Human cytomegalovirus interactome analysis identifies degradation hubs, domain associations and viral protein functions
Figures
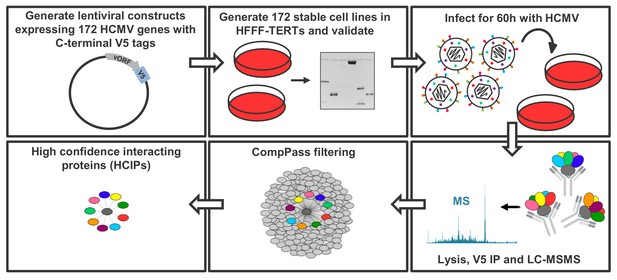
Schematic of the IP strategy.
IP samples were generated and analysed in technical duplicate, using the method originally described in Huttlin et al. (2017); Huttlin et al. (2015) and discussed in detail in the Materials and methods section. For 153 baits with zero or one transmembrane (TM) region predicted by Uniprot, an NP40-based lysis buffer was used; for 18 baits with >1 TM region, a digitonin-based buffer was used, as this has previously been demonstrated to improve identifications of interacting proteins (‘prey’) (Babu et al., 2012) (Supplementary file 1B). Each dataset was scored separately using the CompPASS algorithm (Huttlin et al., 2015; Sowa et al., 2009) to better model detergent-specific variation in IP-MS background. Data reported for each prey protein in every IP include: (a) the number of peptide spectral matches (PSMs), averaged between technical replicates; (b) an entropy score, which compares the number of PSMs between replicates to eliminate proteins that are not detected consistently; (c) a z-score, calculated in comparison to the average and standard deviation of PSMs observed across all IPs; and (d) a normalised WD (NWD) score. The NWD score addresses whether (i) the protein is detected across all IPs, and (ii) whether it is detected reproducibly among replicates. It was calculated as described in Behrends et al. (2010) using the fraction of runs in which a protein was observed, the observed number of PSMs, the average and standard deviation of PSMs observed for that protein across all IPs, and the number of replicates (1 or 2) containing the protein of interest. NWD scores were normalised so that the top 2% earned scores of ≥1.0. Stringent filters were applied to remove inconsistent and low-confidence protein identifications across all IPs and thus minimise both false protein identifications and associations (Huttlin et al., 2015). These included: (a) a minimum PSM score of 1.5 (i.e. ≥3 peptides per protein across both replicates); (b) an entropy score of ≥0.75; and (c) an NWD or z-score in the top 2%. Previous studies have estimated a 5% false discovery rate when employing a similar strategy with a top NWD score cutoff of 2% (Sowa et al., 2009). Interactions passing these criteria are named ‘high confidence interacting proteins’ (HCIPs) (Supplementary file 2B), and were used in all subsequent analyses. For added stringency, the supervised learning algorithm CompPass Plus was employed. This additionally assessed batch variations, overall spectral counts, unique peptide counts and protein detection frequency. Shannon entropy quantified a protein’s consistency of detection across technical duplicate LC-MS analyses, removing inconsistent protein identifications (Huttlin et al., 2017). CompPass plus was developed for interactomes with ≥96 baits and in the present study was only applied to the 153 baits solubilized in NP40. Interactions that passed CompPass filters, had CompPass Plus p(Interactor)>0.75 and in which the prey was identified by ≥2 unique peptides were considered as very high confidence interacting proteins (VHCIPs). These are indicated in green shading in Supplementary file 2B. To facilitate global analysis of all data, and because digitonin-solubilised interactions were not analysed using CompPass plus, HCIPs as opposed to VHCIPs were examined for the remainder of this study. The identification of an interacting protein as a VHCIP nevertheless adds additional confidence that the interaction observed is likely to be genuine.
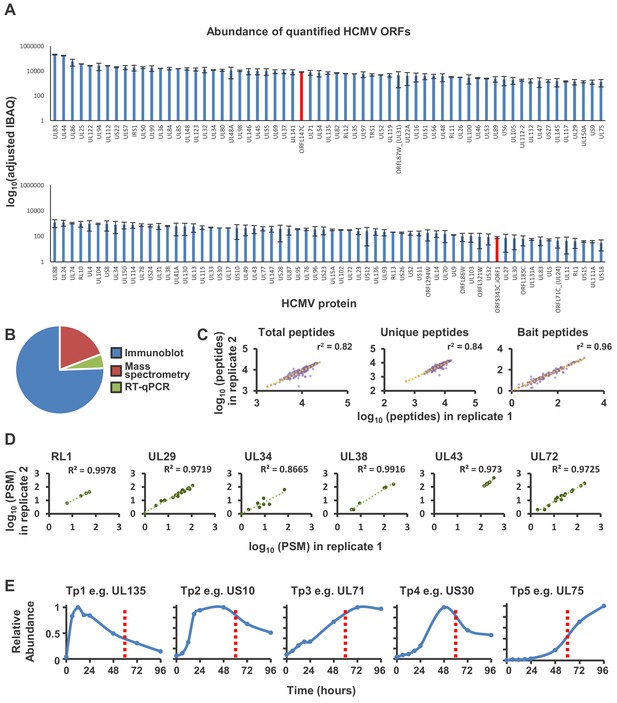
Further details of the interactome.
