Intestinal infection regulates behavior and learning via neuroendocrine signaling
Figures
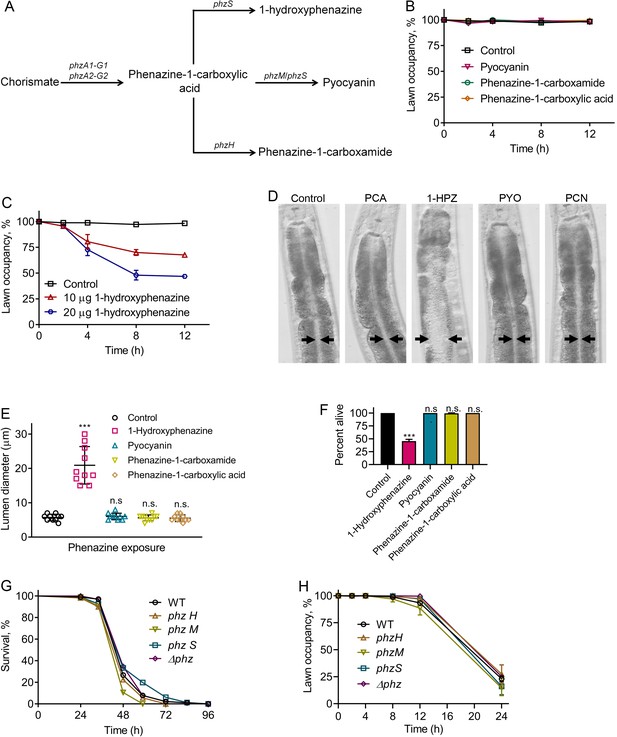
Phenazine-independent elicitation of C. elegans avoidance of P. aeruginosa during infection.
(A) Phenazine synthesis pathway of P. aeruginosa. (B) Time course of the percent occupancy of N2 animals on E. coli lawns containing 20 µg of pyocyanin, phenazine-1-carboxamide, and phenazine-1-carboxylic acid. For the control, the animals were exposed to solvent mock E. coli lawns. (C) Time course of the percent occupancy of N2 animals on E. coli lawns containing 1-hydroxyphenazine. For the control, the animals were exposed to solvent mock E. coli lawns. (D) Representative photomicrographs of N2 animals exposed for 8 hr to E. coli lawns containing 20 µg of phenazine-1-carboxylic acid (PCA), 1-hydroxyphenazine (1-HPZ), pyocyanin (PYO), and phenazine-1-carboxamide (PCN). For the control, the animals were exposed for 8 hr to solvent mock E. coli lawns. Arrows point to the border of the intestinal lumen. (E) Quantification of the diameter of the intestinal lumen of N2 animals exposed for 8 hr to E. coli lawns containing 20 µg of different phenazines. ***p<0.001 via the t test. n.s., non-significant. (F) Percent of animals alive after 24 hr of exposure to E. coli lawns containing 20 µg of different phenazines. The bars show the means ± SD from three independent experiments. ***p<0.001 via the t test. n.s., non-significant. (G) Representative survival plots of N2 animals on different phenazine synthesis pathway mutants of P. aeruginosa. p-value relative to WT, n.s., non-significant. (H) Time course of the percent occupancy of N2 animals on lawns of different phenazine synthesis pathway mutants of P. aeruginosa.
