Axon-like protrusions promote small cell lung cancer migration and metastasis
Figures
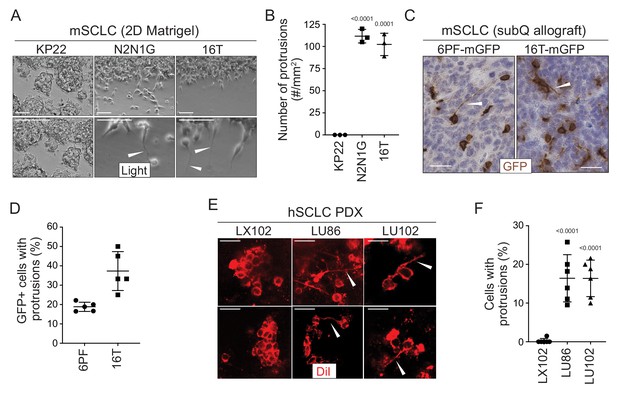
SCLC cells grow protrusions in culture and in vivo.
(A) Representative bright field images of three murine SCLC (mSCLC) cell lines (KP22, N2N1G, and 16T). Cells extend protrusions into a cell-free scratch generated in monolayer cultures. Protrusions are indicated by white arrowheads. Scale bars, 100 μm. N = 3 replicates. (B) Quantification of the number of protrusions that form from each mSCLC cell line as cultured in (A). Each symbol corresponds to the average of two technical replicates of an independent experiment. Mean + /- s.d. is shown, unpaired t-test. (C) Representative images of mSCLC cells (6PF and 16T) growing as subcutaneous tumors. At the time of injection, 10% SCLC cells stably expressing membrane-GFP (mGFP) were mixed with 90% GFP-negative SCLC cells. Immunostaining for GFP generates a brown signal. Examples of protrusions are indicated by white arrowheads. Hematoxylin (blue) stains the nuclei of the cells. (N = 5/allograft, from one biological replicate). Scale bar, 20 μm. (D) Quantification of (C). Each symbol represents an allograft tumor (N = 4/allograft, from one biological replicate). Mean + /- s.d. is shown. (E) Representative images of human SCLC (hSCLC) patient-derived xenografts growing subcutaneously (LX102, LU86, and LU102 models). Tumors were injected with the red fluorescent tracer DiI. Protrusions are indicated by white arrowheads. Scale bar, 20 μm. (F) Quantification of (E). Each symbol represents a xenograft tumor (N = 6/xenograft, from one biological replicate). Mean + /- s.d. is shown, unpaired t-test.
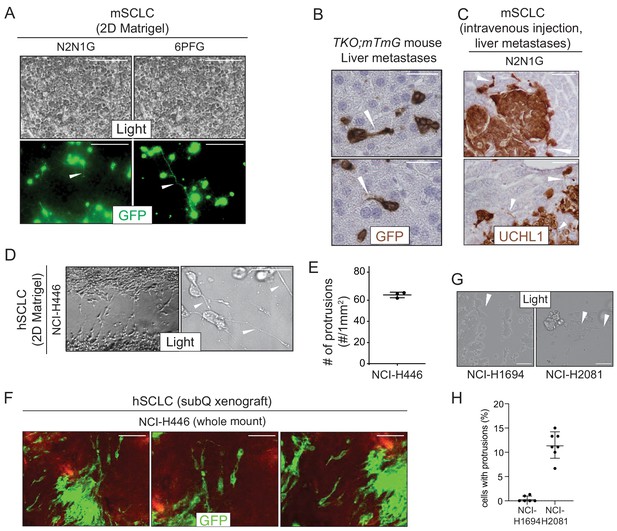
SCLC cells grow protrusions in culture and in vivo.
