Earliest infections predict the age distribution of seasonal influenza A cases
Figures
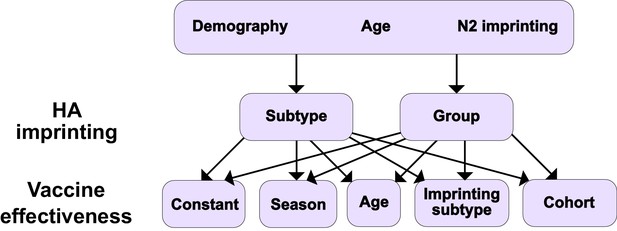
Summary of models tested.
Ten different models result from considering different combinations of HA imprinting and VE. We also tested one additional model excluding the effects of N2 and HA imprinting (Materials and methods: ‘Modeling approach’).
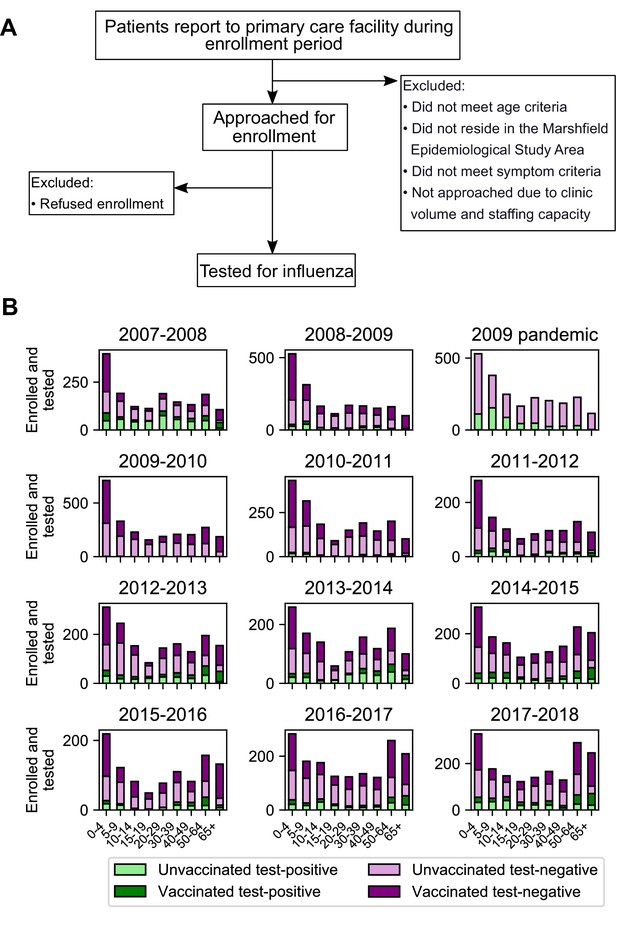
Sample collection and final study population.
Flowchart of sample collection (A) and final study population stratified by season, age, test status, and vaccination status (B). ‘Test-positive’ is defined as testing positive for the dominant circulating influenza A subtype in that season.
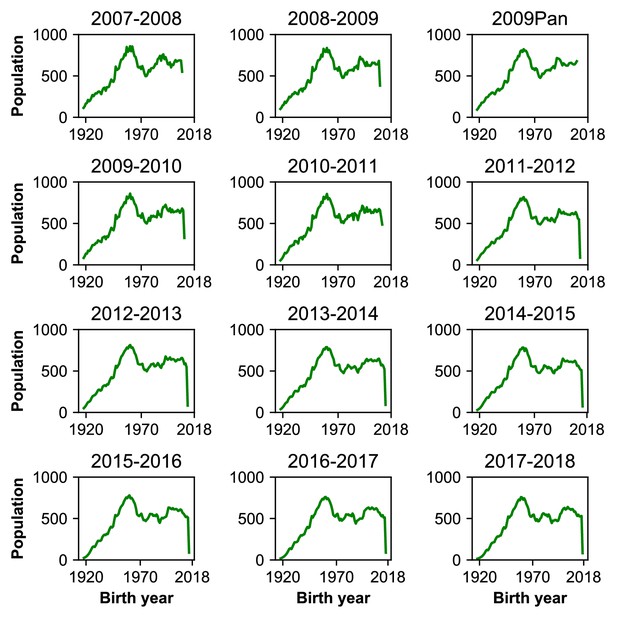
Birth year distribution of population.
Each panel shows the population distribution of all individuals in the study area who met the age criteria for study enrollment. People under 6 months old at the start of the sampling period in a season were not eligible to participate.
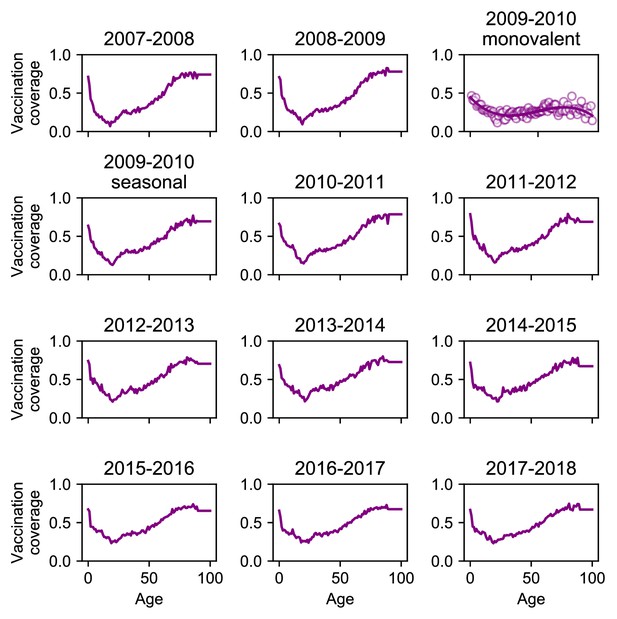
Vaccination coverage.
We estimated monovalent vaccination coverage in 2009–2010 by measuring vaccination coverage among enrolled people and fitting a smoothing spline to the data (solid line).
