MouseBytes, an open-access high-throughput pipeline and database for rodent touchscreen-based cognitive assessment
Figures
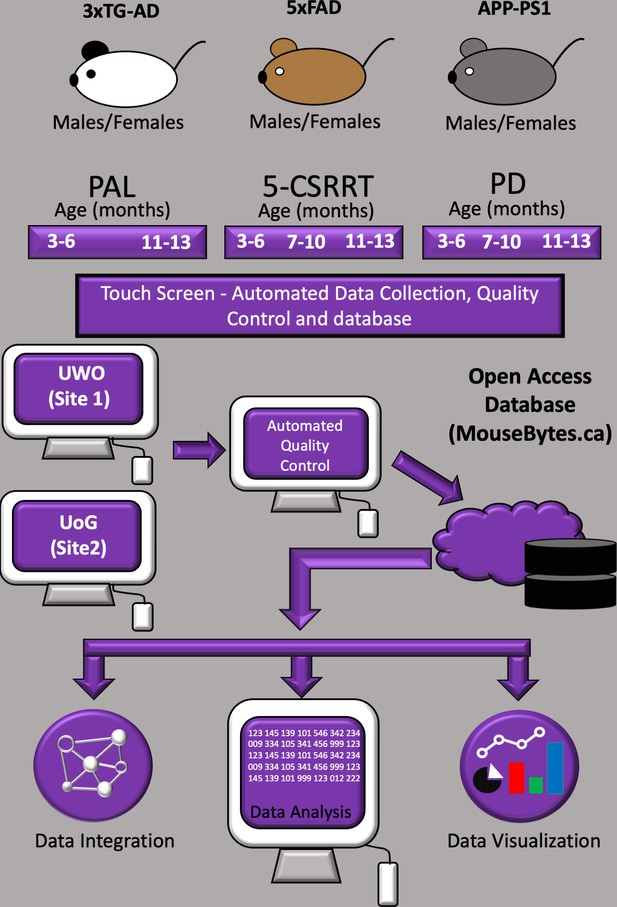
Schematic overview of the automated touchscreen cognition platform.
Males and females of three different AD mouse lines were each evaluated in three different touchscreen tasks. The mice were food restricted and tested longitudinally and at two different sites (The University of Western Ontario (UWO) and University of Guelph (UoG) - Canada). Data were submitted to an automated QC process. Following automated QC, data were uploaded to an open-access database (mousebytes.ca) for post-processing analysis and visualisation using the analytics tool TIBCO Spotfire.
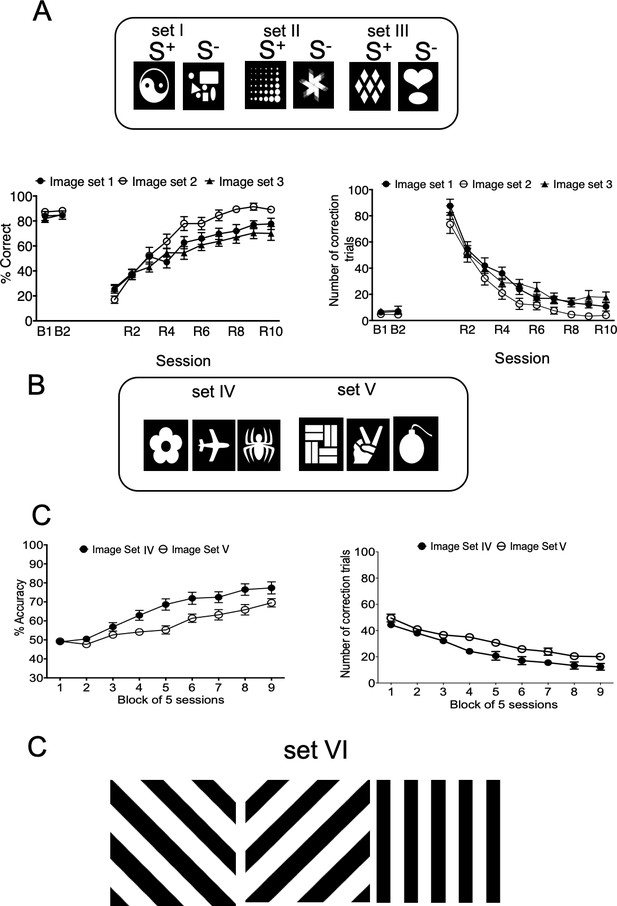
Validation of image sets used during longitudinal testing.