(A) Representative images of mSCLC N2N1G and 6PFG cells growing in dense culture from N = 3 independent experiments. At the time of plating, 3–5% cells expressing membrane-GFP (mGFP, green fluorescence) were mixed and co-cultured with 95–97% SCLC cells that do not expressing GFP. Examples of protrusions are shown with white arrowheads. Scale bars, 100 μm. (B) Representative images of mSCLC cells in the liver from the autochthonous TKO;mTmG model from N = 2 mice. Images were taken from micro-metastases. Immunostaining for GFP generates a brown signal. Protrusions are shown with white arrowheads. Hematoxylin (blue) stains the nucleus of cells. Scale bar, 20 μm. (C) Representative images of liver sections from mice after intravenous injection of mSCLC N2N1G cells from N = 3 mice. Immunostaining for the neuroendocrine marker UCHL1 (brown) outlines the shape of cells. Protrusions are shown with white arrowheads. Scale bars, 50 μm. (D) Representative bright field images of hSCLC NCI-H446 cells when cells are allowed to grow into a cell-free scratch generated in monolayer cultures under Matrigel. Protrusions are shown with white arrowheads. Scale bars, 40 μm. (E) Quantification of (D). N = 3 independent experiments. Mean + /- s.d. is shown. (F) Representative whole mount images of hSCLC NCI-H446 cells growing as a subcutaneous tumor from N = 4 independent xenografts from one experiment. At the time of injection, 10% of the SCLC NCI-H446 cells expressing membrane-GFP (mGFP) were mixed with 90% SCLC NCI-H446 cells not expressing GFP. Scale bars, 100 μm. (G–H) As in D-E for NCI-H1694 and NCI-H2081 hSCLC cells. N = 6–7 independent experiments. Scale bars, 20 μm.
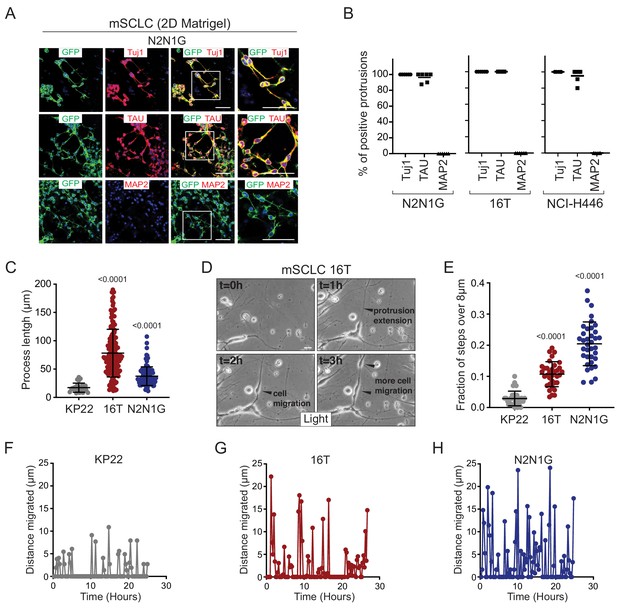
SCLC cells with protrusions migrate in a saltatory fashion similar to neuroblasts.
(A) Representative immunofluorescence images of N2N1G mSCLC cells expressing membrane-GFP (GFP, green) and stained (red) for expression of the neuronal marker Tuj1, the axonal marker TAU, or the dentritic marker MAP2. DAPI marks the nucleus of cells in blue. Scale bars, 50 μm. (B) Quantification of (A) for two mouse SCLC cell lines (16T, N2N1G) and one human SCLC cell line (NCI-H446). Images for 16T and NCI-H446 are shown in Figure 2—figure supplement 1B–C. N = 5/cell line. The bar is the mean. (C) Quantification of the length of protrusions in three mSCLC cell lines (KP22, no visible protrusions, 16T and N2N1G with protrusions). The average cell size in these experiments was ~8 μm. Each dot represents a cell. N > 10 fields were quantified in one biological replicate. Mean + /- s.d. is shown, Mann-Whitney test. (D) Representative still images from time-lapse videomicroscopy analysis of 16T SCLC cells showing the dynamic nature of the protrusions (from Video 1). (E) Quantification of the saltatory movements of three mSCLC cell lines as indicated. Note the correlation between the presence of protrusions and the ability of making longer steps (longer than the average cell size). Each dot represents a cell. N > 10 fields were quantified in one biological replicate. Mean + /- s.d. is shown, Mann-Whitney test. (F–H) Example of single cell movement over time for each of the three mSCLC cell lines.