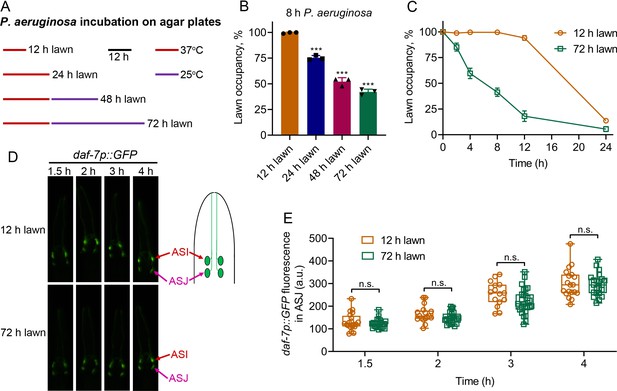
P. aeruginosa-induced daf-7 expression in ASJ neurons is insufficient to elicit avoidance behavior.
(A) Scheme for obtaining P. aeruginosa preparations with varying times and temperatures of incubation on SK plates. (B) Percent lawn occupancy of N2 animals after 8 hr of incubation on different preparations of P. aeruginosa. The black symbols represent individual data points. The bars show the means ± SD from three independent experiments. ***p<0.001 via the t test. (C) Time course of the percent occupancy of N2 animals on 12 and 72 hr lawns of P. aeruginosa. (D) Time course of induction of daf-7p::GFP on 12 and 72 hr lawns of P. aeruginosa. The ASI and ASJ chemosensory neurons are labeled. The drawing depicts the arrangement of the ASI and ASJ neurons in C. elegans head. (E) Quantification of induction of daf-7p::GFP in the ASJ chemosensory neuron on 12 and 72 hr lawns of P. aeruginosa over time. n.s., non-significant via the t test.
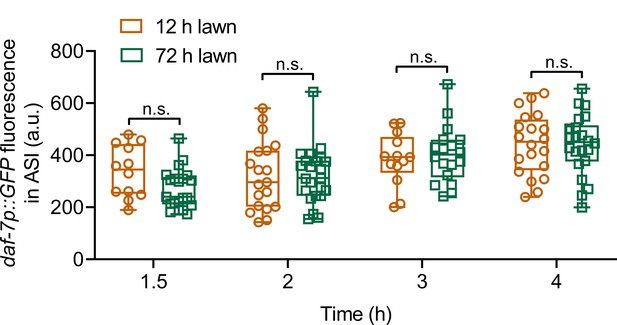
P. aeruginosa-induced daf-7 expression in ASI neurons is indistinguishable on 12 and 72 hr lawns.
Quantification of induction of daf-7p::GFP in the ASI chemosensory neuron on 12 and 72 hr lawns of P. aeruginosa over time. n.s., non-significant via the t test.
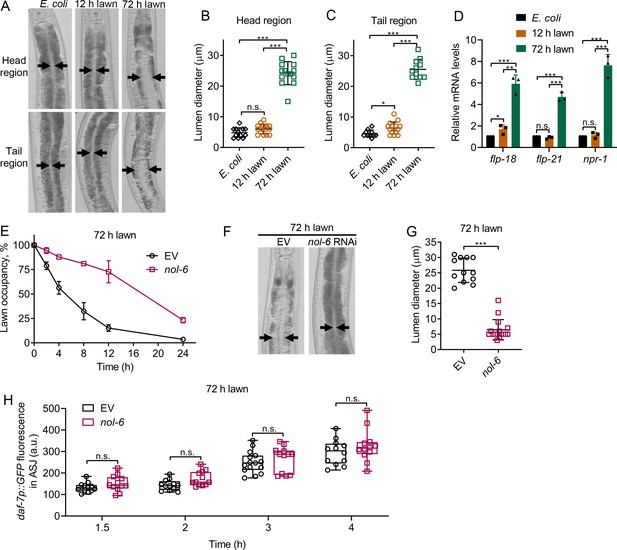
Intestinal lumen bloating underlies the avoidance behavior towards P. aeruginosa.
(A) Representative photomicrographs of N2 animals exposed for 8 hr to E. coli lawns, and 12 and 72 hr lawns of P. aeruginosa. Representative photomicrographs of the head and tail regions are shown. Arrows point to the border of the intestinal lumen. (B and C) Quantification of the diameter of the intestinal lumen of N2 animals exposed for 8 hr to E. coli lawns, and 12 and 72 hr lawns of P. aeruginosa from the head (B) and tail (C) regions. ***p<0.001 and *p<0.05 via the t test. n.s., non-significant. (D) Gene expression analysis of N2 animals grown on E. coli until the young adult stage, followed by incubation for 8 hr on E. coli lawns, and 12 and 72 hr lawns of P. aeruginosa. The black symbols represent individual data points. ***p<0.001, **p<0.01, and *p<0.05 via the t test. n.s., non-significant. (E) Time course of the percent occupancy of the control (EV) as well as nol-6 RNAi animals on 72 hr lawns of P. aeruginosa. (F) Representative photomicrographs of N2 animals grown on control and nol-6 RNAi exposed for 8 hr to 72 hr lawns of P. aeruginosa. Arrows point to the border of the intestinal lumen. (G) Quantification of the diameter of the intestinal lumen of N2 animals grown on control and nol-6 RNAi exposed for 8 hr to 72 hr lawns of P. aeruginosa. ***p<0.001 via the t test. (H) Time course of induction of daf-7p::GFP in ASJ neurons in animals grown on control and nol-6 RNAi and exposed to 72 hr lawns of P. aeruginosa. n.s., non-significant via the t test.
