Movement of accessible plasma membrane cholesterol by the GRAMD1 lipid transfer protein complex
Figures
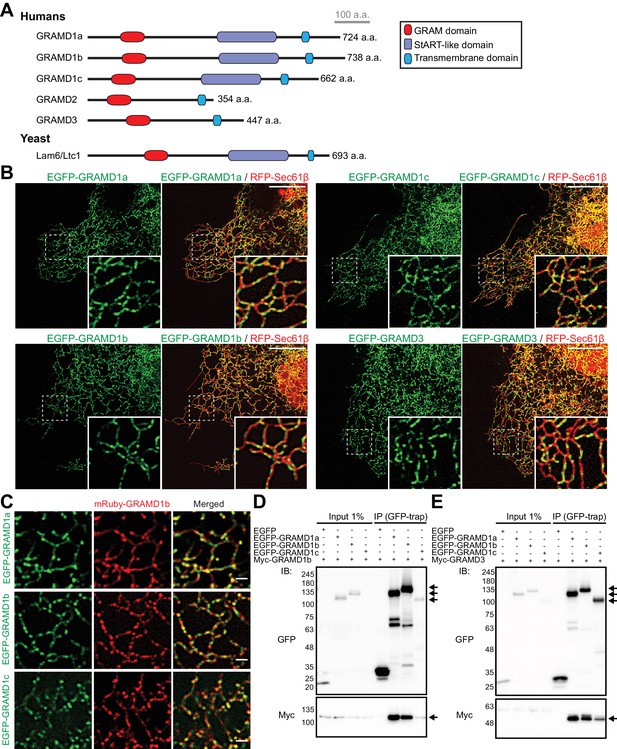
GRAMD proteins form homo- and heteromeric complexes.
(A) Domain structure of GRAMD proteins in comparison to yeast Lam6/Ltc1. (B) Confocal images of live COS-7 cells expressing the ER membrane marker RFP-Sec61β and EGFP–GRAMD protein constructs as indicated. Insets show at higher magnification the regions indicated by white dashed boxes. Note the presence of the patches of EGFP–GRAMDs throughout the tubular ER. Scale bars, 10 µm. (C) Confocal images of live COS-7 cells expressing mRuby-GRAMD1b and EGFP–GRAMD1s as indicated. Note the presence of mRuby–GRAMD1b patches that partially overlap with the patches of EGFP–GRAMD1s. Scale bars, 1 µm. (D, E) Extracts of HeLa cells transfected with the indicated constructs were subjected to anti-GFP immunoprecipitation (IP) and then processed for SDS-PAGE and immunoblotting (IB) with anti-GFP and anti-Myc antibodies. Inputs are 1% of the total cell lysates. Note the strong biochemical interaction between GRAMD1b and GRAMD1s (D) and between GRAMD3 and GRAMD1s (E). Immunoprecipitated EGFP-GRAMD1s, Myc-GRAMD1b and Myc-GRAMD3 are indicated by arrows.
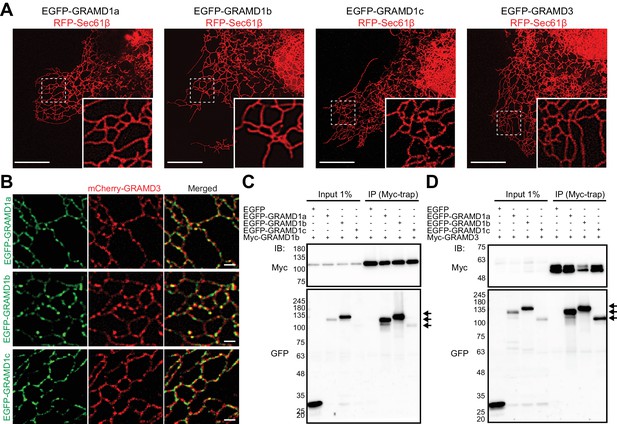
GRAMD proteins form homo- and heteromeric complexes.
(A) Confocal images of live COS-7 cells expressing the ER membrane marker RFP-Sec61β and EGFP–GRAMD protein constructs as indicated (compare with Figure 1B). Insets show at higher magnification the regions indicated by white dashed boxes. Note that the overexpression of EGFP–GRAMD1s does not affect tubular ER structure. Scale bars, 10 µm. (B) Confocal images of live COS-7 cells expressing mCherry–GRAMD3 and EGFP–GRAMD1s as indicated. Note the presence of mCherry–GRAMD3 patches that partially overlap with the patches of EGFP–GRAMD1s. Scale bars, 1 µm. (C, D) Extracts of HeLa cells transfected with the constructs as indicated were subjected to anti-Myc immunoprecipitation (IP) and then processed for SDS-PAGE and immunoblotting (IB) with anti-Myc and anti-GFP antibodies. Inputs are 1% of the total cell lysates. Note the strong biochemical interaction between GRAMD1b and GRAMD1s (C) and between GRAMD3 and GRAMD1s (D). EGFP–GRAMD1s that are present in the anti-Myc immunoprecipitates are indicated by arrows.
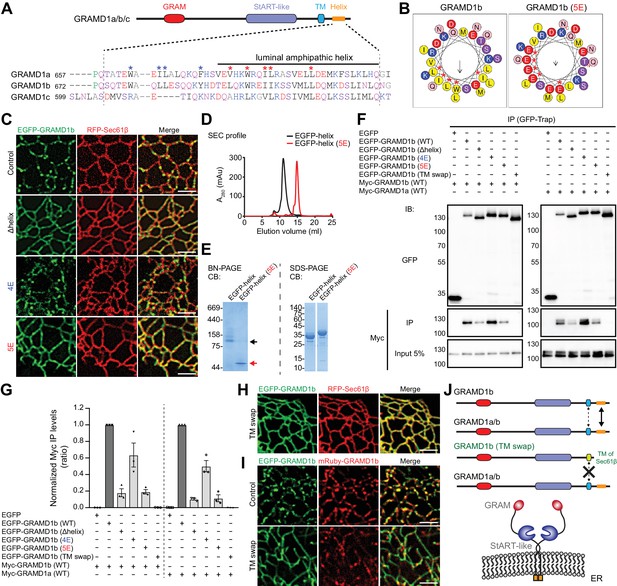
Luminal helix and transmembrane domain of GRAMD1b are important for homo- and heteromeric interaction.