4 month old B6129SF2/J wild-type mice tested in PD (A) and PAL (B) with different image sets. (C) Image set used for PAL task (5xFAD and APP/PS1 mice at 10–11 months of age).
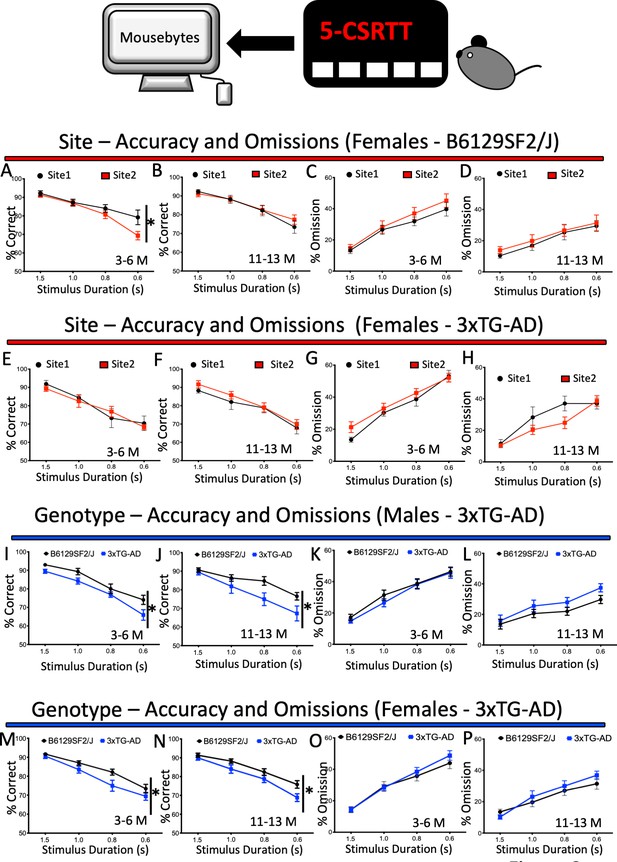
Performance and response measures of Male and Female mice during 5-CSRTT probe trials.
Mice were subjected to a series of probe trials and the averages of accuracy (% correct), omissions (%) and premature responses (number) were plotted at different ages. The plots were generated with data downloaded from MouseBytes and the links (datasets) for the individual analysis can be found in the results section. (A-D), longitudinal site comparison of the performance (accuracy and omissions) of female Wild-type controls (B6129SF2/J) at 3–6 and 11 to 13 months of age; (E-H) longitudinal site comparison of the performance (accuracy and omissions) of female 3xTG-AD at 3–6 and 11 to 13 months of age respectively; (I-L) comparison of the performance (accuracy and omissions) of 3xTG-AD male and their Wild-type controls (B6129SF2/J) at 3–6 and 11 to 13 months of age; (M-P) comparison of the performance (accuracy and omissions) of 3xTG-AD female mice and Wild-type controls (B6129SF2/J) at 3–6 and 11 to 13 months of age. Results are presented as means ± s.e.m.; data were analysed and compared using Repeated measure Two-Way ANOVA and Bonferroni multiple comparisons post-hoc test; *p<0.05, compared with control.
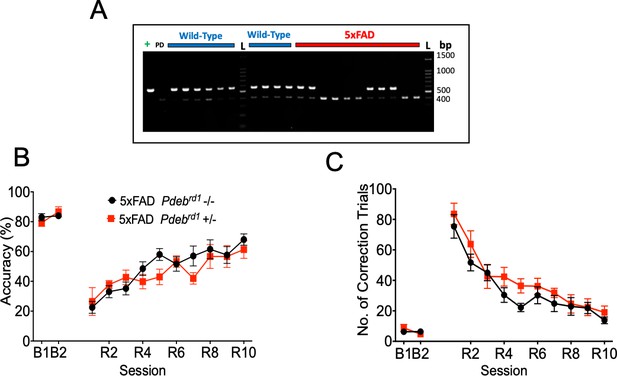
PDEBrd1 mutation effect in the behaviour of 5xFAD mice.