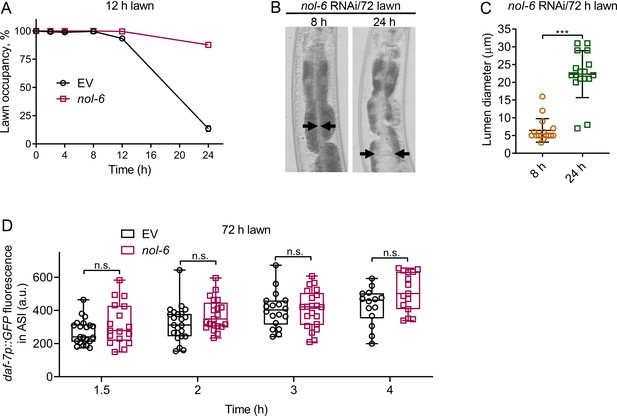
Intestinal lumen bloating underlies the avoidance behavior towards P. aeruginosa.
(A) Time course of the percent occupancy of control (EV) as well as nol-6 RNAi animals on 12 hr lawns of P. aeruginosa. (B) Representative photomicrographs of nol-6 RNAi animals exposed for 8 and 24 hr to 72 hr lawns of P. aeruginosa. Arrows point to the border of the intestinal lumen. (C) Quantification of the diameter of the intestinal lumen of nol-6 RNAi animals exposed for 8 and 24 hr to 72 hr lawns of P. aeruginosa. ***p<0.001 via the t test. (D) Time course of induction of daf-7p::GFP in ASI neurons in animals grown on control and nol-6 RNAi and exposed to 72 hr lawns of P. aeruginosa. n.s., non-significant via the t test.
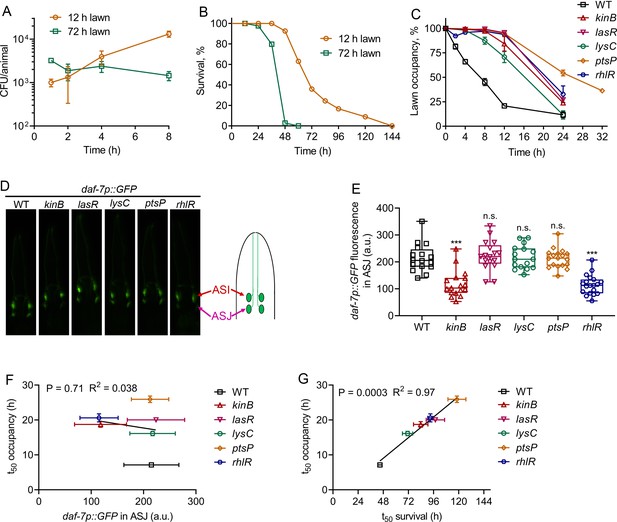
P. aeruginosa virulence correlates with the avoidance behavior.
(A) Time course of colony-forming units (CFU) per animal of N2 animals exposed to 12 and 72 hr lawns of P. aeruginosa-GFP. (B) Representative survival plots of N2 animals on 12 and 72 hr lawns of P. aeruginosa. p<0.0001. (C) Time course of the percent occupancy of N2 animals on 72 hr lawns of different mutants of P. aeruginosa. (D) Representative photomicrographs of daf-7p::GFP expressing animals exposed for 4 hr to lawns of different mutants of P. aeruginosa. The drawing depicts the arrangement of the ASI and ASJ neurons in C. elegans head. (E) Quantification of daf-7p::GFP in the ASJ chemosensory neuron pair in animals exposed for 4 hr to lawns of different mutants of P. aeruginosa. ***p<0.001 via the t test. n.s., non-significant. (F) Correlation of the mean lawn occupancy time (t50 occupancy) to the corresponding levels of daf-7p::GFP in the ASJ chemosensory neuron pair in animals exposed to different P. aeruginosa mutants. (G) Correlation of the mean lawn occupancy time (t50 occupancy) to the corresponding mean survival time (t50 survival) in animals exposed to different P. aeruginosa mutants.