(A) Sequence alignment of the luminal region of GRAMD1s. This region is predicted by Phyre2 to contain an amphipathic helix (Kelley et al., 2015) as indicated. Blue and red asterisks mark hydrophobic amino acid residues that are partially conserved in GRAMD1s. The shared identities of the amino acid sequences of the amphipathic helices predicted by BLAST analysis were: 75% (GRAMD1a vs. GRAMD1b); 75% (GRAMD1a vs. GRAMD1c); and 80% (GRAMD1b vs. GRAMD1c). The effects of the mutations of these residues to glutamic acid (4E in the case of blue marks; 5E in the case of red marks) were tested in GRAMD1b. Black, red, blue, and pink/purple colors denote hydrophobic, acidic, basic, and hydrophilic amino acid residues, respectively. (B) Predicted luminal amphipathic helix region of wild-type GRAMD1b (left panel) and that with L693E, W696E, I699E, I700E and L707E (5E) mutations (right panel) are shown as helical wheel representations. Predictions were made with the Heliquest server (Gautier et al., 2008). (C) Confocal images of live COS-7 cells expressing RFP–Sec61β and EGFP fusions of various GRAMD1b constructs [control, wild-type GRAMD1b; Δhelix, GRAMD1b lacking the predicted luminal amphipathic helix; 4E, GRAMD1b with 4E mutations in the luminal region (W678E, L681E, L682E, Y688E); 5E, GRAMD1b with 5E mutations in the predicted luminal amphipathic helix]. Note the reduced formation of GRAMD1b patches in Δhelix and 5E mutants but not in the 4E mutant. Scale bars, 2 µm. (D) Overlay of the size exclusion chromatography (SEC) profiles of the recombinant EGFP-tagged luminal helix region of wild-type GRAMD1b (EGFP–helix) and EGFP–helix with the 5E mutations [EGFP–helix (5E)]. Note the difference in elution volumes, indicating the formation of complexes mediated by the wild-type luminal helix. (E) Blue native (BN)-PAGE analysis (left panel) and SDS-PAGE analysis (right panel) of SEC-purified EGFP–helix and EGFP–helix (5E). Black and red arrows indicate the major bands for EGFP–helix and EGFP–helix 5E, respectively. Note the difference in their migration pattern in BN-PAGE. CB, Colloidal blue staining. (F) Extracts of HeLa cells transfected with the constructs as indicated were subjected to anti-GFP immunoprecipitation (IP) and then processed for SDS-PAGE and immunoblotting (IB) with anti-GFP and anti-Myc antibodies. Inputs are 5% of the total cell lysates. Note that the interaction of GRAMD1b or GRAMD1a is much reduced in GRAMD1b Δhelix or 5E mutants and abolished in the GRAMD1b (TM swap) mutant (GRAMD1b with its transmembrane domain and luminal region replaced with those of Sec61β) when compared to the levels of interactions seen in cells with wild-type GRAMD1b. This reduction is smaller in the GRAMD1b 4E mutant. (G) Quantification of the co-immunoprecipitation experiments shown in (F). The ratio of the band intensity of the co-immunoprecipitated Myc–GRAMD1b (left) or Myc–GRAMD1a (right) over that of the indicated immunoprecipitated EGFP-tagged proteins were calculated. The values were then normalized by the ratio of the band intensity of Myc–GRAMD1b over that of EGFP–GRAMD1b (WT) (left) or by the ratio of the band intensity of Myc–GRAMD1a over that of EGFP–GRAMD1b (WT) (right) [mean ± SEM, n = 3 IPs for each sample]. (H) Confocal images of a live COS-7 cell expressing RFP-Sec61β and EGFP-tagged GRAMD1b (TM swap). Scale bars, 2 µm. (I) Confocal images of live COS-7 cells expressing mRuby–GRAMD1b and EGFP fusions of GRAMD1b constructs [Control, wild-type GRAMD1b; TM swap, GRAMD1b (TM swap)]. Note the abolished formation of GRAMD1b patches in TM swap mutants. Scale bars, 2 µm. (J) Model of the homo- and heteromeric interactions of GRAMD1a/b. Their complex formation is facilitated primarily by their luminal amphipathic helices and additionally mediated by their transmembrane domains. These regions are important for the ability of GRAMD1s to form complexes and patches on the tubular ER network.
-
Figure 2—source data 1
Dataset for Figure 2.
- https://cdn.elifesciences.org/articles/51401/elife-51401-fig2-data1-v2.xlsx
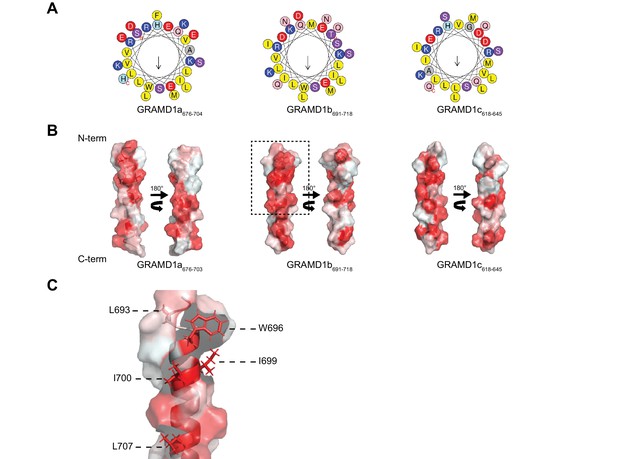
Luminal helix of GRAMD1b is important for homo- and heteromeric interaction.
(A) Predicted luminal amphipathic helix regions of GRAMD1s are shown as helical wheel representations. Predictions were made with the Heliquest server (Gautier et al., 2008). (B) Hydrophobic surface representation of the predicted luminal amphipathic helices of GRAMD1s (see Materials and methods). The red color indicates the hydrophobicity of the amino acid residues. (C) A magnified image of the region of the predicted amphipathic helix of GRAMD1b indicated in a dashed box in (B), showing five amino-acid residues (L693, W696, I699, I700 and L707) that were mutated to glutamic acid in the 5E mutant. Images were generated using the PyMol Molecular Graphics System.
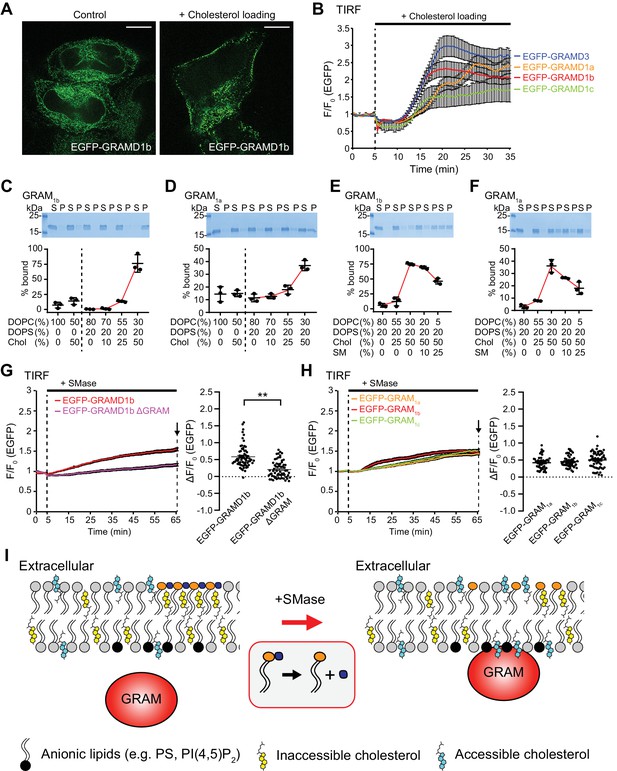
The GRAM domain of GRAMD1s acts as a coincidence detector of unsequestered/accessible cholesterol and anionic lipids, and senses a transient expansion of the accessible pool of cholesterol in the PM.