(A) Representative genotyping results. 5xFAD mice (red) and WT controls (blue) were genotyped as described. In the first lane, there is a positive control (+) for the 550 bp Pdebrd1 mutant allele obtained from FVB/NJ mice, followed by a WT control for the 400 bp wild-type PDEB allele (WT). Ladder (L). (B) 10 months old 5xFADPdebrd1-/- and 5xFADPdebrd1+/- evaluated in the PVD task.
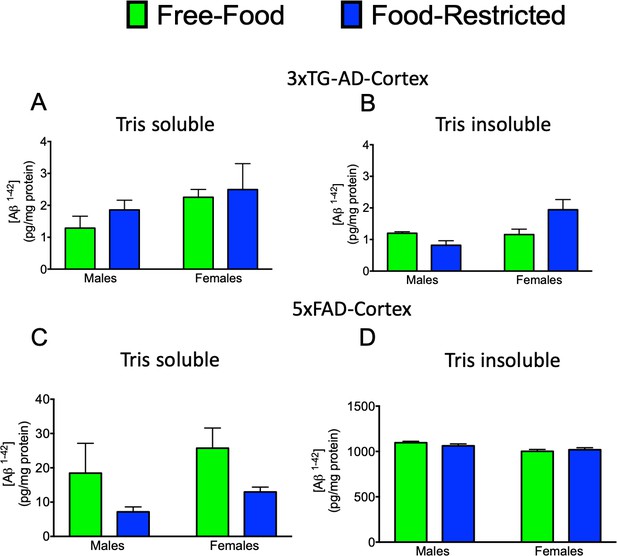
The effect of mild caloric restriction on Aβ(1–42) levels in male and female 3xTG and 5xFAD mice at 6 months of age.
(A) 3xTG Tris-soluble (p=0.921 for males and 0.999 for females); (B) 3xTG-AD Tris-insoluble (p=0.965 for males and 0.1512 for females); (C) 5xFAD Tris-soluble (p=0.163 for males and 0.367 for females); (D) 5xFAD Tris-insoluble (p=0.8271 for males and 0.991 for females);. At least three extracts obtained from each genotype/sex were analysed by Aβ(1–42) ELISA kit.
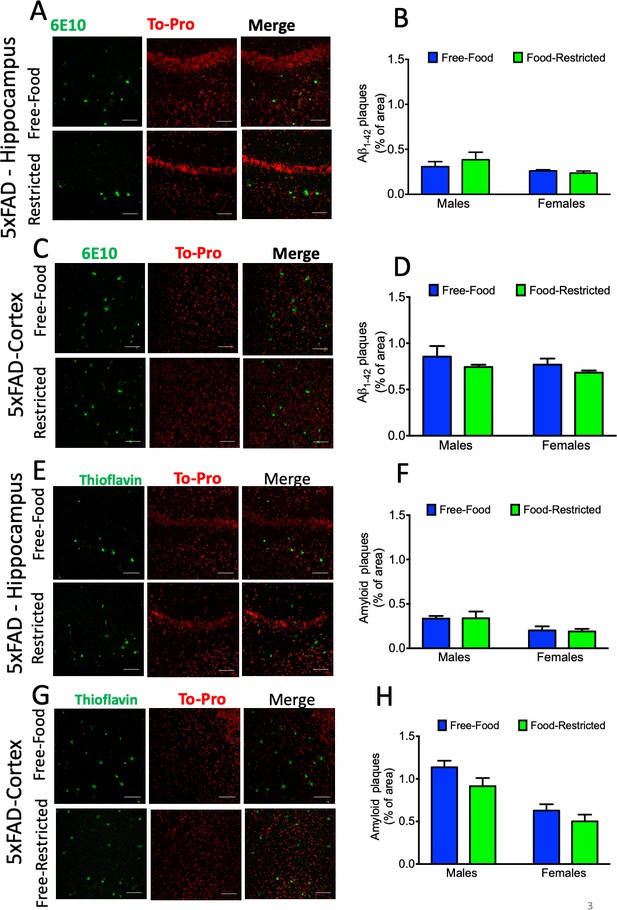
The effect of mild caloric restriction on amyloid pathology in male and female 5xFAD mice at 6 months of age.