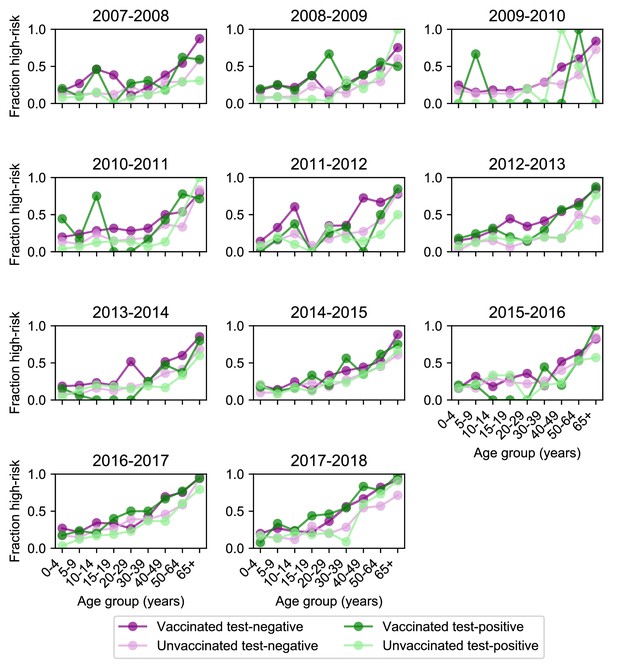
Age distribution of high-risk medical status.
High-risk medical status (Materials and methods, ‘Study cohort’) varies with age and vaccination status but stays relatively consistent across seasons. Each plot shows the fraction of enrolled people who had a high-risk medical condition for each season stratified by age, vaccination status, and test status. High-risk medical condition data was not collected for the 2009 pandemic season.
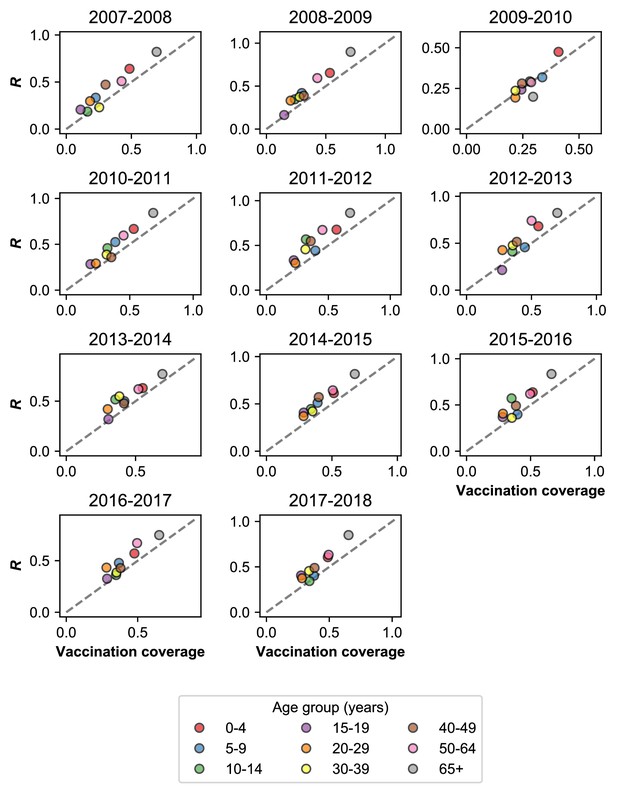
Rate of MAARI in vaccinated and unvaccinated controls.
Vaccinated individuals seek healthcare for MAARI at a higher rate than predicted by vaccination coverage. We measured the fraction of vaccinated people among all who presented with MAARI and tested negative for influenza (; Materials and methods: ‘Vaccination’). This is plotted against vaccination coverage by season for different age groups. The dashed grey line shows where and vaccination coverage are equal. Vaccination coverage for the 2009–2010 season uses monovalent vaccination coverage estimated directly from all individuals with MAARI. We do not show the 2009 pandemic season because the monovalent vaccine was not distributed until the second wave of the pandemic.
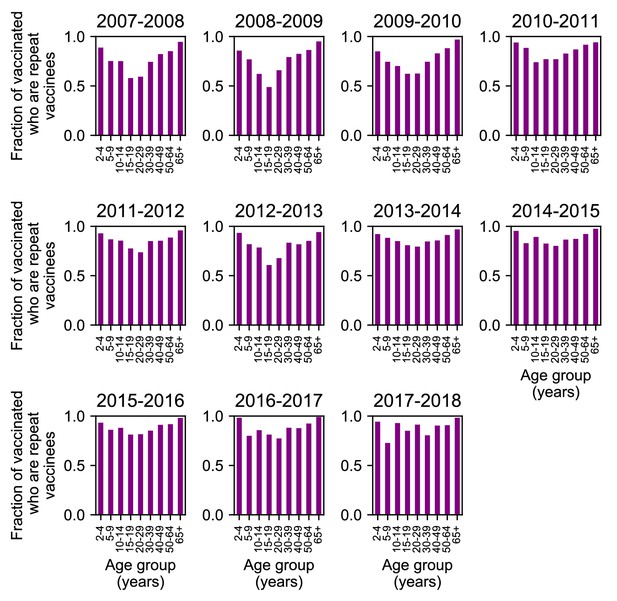
Repeat vaccination by age group and season.
Each bar shows the fraction of individuals who were vaccinated in that season who also received at least one influenza vaccination in the previous two seasons.
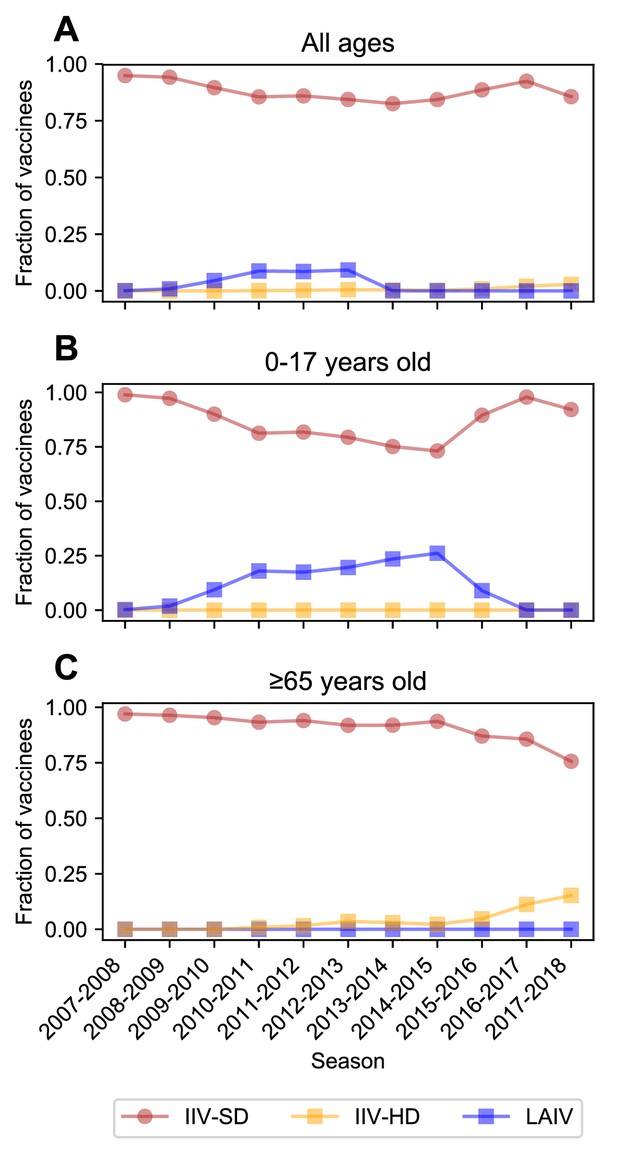
Vaccine type received.