(A) Abundance of 127 quantified canonical and non-canonical HCMV ORFs. The intensity-based absolute quantification (IBAQ) method was adapted for data from two whole cell analyses of HCMV infection at 24, 48 and 72 hr PI (Fielding et al., 2017; Weekes et al., 2014) as described in the Materials and methods section. Normalised average IBAQ values + /- range are plotted for proteins quantified in both analyses. Data is split across two graphs to improve legibility. For proteins only quantified in one of the two analyses, data are shown in Supplementary file 1A. ORFL147C was the most abundant non-canonical ORF, and ORFS343C.iORF1 (referred to in the manuscript as ORFS343C) was one of the least abundant non-canonical ORFs (shown in red). Both were included as baits in the interactome. (B) Proportion of baits whose expression was verified by IB, MS or RT-qPCR (see also Supplementary file 1B). Experiments were performed in advance of the interactome study, using uninfected lysates to validate bait expression. MS was used if the bait was not detected by IB, and RT-qPCR was used if the bait was not identified by MS. Expression of 78% of baits was further validated in the interactome itself, including 9/18 baits initially only observed by RT-qPCR. These nine baits were included in the ‘detected by MS’ category. For the nine baits whose expression was only ever observed by RT-qPCR, four were small proteins of 47–111 amino acids, with 1–4 theoretically observable peptides. None of these four proteins were previously detected in two systematic proteomic analyses of HCMV infection (Nightingale et al., 2018; Weekes et al., 2014). In the same studies, a further 4/9 viral proteins were only detected by a median of 0–2 peptides, and 2/4 had multiple transmembrane domains, suggesting that detection may have been limited by protein abundance or hydrophobicity. Nevertheless, known protein prey of these nine viral baits were detected in the interactome, for example the interaction between the small capsomere-interacting protein UL48A and major capsid protein UL86, and membrane protein US18 and natural killer cell cytotoxicity receptor 3 ligand 1 (NCR3LG1) (Supplementary file 3, (Fielding et al., 2017). Data for the nine baits that were only validated by RT-qPCR was therefore included in this interactome. Only expression of UL136 could not be validated by any method, despite the use of primers that recognized both a short and full-sized amplicon (Supplementary file 1E) and this bait was excluded from further analysis. (C) Correlation of the number of total, unique and bait peptides from each protein identified in replicates 1 and 2. ‘Unique peptides’ refers to peptides that exist only in one protein from the human or HCMV proteomes. ‘Bait peptides’ refer to peptides derived from the bait protein for each IP (i.e. from US1 in the US1 IP; from UL29 in the UL29 IP etc). All data for this figure are also shown in Figure 1—figure supplement 1—source data 1. (D) Reproducibility of interactome analyses. In this interactome, as previously described (Huttlin et al., 2015), biological replicates were pooled and samples analysed in technical duplicate to solve certain technical issues. Specifically, due to the potential for carry-over of peptides between adjacent injections of different IP samples (even with two washes between each sample), it was necessary to use consistency of detection of prey as a measure of confidence in bait-prey interaction. To electronically filter out carry-over contaminants, an entropy score (described in ‘Data analysis’ in Materials and methods) compared the number of peptide-spectrum matches (PSM) between replicate injections and eliminated prey that were not detected consistently. It was therefore important that replicate injection material was as similar as possible to ensure that this filter was efficacious (see also Materials and methods). The CompPass algorithm (Huttlin et al., 2015; Sowa et al., 2009) was developed based on this specific protocol. To gain a measure of variability between biological replicate samples, six IP-MS experiments were repeated with independent analysis of each biological replicate. In this figure, PSM are shown for HCIPs predicted by the interactome for each of the baits. For example, the interactome predicted four HCIPs for RL1: CUL4A, CUL4B, DDA1 and RL1 itself (Figure 2, Supplementary File 2B). All data for this figure are also shown in Figure 1—figure supplement 1—source data 2. (E) Viral proteins expressed at 60 hr PI. Profiles of typical viral proteins from each of five temporal classes Tp1-Tp5 are shown (adapted from Weekes et al., 2014). At 60 hr PI, peptides from 139/139 quantified canonical HCMV proteins and 14/14 quantified non-canonical ORFs were detected.
-
Figure 1—figure supplement 1—source data 1
Correlation of the number of total, unique and bait peptides from each protein identified in replicates 1 and 2.
- https://cdn.elifesciences.org/articles/49894/elife-49894-fig1-figsupp1-data1-v2.xlsx
-
Figure 1—figure supplement 1—source data 2
Reproducibility of interactome analyses.
- https://cdn.elifesciences.org/articles/49894/elife-49894-fig1-figsupp1-data2-v2.xlsx
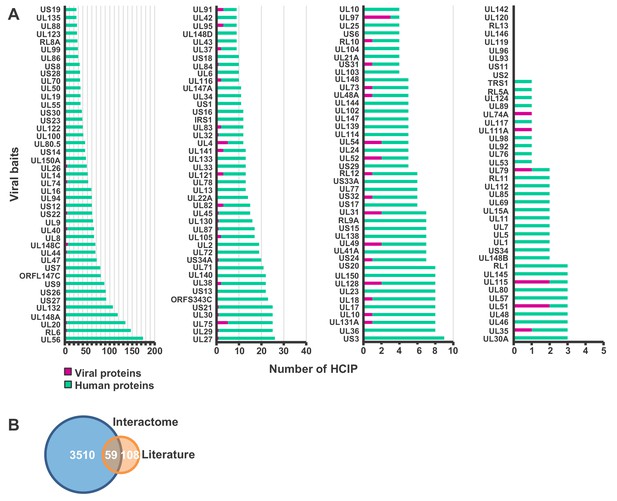
Further details of interactions.
(A) Number of HCIPs per bait excluding bait-bait interactions. Four graphs with different x-axis scales illustrate the range of interacting viral or cellular proteins per bait. Gridlines are displayed at each log interval. Full data is shown in Supplementary file 2A. (B) Validation of the interactome data from BioGRID, IntAct, Uniprot, MINT, and Virus Mentha (Calderone et al., 2015; Chatr-Aryamontri et al., 2013; Licata et al., 2012; Orchard et al., 2014). 167 virus-virus and virus-host interactions were identified from these databases. Of these, 127 were identified in unfiltered data from the HCMV interactome, and 59 passed the stringent scoring thresholds employed (see also Supplementary file 3). Full details of all HCIPs are given in Supplementary file 2.
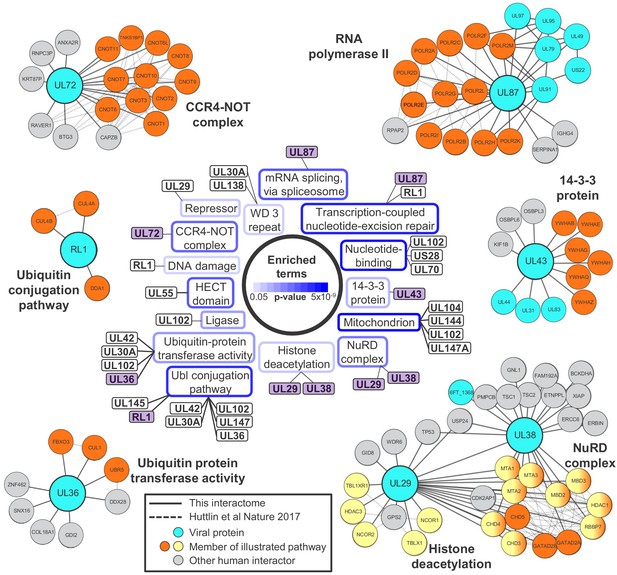
Systematic analysis of interactome data predicts novel functions for viral proteins.
DAVID software with default settings (Huang et al., 2009) was applied to determine which pathways were enriched amongst all HCIPs in the interactome, in comparison to all human proteins as background. Benjamini-Hochberg adjusted p-values are shown as blue surrounds to each pathway enriched at p<0.05. Viral baits are linked to enriched pathways where > 33% of human interacting proteins belonged to a given pathway, and examples are shown around the outside of the figure. These examples are indicated in the central part of the figure by purple shading. For example, 6/9 (67%) human HCIPs for UL43 were part of the 14-3-3 protein family. Viral baits are shown as large turquoise circles, and interacting viral proteins as smaller turquoise circles. Members of enriched pathways are shown in orange or yellow (for NuRD complex and histone deacetylation, protein membership of both pathways is indicated by half-orange, half-yellow circles). Solid lines indicate interactions identified by this interactome, and dashed lines indicated interactions derived from human Bioplex 2.0 and subsequent unpublished data (Huttlin et al., 2017 and http://bioplex.hms.harvard.edu/downloadInteractions.php). Full data are shown in Supplementary file 4. As an alternative approach to highlight cellular functions that predominantly related to individual viral proteins, Figure 2—figure supplement 1 shows pathways with p<0.05 (after Benjamini-Hochberg adjustment) and for which > 33% of the identified cellular protein members of the pathway interacted with a given viral bait.