(A) Confocal images of live HeLa cells expressing EGFP–GRAMD1b with or without cholesterol loading [the treatment with cholesterol/MCD complex (200 µM) for 30 min at 37°C]. Note the extensive recruitment of GRAMD1b to the PM upon cholesterol loading. Scale bars, 10 µm. (B) Time course of normalized EGFP signal, as assessed by total internal reflection fluorescence (TIRF) microscopy, from HeLa cells expressing EGFP–GRAMD protein constructs as indicated. Cholesterol loading [the treatment with cholesterol/MCD complex (200 µM)] is indicated. [mean ± SEM, n = 24 cells (EGFP–GRAMD1a), n = 29 cells (EGFP–GRAMD1b), n = 25 cells (EGFP–GRAMD1c), n = 28 cells (EGFP–GRAMD3); data are pooled from one experiment for GRAMD1a and two experiments for GRAMD1b, GRAMD1c and GRAMD3.] (C–F). Liposome sedimentation assays of the GRAM domain of GRAMD1b (GRAM1b) and GRAMD1a (GRAM1a). Liposomes containing the indicated mole% lipids were incubated with purified GRAM1b proteins (C, E) or purified GRAM1a proteins (D, F). Bound proteins [pellet, (P)] were separated from the unbound proteins [supernatant, (S)], run on SDS-PAGE and visualized by colloidal blue staining (mean ± SEM, n = 3 independent experiments for all the conditions). DOPC, phosphatidylcholine (1,2-dioleoyl-sn-glycero-3-phosphocholine); DOPS, phosphatidylserine (1,2-dioleoyl-sn-glycero-3-phospho-L-serine); Chol, cholesterol; SM, sphingomyelin (N-oleoyl-D-erythro-sphingosylphosphorylcholine). (G) Left: time course of normalized EGFP signal in response to sphingomyelinase (SMase), as assessed by TIRF microscopy of HeLa cells expressing EGFP–GRAMD1b or EGFP–GRAMD1b ΔGRAM. The treatment with SMase (100 mU/ml) is indicated. Right: values of ΔF/F0 corresponding to the end of the experiment as indicated by the arrow [mean ± SEM, n = 72 cells (EGFP–GRAMD1b), n = 64 cells (EGFP–GRAMD1b ΔGRAM); data are pooled from three independent experiments for each condition; two-tailed unpaired Student’s t-test, **p<0.0001]. (H) Left: time course of normalized EGFP signal in response to SMase, as assessed by TIRF microscopy of HeLa cells expressing the indicated EGFP-tagged GRAM domain of GRAMD1s. The treatment with SMase (100 mU/ml) is indicated. Right: values of ΔF/F0 corresponding to the end of the experiment as indicated by the arrow [mean ± SEM, n = 48 cells (EGFP–GRAM1a), n = 50 cells (EGFP–GRAM1b), n = 58 cells (EGFP–GRAM1c); data are pooled from two to three independent experiments for each condition]. (I) Schematics showing the interaction of the GRAM domain of GRAMD1s with the plasma membrane (PM) before and after sphingomyelinase (SMase) treatment. Left: at rest, subthreshold levels of accessible cholesterol in the PM are not sufficient to induce interaction of the GRAM domain with the PM. Right: liberation of the sphingomyelin-sequestered pool of cholesterol by SMase treatment leads to an increase in accessible cholesterol in the PM beyond the threshold, and induces PM recruitment of the GRAM domain as it senses both increase in unsequestered/accessible cholesterol and the presence of anionic lipids in the PM.
-
Figure 3—source data 1
Dataset for Figure 3.
- https://cdn.elifesciences.org/articles/51401/elife-51401-fig3-data1-v2.xlsx
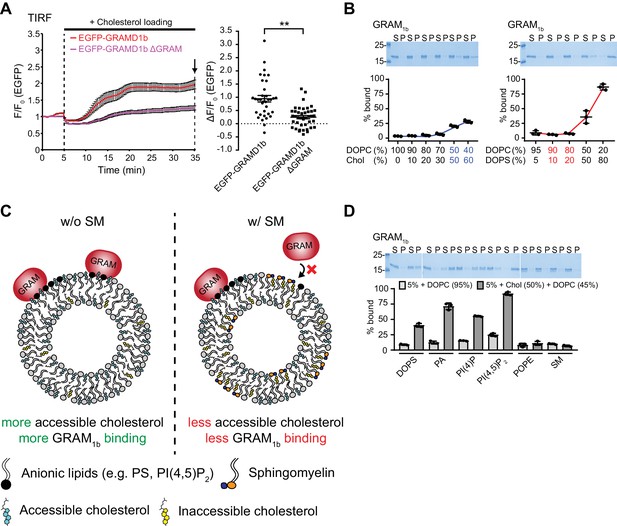
The GRAM domain of GRAMD1s acts as a coincidence detector of unsequestered/accessible cholesterol and anionic lipids.
(A) Left: time course of normalized EGFP signal, as assessed by TIRF microscopy, from HeLa cells expressing EGFP–GRAMD1b or EGFP–GRAMD1b ΔGRAM. Cholesterol loading [the treatment with cholesterol/methyl-β-cyclodextrin (MCD) complex (200 µM)] is indicated. Right: values of ΔF/F0 corresponding to the end of the experiment as indicated by the arrow [mean ± SEM, n = 35 cells (EGFP–GRAMD1b), n = 41 cells (EGFP–GRAMD1b ΔGRAM); data are pooled from two independent experiments for each condition; two-tailed unpaired Student’s t-test, **p<0.0001]. (B , D) Liposome sedimentation assays of the GRAM domain of GRAMD1b (GRAM1b). Liposomes containing the indicated mole% lipids were incubated with purified GRAM1b proteins. Bound proteins [pellet, (P)] were separated from the unbound proteins [supernatant, (S)], run on SDS-PAGE and visualized by colloidal blue staining. Note that the GRAM domain binds only weakly to liposomes when the liposomes contain high levels of cholesterol (50% or 60%, indicated by blue). Note also that the GRAM domain does not bind liposomes that contain physiological levels of PS (10% or 20%, indicated by red) (mean ± SEM, n = 3 independent experiments for all the conditions). DOPC, phosphatidylcholine (1,2-dioleoyl-sn-glycero-3-phosphocholine); DOPS, phosphatidylserine (1,2-dioleoyl-sn-glycero-3-phospho-L-serine); PA, phosphatidic acid (1,2-dioleoyl-sn-glycero-3-phosphate); POPE, phosphatidylethanolamine (1-palmitoyl-2-oleoyl-sn-glycero-3-phosphoethanolamine); SM, sphingomyelin (N-oleoyl-D-erythro-sphingosylphosphorylcholine); Chol, cholesterol; PI(4)P, phosphatidylinositol 4-phosphate; PI(4,5)P2, phosphatidylinositol 4,5-bisphosphate. (C) Schematics showing the interaction of the GRAM domain of GRAMD1s to artificial membranes with or without sphingomyelin (SM) incorporation in our liposome sedimentation assays. Left: the presence of both free cholesterol and anionic lipids in artificial membranes results in strong binding of the GRAM domain when mole% of free cholesterol exceeds a certain threshold. Right: incorporation of sphingomyelin into the artificial membranes reduces the accessibility of cholesterol in dose-dependent manner (due to complex formation between sphingomyelin and cholesterol) and suppresses the binding of the GRAM domain to the artificial membranes.
-
Figure 3—figure supplement 1—source data 1
Dataset for Figure 3—figure supplement 1.
