Delta-like 1 and Delta-like 4 differently require their extracellular domains for triggering Notch signaling in mice
Figures
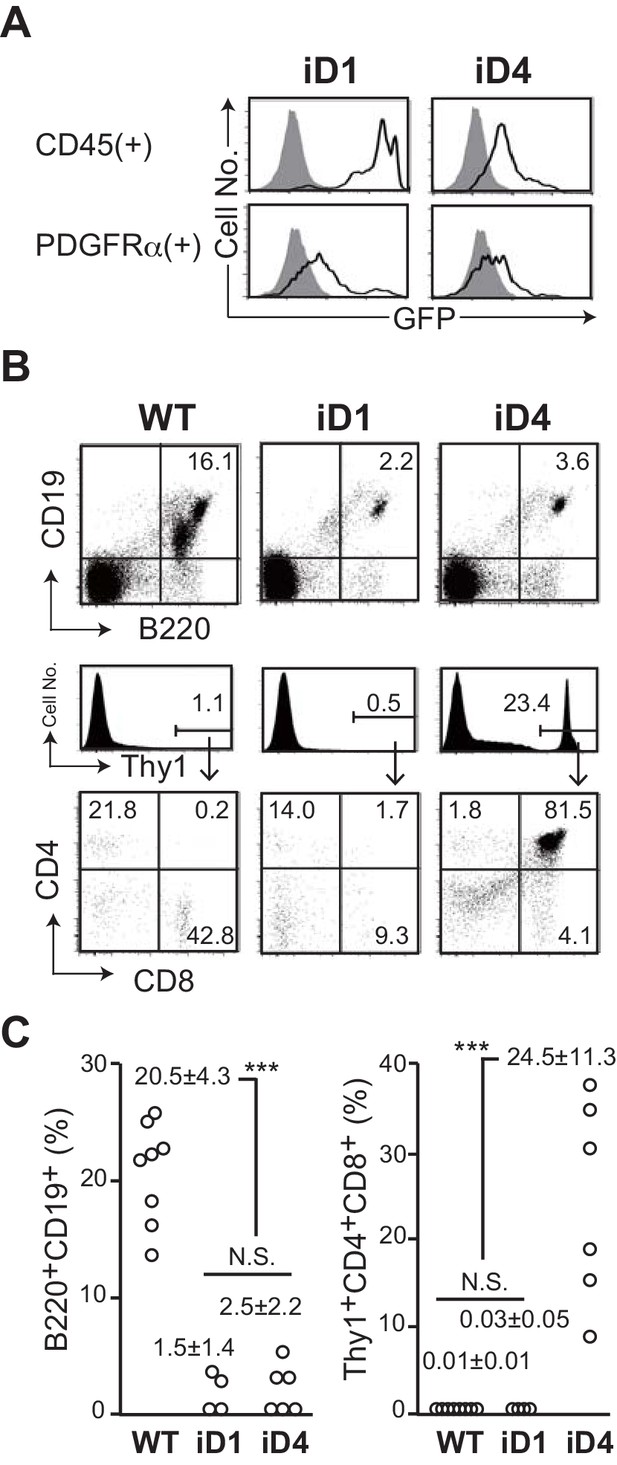
Effect of ectopic expression of Dll1 and Dll4 on the lymphopoiesis in the bone marrow.
(A) GFP expression, which transcripts is driven by CAG promoter at the Rosa26 locus of iD1 or iD4 mice, is detected in CD45+ hematopoietic or PDGFRα+ mesenchymal cell lineages in the bone marrow (BM) by flow cytometry. Open histograms indicate GFP expression of iD1 or iD4 mice, and filled histograms indicate the intrinsic fluorescence of the identical cell population of WT control mice. (B) Flow cytometry of the hematopoietic cells in BM obtained from tamoxifen-administrated WT, iD1, or iD4 mice with RosaCreER allele. One month after the last administration of tamoxifen, cells in BM were obtained and stained with mAbs against surface molecules as shown. Numbers in the profiles indicate the relative percentages, in Thy1+ cells (bottom, CD8 vs. CD4), for each quadrant or fraction. (C) The frequencies of B220+CD19+ B-lineage cells and Thy1+CD4+CD8+ immature T cells among the total cells of BM were examined as shown in B (mean ± SD; WT, n = 8; iD1, n = 4; iD4, n = 6; ***, p<0.001; N.S.: not significant; unpaired Student’s t-test).
-
Figure 1—source data 1
Raw data used to generate the graph in Figure 1C.
