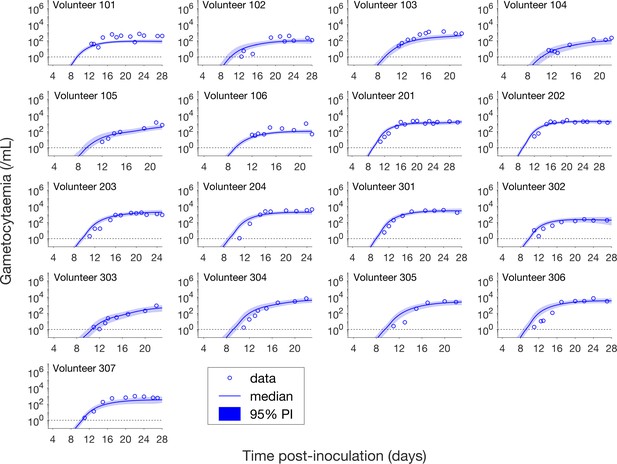
Comparison of model predictions and clinical data for the gametocytemia for all 17 volunteers.
Data are presented by circles. The median of posterior predictions (solid curve) and 95% PI (shaded area) are generated by 5000 model simulations based on 5000 samples from the posterior parameter distribution as described in the Materials and methods. The data points with one parasite/mL (i.e. those points which lie on the dotted line) indicate measurements for which no parasites were detected. The source data and computer code with instructions of implementation to generate Figure 3 are fully publicly available at https://doi.org/10.26188/5cde4c26c8201.
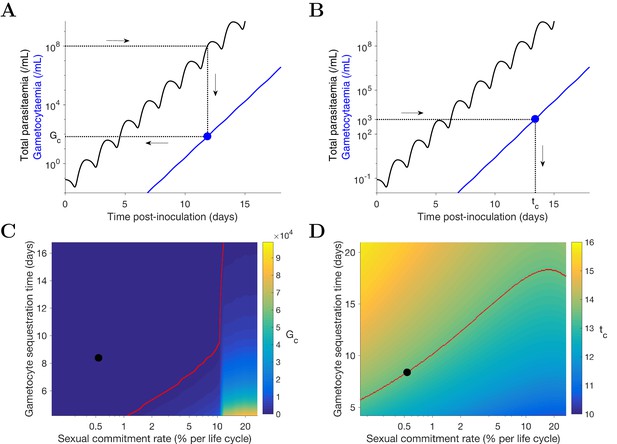
Simulation of two scenarios predicting the dependence of human-to-mosquito transmissibility on the sexual commitment rate and gametocyte sequestration time.
(A) illustration of the first scenario: predicting the critical gametocytemia level (indicated by Gc) at the time when the total parasitemia reaches 108 parasites/mL. (B) illustration of the second scenario: predicting the non-infectious period (indicated by tc), which is defined to be time from inoculation of infected red blood cells to the time when the gametocytemia reaches 103 parasites/mL (a threshold below which human-to-mosquito transmission was not observed [Collins et al., 2018]). (C and D) Heatmaps showing the dependence of the critical gametocytemia Gc and the non-infectious period tc on the sexual commitment rate and gametocyte sequestration time. The black dots represent the value obtained by simulating the gametocyte dynamics model using the median estimates of the posterior samples of the population mean parameters as described in the Materials and methods. The red curve in C is the level curve for Gc = 103 parasites/mL. The red curve in D is the level curve for tc = 13.42 days which is the non-infectious period obtained by model simulation using the posterior estimates of the population mean parameters. The source data and computer code with instruction of implementation to generate Figure 4 are fully publicly available at https://doi.org/10.26188/5cde4c26c8201.
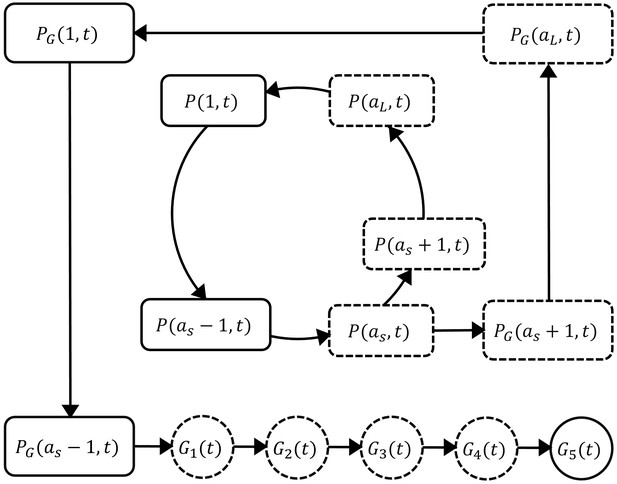
Schematic diagram showing the model compartments and transitions.