- https://cdn.elifesciences.org/articles/51401/elife-51401-fig3-figsupp1-data1-v2.xlsx
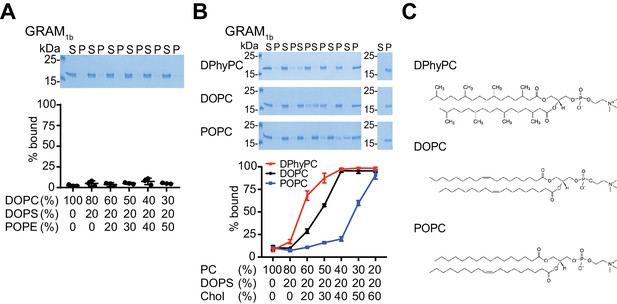
The GRAM domain of GRAMD1s binds to membranes by sensing cholesterol accessibility.
(A, B) Liposome sedimentation assays of the GRAM domain of GRAMD1b (GRAM1b). Liposomes containing the indicated mole% lipids were incubated with purified GRAM1b proteins. Bound proteins [pellet, (P)] were separated from the unbound proteins [supernatant, (S)] and run on SDS-PAGE and visualized by colloidal blue staining. Note that the GRAM domain no longer binds to phosphatidylserine-containing liposomes when cholesterol was replaced with phosphatidylethanolamine (compare with Figure 3C). Note also that the acyl chain diversity of phosphatidylcholine (PC) affects the binding of GRAM1b to liposomes containing phosphatidylserine and cholesterol. POPC has the strongest cholesterol sequestration effect in the membranes, followed by DOPC and DPhyPC (Sokolov and Radhakrishnan, 2010) (mean ± SEM, n = 3 independent experiments for all the conditions). DOPC, 1,2-dioleoyl-sn-glycero-3-phosphocholine; DPhyPC, 1,2-diphytanoyl-sn-glycero-3-phosphocholine; POPC, 1-palmitoyl-2-oleoyl-glycero-3-phosphocholine; DOPS, 1,2-dioleoyl-sn-glycero-3-phospho-L-serine; Chol, cholesterol; POPE, 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphoethanolamine. (C) Chemical structures of the phosphatidylcholines tested in the assay. Note the differences in acyl chain saturation and branching.
-
Figure 3—figure supplement 2—source data 1
Dataset for Figure 3—figure supplement 2.
- https://cdn.elifesciences.org/articles/51401/elife-51401-fig3-figsupp2-data1-v2.xlsx
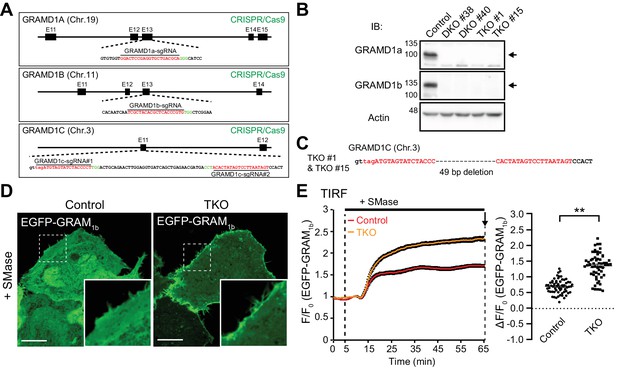
Deletion of GRAMD1s results in exaggerated accumulation of the accessible pool of cholesterol in the PM.
(A) Schematics of the Cas9/sgRNA targeting sites in human GRAMD1A, GRAMD1B and GRAMD1C loci. The targeting sequences are highlighted in red. The protospacer-adjacent motifs (PAMs) are labeled in green. (B) Lysates of control HeLa cells, two independently isolated GRAMD1a/1b DKO cell lines, and two independently isolated GRAMD1 TKO cell lines were processed by SDS–PAGE and immunoblotting (IB) with anti-GRAMD1a, anti-GRAMD1b and anti-actin antibodies. The arrows indicate the specific bands for GRAMD1a and GRAMD1b. (C) Nucleotide sequence analysis of the GRAMD1C gene of the GRAMD1 TKO cell lines. Guide RNA-targeting sites are highlighted in red. (D) Confocal images of live wild-type (Control) and GRAMD1 TKO (TKO) HeLa cells expressing the EGFP-tagged GRAM domain of GRAMD1b (EGFP–GRAM1b) with SMase treatment (100mU/ml for 1 hr at 37°C). Insets show at higher magnification the regions indicated by white dashed boxes. Note the strong recruitment of EGFP–GRAM1b to the PM of GRAMD1 TKO cells compared to that of the control cells. Scale bars, 10 µm. (E) Left: time course of normalized EGFP signal, as assessed by TIRF microscopy, from wild-type (Control) and GRAMD1 TKO (TKO) HeLa cells expressing EGFP–GRAM1b. SMase treatment (100 mU/ml) is indicated. Right: values of ΔF/F0 corresponding to the end of the experiment as indicated by the arrow [mean ± SEM, n = 62 cells (Control), n = 58 cells (TKO); data are pooled from three independent experiments for each condition; two-tailed unpaired Student’s t-test with equal variance, **p<0.0001].
-
Figure 4—source data 1
Dataset for Figure 4.
- https://cdn.elifesciences.org/articles/51401/elife-51401-fig4-data1-v2.xlsx
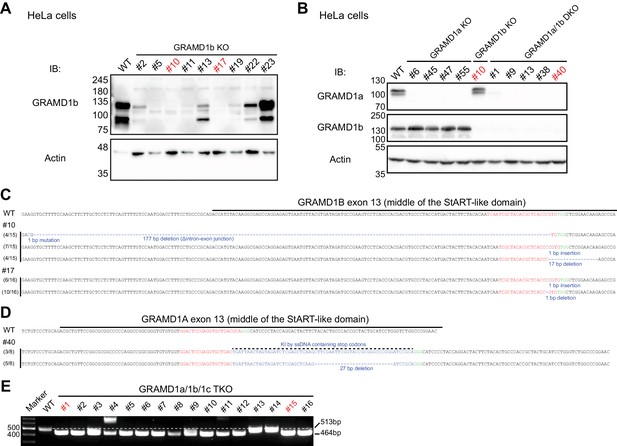
Generation of GRAMD1 triple knockout (TKO) HeLa cells.
(A) Lysates of control HeLa cells (WT) and of nine independently isolated GRAMD1b knockout (KO) cell lines were processed by SDS–PAGE and immunoblotting (IB) with anti-GRAMD1b and anti-actin antibodies. Lines #10 and #17 were further analyzed by nucleotide sequencing [see panel (C)]. (B) Lysates of control HeLa cells (WT), four independently isolated GRAMD1a KO cell lines, one GRAMD1b KO cell line, and five GRAMD1a/1b double knockout (DKO) cell lines were processed by SDS–PAGE and IB with anti-GRAMD1a, anti-GRAMD1b and anti-actin antibodies. Line #40 was further analyzed by nucleotide sequencing [see panel (D)]. (C) Nucleotide sequence analysis of the GRAMD1B gene of the GRAMD1b KO cell lines (#10 and #17). Guide RNA-targeting sites are highlighted in red. GRAMD1b KO #10 was used to generate GRAMD1a/1b DKO cell lines and subsequently to generate GRAMD1 triple knockout (TKO) cell lines. (D) Nucleotide sequence analysis of the GRAMD1A gene of the GRAMD1a/1b DKO cell line (#40). Guide RNA-targeting sites are highlighted in red. GRAMD1a/1b DKO #40 was used to generate GRAMD1 TKO cell lines. (E) Electrophoresis analysis of genomic DNA fragments of the GRAMD1C gene of the GRAMD1 TKO cell lines after genomic PCR. WT control cells show 513 bp bands, whereas TKO cells, such as #1 and #15, show smaller bands (464 bp).