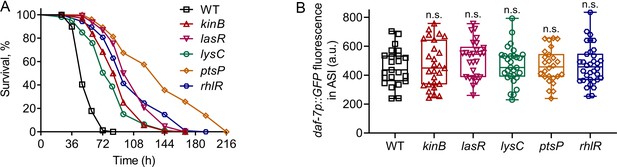
P. aeruginosa mutants with reduced virulence induce daf-7p::GFP in ASI neurons.
(A) Representative survival plots of N2 animals on different mutants of P. aeruginosa. p values for all mutants compared with WT are p<0.0001. (B) Quantification of daf-7p::GFP in the ASI chemosensory neuron pair in animals exposed for 4 hr to lawns of different mutants of P. aeruginosa. n.s., non-significant via the t test.
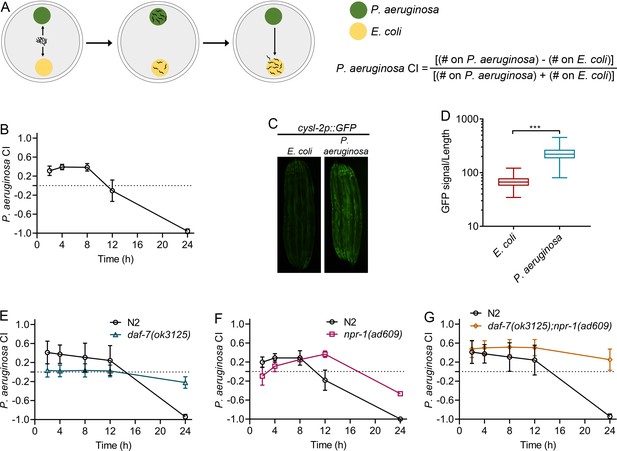
Neuroendocrine signaling involved in the control of aerotaxis behavior regulates associative learning of pathogens.
(A) Schematic representation of the two-choice preference assay. Animals are transferred to the center of plates equidistant from the lawns of P. aeruginosa and E. coli. The number of animals on both lawns is counted at a given time and used to calculate the P. aeruginosa choice index (CI). (B) Time course of the P. aeruginosa CI of N2 animals in a two-choice preference assay containing one lawn of each P. aeruginosa and E. coli. (C) Representative photomicrographs of cysl-2p::GFP-expressing animals exposed for 24 hr to E. coli and P. aeruginosa lawns. (D) Quantification of cysl-2p::GFP levels in animals exposed for 24 hr to E. coli and P. aeruginosa lawns. The quantification was conducted using a COPAS Biosort machine to measure the mean GFP signal and the length of individual animals. The GFP signal of each animal was normalized to its length. The data are plotted as a box and whisker plot from over 100 animals for each condition. ***p<0.001 via the t test. (E) Time course of the P. aeruginosa CI of N2 and daf-7(ok3125) animals in a two-choice preference assay containing one lawn of each P. aeruginosa and E. coli. (F) Time course of the P. aeruginosa CI of N2 and npr-1(ad609) animals in a two-choice preference assay containing one lawn of each P. aeruginosa and E. coli. (G) Time course of the P. aeruginosa CI of N2 and daf-7(ok3125);npr-1(ad609) animals in a two-choice preference assay containing one lawn of each P. aeruginosa and E. coli.
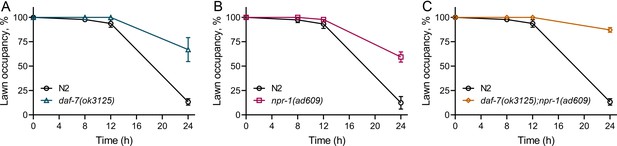
Aerotaxis behavior through neuroendocrine signaling controls microbial avoidance behavior.
(A) Time course of the percent occupancy of N2 and daf-7(ok3125) animals on P. aeruginosa lawns. (B) Time course of the percent occupancy of N2 and npr-1(ad609) animals on P. aeruginosa lawns. (C) Time course of the percent occupancy of N2 and daf-7(ok3125);npr-1(ad609) animals on P. aeruginosa lawns.