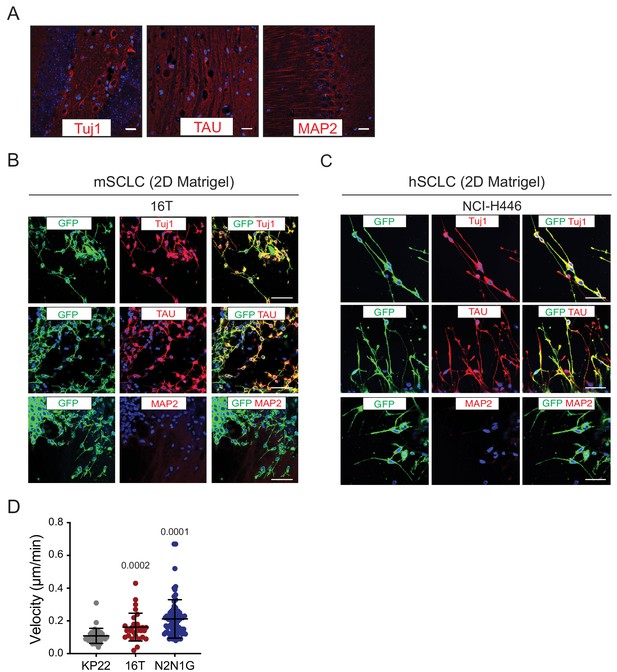
SCLC protrusions resemble axons and enable rapid cell movement.
(A) Representative fluorescence images of a mouse brain section stained with Tuj1, TAU, and MAP2 antibodies (positive controls, red). DAPI marks the nuclei of cells in blue. Scale bars, 50 μm. (B–C) Representative fluorescence images of 16T mSCLC cells (B) and NCI-H446 hSCLC cells (C) expressing membrane-GFP (mGFP) and stained (red) for expression of the neuronal marker Tuj1, the axonal marker TAU, or the dentritic marker MAP2. DAPI marks the nucleus of cells in blue. Quantification is shown in Figure 2B. Scale bars, 50 μm. (D) Quantification of velocity of mSCLC cancer cells from the three mouse SCLC cell lines indicated. Each dot represents a cell. Mean + /- s.d. is shown, Mann-Whitney test.
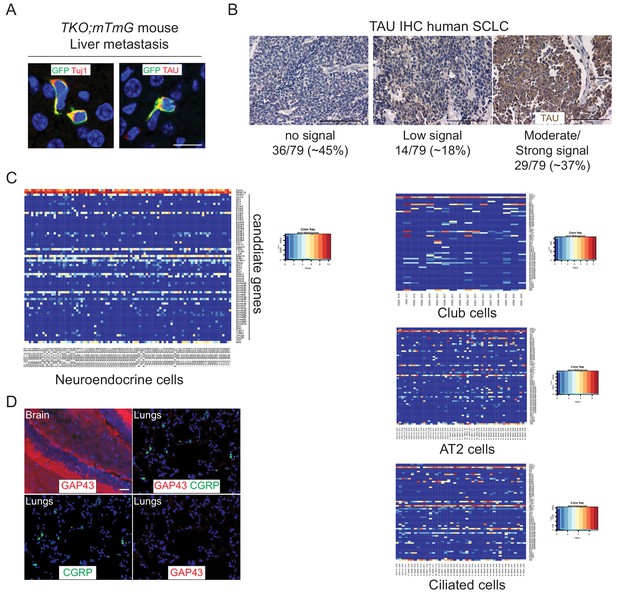
Mouse and human SCLC cells express axonal markers in vivo.