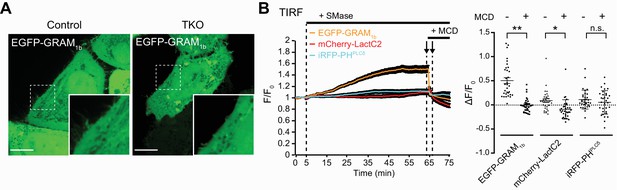
Recruitment of the EGFP-tagged GRAM domain of GRAMD1b (EGFP–GRAM1b) to the PM requires cholesterol.
(A) Confocal images of live wild-type (Control) and GRAMD1 TKO (TKO) HeLa cells expressing the EGFP-tagged GRAM domain of GRAMD1b (EGFP–GRAM1b) before sphingomyelinase (SMase) treatment. Insets show at higher magnification the regions indicated by white dashed boxes. Scale bars, 10 µm. (B) Left: time course of normalized fluorescent signal, as assessed by TIRF microscopy, from GRAMD1 TKO HeLa cells expressing EGFP–GRAM1b, a phosphatidylserine probe (mCherry–LactC2) and a PI(4,5)P2 probe (iRFP–PH-PLCδ). SMase treatment (100 mU/ml) and methyl-β-cyclodextrin (MCD) treatment (10 mM) are indicated. Right: values of ΔF/F0 corresponding to the 65 min and 67 min time points of the experiment as indicated by the arrows (mean ± SEM, n = 32 cells; data are pooled from two independent experiments; Tukey’s multiple comparisons test, **p<0.0001, *p=0.0225, n.s. denotes not significant).
-
Figure 4—figure supplement 2—source data 1
Dataset for Figure 4—figure supplement 2.
- https://cdn.elifesciences.org/articles/51401/elife-51401-fig4-figsupp2-data1-v2.xlsx
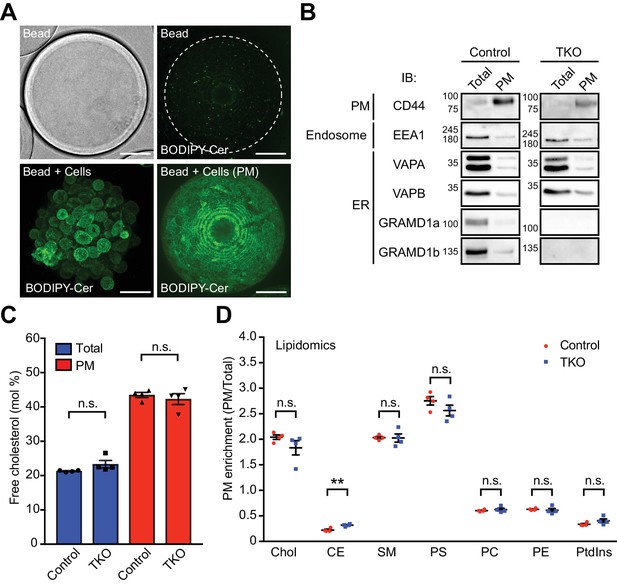
Isolation and characterization of the PM sheets of GRAMD1 triple knockout (TKO) HeLa cells.
(A) Confocal images of HeLa cells attached to a bead. Top: bead without cells. Bottom: bead coated with HeLa cells. Maximum projections of the serial Z-stack of confocal images are shown. The bead-attached PMs were stained with BODIPY TR Ceramide (BODIPY-Cer: green). Scale bars, 50 µm. (B) Bead-attached material [total lysate (Total) and PM sheets (PM)] was processed by SDS–PAGE and immunoblotting (IB) with the indicated antibodies. Note the enrichment of PM proteins and the depletion of ER and endosomal proteins in the PM fractions. (C, D) Quantitative analysis of lipids by mass spectrometry. (C) Comparisons of the PM lipid profiles of wild-type (Control) and GRAMD1 TKO (TKO) HeLa cells. PM enrichment plots the ratio of the indicated lipids present in the PM sheets over those present in the total lysate. (D) Comparisons of the mole% of cholesterol over total lipids for whole cell extracts (Total) and PM extracts (PM). Chol, cholesterol; CE, cholesterol esters; SM, sphingomyelin; PS, phosphatidylserine; PC, phosphatidylcholine; PE, phosphatidylethanolamine; PtdIns, phosphatidylinositol [mean ± SEM, n = 4 extracts (Control), n = 4 extracts (TKO) (four biologically independent samples for each condition), two-tailed unpaired Student’s t-test, **p=0.0036, n.s. denotes not significant].
-
Figure 4—figure supplement 3—source data 1
Dataset for Figure 4—figure supplement 3.
- https://cdn.elifesciences.org/articles/51401/elife-51401-fig4-figsupp3-data1-v2.xlsx
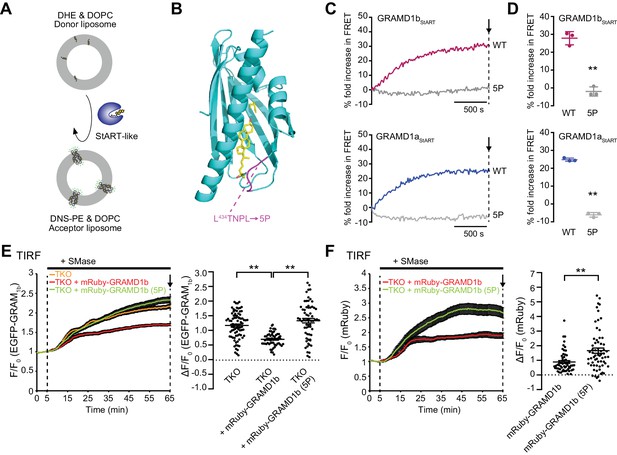
The cholesterol transporting property of the StART-like domain of GRAMD1s is critical for removal of an acutely expanded pool of accessible PM cholesterol.