(A) Representative images of amyloid-beta 6E10 antibody and ToPro-3 (20x magnification; scale bar = 100 µm) and (B) quantification (mean ± SEM) of 6E10 immunoreactivity in the hippocampus (CA1) of mildly food-restricted and free food 5xFAD mice at 6 months of age (p=0.978 for males and >0.999 for females); (C) Representative images of amyloid-beta 6E10 antibody and ToPro-3 (20x magnification; scale bar = 100 µm) and (D) quantification (mean ± SEM) of 6E10 immunoreactivity in the cortices of mildly food-restricted and free food male 5xFAD mice at 6 months of age (p=0.931 for males and 0.976 for females); (E) Representative images of Thioflavin-S (Thio-S) and ToPro-3 (20x magnification; scale bar = 100 µm) and (F) quantification (mean ± SEM) of Thio-S in the hippocampus (CA1b) of mildly food-restricted and free food male 5xFAD mice at 6 months of age (p>0.999 for males and p>0.999 for females); (G) Representative images of Thioflavin-S and ToPro-3 and (H) quantification (mean ± SEM) of Thio-S in the cortices of mildly food-restricted and free food male 5xFAD mice at 6 months of age (p=0.076 for males and p=0.2993 for females). Data was compared by two-way ANOVA followed by Sidak’s multiple comparisons test. At least four samples obtained from each genotype/sex/brain region were analysed.
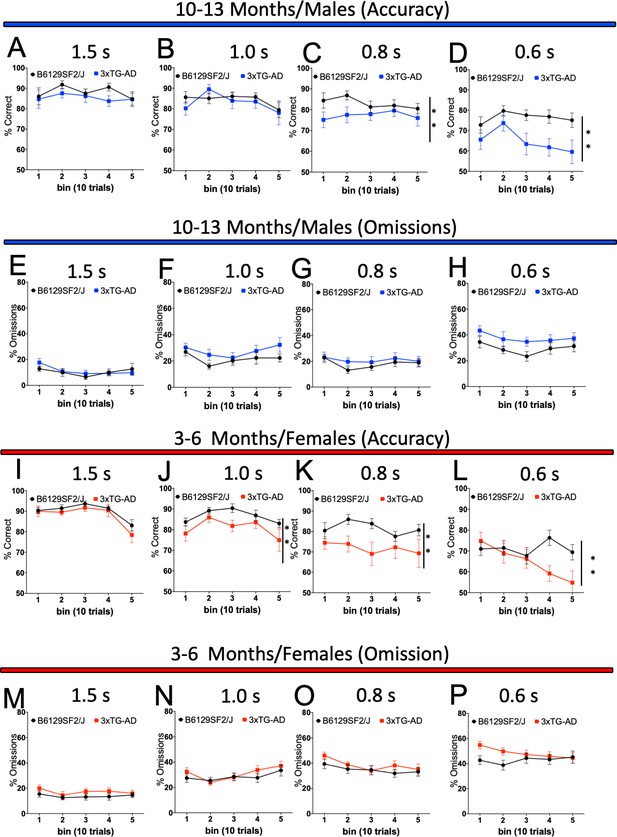
Sustained attention (vigilance) of 3xTG-AD male and female mice during the 5-CSRTT probe trials.
Response accuracy and omissions in Wild-type and 3xTG-AD male (A-D for accuracy and E-H for omission) and female (I-L for accuracy and M-P for omission) mice were analysed across 10-trials blocks within the daily sessions of 50 trials with different stimulus durations. Results are presented as means ± s.e.m.; data were analysed and compared using Repeated measure Two-Way ANOVA and Bonferroni multiple comparisons post-hot test; **p<0.01 compared with control.