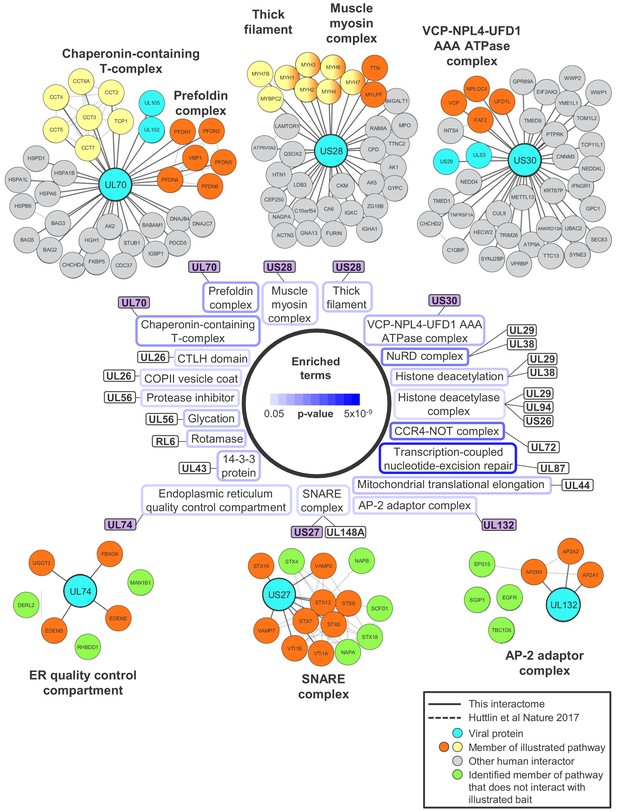
Pathways enriched with p<0.05 (after Benjamini-Hochberg adjustment) and for which > 33% of the identified components interacted with a given viral bait.
For example, all members of the thick filament/muscle myosin complex detected in this interactome interacted with US28 (100%). For the bottom three complexes (UL74, US27 and UL132), each viral bait interacted with a total of 52–107 proteins. For simplicity, only members of the illustrated pathway identified in this interactome are displayed. For example, 14 members of the SNARE complex were enriched in the interactome, of which nine interacted with US27 (64%). Green circles show members of a pathway that were detected in the interactome but did not interact with the bait. Other colouring is shown as described in the legend to Figure 2.
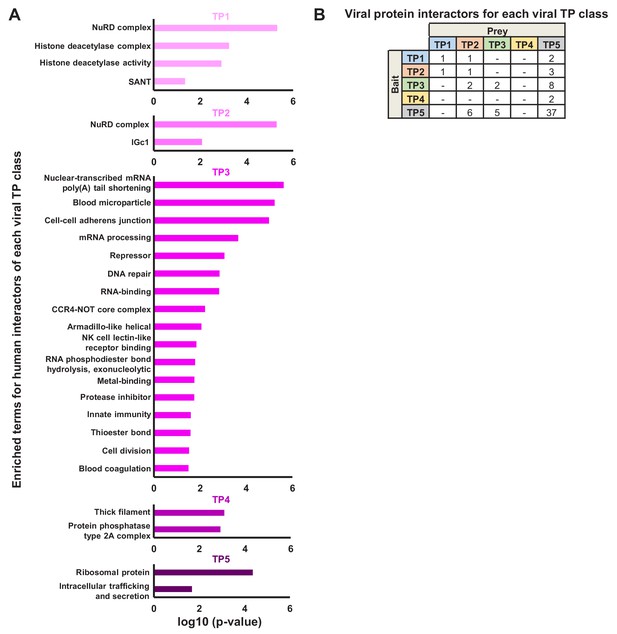
Further details of interactions according to viral protein temporal class.
(A) Functional enrichment of host HCIPs for each temporal class of viral bait. DAVID software with default settings (Huang et al., 2009) was applied to determine which pathways were enriched amongst human HCIPs of each of the Tp1-Tp5 classes of bait, in comparison to all human HCIP as background. Benjamini-Hochberg adjusted p-values are shown on the x-axis. (B) Temporal analysis of viral-viral protein interactions. The contingency table shows 71 viral protein-protein interactions for which both bait and prey had a defined Tp class (from Weekes et al., 2014).
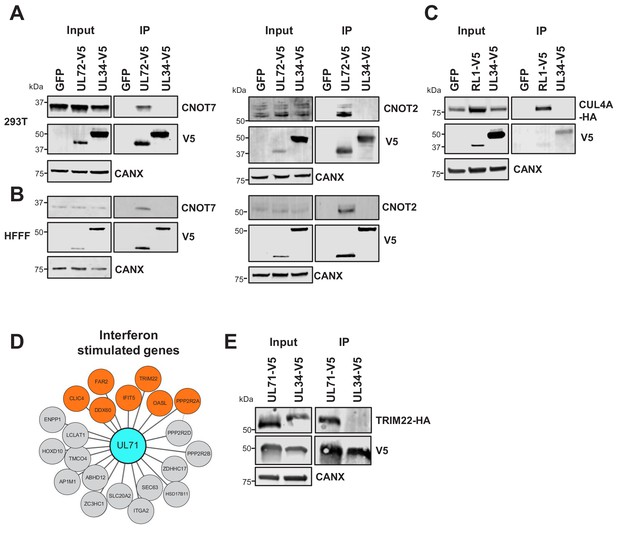
Validation of interactome data by co-IP.
(A) Co-IPs validating that UL72 interacts with CCR4-NOT Transcription Complex Subunits 7 and 2 (CNOT7 and CNOT2), conducted in HEK293T cells. For all experiments in this figure, left panels show an IB of 1–2% of input sample, and right panels shown an anti-V5 co-IP. Cells were transiently transfected with two plasmids, one expressing the C-terminally V5-tagged viral protein and the other expressing the C-terminally HA-tagged cellular prey. Bait proteins were detected with anti-V5, and prey with antibodies against CNOT7 or CNOT2 protein. Controls included GFP or the viral UL34 protein. CANX – calnexin loading control. This figure is representative of n = 1 experiment (CNOT2); n = 2 experiments (CNOT7). Expected sizes: CNOT7: 33 kDa; CNOT2: 52 kDa; CANX: 72 kDa; UL72: 44 kDa; UL34: 45 kDa. (B) Co-IPs validating that UL72 interacts with CNOT7 and CNOT2, conducted in HFFF-TERT cells overexpressing C-terminally V5-tagged UL72. Proteins were detected as described in (A). This figure is representative of n = 2 experiments (CNOT2); n = 1 experiment (CNOT7). Expected sizes: CNOT7: 33 kDa; CNOT2: 52 kDa; CANX: 72 kDa; UL72: 44 kDa; UL34: 45 kDa. (C) Co-IP validating the interaction between RL1 and CUL4A, conducted in HEK293T cells as described in (A), but with detection of CUL4A using anti-HA. This figure is representative of n = 4 experiments. Expected sizes: CUL4A: 77 kDa; RL1: 35 kDa; UL34: 45 kDa; CANX: 72 kDa. (D) HCMV UL71 interacted with multiple interferon-stimulated proteins, including TRIM22. (E) Co-IP validating the interaction between UL71 and TRIM22, conducted as described in (C). This figure is representative of n = 3 experiments. Expected sizes: TRIM22: 56 kDa; UL71: 40 kDa; UL34: 45 kDa; CANX: 72 kDa.