Most vaccinated study participants received the inactivated influenza vaccine. The fraction of vaccinated people who received the standard-dose inactivated influenza vaccine (IIV-SD), the high-dose inactivated influenza vaccine (IIV-HD), or the live attenuated influenza vaccine (LAIV) is shown for all participants (A), children < 18 years old (B), and adults ≥65 years old (C).
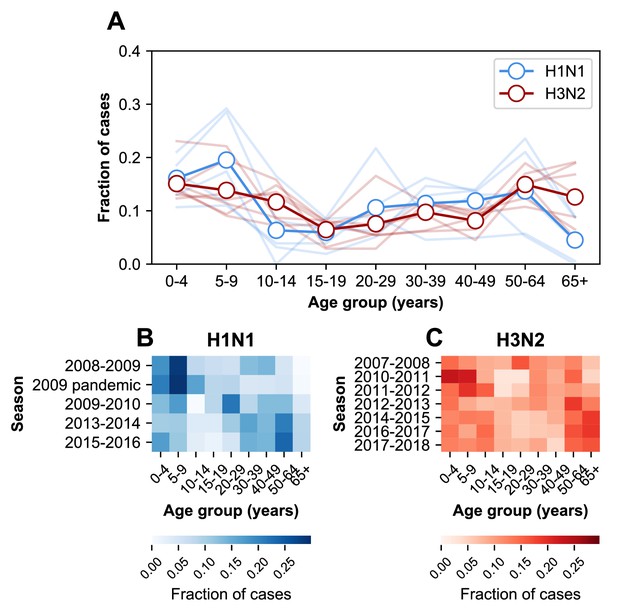
The age distribution of cases.
(A) The age distributions of cases from the 2007–2008 through the 2017–2018 influenza seasons in MESA. Dark lines with open circles indicate the average fraction of cases in each age group. Lighter-colored lines show the age distribution for individual seasons. (B) The age distribution of cases in H1N1-dominated seasons. (C) The age distribution of cases in H3N2-dominated seasons.
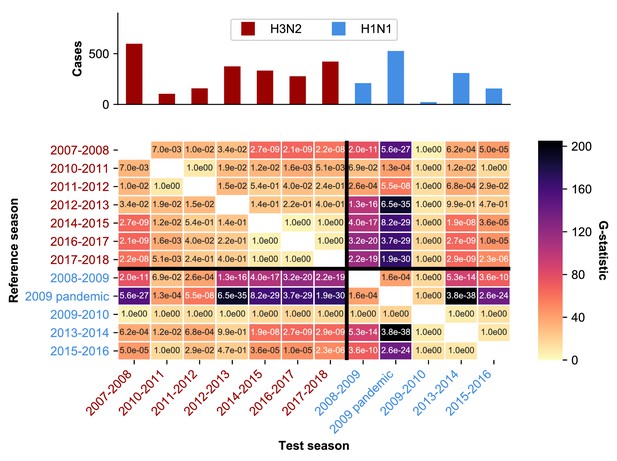
Statistical analysis of age distribution of cases.
Seasons differ in their age distributions. The color intensity of each cell shows the observed G-test statistic, which measures how much the age distributions of two seasons differ from the null expectation that they are drawn from the same distribution (Materials and methods: ‘Calculating differences in the age distribution between seasons'.). The text in each cell shows the Bonferroni-corrected p-value for each G-test. The dominant subtype of each season is indicated by the label color.
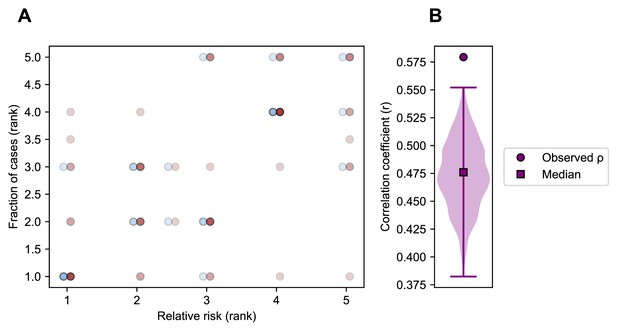
Correlation of relative risk and fraction of cases within an age group.
(A). Each point shows the rank of an age group’s relative risk of infection during the first half compared to the second half of an epidemic period (x-axis) and the rank of the fraction of cases belonging to that age group in the same epidemic period (y-axis) (Appendix 1: ‘Correlation of relative risk and fraction of cases’). Points are colored by the dominant subtype of the season and x-axis values are offset to facilitate visualization. Points with the same x and y values overlap and are indicated by darker shading. (B). To account for potential undersampling of cases at the beginning and end of specific seasons, we simulated 1000 replicate epidemics (Appendix 1 : ‘Sensitivity to sampling effort’) and calculated the same correlation as in panel A. The range is indicated by a vertical line and the median by a square.
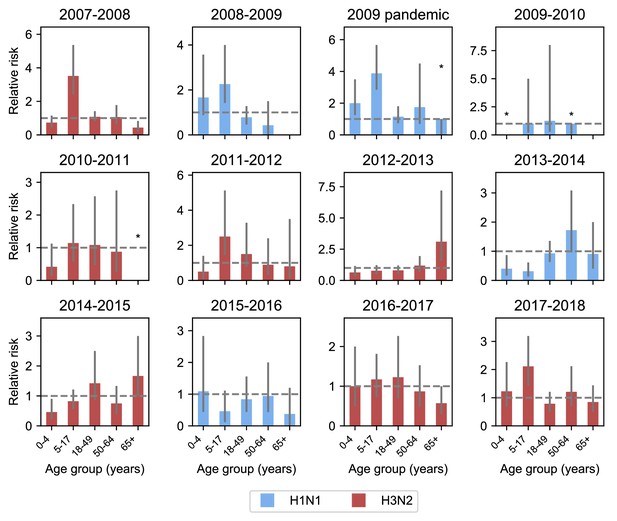
Relative risk among different age groups across seasons.