- https://cdn.elifesciences.org/articles/50979/elife-50979-fig1-data1-v3.xlsx
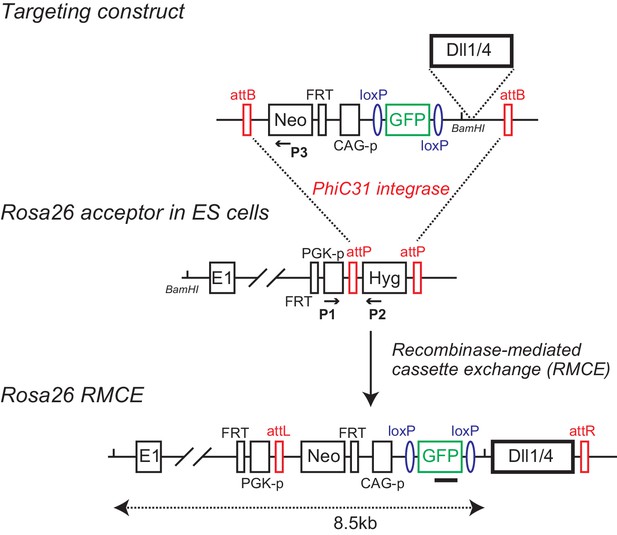
Establishment of conditional Dll transgenic mice by recombinase-mediated cassette exchange (RMCE).
Murine Dll1 or Dll4 cDNAs are placed after the floxed GFP cDNA (and polyA sequence) driven by CAG promoter (CAG-p). The promoterless neomycin-resistant gene (Neo) and FRT sequence were also flanked by attB sites as the targeting construct (Targeting construct). Acceptor ES cells for RMCE harbor a Rosa26 allele where PGK promoter (PGK-p)-driving a hygromycin selection marker (Hyg) is inserted with an FRT site behind the 1st exon, which is flanked by attP sites (Rosa26 acceptor in ES cells). Upon RMCE with ϕC31 integrase, the attP flanked cassette in the acceptor ES cells is replaced by the attB flanked cassette from the targeting construct. FRT sites allow excision of the PGK promoter and neomycin-resistant gene in recombined ES cells or mice. The arrows and filled square depict the primers (P1, P2, and P3) and probe for PCR and Southern blotting, respectively, to detect ES cell clone bearing Rosa26 locus with correct cassette exchange.
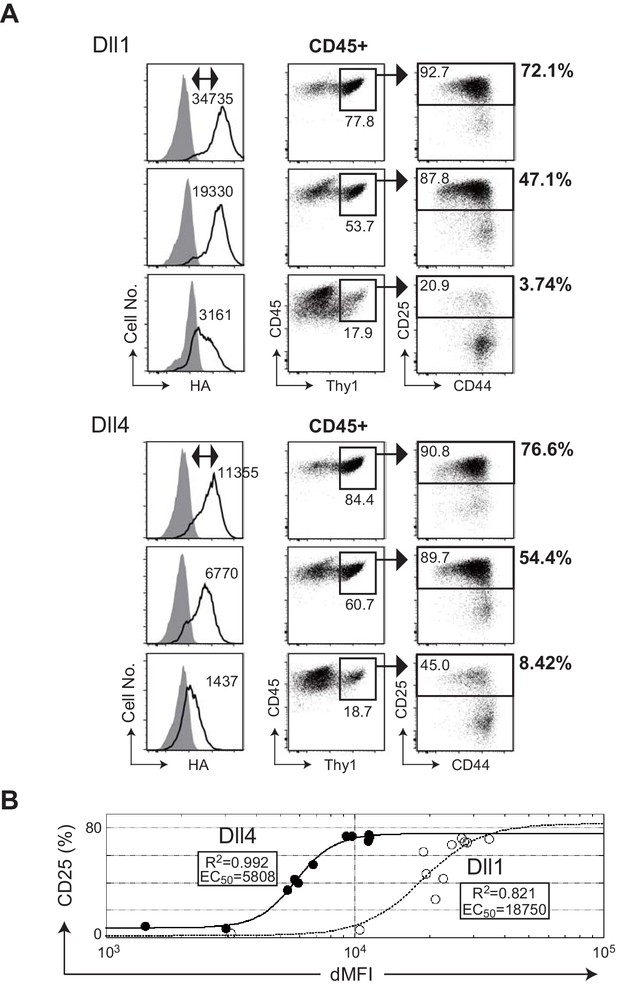
The efficiencies of T cell induction from hematopoietic progenitors by Dll1 or Dll4 on the monolayer cultures with OP9 stromal cells.
(A) Serial induction of exogenous Dll1 or Dll4 labeled at the C-terminus with HA-epitope driven by Tet-off system in OP9 cells was detected by flow cytometry using anti-HA mAb for intracellular staining. OP9 transfectants were treated with doxycycline (0.01, 0.8, and 0.3 ng/mL) for Dll1 or Dll4 to suppress the full activation of their transcription. Open histograms indicate anti-HA mAb staining (Dll1 or Dll4), and filled histograms indicate staining with control rabbit IgG. The difference of each MFI (dMFI) is shown in the left panels. E14.5 fetal liver-derived lineage markers (Gr-1, CD11b, TER119, and CD19)-negative c-kit-positive hematopoietic progenitors were cultured on the Dll1- or Dll4-bearing OP9 cells for 7 days and stained with lineage markers for analysis (Gr-1, CD11b, ST2, and DX5) and (CD45, CD19, Thy1, CD44, and CD25). CD45+ and CD45+Thy1.2+ cells were analyzed in the center and right panels, respectively. The frequencies (%) of lineage markers-negative CD19-Thy1+CD25+ T-lineage cells in CD45+ live cells are shown in the right side of the panels. Numbers in the dot-plot represent the relative percentages for each corresponding fraction. (B) The effectiveness of serial expression (dMFI) of Dll1 (open circle, dotted line) or Dll4 (closed circle, solid line) on OP9 cells for the induction of CD25+T-lineage cells (CD25) shown in A was evaluated by logistic regression analysis. In a 4-parameter logistic equation, EC50 and R2 values were calculated and shown in the graph.