(A) Schematic showing the design of the in vitro lipid transfer assay. Donor liposomes (10% DHE, 90% DOPC) and acceptor liposomes [2.5% Dansyl-PE (DNS-PE), 97.5% DOPC] were incubated with the purified StART-like domain of GRAMD1s (GRAMD1aStART, GRAMD1bStART, or GRAMD1cStART). Transfer of DHE from donor to acceptor liposomes, which results in an increase in fluorescence resonance energy transfer (FRET) between DHE and DNS-PE in acceptor liposomes, was monitored using a fluorometer (see Materials and methods). (B) Design of a mutant StART-like domain that is defective in lipid harboring. Ribbon diagram of the modeled GRAMD1bStART (see Materials and methods) with designed mutations (5P) in the Ω1 loop of GRAMD1b, which is predicted to open and close to capture sterol. (C, D) 5P mutations in the Ω1 loop of the StART-like domain impairs DHE transfer activity. (C) Time course of fold increase in FRET signals. WT GRAMD1bStART and GRAMD1bStART with 5P mutation (2 µM, top panel) and WT GRAMD1aStART and GRAMD1aStART with 5P mutation (0.5 µM, bottom panel) were individually added at time 0. (D) Values of fold increase in FRET signals of acceptor liposomes in the presence of the indicated proteins at the time point corresponding to the end of the experiments [as shown by arrows in (C)] (mean ± SEM, n = 3 independent experiments for all of the conditions; two-tailed unpaired Student’s t-test, GRAMD1bStART**p=0.0003, GRAMD1aStART**p<0.0001). (E, F) Left: time course of normalized (E) EGFP or (F) mRuby signal, as assessed by TIRF microscopy, from GRAMD1 TKO cells expressing EGFP–GRAM1b and mRuby-tagged constructs as indicated. SMase treatment (100 mU/ml) is indicated. Right: values of ΔF/F0 at the time point corresponding to the end of the experiment (as indicated by the arrows). [(E) mean ± SEM, n = 84 cells (TKO), n = 57 cells (TKO + mRuby–GRAMD1b), n = 64 cells (TKO + mRuby–GRAMD1b (5P)); Tukey’s multiple comparisons test, **p<0.0001; (F) mean ± SEM, n = 57 cells (TKO + mRuby–GRAMD1b), n = 64 cells (TKO + mRuby–GRAMD1b (5P)); two-tailed unpaired Student’s t-test, **p=0.0003; data are pooled from three or four independent experiments for each condition.]
-
Figure 5—source data 1
Dataset for Figure 5.
- https://cdn.elifesciences.org/articles/51401/elife-51401-fig5-data1-v2.xlsx
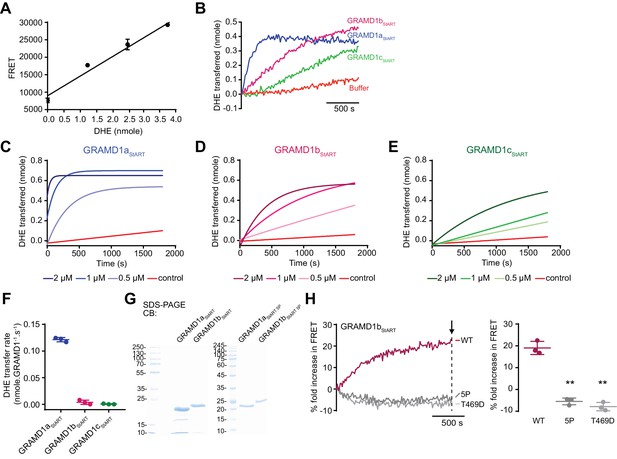
Characterization of the cholesterol transporting property of the StART-like domain of GRAMD1s.
(A) Calibration curve showing the linear relationship between FRET signals and the amount of DHE present in liposomes that contain 2.5 mole % of DNS-PE. This calibration curve was used to measure quantitatively the amount of DHE transferred from donor to acceptor liposomes over time (mean ± s.d., n = 3 for all different DHE concentrations). (B–E). Time course of DHE transfer from donor to acceptor liposomes with the StART-like domains of GRAMD1s as indicated. Purified StART-like domains of GRAMD1s (0.5 µM, 1 µM or 2 µM) were added at time 0. Addition of buffer only was used as a control. Data were fit to a one-phase association equation that was built in the GraphPad Prism 7. (F) Values of DHE transfer rate of GRAMD1aStART, GRAMD1bStART, and GRAMD1cStART, as estimated by the FRET-based lipid transfer assay (see Materials and methods). (G) SDS-PAGE analysis of purified StART-like domains from wild-type GRAMD1a and GRAMD1b [GRAMD1aStART, GRAMD1bStART] and those with 5P mutations [GRAMD1aStART (5P), GRAMD1bStART(5P)]. CB, Colloidal blue staining. (H) Left: time course of fold increase in FRET signals. Wild-type GRAMD1bStART (WT), GRAMD1bStART with 5P mutation (5P) and GRAMD1bStART with T469D mutation (T469D) were individually added (2 μM) at time 0. Right: values of fold increase in FRET signals in the presence of the indicated proteins , at a time point corresponding to the end of the experiment (as indicated by an arrow) are shown (mean ± SEM, n = 3 independent experiments for all of the conditions; Tukey’s multiple comparisons test, WT vs. 5P, **p<0.0001; WT vs. T469D, **p<0.0001).
-
Figure 5—figure supplement 1—source data 1
Dataset for Figure 5—figure supplement 1.
- https://cdn.elifesciences.org/articles/51401/elife-51401-fig5-figsupp1-data1-v2.xlsx
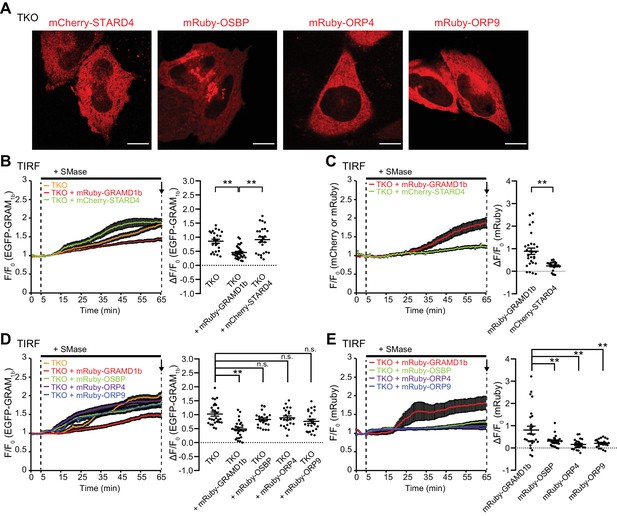
Overexpression of STARD4 and selected ORPs does not rescue the exaggerated accumulation of the accessible pool of PM cholesterol in GRAMD1 TKO cells upon sphingomyelinase treatment.
(A) Confocal images of live GRAMD1 TKO (TKO) HeLa cells expressing the indicated constructs. Scale bars, 10 µm (B, C) Left: time course of normalized (B) EGFP or (C) mCherry or mRuby signal, as assessed by TIRF microscopy, from GRAMD1 TKO cells co-expressing EGFP-GRAM1b and either mCherry-STARD4 or mRuby-GRAMD1b, respectively. Sphingomyelinase (SMase) treatment (100 mU/ml) is indicated. Right: values of ΔF/F0 corresponding at a time point to the end of the experiment (as indicated by the arrows) [(B) mean ± SEM, n = 24 cells (TKO), n = 28 cells (TKO + mRuby–GRAMD1b), n = 25 cells (TKO + mCherry–STARD4); Tukey’s multiple comparisons test, TKO vs. TKO + mRuby–GRAMD1b **p=0.0002; TKO + mRuby–GRAMD1b vs. mCherry–STARD4, **p<0.0001; (C) mean ± SEM, n = 28 cells (TKO + mRuby–GRAMD1b), n = 25 cells (TKO + mCherry–STARD4); two-tailed unpaired Student’s t-test, **p<0.0001; all data are pooled from two independent experiments]. (D, E) Left: time course of normalized (D) EGFP or (E) mRuby signal, as assessed by TIRF microscopy, from GRAMD1 TKO cells expressing EGFP–GRAM1b and mRuby-tagged constructs as indicated. SMase treatment (100 mU/ml) is indicated. Right: values of ΔF/F0 at a time point corresponding to the end of the experiment (as indicated by the arrows) [(D) mean ± SEM, n = 27 cells (TKO), n = 27 cells (TKO + mRuby–GRAMD1b), n = 24 cells (TKO + mRuby–OSBP), n = 22 cells (TKO + mRuby–ORP4), n = 23 cells (TKO + mRuby–ORP9); Tukey’s multiple comparisons test, **p<0.0001, n.s. denotes not significant; (E) mean ± SEM, n = 27 cells (TKO + mRuby–GRAMD1b), n = 24 cells (TKO + mRuby–OSBP), n = 22 cells (TKO + mRuby–ORP4), n = 23 cells (TKO + mRuby–ORP9); Dunnett’s multiple comparisons test, TKO + mRuby–GRAMD1b vs. TKO + mRuby–OSBP, **p=0.0031; TKO + mRuby–GRAMD1b vs. TKO + mRuby–ORP4, **p=0.0001; TKO + mRuby–GRAMD1b vs. TKO + mRuby–ORP9, **p=0.0004; all data are pooled from two independent experiments].