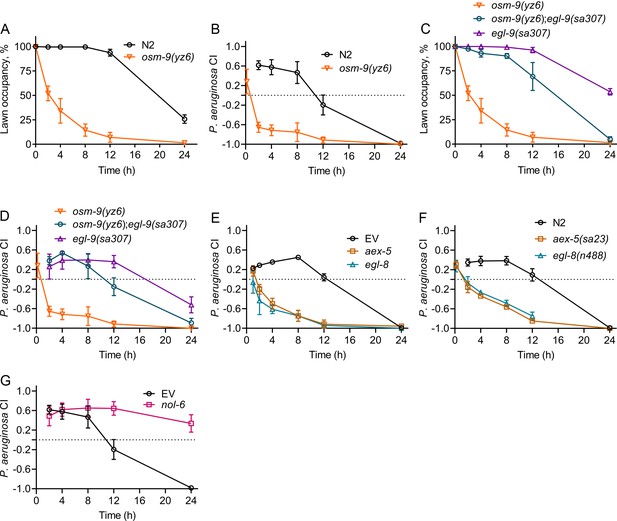
Modulation of aerotaxis behavior by intestinal bloating underlies the change in microbial preference upon infection.
(A) Time course of the percent occupancy of N2 and osm-9(yz6) animals on P. aeruginosa lawns. (B) Time course of the P. aeruginosa CI of N2 and osm-9(yz6) animals in a two-choice preference assay containing one lawn of each P. aeruginosa and E. coli. (C) Time course of the percent occupancy of osm-9(yz6), osm-9(yz6);egl-9(sa307), and egl-9(sa307) animals on P. aeruginosa lawns. (D) Time course of the P. aeruginosa CI of osm-9(yz6), osm-9(yz6);egl-9(sa307), and egl-9(sa307) animals in a two-choice preference assay containing one lawn of each P. aeruginosa and E. coli. (E) Time course of the P. aeruginosa CI of N2 animals grown on RNAi control bacteria, as well as bacteria for RNAi against aex-5 and egl-8 in a two-choice preference assay containing one lawn of each P. aeruginosa and E. coli. EV, empty vector RNAi control. (F) Time course of the P. aeruginosa CI of N2, aex-5(sa23), and egl-8(n488) animals in a two-choice preference assay containing one lawn of each P. aeruginosa and E. coli. (G) Time course of the P. aeruginosa CI of N2 animals grown on nol-6 RNAi as well as control bacteria in a two-choice preference assay containing one lawn of each P. aeruginosa and E. coli. EV, empty vector RNAi control.
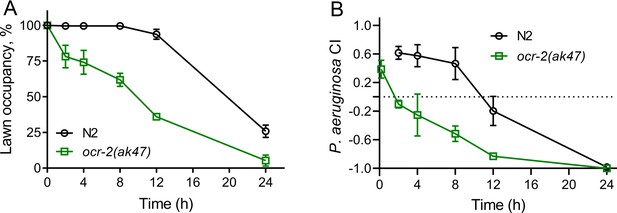
Modulation of aerotaxis behavior alters microbial preference.
(A) Time course of the percent occupancy of N2 and ocr-2(ak47) animals on P. aeruginosa lawns. (B) Time course of the P. aeruginosa choice index (CI) of N2 and ocr-2(ak47) animals in a two-choice preference assay containing one lawn of each P. aeruginosa and E. coli.
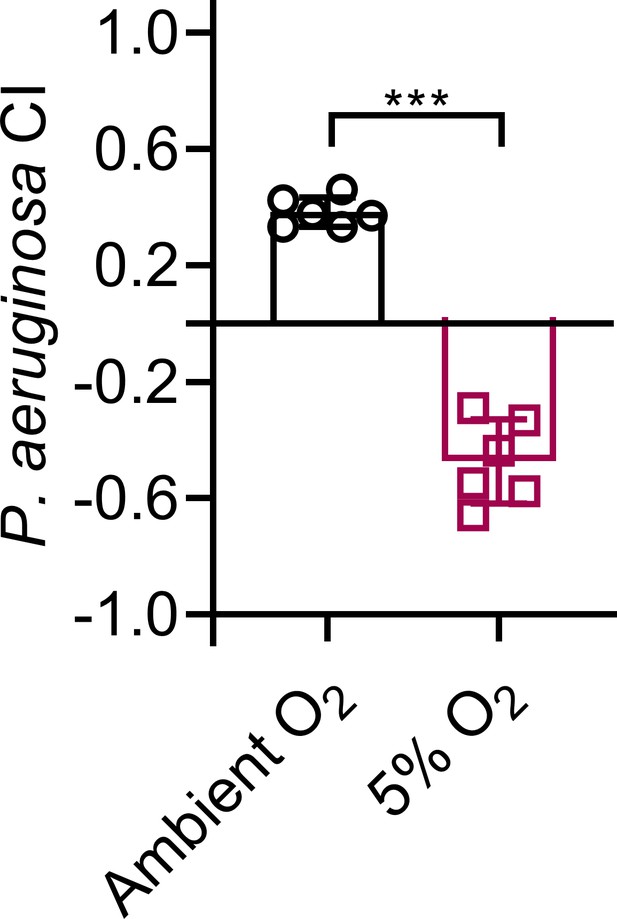
Modulation of aerotaxis alters microbial preference.