-
Figure 2—source data 1
Raw data used to generate the graph in Figure 2B.
- https://cdn.elifesciences.org/articles/50979/elife-50979-fig2-data1-v3.xlsx
-
Figure 2—source data 2
Measurement of the induction efficiencies for the CD25+ T-lineage cells of serial expression of Dll4 or Dll1 as EC50 as shown in Figure 2B.
- https://cdn.elifesciences.org/articles/50979/elife-50979-fig2-data2-v3.xlsx
-
Figure 2—source data 3
Raw data used to generate Figure 2—source data 2.
- https://cdn.elifesciences.org/articles/50979/elife-50979-fig2-data3-v3.xlsx
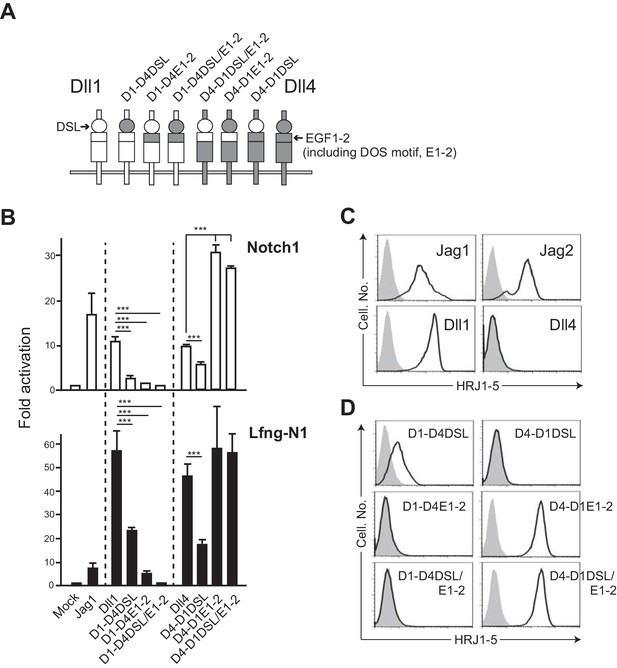
The swapping chimeras of DSL domain and/or the 1st/2nd EGF-like repeats between Dll1 and Dll4.
(A) Schematic structure of the Dll variants. Dll1 and Dll4 were intact and depicted by open (Dll1) or filled (Dll4) columns. The DSL domain and the 1st/2nd EGF-like repeats (E1-2) were represented by circle and top region of square, respectively. D1-D4DSL, D1-D4E1-2, and D1-DSL/E1-2 were Dll1-based chimeras with Dll4-derived DSL domain and/or E1-2. Similarly, Dll4-based chimeras were generated with Dll1-derived domains (D4-D1DSL, D4-D1E1-2, and D4-D1DSL/E1-2). Expression of NotchLs were monitored by GFP expression (Figure 3—figure supplement 2). (B) The swapping chimeras of DSL domain and/or the 1st/2nd EGF-like repeats transduce Notch signaling. Stable transfectants expressing murine Notch1 and control vector (Notch1, open column) or Notch1 and Lfng (Lfng-N1, filled column) were transiently transfected with a TP1-luciferase reporter plasmid, pGa981-6, and a pRL-TK plasmid for internal control. Cells were harvested at 24 hr after transfection, and co-cultured for an additional 40 hr with the transfectants expressing the DSL/E1-2 swapping chimeras. The relative induction of luciferase activity in each sample (mean ± SD, n = 3; ***, p<0.001; unpaired Student’s t-test) was calculated and described as fold activation against the control (value from the culture with mock transfectant not expressing Notch ligand). Data represents three independent experiments. (C) Monoclonal antibody originally established by us, HRJ1-5, broadly reacted with murine Jag1, Jag2, and Dll1, but not with Dll4. The transfectants of BM-derived mesenchymal cell line, originally not expressing any Notch ligands, established in our lab, expressing murine Jag1, Jag2, Dll1, or Dll4, were stained with HRJ1-5, and analyzed by flow cytometry. Open histograms indicate HRJ1-5 staining and filled histograms indicate staining with control hamster IgG. (D) Reactivity of HRJ1-5 with Dll chimeras. Each transfectant shown in the panel was stained with HRJ1-5 and analyzed as in C.
-
Figure 3—source data 1
Raw data (Fold activation) of luciferase activity used to generate the graph in Figure 3B.