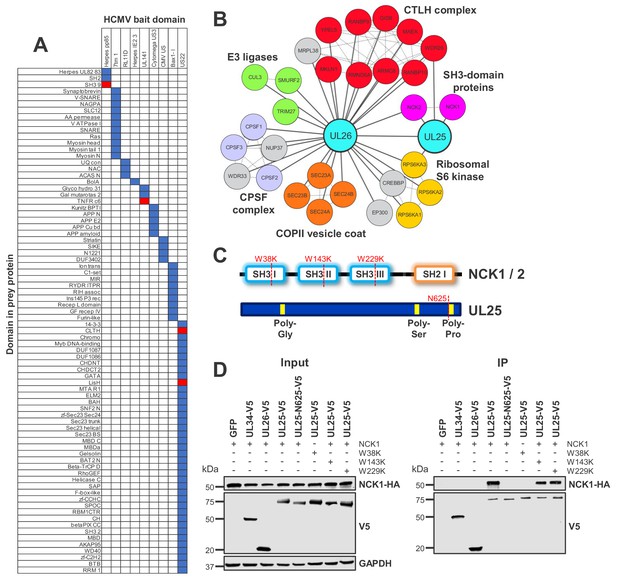
Interaction between UL25 and NCK1 identified by domain association analysis.
(A) Table depicting significant associations between domains present in HCMV baits (top) and human or viral prey (side). Pfam domains were mapped onto every bait and prey protein in the interactome (Finn et al., 2014). The numbers of interactions emanating from proteins containing each domain were tallied individually, along with the numbers of interactions linking each observed domain pair. Contingency tables were then populated to relate domain associations. For each pair, Fisher’s exact test determined the likelihood of a non-random association. p values were adjusted for multiple hypothesis testing (Benjamini and Hochberg, 1995). Coloured boxes identify domain pairs that associate at a 1% false discovery rate (FDR). Red boxes indicate domain pairs from this analysis discussed in the text. Domain associations are only shown for domains occurring in at least two viral proteins. Supplementary file 5 shows the full underlying data. (B) All HCIPs for UL25 and a subset of HCIPs for UL26 (full data are shown in Figure 4—figure supplement 1). DAVID analysis identified that members of the C-terminal to LisH (CTLH) complex and COPII vesicle coat proteins were enriched among UL26 HCIPs (Figure 2—figure supplement 1). Domain association analysis suggested that interaction of UL26 with CTLH components may occur via interaction of the viral US22 domain with either cellular CLTH or LisH domains (Supplementary file 5). Dashed lines represent human-human interactions derived either from Bioplex 2.0 as described in Figure 2 or from curated or experimental data in the STRING database. CPSF - Cleavage and polyadenylation specificity factor. (C) Schematic of NCK1 and UL25 protein structures, indicating the position of point mutations or truncation for (D). (D) Co-IP demonstrating that the UL25 proline-rich C-terminal domain associates with the first NCK1 SH3 domain, conducted as described in Figure 3. HEK293T cells were transiently transfected with the indicated plasmids, one expressing the C-terminally V5-tagged viral protein and the other expressing C-terminally HA-tagged NCK1. These proteins were detected with anti-V5 and anti-HA. Mutations or truncations of each gene are indicated in the figure and in (C). GAPDH – loading control. This figure is representative of n = 3 experiments. Expected sizes: NCK1: 43 kDa; UL25: 74 kDa; UL26: 21 kDa; GAPDH: 36 kDa.
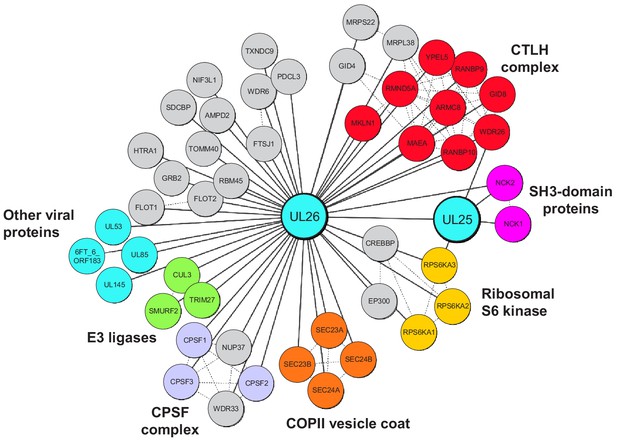
Full interaction data for UL25 and UL26, annotated as described in Figure 4B.
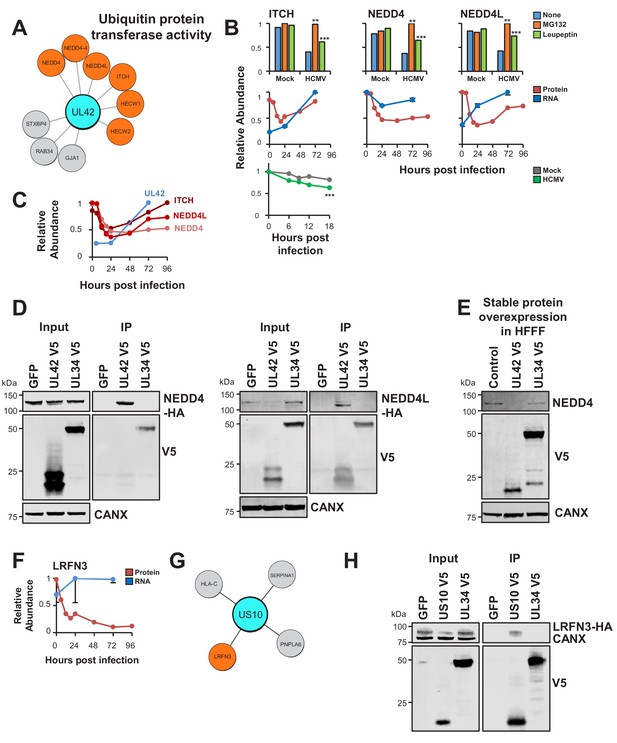
UL42 identified as a hub of E3 destruction by a combination of interactome and degradation data.