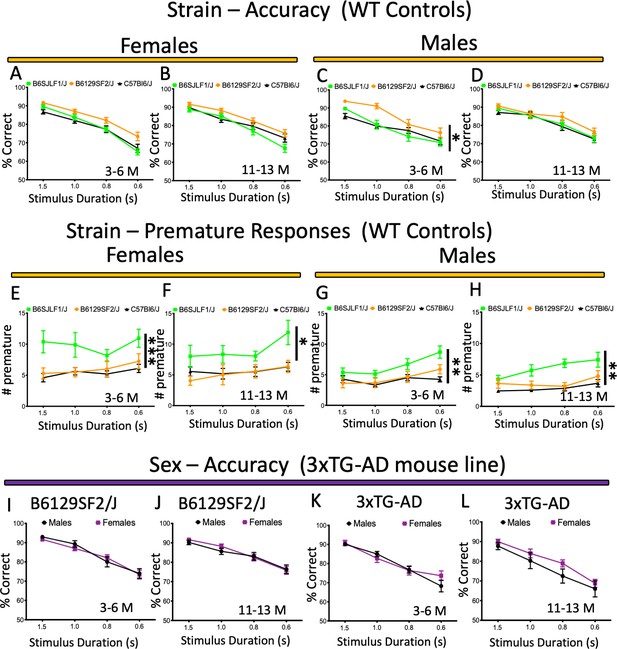
Performance and response measures of male and female mice during the 5-CSRTT probe trials.
(A-D) Strain/mouse background comparison (accuracy) of female and male Wild-type controls (B6129SF2/J, B6SJLF1/J, C57Bl6/J) at 3–6 and 11–13 months of age; (E-H) Strain/mouse background comparison (premature responses) of female and male Wild-type controls (B6129SF2/J, B6SJLF1/J, C57Bl6/J) at 3–6 and 11–13 months of age; (I-L) Sex comparison (accuracy) of B6129SF2/J and 3xTG-AD females and males at 3–6 and 11–13 months of age. Results are presented as means ± s.e.m.; data were analysed and compared using Repeated measure Two-Way ANOVA and Bonferroni multiple comparisons post-hot test; *p<0.05, **p<0.01 and ***p<0.001 compared with control.
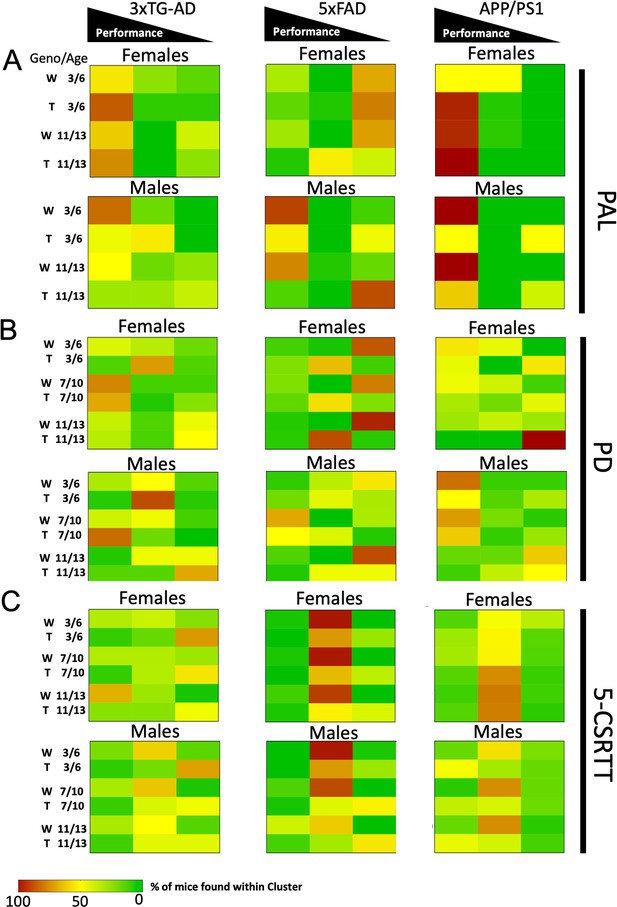
Heatmap visualisations of k-mean group membership across experiments.
The percentage of group representation per strain is shown by the cell color. Cell color closer to red indicates a higher representation of mice in the k-mean grouping. Mice are divided by sex (male and female), genotype (W for wild type and T for transgenic), and age (3–6, 7–10, and 11–13). Analysis of the PAL (A) task data uncovered early group membership variability in the APP/PS1 mouse line, as well as later group membership differences in the male 5xFAD mice. In the PD (B) experiment, only APP/PS1 transgenic mice showed an increase in membership of the low performing group compared to wildtype. See also S5. In the 5-CSRTT (C), transgenic 3xTG-AD and 5xFAD mice were found to shift to the lowest performing group of mice.