The model is comprised of three parts describing three populations of parasites: asexual parasites (), sexually committed parasites () and gametocytes (). P and PG are functions of asexual parasite age and time . Square compartments in the inner loop represent the asexual parasite population which follows a cycle of maturation and replication every hours. Sexual commitment occurs from age and a fraction of asexual parasites become sexually committed (the bigger square compartments in the outer loop) and eventually enter the development of stage I–V gametocytes (G1–G5). The compartments with a dashed boundary are sequestered to tissues and thus not measurable in a blood smear. The notation for each compartment is consistent with those in the model equations and is explained in the main text.
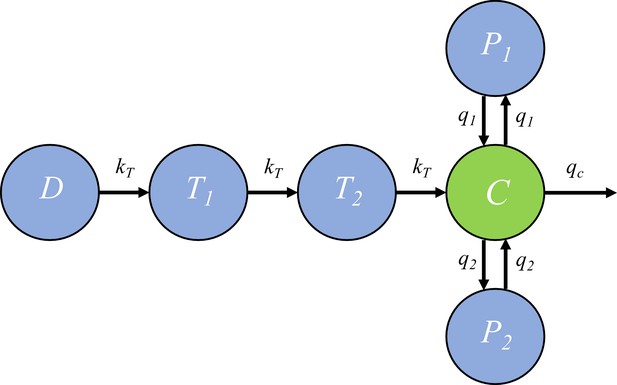
The pharmacokinetic model of piperaquine (PQP).
The model is a three-compartment disposition model with two transit compartments for absorption. State D represents the dose of PQP. T1 and T2 represent the two transit compartments. C is the central compartment and PQP concentration in this compartment was measured (which are shown in Figure 6—figure supplement 1). P1 and P2 represent two peripheral compartments. kT, q1, q2 and qc are the rates of flow into or out of compartments.
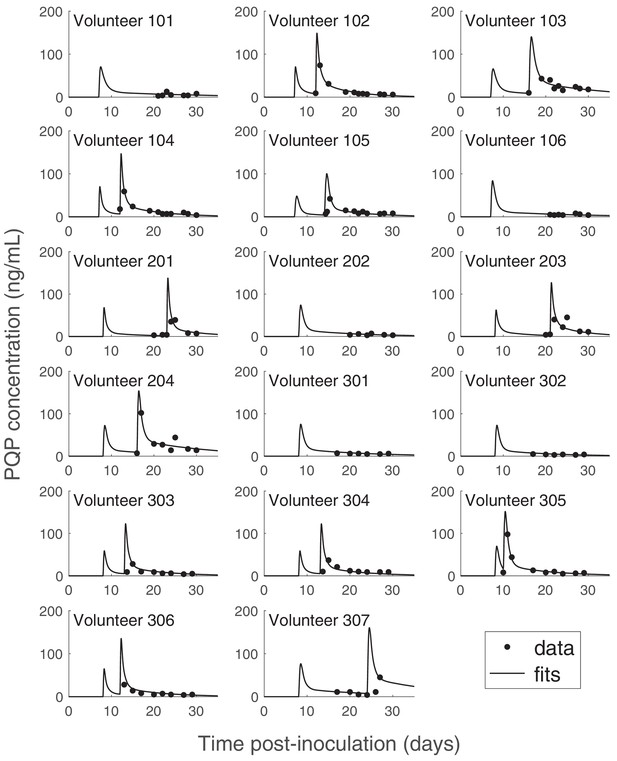
PK data and optimized PK curves (the ‘fits’) of piperaquine (PQP) concentration for all volunteers.
The details of the optimization approach are provided in the Materials and methods in the main text and Appendix 1. Some volunteers have two peaks of PQP concentrations because they had recrudescent asexual parasitemia (see Figure 1 in the main text) and were treated with a second dose of 960 mg PQP. The source data and computer code with instructions of implementation to generate Figure 6—figure supplement 1 are fully publicly available at https://doi.org/10.26188/5cde4c26c8201.
Tables
Estimates of some key biological parameters and comparison with the literature.
The estimates of the biological parameters (middle column) are shown as the median and 95% credible interval (CI) of the marginal posterior parameter distribution (Figure 1—figure supplement 15). Estimates from the literature (third column) are shown in the format of either ‘mean estimate (95% confidence interval)’ or ‘mean estimate [minimum – maximum estimate]’ or simply ‘a low estimate – a high estimate’. Note some quoted estimates are from either in vivo or in vitro studies of P. falciparum. The source data and computer code with instructions of implementation to generate our model estimates (middle column) in Table 1 are fully publicly available at https://doi.org/10.26188/5cde4c26c8201.
| Biological parameters (unit) | Median estimate (95% CI) | Estimates in the literature |
|---|---|---|
| Sexual commitment rate (%/asexual replication cycle) | 0.54 (0.30–1.00) | 11 (6.2–15.8) (Filarsky et al., 2018) (in vitro) 0.64 [0.027–13.5] (Eichner et al., 2001) (in vivo) |
| Gametocyte sequestration time (days) | 8.39 (6.54–10.59) | 7.4 [4 – 12] (Eichner et al., 2001) (in vivo) |
| Circulating gametocyte lifespan (days) | 63.5 (12.7–1513.9) | 16–32 (Gebru et al., 2017) (in vitro) 6.4 [1.3–22.2] (Eichner et al., 2001) (in vivo) |
| Parasite multiplication factor (per asexual replication cycle) | 21.8 (17.6–26.9) | 10–33 (Wockner et al., 2017) (in vivo) 16.4 (15.1–17.8)a (in vivo) |
-
a JS McCarthy, personal communication, May 2019.