(A) Representative immunofluorescence staining of SCLC cells in the liver of a TKO;mTmG mouse (in which SCLC cancer cells express membrane GFP (GFP)). These cancer cells have protrusions positive for TAU and Tuj1. Images represent a merge of the GFP signal (green) and the signal for the TAU or Tuj1 antibodies (red). The nucleus of cells is labeled in blue by DAPI. Scale bar, 20 μm. (B) Representative images of immunohistochemistry (IHC) for TAU (brown) on human SCLC tissue microarrays (N = 79 human samples analyzed). The signal was evaluated by a certified pathologist (K.C.). Scale bars, 100 μm. (C) Gene expression analysis of the 69-gene list signature in single cells from the adult lung. Data are log2(tpm) from Ouadah et al. (2019). Nfib was added to the list of genes. Calca and Resp18 are shown as positive controls for neuroendocrine cells (top rows). Note that the gene signature with the 69 candidate genes is not globally activated in any of these lung epithelial cell types. (D) Representative immunofluorescence images for GAP43 (red) on sections of adult brain or adult lungs. CGRP-positive neuroendocrine cells can be detected in the lungs but these cells are not GAP43-positive (GAP43 is not detected in other lung epithelial cells either). DAPI marks the nucleus of cells in blue in all the sections. Scale bar, 20 µm.
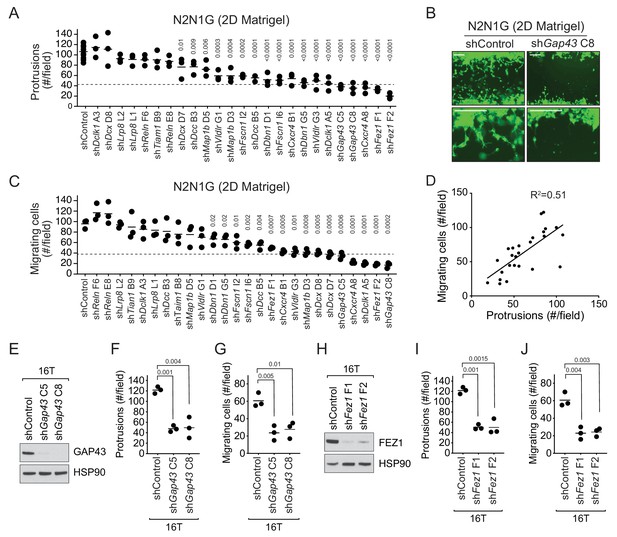
The axonal-like protrusions contribute to the migratory ability of SCLC cells in culture.
(A) Quantification of the number of cells with protrusions when mGFP-labeled N2N1G mSCLC cells were allowed to grow into a cell-free scratch generated in monolayer cultures under Matrigel. N = 3 independent experiments (shControl, N = 3 per experiment, total N = 9 plotted together). An unpaired t-test was used for statistical analysis and p-values are shown. Only significant p-values are shown. The dotted line represents a 60% reduction compared to the mean value of the controls. (B) Representative images of the data quantified in (A) and (C) with knock-down of Gap43. Scale bars, 100 μm. (C) Quantification of the migration of cells with protrusions when mGFP-labeled N2N1G mSCLC cells were allowed to grow into a cell-free scratch generated in monolayer cultures under Matrigel. N = 3 independent experiments. An unpaired t-test was used for statistical analysis and p-values are shown. Only significant p-values are shown. The dotted line represents a 60% reduction compared to the mean value of the controls. (D) Correlation of the data in (A) and (C) using the mean value for each knock-down. Pearson correlation R2 value is shown. (E and H) Immunoblot analysis of GAP43 or FEZ1 levels, respectively, in control and knock-down 16T mSCLC cells. HSP90 is a loading control. (F and I) Quantification of the number of cells with protrusions as in (A) with 16T mSCLC cells and Gap43 or Fez1 knock-down, respectively (N = 3). An unpaired t-test was used for statistical analysis and p-values are shown. (G and J) Quantification of the migration of cells with protrusions as in (B) with 16T mSCLC cells and Gap43 or Fez1 knock-down, respectively (N = 3). An unpaired t-test was used for statistical analysis and p-values are shown.