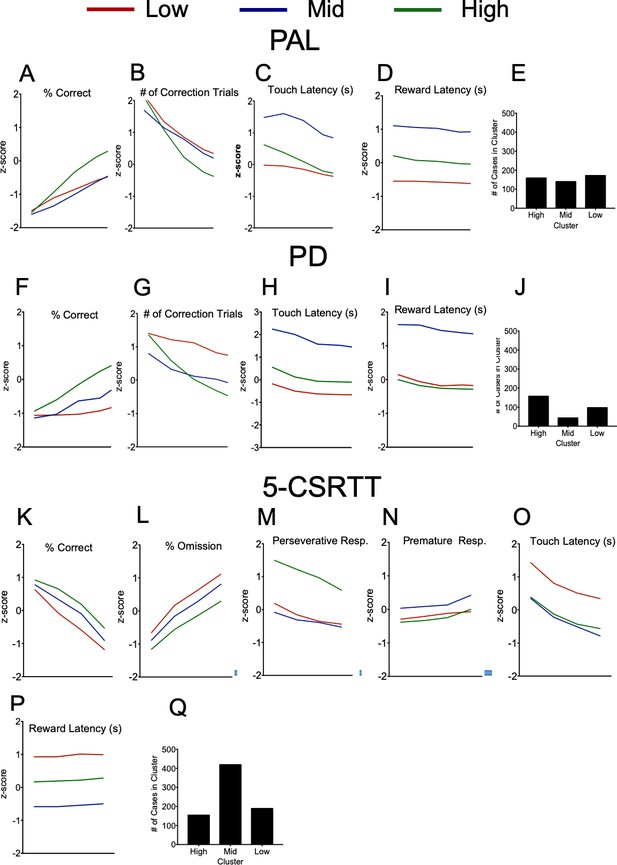
Results of longitudinal clustering analysis of PAL, PD and 5-CSRTT data. z-scores for parameters assessed during the 5-CSRTT, sorted by cluster.
For PAL – (A) Accuracy across blocks by cluster. (B) Number of correction trials across blocks by cluster. (C) Correct Touch latency across blocks by cluster (D) Reward latency across blocks by cluster. (E) Number of mice belonging to each cluster across strain, age, sex and genotype. The high performing cluster is characterised by low errors and high accuracy. The middle performing cluster is characterised by middle accuracy and errors, as well slow response times. The low performing cluster is characterized by low accuracy and high errors. For PD - (F) Response accuracy across trials by cluster. (G). Errors committed across trials by cluster. (H) Response latency across trials by cluster. (I) Reward collection latency across trials by cluster. The high performing cluster is characterised by low errors and high accuracy. The middle performing cluster is characterised by middle accuracy and errors, and slow response times. The low performing cluster is characterised by low accuracy and high errors. (J) Number of mice belonging to each cluster across strain, age, sex and genotype. For 5-CSRTT - (K) Response accuracy across trials as stimulus duration is reduced. (L) Omission errors committed as stimulus duration is reduced. (M) Perseverative responses as stimulus duration is reduced. (N) Premature responses as stimulus duration is reduced. (O) Correct response latency as stimulus duration is reduced. (P) Reward collection latency as stimulus duration is reduced. The high performing cluster is characterised by low omissions, high accuracy, but high number of perseverative responses. The middle performing cluster is characterised by middle accuracy and omission, and increased number of premature responses. The low performing cluster is characterised by low accuracy, high omission, and slow response times (Correct touch latency and reward collection latency) (Q) Number of mice belonging to each cluster across strain, age, sex and genotype.
Tables
5-CSRTT analyses.