Each panel shows the relative risk of infection in the first versus the second half of an epidemic for different age groups in each season (Materials and methods: ‘Calculating relative risk’). Relative risk greater than 1 (indicated by the grey dashed line) means that an age group was more likely to be infected at during the first rather than second half of an epidemic. Age groups with no cases in the latter half of a season are indicated by asterisks and no bar. The dominant subtype of each subtype is indicated by the bar color. 95% binomial confidence intervals are indicated by grey vertical lines. Bars with asterisks over them indicate that the 95% confidence interval includes the scenario where all cases occur in the first half of the season.
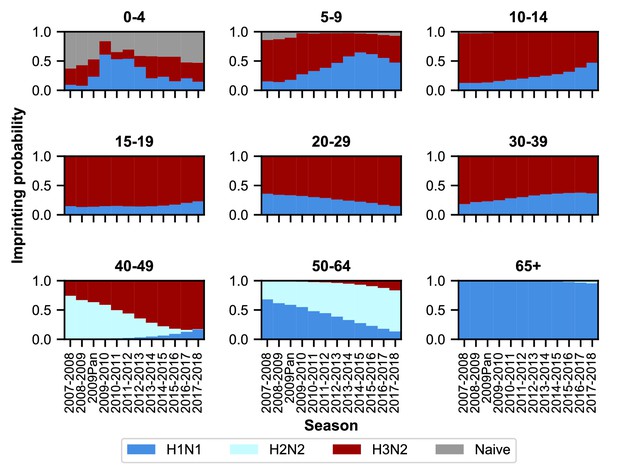
Imprinting probabilities by age group across seasons.
Each panel shows the imprinting probabilities of an age group from the 2007–2008 season through the 2017–2018 season. The color of each bar corresponds to the imprinting subtype or naive individuals, who have not yet been infected.
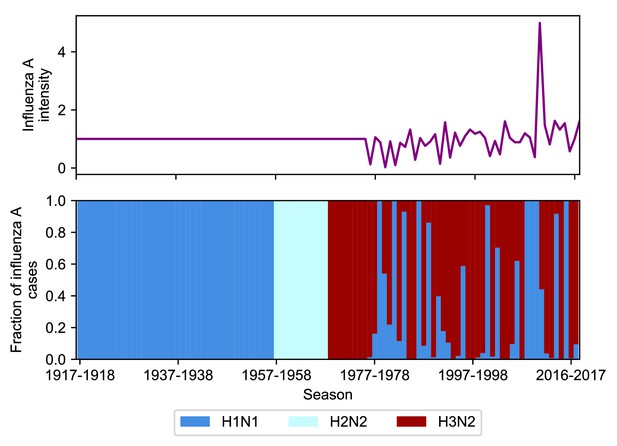
Intensity and subtype frequencies of influenza A.
The intensity (top panel) and subtype frequencies (bottom panel) of influenza A seasons in the United States. Intensity is measured as the product of influenza-like illness (ILI) and the fraction of respiratory specimens testing positive for influenza A in national surveillance data (Appendix 1: ‘Seasonal intensity’). This is normalized to the average intensity value between 1977 and 2017–2018. Seasons before 1977 where United States ILI surveillance data are unavailable are assumed to have an intensity score of 1 (i.e., the average score over all other seasons). Subtype frequencies were obtained from national surveillance data before the 2007–2008 season and directly from the MESA studies afterwards.
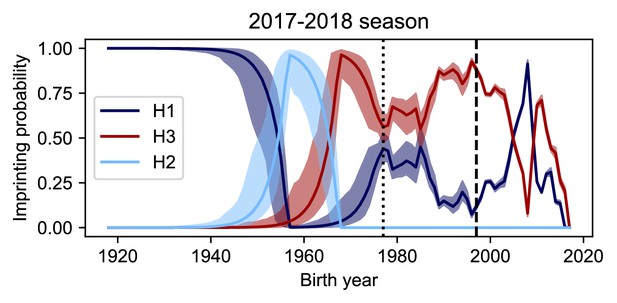
Imprinting probabilities with random sampling of seasonal intensity.
Uncertainty in ILI and the frequency of A have a small impact on imprinting probabilities. We simulated 10000 datasets to represent the range of possible epidemic sizes for seasons where we did not have data on either ILI or the frequency of influenza A (Appendix 1: ‘Sensitivity to uncertainty in ILI and the frequency of influenza A’). The vertical dashed line shows the point at which data on ILI and the frequency of influenza A are available while the vertical dotted line shows the point at which data on only the frequency of A is available. The median imprinting probabilities for those simulations is shown as a solid line with the maximum and minimum imprinting probabilities shown by the bounds of the shaded area.
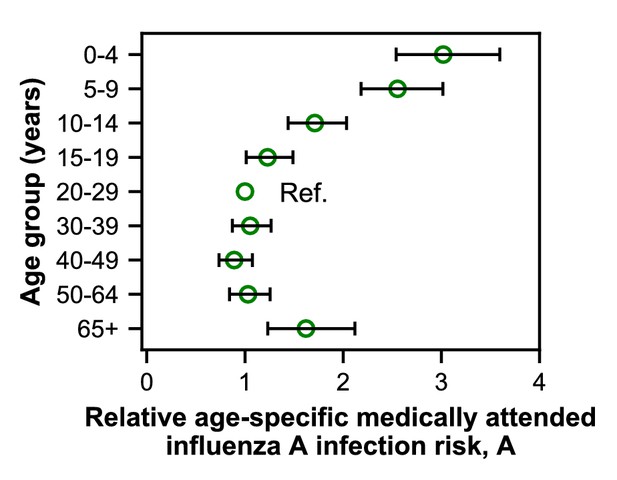
Estimates of relative age-specific medically attended influenza infection risk.
Open circles represent the maximum likelihood estimates of parameters describing age-specific differences in the relative risk of medically attended influenza A infection. Lines show the 95% confidence interval.