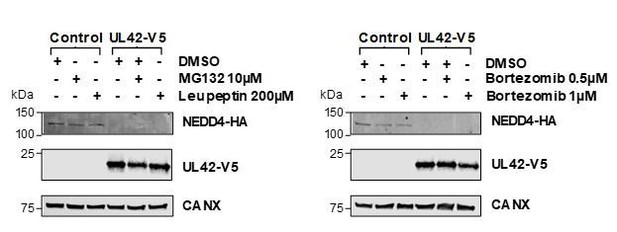
US10 interacts with LRFN3, which is rapidly downregulated from the PM during HCMV infection. (A) High-confidence cellular interactors of UL42. 57% of UL42 interactors exhibited ubiquitin protein transferase activity (Figure 2, counting NEDD4 only once). UL42 interacted with NEDD4, NEDD4 isoform four and NEDD4L, in addition to HECT, C2 and WW Domain Containing E3 Ubiquitin Protein Ligases HECW1 and 2. NEDD4-4: isoform 4 of NEDD4. (B) ITCH, NEDD4 and NEDD4L are degraded during early HCMV infection (data from Nightingale et al., 2018). Protein degradation was measured using three orthogonal tandem mass tag (TMT)-based proteomic screens. The first measured protein abundance throughout early infection in the presence or absence of inhibitors of the proteasome or lysosome. The second compared transcript and protein abundance over time to distinguish between degraded and transcriptionally regulated proteins. The third employed an unbiased global pulse-chase to compare the rates of protein degradation during HCMV infection against mock infection (NEDD4 and NEDD4L were not quantified in this latter screen). Benjamini-Hochberg adjusted Significance A values were used to estimate p-values in the top panels; **p<0.005, ***p<0.0005. Mean and SEM are shown for transcript quantitation (n = 3) in the middle panels. A p-value for the difference between rates of degradation is shown in the bottom panel; ***p<0.0005. All calculations and statistics are described in Nightingale et al. (2018). (C) UL42 transcript is expressed contemporaneously with NEDD4 and NEDD4L degradation. Protein profiles from Figure 5B (red colour, Nightingale et al., 2018) are overlaid with a UL42 transcript profile (blue colour, Stern-Ginossar et al., 2012). UL42 transcript was not detected in our previous RNAseq analysis (Nightingale et al., 2018). (D) Validation of interaction between UL42 and NEDD4 (left panel) and NEDD4L (right panel) by co-IP, conducted as described in Figure 3. HEK293T cells were transiently transfected with the indicated plasmids, one expressing the C-terminally V5-tagged viral protein and the other expressing C-terminally HA-tagged NEDD4 or NEDD4L. These proteins were detected with anti-V5 and anti-HA. This figure is representative of n = 2 experiments (NEDD4); n = 1 experiment (NEDD4L). Expected sizes: NEDD4: 104–149 kDa; NEDD4L: 96–111 kDa; UL42: 14 kDa; UL34: 45 kDa; CANX: 72 kDa. (E) UL42 was sufficient to degrade NEDD4. HFFF-TERTs expressing UL42 or controls were lysed and immunoblotted as indicated. Anti-NEDD4 was used to detect endogenous NEDD4. This figure is representative of n = 1 experiment. Expected sizes: NEDD4: 104–149 kDa; UL42: 14 kDa; UL34: 45 kDa; CANX: 72 kDa. (F) LRFN3 was rapidly downregulated from the PM during HCMV infection, in the presence of upregulated transcript (mean and SEM are shown for transcript quantitation (n = 3); data are from Nightingale et al., 2018). (G) HCIPs of US10, including LRFN3. (H) Validation of the interaction between US10 and LRFN3 by co-IP, conducted as described in Figure 3. Prey were detected using anti-HA. This figure is representative of n = 2 experiments. Expected sizes: LRFN3: 66 kDa; US10: 21 kDa; UL34: 45 kDa; CANX: 72 kDa.
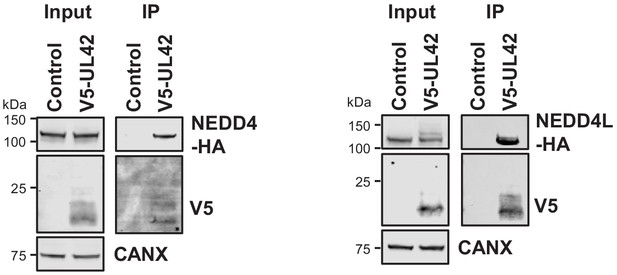
Validation of interaction between UL42 and NEDD4 (left panel) and NEDD4L (right panel) by co-IP, conducted as described in Figure 3.
HEK293T cells were transiently transfected with the indicated plasmids, one expressing N-terminally V5-tagged UL42 and the other expressing C-terminally HA-tagged NEDD4 or NEDD4L. These proteins were detected with anti-V5 and anti-HA. This figure is representative of n = 1 experiments. Expected sizes: NEDD4: 104–149 kDa; NEDD4L: 96–111 kDa; UL42: 14 kDa; CANX: 72 kDa.
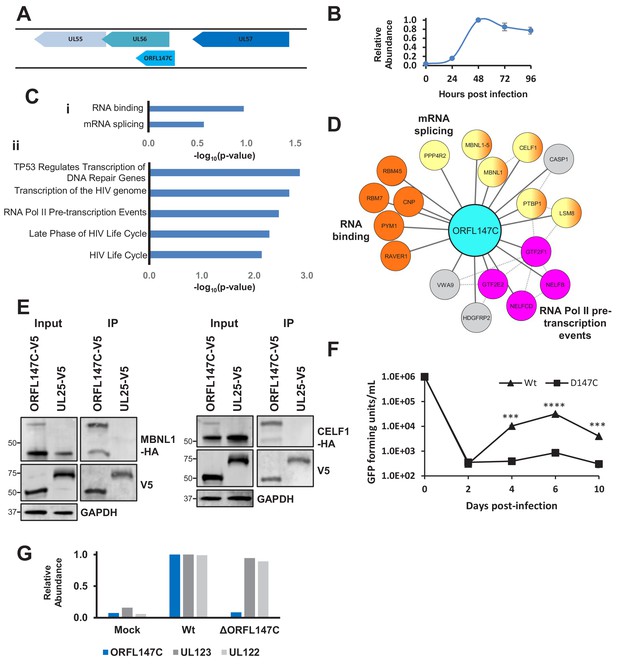
HCMV ORFL147C interactors function in RNA binding, splicing and transcription.