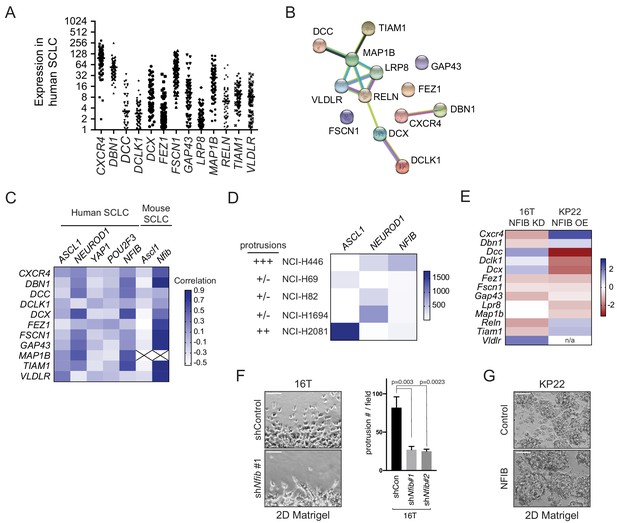
The expression of the 13 genes selected for their possible role in the formation of protrusions is in part regulated by NFIB.
(A) mRNA levels of candidate genes in human primary SCLC tumors (RNA-seq from George et al., 2015). (B) Network representation of the 13 candidates. Edges in the STRING analysis represent protein-protein associations but do not necessarily mean that they physically bind to each other. Blue edges represent known interactions from curated databases. Pink edges represent known experimentally-validated interactions. Others are predicted interactions, including text mining and co-expression (see string-db.org). (C) Spearman correlation of the 13 genes with key genetic drivers of SCLC phenotypes. The numbers for these analyses are from George et al. (2015), and Yang et al. (2018) (see also Supplementary file 2–tables 2-3). Note the higher correlation overall with NFIB expression. MAP1B was not identified in the mouse RNA data. (D) Gene expression analysis of three key genetic drivers of SCLC phenotypes in five human SCLC cell lines, along with the protrusions phenotype (see Figure 1—figure supplement 1 and data not shown). YAP1 and POU2F3 are not detected in these cells lines (data from CCLE RNA-seq analyses, https://www.ebi.ac.uk/gxa/experiments/E-MTAB-2770/Results). Note that the growth of protrusions does not correlate with ASCL1 or NEUROD1 expression. (E) Gene expression analysis (log two fold-change) of the 13 genes following NFIB knock-down in NFIB-high 16 T cells (which grow protrusions) and NFIB over-expression in NFIB-low KP22 cells (which don’t grow protrusions). RNA-seq data and immunoblot validation of the immunoblots for NFIB are from Denny et al. (2016). (F) Representative bright field image and quantification of the number of protrusions when 16T mSCLC cells were allowed to grow into a cell-free scratch generated in monolayer cultures under Matrigel with control shRNA or NFIB knock-own (N = 3). Scale bars, 100 μm. *p<0.05, **p<0.01, p<0.005, student t-test. (G) Representative bright field image of KP22 mSCLC cells with ectopic expression of GFP (control) of NFIB. Note the absence of protrusions (as quantified in Figure 1B) (N = 1 with three technical replicates). For (F) and (G), the immunoblot analysis of NFIB knock-down and ectopic expression in 16T and KP22 cells can be found in Denny et al. (2016).
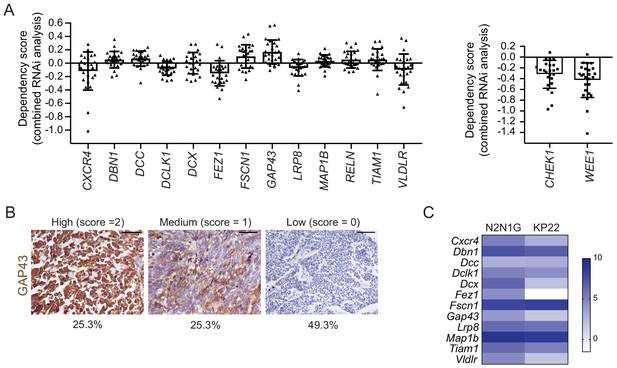
The 13 genes selected for their possible role in the formation of protrusions are expressed in human SCLC but do not play a key role in the expansion of SCLC cell populations.