- https://cdn.elifesciences.org/articles/50979/elife-50979-fig3-data1-v3.xlsx
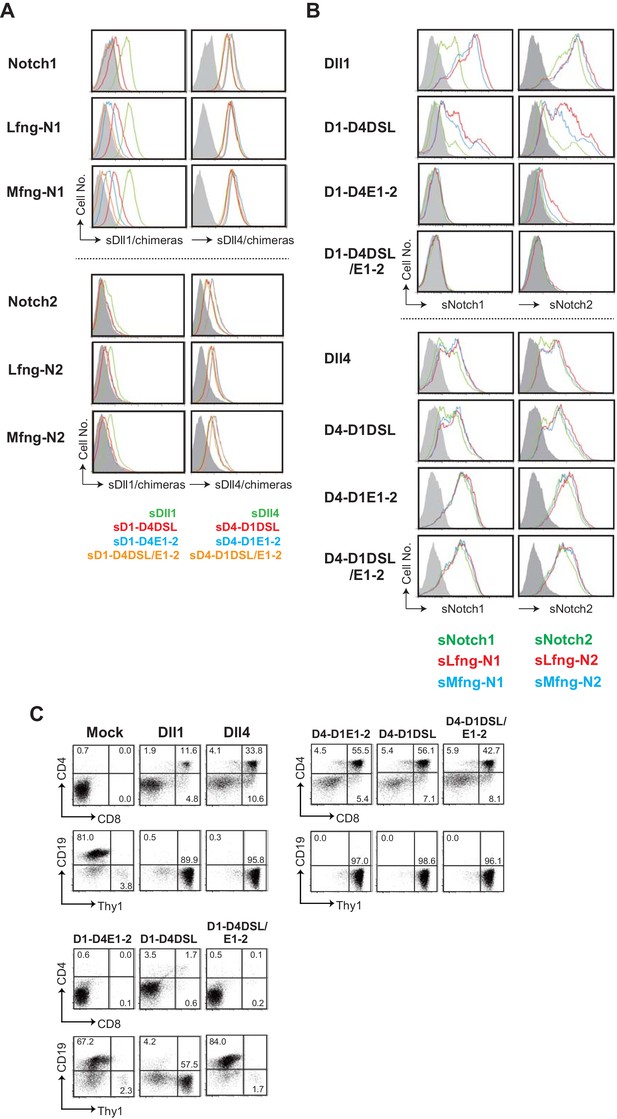
Characterization of the swapping chimeras of DSL and/or the 1st/2nd EGF-like repeats (EGF1-2) between Dll1 and Dll4.
(A) Binding activity of soluble swapping chimeras based on Dll1 or Dll4 to Notch1 or Notch2 expressed on the surface of BM-derived mesenchymal cells with or without the glycosylation by lunatic or manic fringe (Lfng, Mfng) is measured by flow cytometry. Soluble Dll chimeras, depicted in Figure 3A, are composed of the extracellular region of Dll1, Dll4, or their chimeras and Fc domain of human IgG1. Soluble Dll1 (green), D1-D4DSL (red), D1-D4DOS (blue), and D1-D4DSL/DOS (orange), shown in the right side of panels, were prepared and incubated with Notch1 transfectants. Their bindings were detected by anti-human IgG Ab and shown in the first lane of upper panels. Soluble Dll4 (green), D4-D1DSL (red), D4-D1DOS (blue), and D4-D1DSL/DOS (orange) were also used in the second lane of upper panels. Similarly, soluble Dll-derived molecules were used to check their binding activities to the Notch2 transfectants with or without glycosylation in the lower panels. (B) Binding activity of the DSL/EGF1-2 swapping chimeras with soluble Notch1 and Notch2. Soluble Notch receptors are composed of the N-terminus to 15th EGF repeats and the Fc domain of human IgG1. The transfectants expressing Dll1, Dll4, or their chimeras were incubated with soluble Notch1 (sNotch1) or Notch2 (sNotch2), and their binding were detected. The soluble Notchs were produced by CHO cells expressing murine Lfng (red line), Mfng (blue line), or vector control (green line), and used in this experiment. (C) Induction of T-lineage cells by the swapping chimeras in vitro. Fetal liver-derived lineage markers-negative c-kit-positive hematopoietic progenitors were cultured on the OP9 cells expressing Dll1, Dll4, or their chimeras for 13 days. After the cultures, live cells were stained with CD19, Thy1, CD4, and CD8, and analyzed by flow cytometry. Numbers in the dot-plot represent the relative percentages for each corresponding quadrants.