(A) Diagram of the ORFL147C coding sequence and relation to neighbouring viral genes. (B) Expression kinetics of ORFL147C, taken from Weekes et al. (2014). Data was taken from experiments WCL2 and WCL3, enabling assessment of 24, 48, 72 and 96 hr time points in biological duplicate. Error bars show range. Mean expression was normalised to a maximum of 1. (C) Enrichment analysis of 80 human HCIPs interacting with ORFL147C. (i) DAVID analysis using all human proteins as background. Benjamini-Hochberg adjusted p-values are shown. (ii) Reactome database analysis (Fabregat et al., 2018) showing results with a minimum of 4 entities per enriched pathway. Full details of interacting proteins are given in Supplementary file 7A-B. (D) A subset of HCIPs for ORFL147C (full data are shown in Figure 6—figure supplement 1). Dashed lines represent human-human interactions derived from Bioplex 2.0 as described in Figure 2, in addition to known interactions that had been experimentally determined or derived from curated data as part of the STRING database. (E) Validation of interaction between ORFL147C and MBNL1 and CELF1 by co-IP, conducted as described in Figure 3. HEK293T cells were transiently transfected with the indicated plasmids, one expressing the C-terminally V5-tagged viral protein and the other expressing C-terminally HA-tagged MBNL1 or CELF1. These proteins were detected with anti-V5 and anti-HA. GAPDH – calnexin loading control. This figure is representative of n = 1 experiment. Expected sizes: MBNL1: 33–42 kDa; CELF1: 50–55 kDa; ORFL147C: 50 kDa; UL25: 74 kDa; GAPDH: 36 kDa. (F) Growth analysis of an ORFL147C-deficient recombinant. The ORFL147C and wild-type viruses were HCMV strain Merlin recombinants in which the enhanced GFP (eGFP) gene was cloned as a 3’-terminal fusion with immediate-early gene UL36, with a self-cleaving P2A peptide releasing the reporter following synthesis. Insertion of GFP does not impede UL36 function in such recombinants (Nightingale et al., 2018). Cells were infected at a MOI of 1, and supernatants harvested and titred every two days. Cells were infected in biological duplicates, and each supernatant was titred in technical duplicates. Mean values are shown, and error bars represent SD. p-values for a difference between wild-type and ORFL147C-deficient virus were estimated using a two-tailed Student’s t-test. ***p<0.001, ****p<0.0001. This figure is representative of n = 2 experiments. All data for this figure are also shown in Figure 6—source data 1. (G) ORFL147C protein is not expressed during infection with the ORFL147C-deficient recombinant (MOI = 2, 48 hr post infection). Viral protein expression was analysed using tandem mass tag-based proteomics as previously described (Nightingale et al., 2018). ORFL147C protein was measured at the same level as during mock infection in cells infected with the ORFL147C-deficient recombinant, attributable to noise. All data for this figure are also shown in Figure 6—source data 2.
-
Figure 6—source data 1
Growth analysis of an ORFL147C-deficient recombinant.
- https://cdn.elifesciences.org/articles/49894/elife-49894-fig6-data1-v2.xlsx
-
Figure 6—source data 2
Tandem mass tag-based proteomics analysis of ORFL147C protein expression.
- https://cdn.elifesciences.org/articles/49894/elife-49894-fig6-data2-v2.xlsx
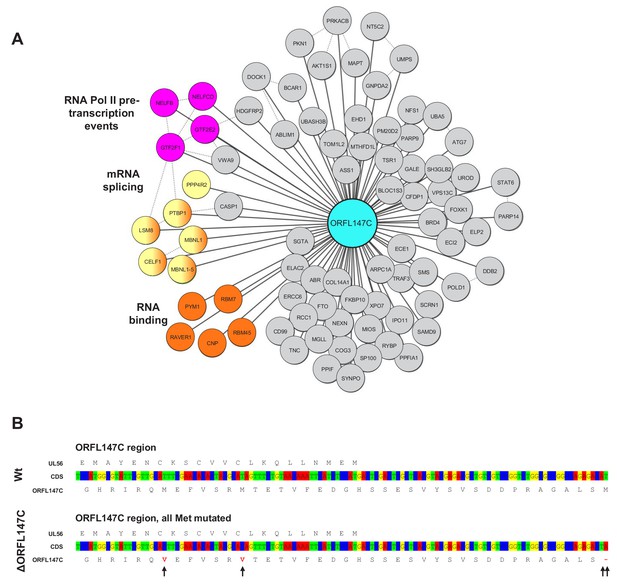
Further details of ORFL147C interactions, and construction of the ΔORFL147C virus.
(A) Full interaction data for ORFL147C, annotated as described in Figure 4B. (B) Construction of a viral ORFL147C deletion mutant. The three most N-terminal methionines in ORF147C were mutated without affecting the amino acid sequences coded by UL56. The N-terminal protein sequences of ORFL147C and UL56 are shown, in addition to corresponding coding sequences in the region of the mutations.
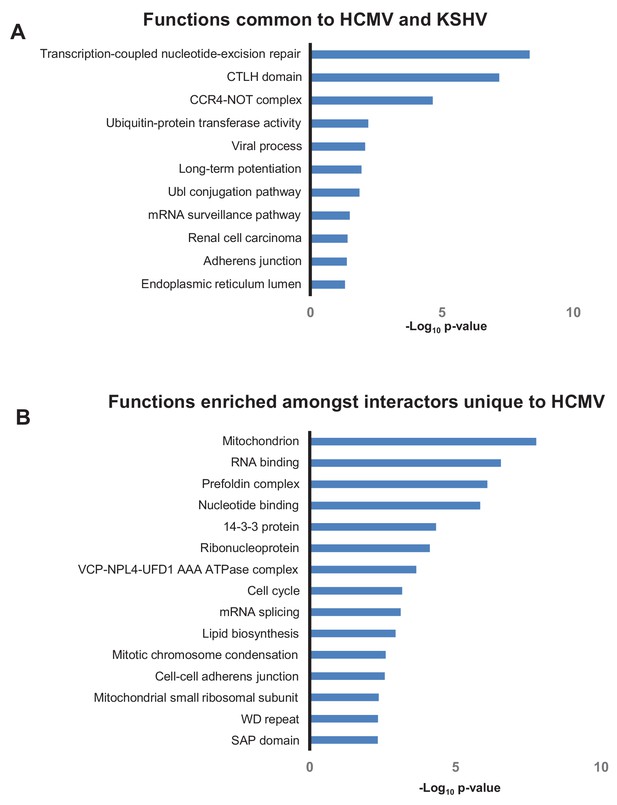
Overlap in functions targeted by different viruses.
(A) DAVID analysis of pathway enrichment among 176 HCIPs that interacted both with HCMV baits (this study) and KSHV baits (Davis et al., 2015), in comparison to all human proteins as background. Benjamini-Hochberg adjusted p-values are shown for each pathway. Full details of interacting viral and host proteins are given in Supplementary file 7A. (B) DAVID analysis of pathway enrichment among HCIPs that only interacted with HCMV but not KSHV baits, in comparison to all human proteins as background. As the KSHV interactome was performed in HEK293T cells as opposed to HFFFs, the list of HCMV HCIPs was first filtered to include proteins that were clearly detectable in HEK293Ts, using the list of ~50,000 unfiltered bait-prey interactions from KSHV to indicate protein expression (Davis et al., 2015). Subsequently, both high confidence interacting prey of KSHV baits, and first degree interactors of these prey from the human interactome, were excluded (Huttlin et al., 2017), to leave a list of proteins that only interacted with HCMV. Benjamini-Hochberg adjusted p-values are shown for each pathway. Full details of interacting viral and host proteins are given in Supplementary file 7B.