(A) DepMap analysis (depmap.org) of the requirement for the 13 candidate genes in 25 human SCLC cell lines (RNAi combined analysis). Note that in a number of cell lines, the knock-down of candidate genes results in a positive score, indicative of a better expansion upon knock-down. Even in cases where the scores are negative, the negative values are small (the data for the genes coding for the CHK1 and WEE1 kinases, which are considered therapeutic targets in SCLC, are shown on the right hand side). (B) Representative images of immunohistochemistry (IHC) for GAP43 (brown) on human SCLC tissue microarrays (N = 79 human samples analyzed). The signal was evaluated by a certified pathologist (K.C.) and the scores are indicated. Hematoxylin (blue) stains the nuclei of the cells. Scale bars, 50 μm. (C) Heat map gene expression analysis of 12/13 genes (Reln was not detected) in N2N1G and KP22 cells. Note that the growth of protrusions in N2N1G cells correlates with an overall increase in the expression of the gene signature. Data are log2 values from RNA-seq.
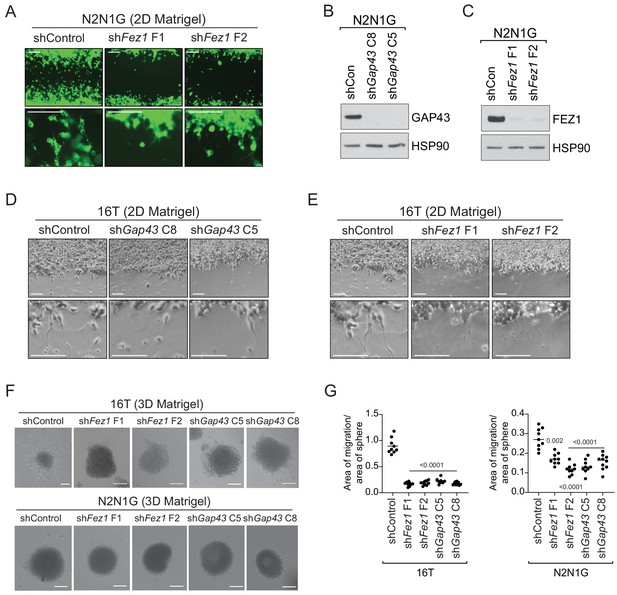
Knock-down of GAP43 and FEZ1 disrupts the formation of protrusions and cell migration in mouse SCLC cell lines in culture.
(A) Representative images of the data quantified in Figure 3A and C with knock-down of Fez1. Scale bars, 100 μm. (B–C) Immunoblot analysis of GAP43 (B) or FEZ1 (C) levels in control and knock-down N2N1G mSCLC cells. HSP90 is a loading control. (D–E) Representative images of the data with knock-down of Gap43 (D) or Fez1 (E) in 16 T cells. These data are quantified in Figure 3F–G (for GAP43) and Figure 3I–J (for FEZ1). The shControl targets GFP. Scale bars, 100 μm. (F) Representative images of 16T and N2N1G mouse SCLC cells in spheroids growing in 3D Matrigel with control shRNA or knock-down of Gap43 or Fez1. Scale bars, 50 μm. (G) Quantification of (F), determining the migration of cells out of the spheroids by measuring the area covered by cells outside of the spheroids relative to the size of each spheroid analyzed, 48 hr after plating the spheroids into the Matrigel. An unpaired t-test was used for statistical analysis. All p-values<0.0001 except one, as indicated.
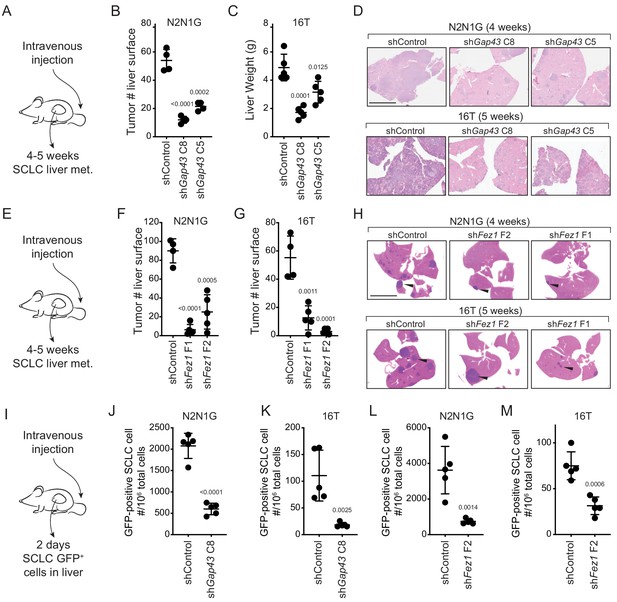
Genes involved in the generation of protrusions also control the formation of metastases.