Summary of conventional genotype analyses on the 5-CSRTT task. Summary results were based on simple 2 (genotype) x 4 (stimulus duration) split-plot ANOVA. Impairment or Facilitation was determined by looking for a significant genotype effect or interaction. (3x – 3xTG-AD, 5x – 5xFAD and APP – APP/PS1) mouse lines. Impairment (↓), Improvement (↑) No Effect (-). See also Supplementary file 2 and 5.
| Accuracy | Omissions | Premature Responses | Perseverative Responses | Touch Latency | Reward Latency | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sex | Age (months) | 3x | 5x | APP | 3x | 5x | APP | 3x | 5x | APP | 3x | 5x | APP | 3x | 5x | APP | 3x | 5x | APP | ||||||
| Female | 3-6 | ↓ | - | ↑ | ↓ | - | ↑ | ↑ | - | - | ↑ | - | ↓ | ↓ | - | ↑ | - | ↓ | - | ||||||
| 7-10 | ↓ | ↓ | - | - | - | ↑ | - | - | - | ↑ | ↑ | - | ↓ | - | - | - | ↓ | - | |||||||
| 11-13 | ↓ | ↓ | - | - | - | - | - | - | - | ↑ | ↑ | - | ↓ | - | - | - | ↓ | - | |||||||
| Male | 3-6 | ↓ | - | - | - | - | - | - | - | - | - | - | ↓ | ↓ | - | - | ↓ | ↓ | - | ||||||
| 7-10 | ↓ | - | - | - | - | - | - | - | - | - | ↑ | ↓ | ↓ | ↓ | - | ↓ | ↓ | - | |||||||
| 11-13 | ↓ | ↓ | ↑ | ↓ | ↓ | - | - | - | - | - | ↑ | ↓ | ↓ | ↓ | - | - | ↓ | ↓ | |||||||
PD and PAL analyses.
Summary of conventional genotype analyses on the PD and PAL tasks. Summary results were based on simple 2 (genotype) x 4 (stimulus duration) split-plot ANOVA. Impairment or Facilitation was determined by looking for a significant genotype effect or interaction. (3x – 3xTG-AD, 5x – 5xFAD and APP – APP/PS1) mouse lines. Impairment (↓), Improvement (↑) No Effect (-). See also Supplementary file 3, 4, 6, 7.
| Accuracy | Correction Trials | Touch Latency | Reward Latency | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Task | Sex | Age (months) | 3x | 5x | APP | 3x | 5x | APP | 3x | 5x | APP | 3x | 5x | APP | |
| PD | Female | 3-6 | - | - | ↓ | - | ↓ | ↓ | ↓ | ↓ | ↑ | - | ↓ | ↑ | |
| 7-10 | - | ↓ | - | - | ↓ | ↓ | ↓ | ↓ | ↑ | - | ↓ | ↑ | |||
| 11-13 | - | - | - | - | - | - | - | ↓ | ↑ | - | ↓ | - | |||
| Male | 3-6 | ↑ | - | ↓ | ↑ | - | - | - | - | - | - | - | - | ||
| 7-10 | - | - | - | - | - | - | - | - | - | - | ↓ | - | |||
| 11-13 | - | - | - | - | - | - | ↑ | ↓ | - | - | ↓ | - | |||
| PAL | Female | 3-6 | - | - | ↑ | - | ↓ | ↑ | - | - | ↑ | ↑ | ↓ | ↑ | |
| 11-13 | - | - | - | - | ↓ | - | - | ↓ | - | ↑ | ↓ | ↓ | |||
| Male | 3-6 | ↓ | - | ↓ | ↓ | ↓ | ↓ | ↓ | ↑ | ↑ | ↓ | ↓ | - | ||
| 11-13 | - | ↓ | - | - | ↓ | - | - | - | - | ↓ | ↓ | - | |||
p-values from Fisher Exact Test for K-Mean Group.
Group Differences between wildtype and Transgenic mice across behavioural experiments. Fisher’s exact test was conducted to compare the % membership of the high, mid, and low k-mean groups between wildtype and transgenic mice for each strain, sex, and age. All p-values have been adjusted with the Benjamini and Hochberg Correction.