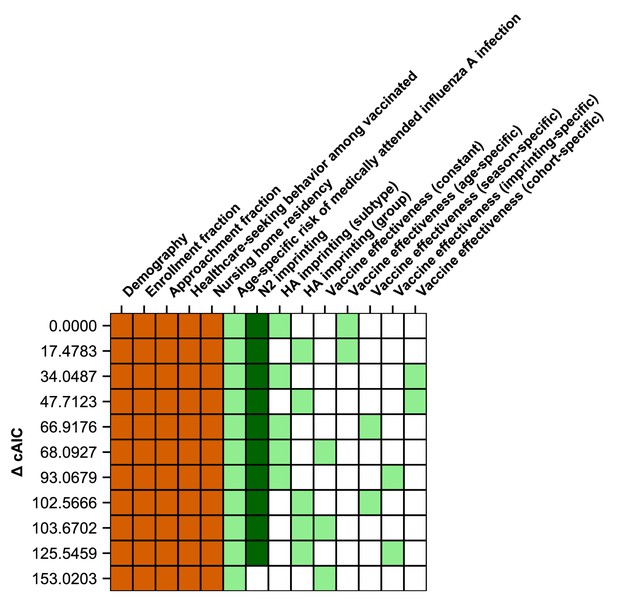
Ranking of models fitted to all ages.
The best-fitting model includes age-specific risk of medically attended influenza A infection, HA subtype imprinting, and age-specific VE. The 11 main models are shown as rows with colored squares indicating whether that model included parameters indicated by the columns. Orange squares indicate covariates that were not estimated. Light green squares mean that a given estimated parameter was supported. Dark green squares mean that the model did not support the inclusion of the parameters indicated by the column (i.e., the CI includes 0). Models are sorted by their cAIC relative to the best-fitting model.
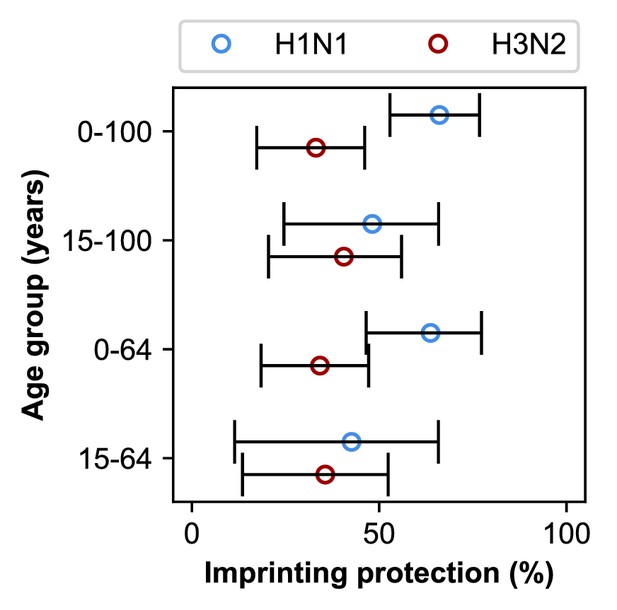
Estimates of imprinting strength.
Imprinting is more protective against H1N1 infection than H3N2 infection. Open circles represent the maximum likelihood estimates of imprinting parameters from the model including HA subtype imprinting and age-specific VE fitted to the indicated age group (y-axis). Black lines show 95% confidence intervals.
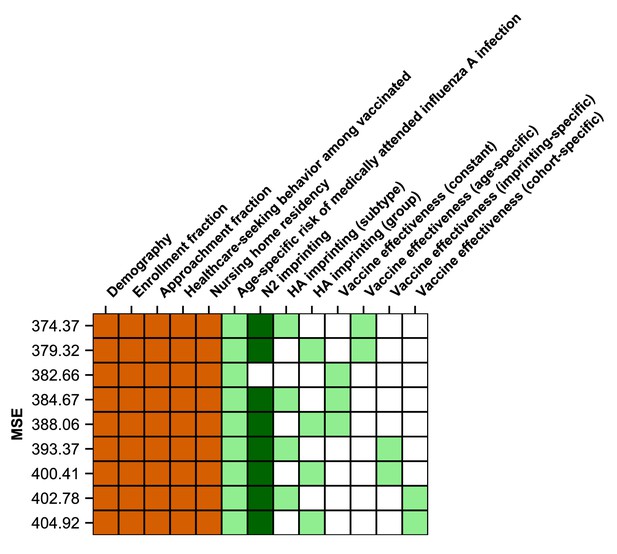
Ranking of models by predictive power.
The model which best-predicts excluded seasons includes HA subtype imprinting and age-specific VE. Models are shown as rows with colored squares indicating whether that model included parameters indicated by the columns. Orange squares indicate covariates that were not estimated. Light green squares mean that a given estimated parameter was supported. Dark green squares mean that the model did not support the inclusion of the parameters indicated by the column (i.e., the CI includes 0). Models are sorted by their MSE in predicting excluded seasons (Appendix 1: ‘Evaluation of predictive power’).
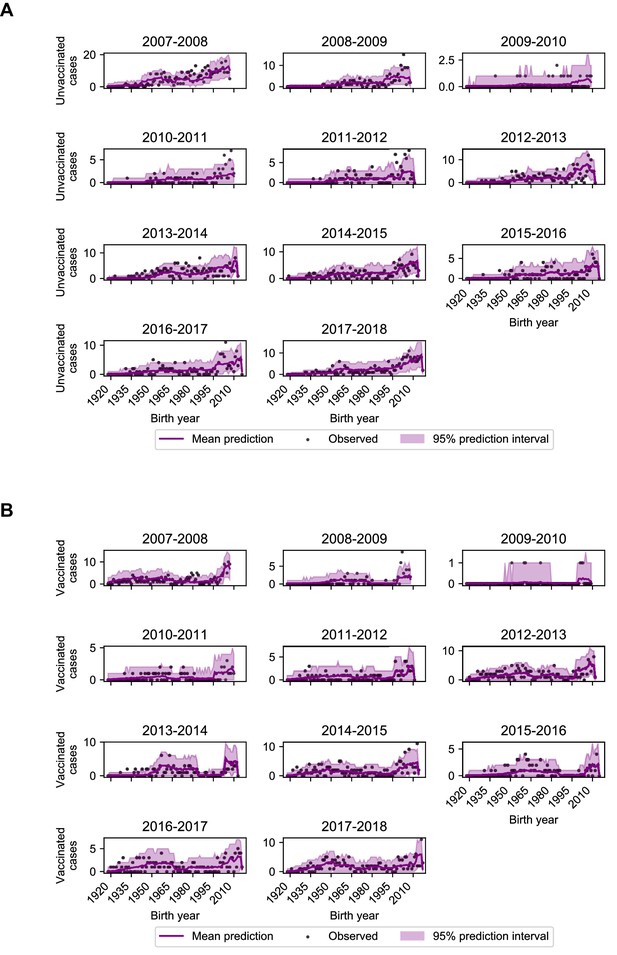
Model performance on excluded seasons.