(A) Diagram of the approach to investigate the formation of liver metastases (met.) after intravenous injection of SCLC cells. (B–C) Quantification of the number of metastases 4 and 5 weeks after intravenous injection of N2N1G and 16T mSCLC cells, respectively, with control knock-down or knock-down of Gap43 with two independent shRNAs. For N2N1G, tumors at the surface of the liver were quantified on the liver surface, as shown in Figure 4—figure supplement 2D. Too many tumors were present with the 16 T cell line and the control shRNA, and quantification was thus performed by measuring liver weight. N = 4–5 mice per condition in one biological replicate. Mean + /- s.d. unpaired t-test. (D) Representative hematoxylin and eosin (H and E) images of liver sections of mice in (B–C). Scale bars, 5 mm. (E–H) As shown in (A–D) for Fez1 knock-down. See Figure 4—figure supplement 2E for representative images with N2N1G cells for the quantification in (F–G) of tumors at the surface of the liver. Arrows point to metastases. N = 4–5 mice per condition in one biological replicate. Mean + /- s.d. is shown, unpaired t-test. (I) Diagram of the approach to investigate early steps in liver metastasis, 2 days after intravenous injection. (J–M) Quantification of the number of GFPpositiveN2N1G and 16T mSCLC cells 2 days after intravenous injection. See Figure 4—figure supplement 2F-ID for representative flow cytometry. N = 5 mice per condition in one biological replicate. Mean + /- s.d., unpaired t-test.
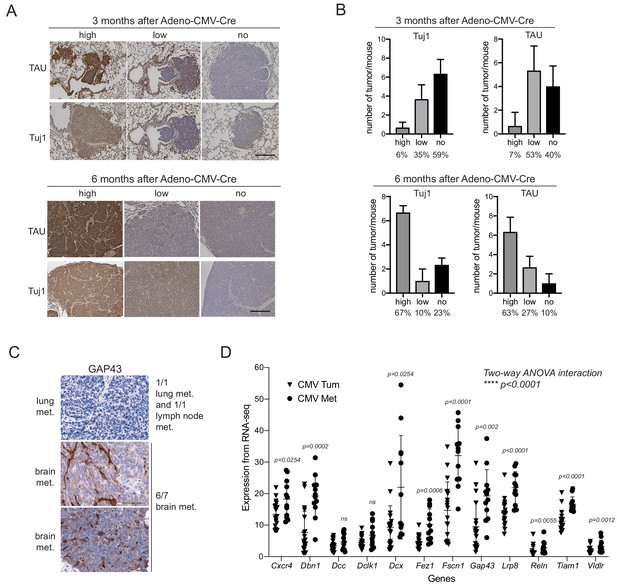
Increased expression of axonal markers in metastatic SCLC.
(A) Representative images of immunohistochemistry experiments on lung sections from TKO mice 3 months and 6 months after SCLC initiation with Ad-CMV-Cre. None of the mice had metastases at the 3 month time point while all the mice analyzed had evidence of metastasis at the 6 month time point. The Tuj1 antibody marks neuronal tubulin and TAU is a marker of axons. Hematoxylin was used as a counterstain (purple). Scale bar, 100 μm. (B) Quantification of (A), with N = 30–32 tumors analyzed from N = 3 mice at the 3 month time point and N = 30 tumors analyzed from N = 3 mice at the 6 month time point. Percentages are indicated. (C) Images of GAP43 immunohistochemistry experiments for representative metastases. Hematoxylin was used as a counterstain (purple). Out of 9 metastases analyzed, six showed positive staining for GAP43. Scale bar, 100 μm. (D) Analysis of the expression of 12/13 genes in the selected list of genes involved in axonogenesis and neuronal migration comparing metastases to primary tumors in the Adeno-CMV-Cre TKO mouse model (from Supplementary file 2–table 2 and from Denny et al., 2016, and Yang et al., 2018). The two groups (primary tumors ‘CMV Tum’ and metastases ‘CMV Met’) were first compared by two-way ANOVA and then each gene was compared by t-test.