Details of the gametocyte dynamics model parameters.
The table includes the unit, description and prior distribution for each model parameter. For the uniform prior distributions (U), the lower bounds are non-negative based on the definitions of the model parameters and the upper bounds for the prior distributions were chosen to be sufficiently wide in order to accommodate all biologically plausible values from the literature (Zaloumis et al., 2012). We assumed parasites younger than 25h are circulating and thus fix to be 25h. For 3D7 strain, the asexual replication cycle is approximately 39–45h (based on in vitro estimates [Duffy and Avery, 2017] and personal communication [JS McCarthy, personal communication, May 2019]) and we fix to be 42h.
| Parameter | Unit | Description | Prior distribution |
|---|---|---|---|
| parasites/mL | inoculation size | U(0, 10) | |
| h | mean of the initial parasite age distribution | U(0, 35) | |
| h | SD of the initial parasite age distribution | U(0, 20) | |
| (unitless) | parasite replication rate | U(0, 100) | |
| h−1 | maximum rate of parasite killing by PQP | U(0, 1) | |
| ng/mL | half-maximum effective PQP concentration | U(1, 100) | |
| (unitless) | Hill coefficient for PQP | U(0, 20) | |
| (unitless) | the fraction of parasites entering sexual development per asexual replication cycle | U(0, 1) | |
| h−1 | death rate of asexual and sexual parasites | U(0, 0.2) | |
| h−1 | maturation rate of gametocytes | U(0, 0.1) | |
| h−1 | death rate of sequestered gametocytes | U(0, 0.1) | |
| h−1 | death rate of circulating gametocytes | U(0, 0.1) | |
| h | sequestration age of asexual parasites | fixed to be 25 | |
| h | length of life cycle of asexual parasites | fixed to be 42 |
-
SD: standard deviation; h: hour.
Parameter values used to generate the optimized PK curves for all 17 volunteers.
The units of the model parameters are given in the parentheses. The optimized PK curves are shown in Figure 6—figure supplement 1.
| PK parameter (unit) | kT(h-1) | qc(L/h) | Vc(L) | q1(L/h) | V1(L) | q2(L/h) | v2(L) |
|---|---|---|---|---|---|---|---|
| Volunteer 101 | 1/2.43 | 53.7 | 831 | 2240 | 3958 | 118 | 17177 |
| Volunteer 102 | 1/2.32 | 123.4 | 582 | 849 | 3985 | 163 | 10355 |
| Volunteer 103 | 1/3.07 | 61.3 | 756 | 2686 | 3910 | 127 | 14346 |
| Volunteer 104 | 1/2.32 | 134.7 | 488 | 746 | 3986 | 190 | 15007 |
| Volunteer 105 | 1/3.25 | 160 | 1342 | 5034 | 3986 | 190 | 17546 |
| Volunteer 106 | 1/2.40 | 60.7 | 667 | 2992 | 3043 | 123 | 16374 |
| Volunteer 201 | 1/2.32 | 160 | 469 | 746 | 3986 | 190 | 15557 |
| Volunteer 202 | 1/2.79 | 71.4 | 804 | 2020 | 3010 | 149 | 13300 |
| Volunteer 203 | 1/2.32 | 141.9 | 1243 | 803 | 3973 | 190 | 11825 |
| Volunteer 204 | 1/2.32 | 63.3 | 895 | 2169 | 3327 | 165 | 16205 |
| Volunteer 301 | 1/2.79 | 61.4 | 804 | 2020 | 3010 | 149 | 13300 |
| Volunteer 302 | 1/2.79 | 81.4 | 804 | 2020 | 3010 | 149 | 13300 |
| Volunteer 303 | 1/2.45 | 159.4 | 1332 | 779 | 3970 | 190 | 17355 |
| Volunteer 304 | 1/2.48 | 158.9 | 1339 | 762 | 3981 | 190 | 17290 |
| Volunteer 305 | 1/2.53 | 120.9 | 1152 | 1905 | 2979 | 123 | 13896 |
| Volunteer 306 | 1/2.32 | 158.8 | 870 | 761 | 3983 | 190 | 17488 |
| Volunteer 307 | 1/2.79 | 51.4 | 804 | 2020 | 3010 | 149 | 13300 |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.49058.026