Each panel shows the number of observed and predicted cases by birth year among unvaccinated (A) and vaccinated (B) study participants. Predictions and 95% prediction intervals were generated by fitting the model including age-specific risk of medically attended influenza A infection, HA subtype imprinting, and age-specific VE fitted to all seasons except the season in the panel (Appendix 1: ‘Evaluation of predictive power’).
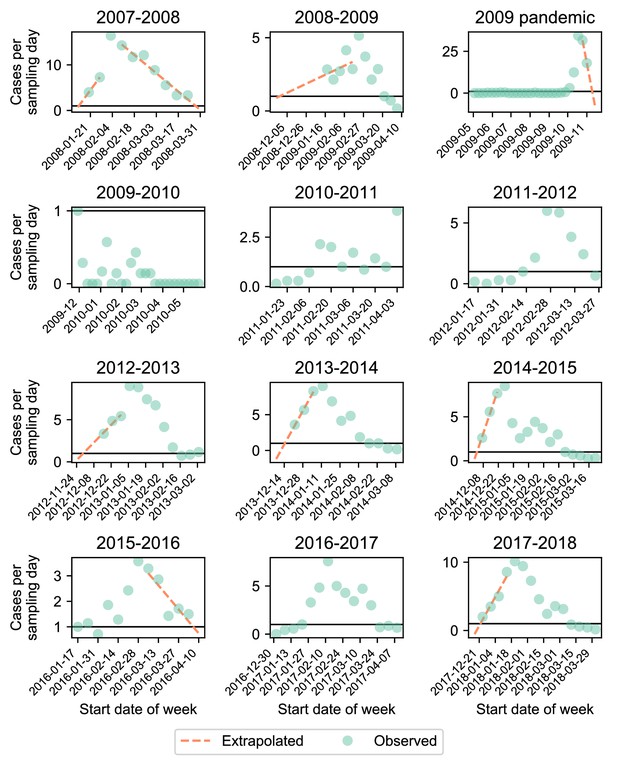
Cases per sampling day.
Each panel shows the number of cases per sampling day (green circles). We extrapolated cases at the start and end of the season (orange dashed line) if the observed number of cases per day exceeded 1 (black line) at the start and end of that season (Appendix 1: ‘Sensitivity to sampling effort’).
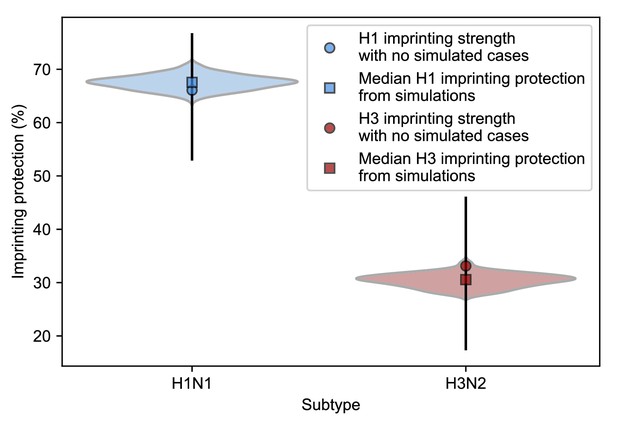
Estimates of imprinting protection with added simulated cases.
We fitted the model including HA subtype imprinting and age-specific VE to simulated cases in seasons where the enrollment period does not fully overlap the epidemic period and recorded the maximum likelihood estimates for H1N1 and H3N2 imprinting protection (Appendix 1: ‘Sensitivity to sampling effort’). The distributions of these values are shown as violin plots and the medians are shown as squares. Estimates of imprinting protection from the best-fitting model without simulated data with a 95% confidence interval are shown as circles with error bars.

Correlation of excess cases between seasons.
We tested whether excess cases in each birth cohort were negatively correlated with excess cases in the same birth cohort in the next season of the same subtype (Appendix 1: ‘Calculating excess cases’). We find a weak positive correlation for cases of H1N1 (Spearman’s =0.12, 95% CI 0.02–0.22) and H3N2 (Spearman’s =0.05, 95% CI −0.03–0.14).
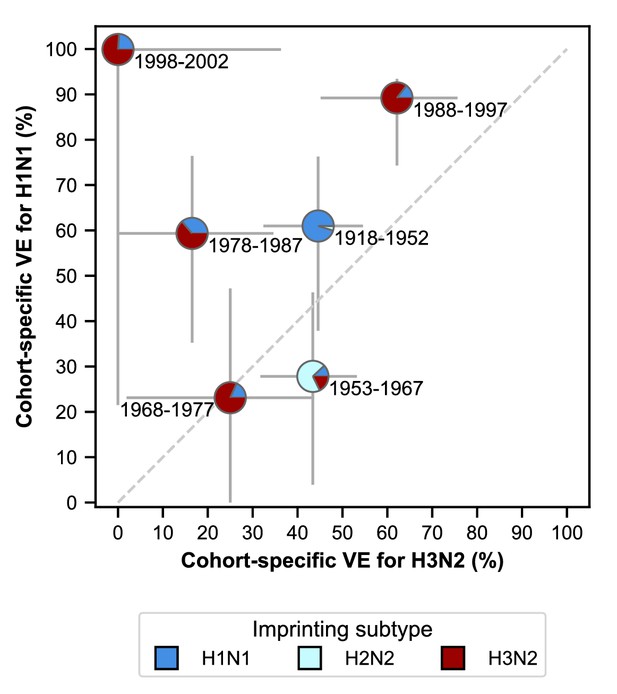
Estimates of birth-cohort-specific VE.
Birth-cohort-specific VE differs significantly between subtypes and birth cohorts. The location of each pie chart represents the H3N2 (x-axis) and H1N1 (y-axis) VE estimates for a birth cohort (indicated by text) obtained from our model fitted to people ≥15 years old. Pie charts are colored by the probability of first infection by each subtype (i.e, imprinting probability). 95% confidence intervals of the VE estimates are indicated by light grey solid lines. The dashed grey line shows the diagonal where the VE estimate for H1N1 is equal to the VE estimate for H3N2.