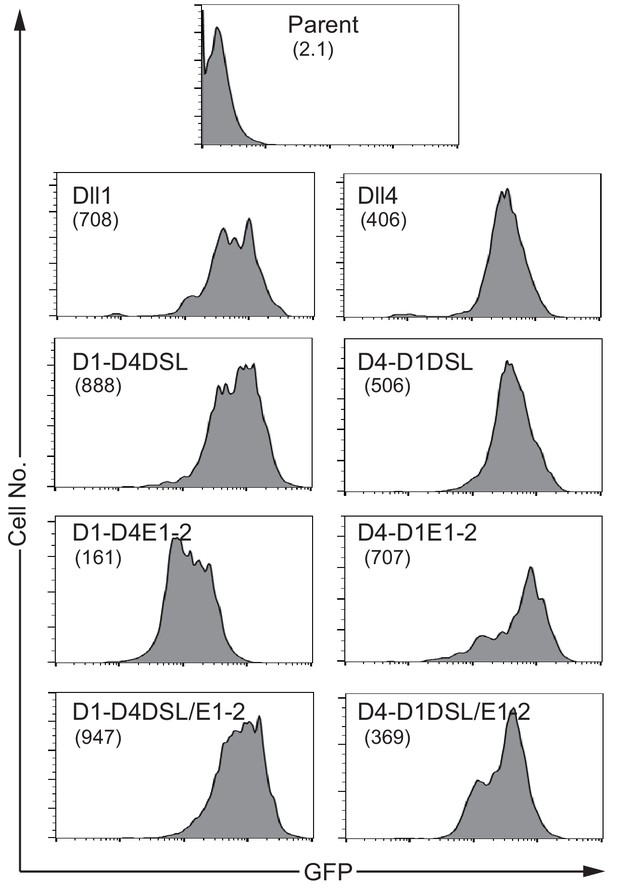
NotchLs transfectants used in Figure 3 were established via infection with a retrovirus encoding the respective NotchL or their chimeric molecules of parental fibroblast cell lines (parent), and were monitored via GFP expression by flow cytometry.
The numbers in parenthesis represent the mean fluorescence intensity (MFI).
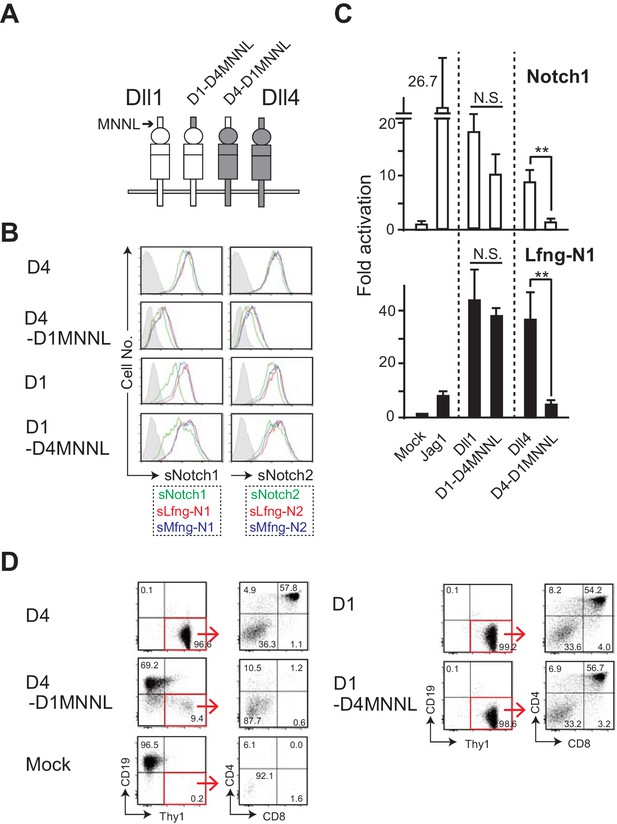
The swapping chimera of MNNL domain between Dll1 and Dll4.
(A) Schematic structure of the Dll variant of MNNL (the N-terminal) domain. D1-D4MNNL and D4-D1MNNL were Dll1- and Dll4-based chimeras with Dll4- and Dll1-derived MNNL domains, respectively. Expression of NotchLs were monitored by the GFP expression, the intracellular staining with anti-HA mAb and the surface staining with HRJ1-5 or anti-Dll4 mAbs (Figure 4—figure supplement 1). (B) Binding activity of the MNNL swapping chimeras with soluble Notch1 and Notch2. Soluble Notch receptors comprised the N-terminus to the 15th EGF repeats and the Fc domain of human IgG1. The transfectants of BM-derived mesenchymal cell line expressing Dll1 (D1), Dll4 (D4), or their MNNL swapping chimeras (D4-D1MNNL and D1-D4MNNL) were incubated with soluble Notch1 (sN1) or Notch2 (sN2), and their binding was detected by anti-human IgG Ab as described in the Materials and methods. Soluble Notchs were produced by CHO cells expressing murine Lfng (red line), Mfng (blue line), or vector control (green line), and used in this experiment. (C) The MNNL swapping chimeras transduce Notch signaling. Reporter assay was carried out as shown in Figure 3B. The relative induction of luciferase activity in each sample (mean ± SD, n = 3; **, p<0.01; N.S.: not significant; unpaired Student’s t-test) was calculated. Data represents three independent experiments. (D) Induction of T-lineage cells by MNNL swapping chimeras in in vitro. Fetal liver-derived lineage markers-negative c-kit-positive hematopoietic progenitors were cultured on the OP9 cells expressing Dll1, Dll4, or their MNNL chimeras for 13 days. After the cultures, live cells were stained with CD19, Thy1, CD4, and CD8, and analyzed by flow cytometry. Numbers in the dot-plot represent the relative percentages for each corresponding fraction or quadrant in Thy1+CD19- fraction (red square).
-
Figure 4—source data 1
Raw data (Fold activation) of luciferase activity used to generate the graph in Figure 4C.