Tables
| Reagent type | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (HCMV) | HCMV Merlin | Stanton et al., 2010 | RCMV1111 | |
| Strain, strain background (HCMV) | HCMV Merlin UL36-GFP deltaORFL147C | This paper | RCMV2697 | Available from Dr Michael Weekes’ lab, University of Cambridge |
| Strain, strain background (HCMV) | HCMV Merlin UL36-GFP | Nightingale et al., 2018 | RCMV2582 | |
| Strain, strain background (Escherichia coli) | E. coli. (α-Select Silver Competent Cells) | Bioline | Cat#BIO-85026 | |
| Cell line (Homo-sapiens) | HFFF immortalised with human telomerase (HFFF-TERT) | McSharry et al., 2001 | ||
| Cell line (Homo-sapiens) | Human Embryonic Kidney 293 T cells | Menzies et al., 2018 | ATCC Cat#CRL-3216, RRID:CVCL_0063 | |
| Antibody | Anti-V5 Agarose Affinity Gel | Sigma-Aldrich | Cat#A7345; RRID:AB_10062721 | (30 µl/mL) |
| Antibody | Mouse monoclonal anti-GAPDH | R and D Systems | Cat#MAB5718; RRID:AB_10892505 | (1:10.000) |
| Antibody | Rabbit polyclonal anti-Calnexin | LifeSpan Biosciences | Cat#LS-B6881; RRID:AB_11186721 | (1:10.000) |
| Antibody | Rabbit monoclonal anti-HA (C29F4) | Cell Signaling Technologies | Cat#3724S; RRID:AB_1549585 | (1:1000) |
| Antibody | Mouse monoclonal anti-V5 | Thermo | Cat#R960-25; RRID:AB_2556564 | (1:5000) |
| Antibody | Rabbit polyclonal anti-CNOT2 | Novus Biologicals | Cat#NBP2-56034; RRID:AB_2801658 | (1:1000) |
| Antibody | Rabbit monoclonal anti-CNOT7 | Abcam | Cat#ab195587; RRID:AB_2801659 | (1:1000) |
| Antibody | Mouse monoclonal anti-NEDD4 | R and D Systems | Cat#MAB6218; RRID:AB_10920762 | (1:1000) |
| Antibody | IRDye 680RD goat anti-mouse IgG | LI-COR | Cat#925–68070, RRID:AB_2651128 | (1:10.000) |
| Antibody | IRDye 800CW goat anti-rabbit IgG | LI-COR | Cat#925–32211, RRID:AB_2651127 | (1:10.000) |
| Antibody | IRDye 680RD goat anti-rabbit IgG | LI-COR | Cat#926–68071; RRID:AB_10956166 | (1:10.000) |
| Antibody | IRDye 800CW goat anti-mouse IgG | LI-COR | Cat#926–32210; RRID:AB_621842 | (1:10.000) |
| Antibody | Human TruStain FcX | BioLegend | Cat#422302; RRID:AB_2818986 | 1:20 |
| Recombinant DNA reagent | pHAGE-pSFFV | Nightingale et al., 2018 | ||
| Recombinant DNA reagent | pDONR223 | Nightingale et al., 2018 | ||
| Recombinant DNA reagent | pDONR221-MBLN1 | Harvard PlasmID | Cat#HsCD00079833 | |
| Recombinant DNA reagent | pDONR221-CUGBP1 | Harvard PlasmID | Cat#HsCD00039403 | |
| Recombinant DNA reagent | pOTB7-CUL4A | Harvard PlasmID | Cat#HsCD00325140 | |
| Recombinant DNA reagent | pCMV-SPORT6-NEDD4L | Harvard PlasmID | Cat#HsCD00337956 | |
| Recombinant DNA reagent | pENTR223-NCK1 | Harvard PlasmID | Cat#HsCD00370605 | |
| Recombinant DNA reagent | pDONR223-CNOT2 | Harvard PlasmID | Cat#HsCD00080019 | |
| Recombinant DNA reagent | pHAGE-CNOT7 | Harvard PlasmID | Cat#HsCD00453329 | |
| Recombinant DNA reagent | PHAGE-P-CMVt-N-HA Nedd4 wt | Addgene | Cat#24124 | |
| Recombinant DNA reagent | pDONR221-LRFN3 | Harvard PlasmID | Cat#HsCD00041564 | |
| Sequence-based reagent | M13-F | GENEWIZ | PCR primers | GTAAAACGACGGCCAG |
| Sequence-based reagent | M13-R | GENEWIZ | PCR primers | CAGGAAACAGCTATGAC |
| Sequence-based reagent | pHAGE-pSFFV-Seq | This paper | PCR primers | CGCGCCAGTCCTCCGATTG |
| Sequence-based reagent | GAW-CMVp-F | This paper | PCR primers | GGGACAAGTTTGTACAAAAAAGCAGCTGAAGACACCGGGACCGATC |
| Sequence-based reagent | attB2-V5-R | This paper | PCR primers | GGGGACCACTTTGTACAAGAAAGCTGGGTTTACGTAGAATCAAGACCTAGGAGC |
| Peptide, recombinant protein | V5 Epitope Tag | Alpha Diagnostic International | Cat#SP-59199–5 | |
| Peptide, recombinant protein | Trypsin | Promega | Cat#V5111 | |
| Commercial assay or kit | BCA Protein Assay Kit | Thermo Fisher | Cat#23227 | |
| Commercial assay or kit | Micro BCA Protein Assay Kit | Thermo Fisher | Cat#23235 | |
| Commercial assay or kit | RNeasy Mini Kit | Qiagen | Cat#74104 | |
| Commercial assay or kit | Empore SPE Disks | Supelco | Cat#66883 U | |
| Commercial assay or kit | GoScript Reverse Transcriptase kit | Promega | Cat#A5001 | |
| Commercial assay or kit | Power SYBR Green PCR Master Mix | Thermo Fisher | Cat#4367659 | |
| Commercial assay or kit | Gateway BP Clonase II Enzyme Mix | Invitrogen | Cat#56481 | |
| Commercial assay or kit | Gateway LR Clonase Enzyme Mix | Invitrogen | Cat#56484 | |
| Chemical compound, drug | Dexamethasone | Sigma-Aldrich | Cat#D4902 | |
| Chemical compound, drug | DL-Dithiothreitol | Sigma-Aldrich | Cat#43815–1G | |
| Software, algorithm | ‘MassPike’, a Sequest-based software pipeline for quantitative proteomics. | Professor Steven Gygi’s lab, Harvard Medical School, Boston, USA. | ||
| Software, algorithm | SEQUEST | Eng et al., 1994 | ||
| Software, algorithm | DAVID software | https://david.ncifcrf.gov/ | DAVID, RRID:SCR_001881 | |
| Software, algorithm | Reactome software | https://reactome.org/ | Reactome, RRID:SCR_003485 | |
| Software, algorithm | Image Studio Lite | LI-COR | Ver. 5.2; Image Studio Lite, RRID:SCR_013715 | |
| Software, algorithm | Cytoscape | The Cytoscape Consortium | Ver 3.7.1; Cytoscape, RRID:SCR_003032 | |
| Software, algorithm | DNASTAR Lasergene - SeqBuilder | DNASTAR, Inc | Ver. 12; DNASTAR: Lasergene Core Suite, RRID:SCR_000291 | |
| Software, algorithm | FlowJo | FlowJo | Ver. 10; FlowJo, RRID:SCR_008520 | |
| Software, algorithm | CompPass | Sowa et al., 2009 | ||
| Software, algorithm | CompPass Plus | Huttlin et al., 2015 | ||
| Other | Orbitrap Fusion Mass Spectrometer | ThermoFisher Scientific | Cat#IQLAAEGAAP FADBMBCX | Instrument |
| Other | Orbitrap Fusion Lumos Mass Spectrometer | ThermoFisher Scientific | Cat#IQLAAEGAAP FADBMBHQ | Instrument |
| Other | Raw Mass Spectrometry Data Files | This paper | ProteomeXchange Consortium via the PRIDE partner repository with dataset identifier PXD014845. | Raw data |
Additional files
-
Supplementary file 1
Details of the interactome.