The P. aeruginosa choice index (CI) of N2 animals in a two-choice preference assay containing one lawn of each P. aeruginosa and E. coli after 2 hr of incubation in a hypoxia chamber purged with 5% oxygen. The control animals were incubated for 2 hr at ambient oxygen. ***p<0.001 via the t test.
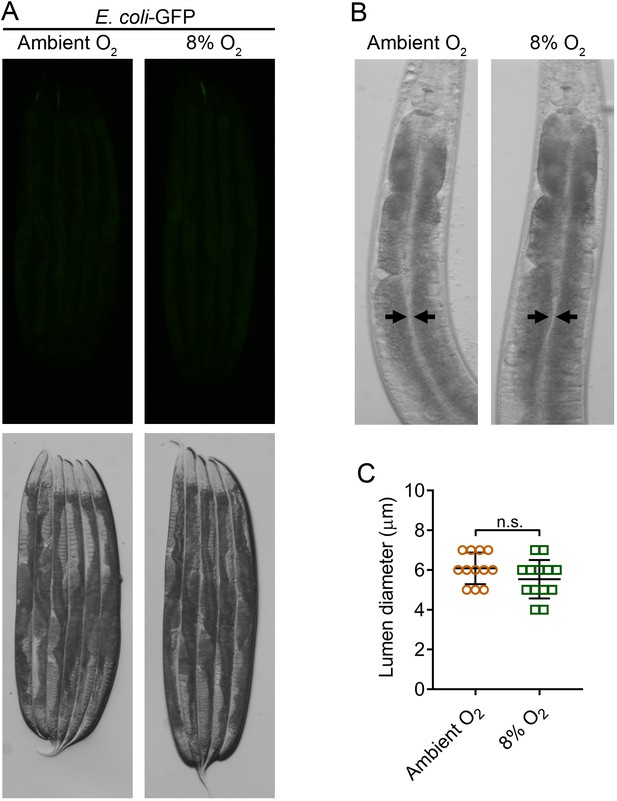
Low oxygen levels do not affect intestinal colonization and bloating.
(A) Representative photomicrographs of N2 animals that were incubated on E. coli-GFP lawns for 24 hr either at ambient oxygen levels or 8% oxygen levels in a hypoxia chamber. (B) Representative photomicrographs of the head regions of N2 animals that were incubated on E. coli-GFP lawns for 24 hr either at ambient oxygen levels or 8% oxygen levels in a hypoxia chamber. Arrows point to the border of the intestinal lumen. (C) Quantification of the diameter of the intestinal lumen of N2 animals that were incubated on E. coli-GFP lawns for 24 hr either at ambient oxygen levels or 8% oxygen levels in a hypoxia chamber. n.s., non-significant via the t test.
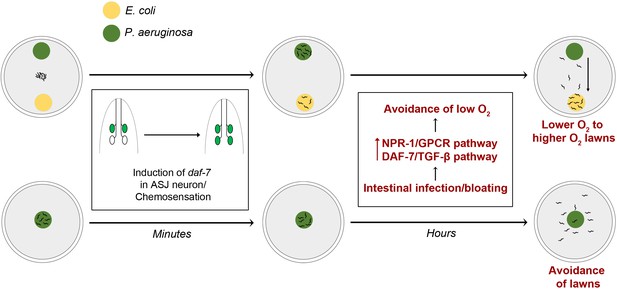
Model for intestinal infection-regulated microbial perception.