- https://cdn.elifesciences.org/articles/50979/elife-50979-fig4-data1-v3.xlsx
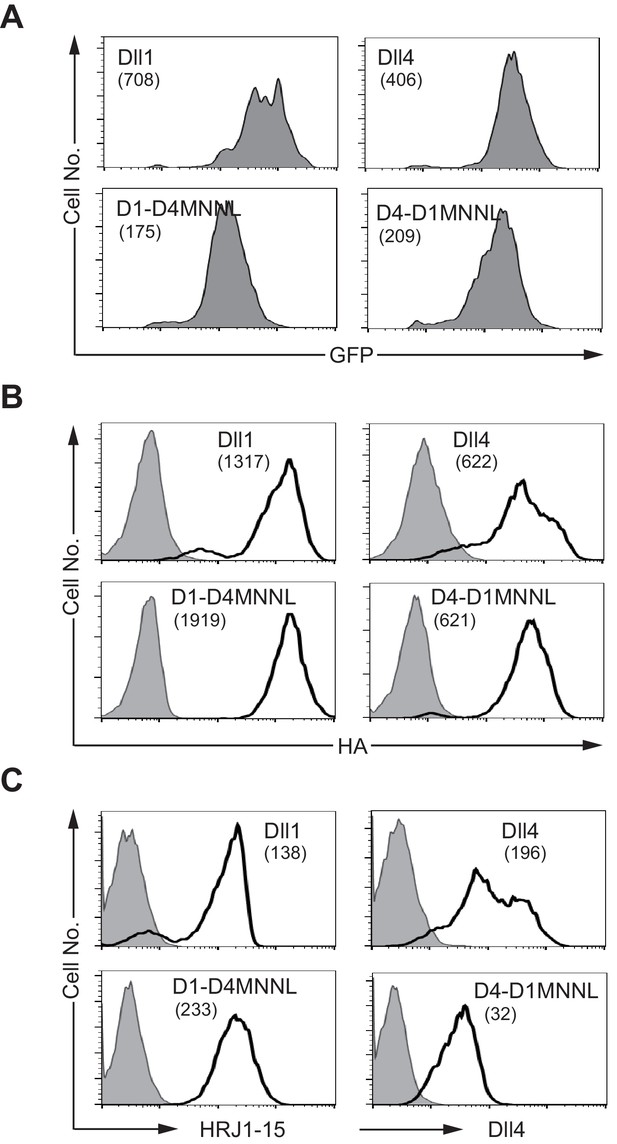
NotchLs transfectants used in Figure 4 were monitored by the expression of GFP and the staining with anti-HA , anti-Dll4 or HRJ1-5 mAbs.
(A) Intracellular staining with anti-HA mAb was used to detect the total amount of NotchL (B, open histogram) or control rabbit IgG (B, filled histogram). The surface staining with HRJ1-5 mAb marks the DOS motif at the E1-2 region (C, open histogram, left panels). The anti-Dll4 mAb recognizes mainly the DSL region of Dll4 (C, open histogram, right panels). Hamster IgG is used as a control (C, filled histogram). These were monitored by flow cytometry. The numbers in parenthesis represent the mean fluorescence intensity (MFI) of GFP (A), or the difference of each MFI (B and C, dMFI).
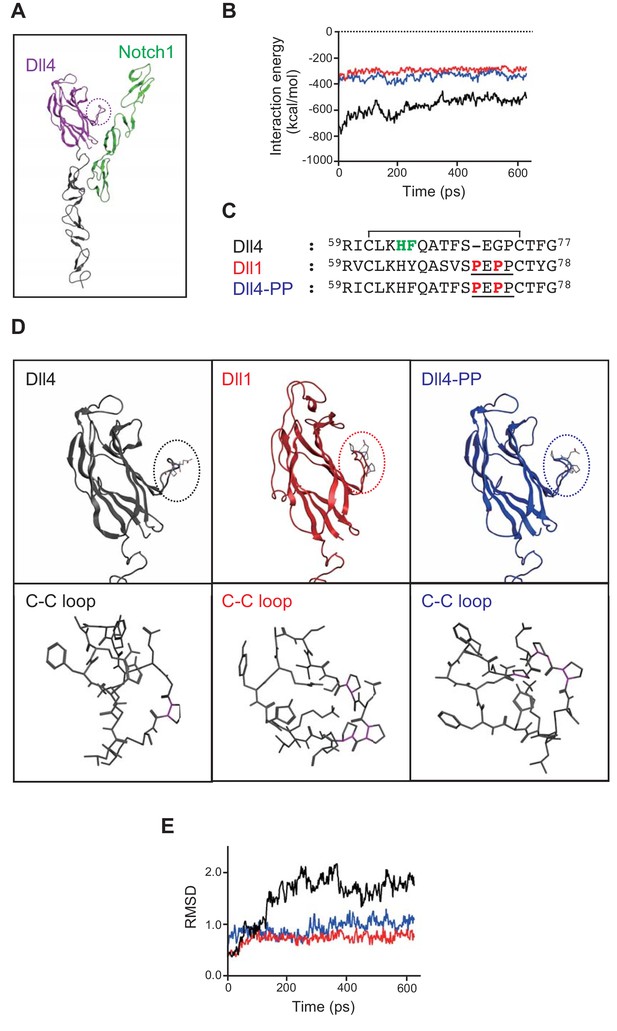
In silico analysis of Dll/Notch1 complexes.