-
Figure 5—figure supplement 2—source data 1
Dataset for Figure 5—figure supplement 2.
- https://cdn.elifesciences.org/articles/51401/elife-51401-fig5-figsupp2-data1-v2.xlsx
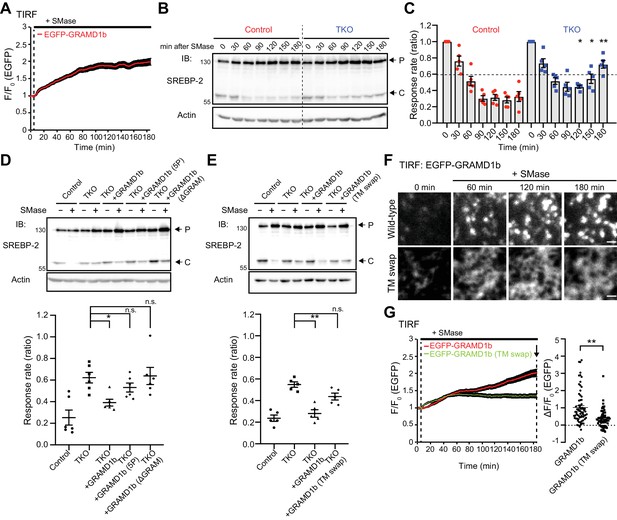
GRAMD1s-mediated PM to ER cholesterol transport plays a role in the suppression of SREBP-2 cleavage upon sphingomyelinase treatment.
(A) Time course of normalized EGFP signal, as assessed by TIRF microscopy, from HeLa cells expressing EGFP–GRAMD1b in response to sphingomyelinase (SMase) treatment (100 mU/ml). Note the sustained recruitment of EGFP–GRAMD1b to the PM even after 3 hr of the SMase treatment (mean ± SEM, n = 74 cells; data are pooled from four independent experiments). (B) Wild-type (Control) and GRAMD1 TKO (TKO) HeLa cells were cultured in the medium supplemented with 10% lipoprotein-deficient serum (LPDS) and mevastatin (50 µM) for 16 hr and then treated with SMase (100 mU/ml) for the indicated time at 37°C. Lysates of the cells were processed for SDS-PAGE and immunoblotting (IB) with anti-SREBP-2 and anti-Actin antibodies. Arrows indicate precursor (P) and cleaved (C) forms of SREBP-2, respectively. (C) Quantification of the response rate of the suppression of SREBP-2 cleavage upon SMase treatment from the experiment shown in (B). For each time point, the ratio of the band intensity of the cleaved SREBP-2 over the total band intensity of cleaved and precursor forms of SREBP-2 was normalized by the ratio obtained from time 0, and plotted as response rate. Note that the suppression of SREBP-2 cleavage is attenuated in GRAMD1 TKO cells [mean ± SEM, n = 5 lysates (independent experiments) for each time point; multiple comparisons were made using the Holm-Sidak method, *p=0.0461 (120 min), *p=0.0238 (150 min), **p=0.0052 (180 min)]. (D, E) Wild-type (Control) and GRAMD1 TKO (TKO) HeLa cells, transfected with the EGFP-tagged GRAMD1s constructs as indicated, were cultured in the medium supplemented with 10% lipoprotein-deficient serum (LPDS) and mevastatin (50 µM) for 16 hr and then treated with SMase (100 mU/ml) for 3 hr at 37°C. Top: lysates of the cells were processed for SDS-PAGE and IB with anti-SREBP-2 and anti-actin antibodies. Arrows indicate precursor (P) and cleaved (C) forms of SREBP-2. Bottom: the response rate was calculated as in panel (C) except that the ratio obtained from the cells with SMase treatment was normalized by the ratio obtained from the cells without SMase treatment for each condition. Note the rescue by expression of EGFP–GRAMD1b but not by mutant versions of EGFP–GRAMD1b (5P, ΔGRAM, TM swap) [(D) mean ± SEM, n = 6 lysates (independent experiments) for each condition, Dunnett’s multiple comparisons test, *p=0.0162; (E): mean ± SEM, n = 5 lysates (independent experiments) for each condition, Dunnett’s multiple comparisons test, **p<0.0001; n.s. denotes not significant]. (F) Representative TIRF images of live HeLa cells expressing EGFP–GRAMD1b (Wild-type) and EGFP–GRAMD1b TM swap (TM swap) treated as described for panel (G). Note the differences in how these proteins are recruited to the PM. Wild-type GRAMD1b accumulated progressively at ER–PM contacts, forming patches by the end of the 3 hr imaging period, whereas the GRAMD1b TM swap mutant remained diffuse on the tubular ER, even at the end of the 3 hr imaging period. Scale bars, 1 µm. (G) Left: time course of normalized EGFP signal, as assessed by TIRF microscopy, from HeLa cells expressing EGFP–GRAMD1b or EGFP–GRAMD1b TM swap in response to SMase treatment (100 mU/ml). Note the reduced recruitment of EGFP–GRAMD1b TM swap to the PM compared to EGFP–GRAMD1b after 3 hr of the SMase treatment. Right: values of ΔF/F0 at a time point corresponding to the end of the experiment (as indicated by the arrow) [mean ± SEM, n = 72 cells (GRAMD1b), n = 69 cells (GRAMD1b TM swap); data are pooled from three independent experiments for each condition; two-tailed unpaired Student’s t-test, **p<0.0001].
-
Figure 6—source data 1
Dataset for Figure 6.
- https://cdn.elifesciences.org/articles/51401/elife-51401-fig6-data1-v2.xlsx
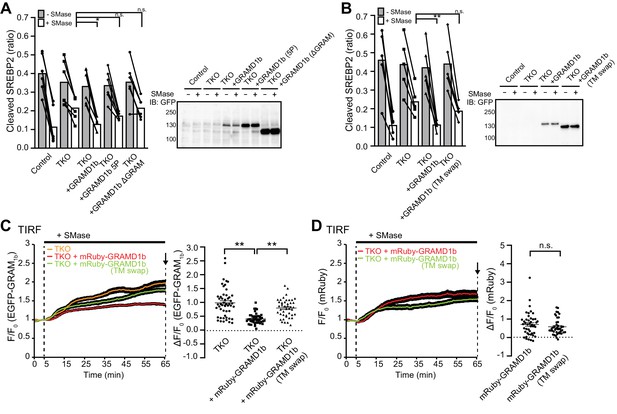
Transport of cholesterol from the PM to the ER by GRAMD1 proteins requires their StART-like and GRAM domains as well as their complex formation.