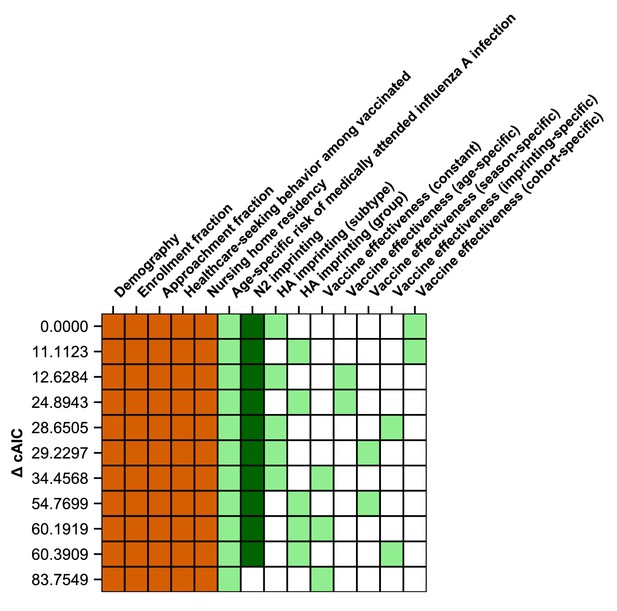
Ranking of models fitted to people ≥15 years old.
A model including age-specific risk of medically attended influenza A infection, HA subtype imprinting, and birth-cohort-specific VE best fits cases of people ≥15 years old. The 11 main models are shown as rows with colored squares indicating whether that model uses parameters indicated by the columns. Orange squares indicate covariates that were not estimated. Light green squares mean that a given estimated parameter was supported. Dark green squares mean that the model did not support the inclusion of the parameters indicated by the column (i.e, the CI includes 0). Models are sorted by their cAIC relative to the best-fitting model.
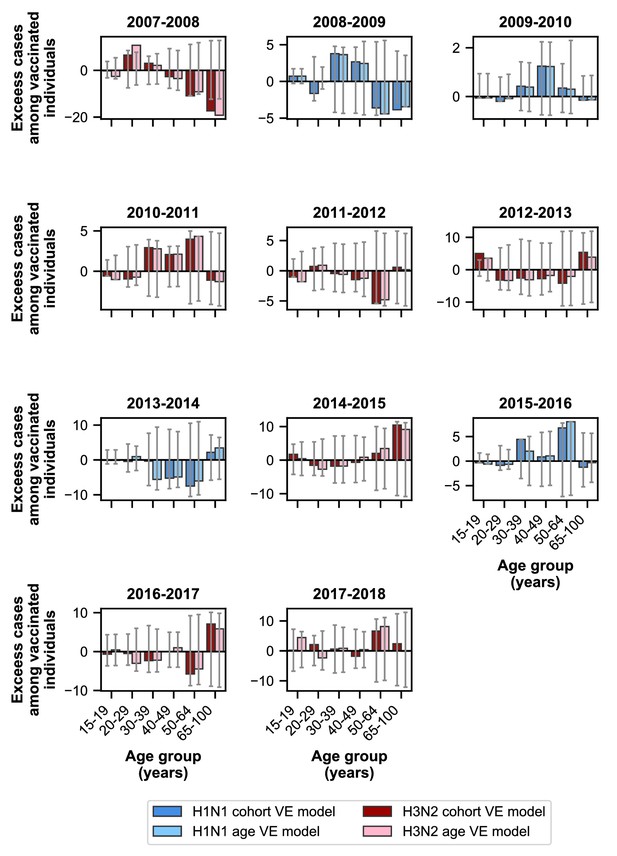
Excess cases for models using birth-cohort-specific VE and age-specific VE.
The birth-cohort-specific VE model predicts observed cases better than the age-specific VE model for people ≥15 years old. Bars show the excess cases in vaccinated individuals relative to the birth-cohort-specific VE model (dark colors) and the age-specific VE model (light colors) for age groups ≥15 years old. Colors indicate the dominant subtype of a given season. 95% prediction intervals are shown as grey error bars.
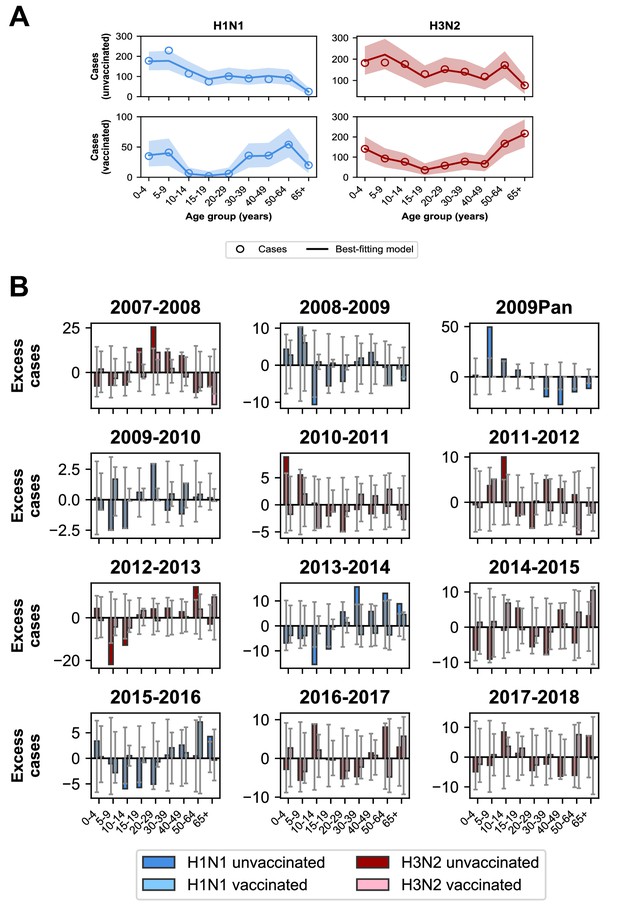
Model predictions compared to observed case counts.
(A) The model including age-specific VE and subtype-specific HA imprinting accurately predicts the overall age distribution of cases across seasons and age groups. Each row depicts the age distribution of cases among unvaccinated (top) and vaccinated (bottom) individuals over all sampled seasons (2007–2008 through 2017–2018). Each column indicates H1N1 cases (left, blue) and H3N2 cases (right, red). Open circles represent observed cases, solid lines represent the predicted number of cases from the best-fitting model, the shaded area represents the 95% prediction interval of the best-fitting model. (B) Excess cases of dominant subtype for each season. Excess cases are defined as the predicted number of cases from the best-fitting model - observed cases (Appendix 1: ‘Calculating excess cases’). Each panel shows the excess cases of the dominant subtype for each season for each age group among unvaccinated (dark bars) and vaccinated (light bars) individuals. Grey error bars show the 95% prediction interval.