(A) Relative abundance of all canonical and non-canonical viral proteins quantified in experiment whole cell lysate 3 (WCL3) from Weekes et al. (2014) and whole cell lysate series three from Fielding et al. (2017). Further details of the calculations employed are given in Figure 1—figure supplement 1A and the Materials and methods section. (B) Details of all 172 baits. Bait expression was verified by IB, MS or RT-qPCR (Figure 1—figure supplement 1B). (C) Relative abundance of all human proteins expressed in HFFFs, calculated as described in (A). The ‘rank’ column indicates the ranked average IBAQ abundance. The most abundant protein calculated by this method was ranked 1, and least abundant ranked 8129. (D) Coding sequences of all viral genes used in this study. A six base-pair linker region, a V5 tag then a stop codon directly followed each sequence (Key Resources Table). Codon usage was optimised for expression for US14, US17 and UL74. (E) Oligonucleotides and templates employed in the generation and RT-qPCR of each viral vector. (F) Oligonucleotides and templates employed in the generation and RT-qPCR of each human overexpression vector.
- https://cdn.elifesciences.org/articles/49894/elife-49894-supp1-v2.xlsx
-
Supplementary file 2
Full interactome data.
(A) Numbers of HCIPs per bait, excluding bait-bait interactions. (B) HCIPs for each bait (see Figure 1 and the Materials and methods section for details of the filtering employed, and the scores shown in this table). For baits solubilised in NP40, VHCIPs are shown in green. The ‘Prey IBAQ rank’ column shows the ranked IBAQ abundance from Supplementary file 1C, and gives an indication of how abundant each prey protein was in infected HFFFs. A range of ranks is shown where more than one isoform of a protein could be detected, in order to reflect data for all isoforms of that protein. Abundantly expressed prey may be more easily validated using IB with antibodies against an endogenous protein; less abundant proteins may require overexpression to enable detection. (C) All detected interacting proteins for each bait, without filtering.
- https://cdn.elifesciences.org/articles/49894/elife-49894-supp2-v2.xlsx
-
Supplementary file 3
Validation of the interactome data from BioGRID, IntAct, Uniprot, MINT and Virus Mentha (Calderone et al., 2015; Chatr-Aryamontri et al., 2013; Licata et al., 2012; Orchard et al., 2014).
Columns give details of the database(s) that included each interaction, the method used, and cell type employed. Interactome scores from the present study are shown in columns H-K. Column L shows whether a given interaction was validated in this interactome. A value of 1 indicates validation; 0 indicates detection of the interaction but failure to pass stringent scoring thresholds; ‘ND’ indicates the interaction was not detected by the interactome. Column M shows the ranked abundance of each human prey protein from Supplementary file 1C. Interactions that were not detected in this study included a number of prey proteins that could not be detected in HFFFs. Further details are given in the Materials and methods section.
- https://cdn.elifesciences.org/articles/49894/elife-49894-supp3-v2.xlsx
-
Supplementary file 4
Enriched functional pathways, protein components and interacting viral baits.
(A) All enriched functional pathways amongst all human HCIPs (p<0.05, after Benjamini-Hochberg adjustment). Column D shows the bait(s) interacting with each pathway component. (B) Further details of viral baits interacting with components of each pathway. Two values are shown: ‘% interaction’, the percentage of human interactors of each bait that belonged to the pathway (relates to Figure 2, where viral baits are included if >33% of interactors belonged to a given pathway). ‘% function’ illustrates the percentage of proteins from the pathway that interacted with the bait (relates to Figure 2—figure supplement 1, where viral baits are included if >33% of the pathway components identified interacted with a given viral bait). Values of >33% are coloured in this table. The ‘count’ column shows the total number of interacting pathway members; Figure 2 and Figure 2—figure supplement 1 included data with counts ≥ 2. (C) All enriched functional pathways amongst human HCIPs from each temporal class (p<0.05, after Benjamini-Hochberg adjustment). Column E shows the bait(s) interacting with each pathway component. This data underlies Figure 2—figure supplement 2A. (D) Temporal interactions of viral bait and viral prey proteins. This data underlies Figure 2—figure supplement 2B.
- https://cdn.elifesciences.org/articles/49894/elife-49894-supp4-v2.xlsx
-
Supplementary file 5
Full data underlying the domain-domain association predictions.
(A) HCMV proteins that contain each described Pfam domain. Links are given to additional information on each domain on the Pfam website. Overall 96 domains have been identified in HCMV proteins by Pfam, however only 10 domains were identified in two or more baits. Only this subset was examined to increase confidence in domain association predictions. (B) Subset of Supplementary file 2B illustrating individual protein-protein interactions that underpin data shown in Figure 4A.
- https://cdn.elifesciences.org/articles/49894/elife-49894-supp5-v2.xlsx
-
Supplementary file 6
Proteins degraded early during HCMV infection from Nightingale et al. (2018), using sensitive criteria.
Interactome data identified viral baits for 31 of these degraded proteins.
- https://cdn.elifesciences.org/articles/49894/elife-49894-supp6-v2.xlsx
-
Supplementary file 7
Enrichment of functional pathways among proteins interacting with (A) ORFL147C, using DAVID software and a maximum p-value of 0.3; (B) ORFL147C, using the Reactome database and ≥4 entities per enriched pathway; (C) both HCMV and KSHV (Davis et al., 2015), using DAVID software and a maximum p-value of 0.05; (D) only HCMV as described in Figure 5C, using DAVID software and a maximum p-value of 0.01.
- https://cdn.elifesciences.org/articles/49894/elife-49894-supp7-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/49894/elife-49894-transrepform-v2.pdf