| 3xTG-AD | 5xFAD | APP/PS1 | ||||||
|---|---|---|---|---|---|---|---|---|
| Task | Age | Female | Male | Female | Male | Female | Male | |
| 5-CSRTT | 3-6 | .03 | .01 | .03 | .06 | .17 | .13 | |
| 7-10 | .19 | .06 | .02 | .01 | .77 | .07 | ||
| 11-13 | .01 | .39 | .02 | <0.001 | 1.00 | .19 | ||
| PD | 3-6 | .34 | .25 | <0.001 | .92 | .02 | .69 | |
| 7-10 | .81 | .16 | <0.001 | .06 | .42 | .61 | ||
| 11-13 | 1.00 | .36 | <0.001 | .03 | .18 | .81 | ||
| PAL | 3-6 | .59 | .19 | .33 | .19 | .06 | .03 | |
| 11-13 | .77 | .95 | .03 | .01 | 1.00 | .19 | ||
| Reagent type (species) or Resource | Designation | Source or reference | Identifiers | Additional Information |
|---|---|---|---|---|
| Antibody | 6E10 Primary Antibody (Human Monoclonal) | Covance | RRID:AB_2564652 Lot#: D13EF01399 Cat#: SIG-39320 | IF (1:200) |
| Antibody | 488 Goat Anti-Mouse Secondary Antibody (Mouse Polyclonal) | Invitrogen | RRID:AB_2534069 Cat#: A-11001 | IF(1:1000) |
| Commercial Assay Kit | Amyloid Beta 42 Human ELISA Kit – Ultrasensitive | Invitrogen | Cat#:KHB3544 | |
| Strain | B6.Cg-Tg[APPswe,PSEN1dE9]85Dbo/Mmja (Mouse, Male, Female) | Jackson Laboratories | RRID:MGI:034832-JAX Stock#: 034832-JAX | |
| Strain | (B6;129-Psen1tm1MpmTg[APPSwe,tauP301L-1Lfa 0 (Mouse, Male, Female) | Jackson Laboratories | RRID:MGI:101045-JAX Stock#: 101045-JAX | |
| Strain | B6SJL-Tg(APPSwFlLon,PSEN1*M146L*L286V)6799Vas/Mmja (Mouse, Male, Female) | Jackson Laboratories | RRID:MGI:034840-JAX Stock#: 034840-JAX | |
| Strain | C57BL/6 (Mouse, Male, Female) | Jackson Laboratorie | RRID:MGI:000664-JAX Stock#: 000664 | |
| Strain | B6129SF1/J(Mouse, Male, Female) | Jackson Laboratorie | RRID:MGI:101043 Stock#: 101043 | |
| Strain | B6SJLF1/J(Mouse, Male, Female) | Jackson Laboratorie | RRID:MGI:100012 Stock#: 100012 | |
| Software | ABET II Touch | Lafayette Neuroscience | Model#: 89505 | |
| Software | Spotfire | TIBCO | https://www.tibco.com/products/tibco-spotfir | |
| Software | Touchscreen Quality Control Syste | BrainsCAN | https://github.com/srmemar/Mousebytes-QualityControl |
Additional files
-
Supplementary file 1
The N values for each group of animals used in this study.
- https://cdn.elifesciences.org/articles/49630/elife-49630-supp1-v2.xlsx
-
Supplementary file 2
Summary of split-plot ANOVA of all behavioural measures for each genotype for 5-CSRTT.
- https://cdn.elifesciences.org/articles/49630/elife-49630-supp2-v2.xlsx
-
Supplementary file 3
Summary of split-plot ANOVA of all behavioural measures for each genotype for PAL.
- https://cdn.elifesciences.org/articles/49630/elife-49630-supp3-v2.xlsx
-
Supplementary file 4
Summary of split-plot ANOVA of all behavioural measures for each genotype for PD.
- https://cdn.elifesciences.org/articles/49630/elife-49630-supp4-v2.xlsx
-
Supplementary file 5
Summary of ANOVA analyses characterising genotype specific effects for 5-CSRTT.
- https://cdn.elifesciences.org/articles/49630/elife-49630-supp5-v2.xlsx
-
Supplementary file 6
Summary of ANOVA analyses characterising genotype specific effects for PAL.
- https://cdn.elifesciences.org/articles/49630/elife-49630-supp6-v2.xlsx
-
Supplementary file 7
Summary of ANOVA analyses characterizing genotype specific effects for PD.
- https://cdn.elifesciences.org/articles/49630/elife-49630-supp7-v2.xlsx
-
Supplementary file 8
Vigilance analysis characterising mice performance across blocks of 10 trials in 5-CSRTT.
- https://cdn.elifesciences.org/articles/49630/elife-49630-supp8-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/49630/elife-49630-transrepform-v2.pdf