(A) Structure of Dll4 (MNNL, purple; DSL and EGF1-2, black) bound to Notch1 (EGF11-13, green) is shown in ribbon representation. The dotted circle indicates the loop structure with disulfide bond (C-C loop) in MNNL domain. (B) Molecular dynamic simulation of 600 ps for the interaction energy of Dll4/Notch1 (black), D4-D1MNNL/Nothc1 (red), and Dll4-based mutant with two proline residues (characteristic of Dll1) (Dll4-PP/Notch1, blue). (C) Amino acid (AA) sequence comparison of the C-C loop in MNNL domain between Dll4, Dll1, and Dll4-PP. Numbers on the AA sequences represent the position from the N-terminus. D4-PP is the Dll4-based mutant with the inserted (71st position) and substituted (73rd position) mutations of proline residue (bold red; characteristic of Dll1, underline). The AAs in the C-C loop contributing to the direct binding with Notch receptor are labeled (bold green) described as previously. Line over the sequence represents the disulfide bridge between cysteine residues (61st to 74th). (D) Structure of MNNL domain of Dll4 (black), Dll1 (red), and Dll4-PP (blue) in ribbon representation (upper panels) with enlarged wireframe model of the C-C loop (lower panels). Magenta, C-N bond in proline. (E) Molecular dynamic simulation of 600 ps for the RMSD of the C-C loop in the MNNL domain of Dll4 (black), Dll1 (red), and Dll4-PP (blue).
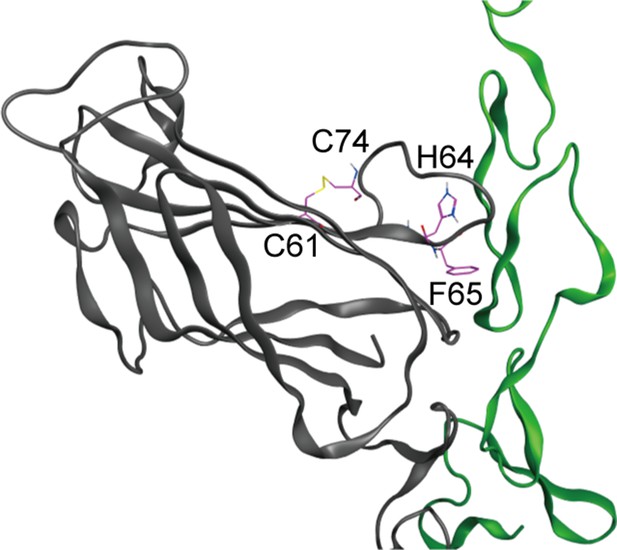
The loop structure with disulfide bond between 61Cys (C61) and 72Cys (C72) includes the key residues, 64His (H64) and 65Phe (F65), filled circle in Figure 5C, that comprise the binding surface of MNNL domain of Dll4 which interacts with Notch1.
Molecular dynamics simulation of Dll4 (Figure 5—video 1, black), Dll1 (Figure 5—video 2, red) and Dll4-PP (Figure 5—video 3, blue) with Notch1 (green) for 600ps were exhibited in the movies.
Molecular simulation of Dll1.
Molecular simulation of Dll4-PP.
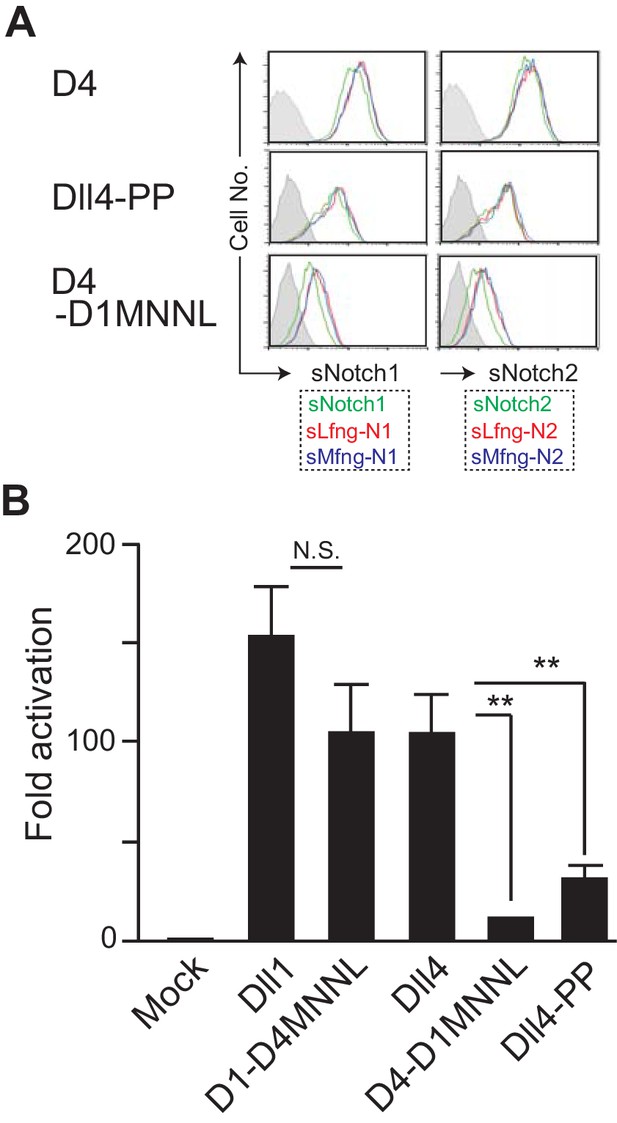
Effect of the mutation in the loop structure of MNNL domain of Dll4.