Rapid chemosensation of P. aeruginosa resulting in the induction of daf-7 expression in the ASJ neuron pair is insufficient for eliciting avoidance behavior. Intestinal infection leading to bloating activates NPR-1/GPCR and DAF-7/TGF-β neuroendocrine pathways, driving the evacuation of low O2 P. aeruginosa lawns and change in preference from relatively lower O2 lawns of P. aeruginosa to relatively higher O2 lawns of E. coli.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Escherichia coli) | OP50 | Caenorhabditis Genetics Center (CGC) | OP50 | |
| Strain, strain background (E. coli) | HT115 | Source BioScience | HT115 | |
| Strain, strain background (E. coli) | DH5α-GFP | Frederick M Ausubel laboratory | DH5α-GFP | |
| Strain, strain background (Pseudomonas aeruginosa) | PA14 | Frederick M Ausubel laboratory | PA14 | |
| Strain, strain background (P. aeruginosa) | PA14-GFP | Frederick M Ausubel laboratory | PA14-GFP | |
| Strain, strain background (P. aeruginosa) | PA14 kinB | Deborah Hung laboratory | PA14 kinB | |
| Strain, strain background (P. aeruginosa) | PA14 lasR | Deborah Hung laboratory | PA14 lasR | |
| Strain, strain background (P. aeruginosa) | PA14 lysC | Jason A Papin laboratory | PA14 lysC | |
| Strain, strain background (P. aeruginosa) | PA14 rhlR | Thomas K Wood laboratory | PA14 rhlR | |
| Strain, strain background (P. aeruginosa) | PA14 phz | Lars Dietrich laboratory | PA14 phz | Lacks both phzA1-G1 and phzA2-G2 operons |
| Strain, strain background (P. aeruginosa) | PA14 phzH | Lars Dietrich laboratory | PA14 phzH | |
| Strain, strain background (P. aeruginosa) | PA14 phzM | Lars Dietrich laboratory | PA14 phzM | |
| Strain, strain background (P. aeruginosa) | PA14 phzS | Lars Dietrich laboratory | PA14 phzS | |
| Strain, strain background (P. aeruginosa) | PA14 ptsP | Meta Kuehn laboratory | PA14 ptsP | |
| Strain, strain background (Caenorhabditis elegans) | N2 Bristol | CGC | N2 | |
| Strain, strain background (C. elegans) | ksIs2 [daf-7p::GFP + rol-6(su1006)] | CGC | FK181 | |
| Strain, strain background (C. elegans) | nIs470 [cysl-2p::GFP + myo-2p::mCherry] | CGC | DMS640 | |
| Strain, strain background (C. elegans) | ocr-2(ak47) | CGC | CX4544 | |
| Strain, strain background (C. elegans) | osm-9(yz6) | CGC | JY190 | |
| Strain, strain background (C. elegans) | aex-5(sa23) | CGC | JT23 | |
| Strain, strain background (C. elegans) | egl-8(n488) | CGC | MT1083 | |
| Strain, strain background (C. elegans) | egl-9(sa307) | CGC | JT307 | |
| Strain, strain background (C. elegans) | npr-1(ad609) | CGC | DA609 | |
| Strain, strain background (C. elegans) | daf-7(ok3125) | CGC | RB2302 | |
| Strain, strain background (C. elegans) | osm-9(yz6);egl-9(sa307) | This study | Materials and methods section | |
| Strain, strain background (C. elegans) | daf-7(ok3125);npr-1(ad609) | This study | Materials and methods section | |
| Software, algorithm | GraphPad Prism 8 | GraphPad Software | RRID:SCR_002798 | https://www.graphpad.com/scientificsoftware/prism/ |
| Software, algorithm | Photoshop CS5 | Adobe | RRID:SCR_014199 | https://www.adobe.com/products/photoshop.html |
| Software, algorithm | ImageJ | NIH | RRID:SCR_003070 | https://imagej.nih.gov/ij/ |
| Software, algorithm | Leica LAS v4.6 | Leica | RRID:SCR_013673 | https://www.leica-microsystems.com/ |
| Other | Hypoxia chamber | STEMCELL Technologies | CAT# 27310 | https://www.stemcell.com/products/hypoxia-incubator-chamber.html |