Tables
Estimates of parameters shared by the age-specific VE and birth-cohort-specific VE models.
| Model with age-specific VE, age ≥6 months (MLE, 95% CI) | Model with age-specific VE, age ≥15 years (MLE, 95% CI) | Model with age-specific VE, age < 65 years (MLE, 95% CI) | Model with age-specific VE, age 15–64 years (MLE, 95% CI) | Model with birth-cohort- specific VE, age ≥15 years (MLE, 95% CI) | |
|---|---|---|---|---|---|
| Imprinting protection (%) | |||||
| H1 | 66 (53, 77) | 48 (25, 66) | 64 (47, 77) | 43 (11, 66) | 49 (24, 67) |
| H3 | 33 (17, 46) | 41 (20, 56) | 34 (18, 47) | 36 (13, 52) | 41 (20, 56) |
| N2 | 0 (0, 7) | 0 (0, 11) | 0 (0, 8) | 0 (0, 10) | 0 (0, 11) |
| Age-specific risk of medically attended influenza A infection | |||||
| 0–4 years | 3.0 (2.5, 3.6) | N.A. | 3.0 (2.5, 3.6) | N.A. | N.A. |
| 5–9 years | 2.6 (2.2, 3.0) | N.A. | 2.5 (2.2, 3.0) | N.A. | N.A. |
| 10–14 years | 1.7 (1.4, 2.0) | N.A. | 1.7 (1.4, 2.0) | N.A. | N.A. |
| 15–19 years | 1.2 (1.0, 1.5) | 1.2 (1.0, 1.5) | 1.2 (1.0, 1.5) | 1.2 (1.0, 1.5) | 1.2 (1.0, 1.5) |
| 30–39 years | 1.1 (0.9, 1.3) | 1.1 (0.9, 1.3) | 1.1 (0.9, 1.3) | 1.1 (0.9, 1.3) | 1.1 (0.9, 1.3) |
| 40–49 years | 0.9 (0.7, 1.1) | 0.9 (0.8, 1.1) | 0.9 (0.7, 1.1) | 0.9 (0.8, 1.1) | 0.9 (0.8, 1.1) |
| 50–64 years | 1.0 (0.8, 1.3) | 1.0 (0.8, 1.2) | 1.0 (0.8, 1.3) | 1.0 (0.8, 1.2) | 0.9 (0.7, 1.1) |
| 65+ years | 1.6 (1.2, 2.1) | 1.4 (1.0, 1.9) | N.A | N.A. | 1.5 (1.1, 1.9) |
Estimates of age-specific VE parameters in models fitted to different age groups.
| Model with age-specific VE, age ≥6 months (MLE, 95% CI) | Model with age-specific VE, age ≥15 years (MLE, 95% CI) | Model with age-specific VE, age < 65 years (MLE, 95% CI) | Model with age-specific VE, age 15–64 years (MLE, 95% CI) | |
|---|---|---|---|---|
| Age-specific VE against H1N1 (%) | ||||
| 0–4 years | 69 (56, 84) | N.A. | 68 (55, 83) | N.A. |
| 5–9 years | 26 (0, 48) | N.A. | 24 (0, 47) | N.A. |
| 10–14 years | 92 (80, 96) | N.A. | 92 (80, 96) | N.A. |
| 15–19 years | 86 (62, 95) | 89 (66, 97) | 86 (61, 95) | 89 (65, 97) |
| 20–29 years | 84 (65, 91) | 86 (69, 91) | 83 (63, 90) | 85 (67, 91) |
| 30–39 years | 8 (0, 37) | 22 (0, 47) | 5 (0, 35) | 19 (0, 45) |
| 40–49 years | 18 (0, 45) | 28 (0, 47) | 14 (0, 42) | 24 (0, 49) |
| 50–64 years | 32 (7, 51) | 39 (16, 56) | 28 (2, 48) | 37 (14, 55) |
| 65+ years | 50 (16, 71) | 64 (39, 83) | N.A. | N.A. |
| Age-specific VE against H3N2 (%) | ||||
| 0–4 years | 58 (48, 67) | N.A. | 58 (48, 67) | N.A. |
| 5–9 years | 45 (31, 58) | N.A. | 45 (30, 57) | N.A. |
| 10–14 years | 23 (0, 41) | N.A. | 22 (0, 41) | N.A. |
| 15–19 years | 31 (3, 53) | 33 (4, 55) | 30 (2, 53) | 32 (1, 54) |
| 20–29 years | 34 (11, 51) | 37 (15, 53) | 33 (11, 51) | 36 (14, 53) |
| 30–39 years | 10 (0, 31) | 15 (0, 35) | 9 (0, 30) | 12 (0, 33) |
| 40–49 years | 36 (15, 52) | 42 (24, 57) | 36 (15, 52) | 42 (23, 57) |
| 50–64 years | 47 (35, 56) | 49 (37, 58) | 47 (35, 57) | 48 (36, 58) |
| 65+ years | 41 (24, 54) | 38 (20, 52) | N.A. | N.A. |
Estimates of imprinting protection fitted to datasets representing upper and lower bounds of imprinting probabilities.
| Dataset | Best-fitting model | H1 imprinting protection (%, 95% CI) | H3 imprinting protection (%, 95% CI) |
|---|---|---|---|
| Lower bound | Demography, age, HA imprinting, age-specific VE | 72 (57, 84) | 32 (17, 44) |
| Upper bound | Demography, age, HA imprinting, age-specific VE | 61 (48, 72) | 37 (20, 51) |
Estimates of imprinting protection for models with different age groups.
| Age groups (years) | Best-fitting model | H1 imprinting protection (%, 95% CI) | H3 imprinting protection (%, 95% CI) |
|---|---|---|---|
| 0–4, 5–17, 18–64, 65+ | Demography, age, HA imprinting, age-specific VE | 56 (40, 68) | 36 (25, 46) |
| 0–8, 9–17, 18–49, 50–64, 65+ | Demography, age, HA imprinting, age-specific VE | 62 (47, 74) | 35 (21, 48) |
Estimates for VE from model with birth-cohort-specific VE fitted to people ≥15 years old.
| Birth cohort | H1N1 VE (%, MLE, 95% CI) | H3N2 VE (%, MLE, 95% CI) |
|---|---|---|
| 1998–2002 | 100 (22, 100) | 0 (0, 36) |
| 1988–1997 | 89 (74, 93) | 62 (45, 76) |
| 1978–1987 | 59 (35, 76) | 17 (0, 35) |
| 1968–1977 | 23 (0, 47) | 25 (2, 44) |
| 1953–1967 | 28 (4, 46) | 43 (32, 53) |
| 1918–1952 | 61 (38, 76) | 45 (32, 55) |