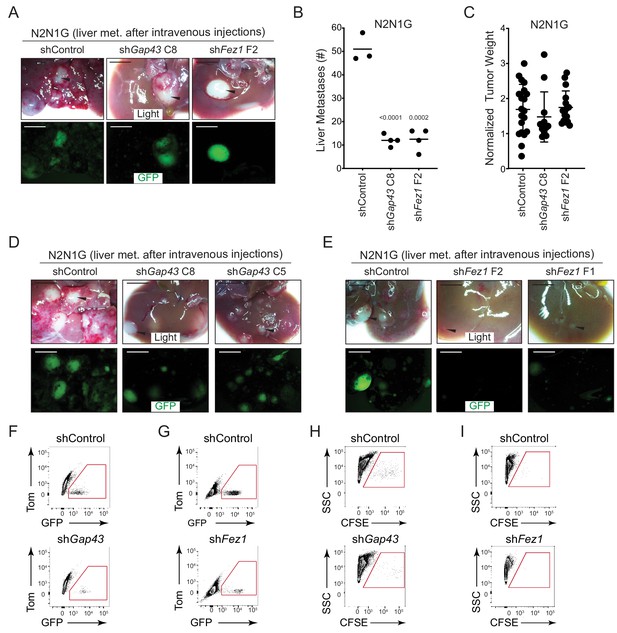
Reduced formation of metastasis upon knock-down of GAP43 and FEZ1 in SCLC cells.
(A) Representative live and epifluorescence images (GFP, green) of liver section of mice 4 weeks after intravenous injection of GFP-positive N2N1G mSCLC cells, with control knock-down or knock-down of Gap43 or Fez1. Arrows point to metastases. Scale bars, 5 mm. (B) Quantification of (A). The bar is the mean, unpaired t-test. (C) Quantification of tumor weight after subcutaneous injection of control and knock-down N2N1G cells. Values are not statistically significant by t-test. (D–E) Representative bright light and epifluorescence images (GFP, green) of livers from mice 4 weeks after intravenous injection of GFP-positive N2N1G mSCLC cells, with control knock-down or knock-down of Gap43 or Fez1. Arrows point to metastases. Scale bars, 5 mm. (F–G) Representative flow cytometry quantification of GFP-positive N2N1G cells in the liver 2 days after intravenous injection. (H–I) Representative flow cytometry quantification of CFSE-labeled 16 T cells in the liver 2 days after intravenous injection.
Videos
Time-lapse video of 16T mouse SCLC cells (images collected every 15 min).
Time-lapse video of KP22 mouse SCLC cells (images collected every 15 min).
Time-lapse video of 16T mouse SCLC cells (images collected every 15 min).
Time-lapse video of N2N1G mouse SCLC cells (images collected every 15 min).
Additional files
-
Supplementary file 1
Key Resources table.
- https://cdn.elifesciences.org/articles/50616/elife-50616-supp1-v2.docx
-
Supplementary file 2
This Excel file contains all the Tables associate with the manuscript.
Table 1: A Summary of all the cell line that are used and tested for protrusion formation. Table 2: Expression of genes involved in axonogenesis and axon guidance in primary human SCLC tumors by RNA-seq Table3: Expression of genes involved in axonogenesis and axon guidance in mouse SCLC tumors and metastases by RNA-seq Table 4: Metascape analysis of the top 20 clusters with their representative enriched terms (one per cluster) for the 69 candidates. Table 5: Biological process GO term analysis for the 13 selected proteins Table 6: 13 candidate genes that may control the growth of axonal-like protrusions on SCLC cells Table 7: Expression levels and dependency scores for the 13 candidates in human SCLC cells Table 8: Knock-down of the 13 candidate genes in mouse SCLC cells Table 9: RNA-seq analysis of N2N1G and KP22 mouse SCLC cells
- https://cdn.elifesciences.org/articles/50616/elife-50616-supp2-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/50616/elife-50616-transrepform-v2.docx