(A) Binding activity with the soluble Notch1 and Notch2 was detected by flow cytometry as shown in Figure 4B. (B) Inducing activity of Notch signaling in vitro was examined by the co-cultures of NotchL and Notch1/Lfng transfectants as shown in Figure 3B. The fold activation against the control was calculated from each sample (mean ± SD, n = 3; **, p<0.01; N.S.: not significant; unpaired Student’s t-test). Data represents three independent experiments. Expression of NotchLs were monitored by the GFP expression, the intracellular staining with anti-HA mAb and the surface staining with HRJ1-5 or anti-Dll4 mAbs (Figure 6—figure supplement 1).
-
Figure 6—source data 1
Raw data (Fold activation) of luciferase activity used to generate the graph in Figure 6B.
- https://cdn.elifesciences.org/articles/50979/elife-50979-fig6-data1-v3.xlsx
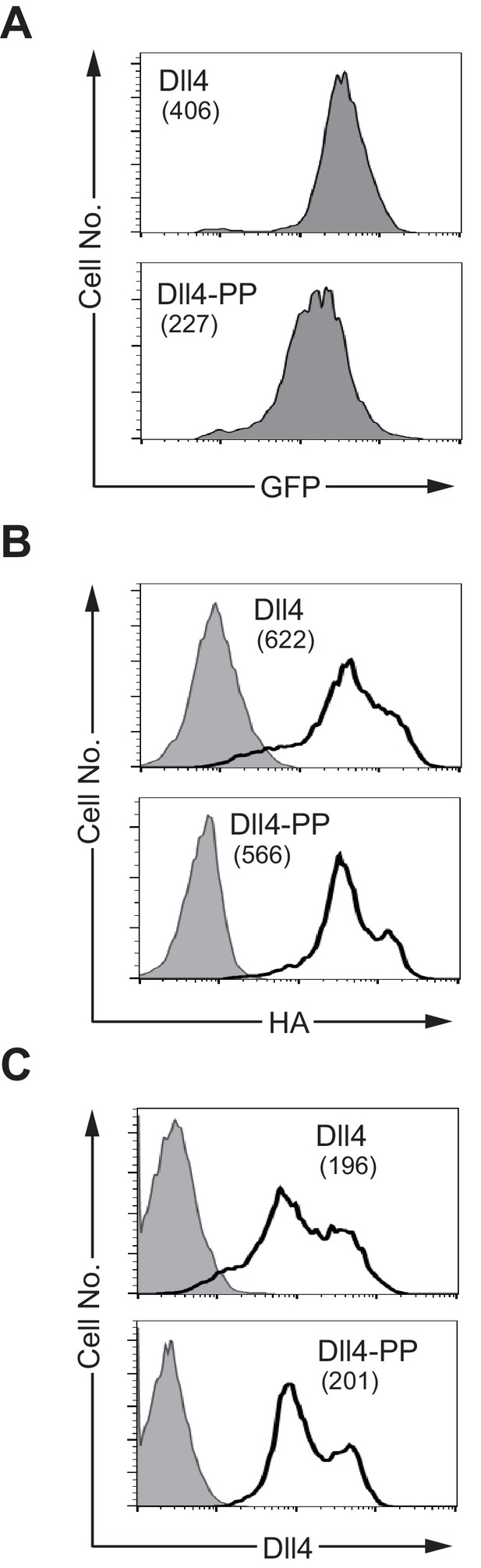
NotchLs transfectants used in Figure 6 were monitored by the expression of GFP and the staining with anti-HA, anti-Dll4 or HRJ1-5 mAbs.
The transfectants of Dll4 and D4-PP were monitored by the expression of GFP (A). The intracellular staining with anti-HA mAb (B) and the surface staining with anti-HRJ1-5 mAb recognizes the DOS motif at the E1-2 region (C, left panels). The anti-Dll4 mAb (C, right panels) recognizes mainly the DSL region of Dll4 as shown in Figure 4—figure supplement 1. The numbers in parenthesis represent the mean fluorescence intensity (MFI) of GFP (A), or the difference of each MFI (B and C, dMFI).
Additional files
-
Supplementary file 1
Key resources table.
- https://cdn.elifesciences.org/articles/50979/elife-50979-supp1-v3.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/50979/elife-50979-transrepform-v3.docx