(A, B) Left: quantification of the ratio of the band intensity of the cleaved SREBP-2 over the total band intensity of cleaved and precursor forms of SREBP-2 from the experiments shown in Figure 6D,E [(A) mean ± SEM, n = 6 lysates (independent experiments) for each condition, Dunnett’s multiple comparisons test, *p=0.0111; (B) mean ± SEM, n = 5 lysates (independent experiments) for each condition, Dunnett’s multiple comparisons test, **p=0.0074; n.s. denotes not significant]. Right: lysates of wild-type control (Control) and GRAMD1 TKO (TKO) HeLa cells expressing the indicated EGFP-tagged constructs were processed by SDS–PAGE and immunoblotting (IB) with anti-GFP antibodies. (C, D) Left: time course of normalized (C) EGFP or (D) mRuby signal, as assessed by TIRF microscopy, from GRAMD1 TKO (TKO) HeLa cells expressing EGFP–GRAM1b and mRuby-tagged constructs as indicated. Sphingomyelinase (SMase) treatment (100 mU/ml) is indicated. Right: values of ΔF/F0 corresponding to the end of the experiment as indicated by the arrow [(C) mean ± SEM, n = 50 cells (TKO), n = 45 cells (TKO + mRuby–GRAMD1b), n = 43 cells (TKO + mRuby–GRAMD1b [TM swap]); data are pooled from three independent experiments for each condition; Tukey’s multiple comparisons test; TKO vs. TKO + GRAMD1b and TKO + GRAMD1b vs. TKO + GRAMD1b (TM swap), **p<0.0001; (D) mean ± SEM, n = 45 cells (TKO + mRuby–GRAMD1b), n = 43 cells (TKO + mRuby–GRAMD1b [TM swap]); data are pooled from three independent experiments for each condition; n.s. denotes not significant].
-
Figure 6—figure supplement 1—source data 1
Dataset for Figure 6—figure supplement 1.
- https://cdn.elifesciences.org/articles/51401/elife-51401-fig6-figsupp1-data1-v2.xlsx
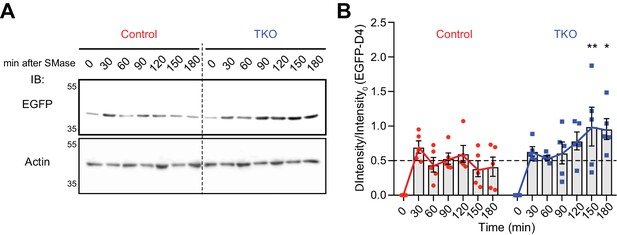
Deletion of GRAMD1s results in sustained D4 binding to the PM upon sphingomyelinase treatment.
(A) Wild-type (Control) and GRAMD1 TKO (TKO) HeLa cells were cultured in the medium supplemented with 10% lipoprotein-deficient serum (LPDS) and mevastatin (50 µM) for 16 hr, and then treated with sphingomyelinase (SMase) (100 mU/ml) for the indicated time at 37°C. Cells were then washed and incubated with phosphate-buffered saline (PBS) containing recombinant EGFP–D4 proteins (10 µg/ml) for 15 min at room temperature. After washing with PBS, lysates of the cells (10 µg protein for each lane) were processed for SDS-PAGE and immunoblotting (IB) with anti-GFP and anti-actin antibodies. (B) Quantification of the binding of EGFP–D4 proteins to the PM upon SMase treatment from the experiment shown in panel (A). For each time point, the ratio of the band intensity of EGFP–D4 over the band intensity of EGFP–D4 at time 0 was calculated, and ΔIntensity/Intensity0 was plotted to show the changes in D4 accessible PM cholesterol over time. Note the sustained increase in the binding of EGFP–D4 in GRAMD1 TKO cells compared to that in control cells [mean ± SEM, n = 5 lysates (independent experiments) for each time point, multiple comparisons with the Holm-Sidak method, **p=0.0082 (150 min), *p=0.0240 (180 min)].
-
Figure 6—figure supplement 2—source data 1
Dataset for Figure 6—figure supplement 2.
- https://cdn.elifesciences.org/articles/51401/elife-51401-fig6-figsupp2-data1-v2.xlsx
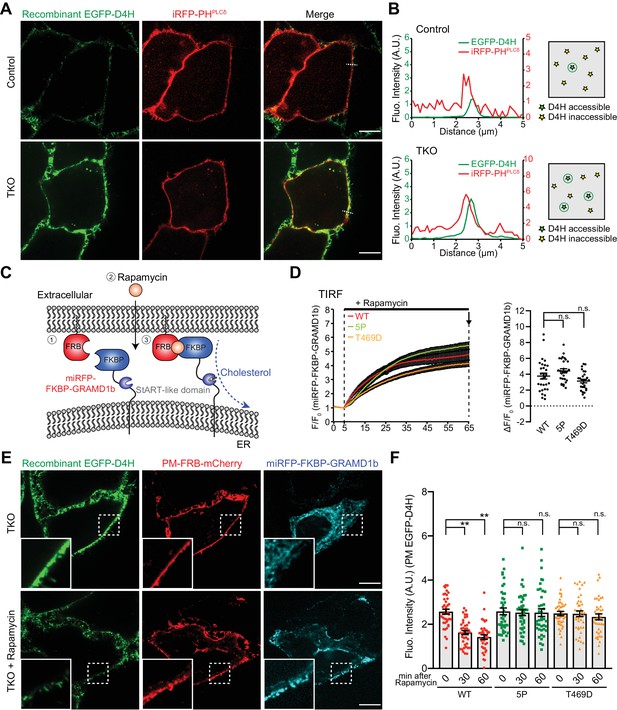
Acute recruitment of GRAMD1b to ER–PM contacts facilitates removal of the expanded pool of accessible PM cholesterol in GRAMD1 triple knockout (TKO) cells.
(A) Left: confocal images of live wild-type (Control) and GRAMD1 TKO (TKO) HeLa cells expressing a PI(4,5)P2 probe/PM marker (iRFP-PH-PLCδ) that are stained with recombinant EGFP–D4H proteins (15 µg/ml) for 15 min at room temperature. Scale bars, 10 µm. Note the increased accumulation of D4H-accessible PM cholesterol in GRAMD1 TKO cells compared to control cells, as detected by the presence of strong EGFP–D4H signals at the PM visualized by iRFP-PH-PLCδ. Scale bars, 10 µm. (B) Left: line scan analysis of the regions indicated by white dotted lines in the images shown in panel (A), showing the increase of EGFP–D4H signals at the PM (near the peak of iRFP-PH-PLCδ signals). Right: schematics showing the D4H-accessible pool of cholesterol on the outer leaflet of the PM (view from extracellular side) in wild-type (Control) and GRAMD1 TKO (TKO) HeLa cells. Green stars indicate D4H-accessible cholesterol, whereas yellow stars indicate D4H inaccessible cholesterol. (C) Schematic representation of the rapamycin-induced GRAMD1b PM recruitment strategy. GRAMD1b was rapidly recruited to the PM by rapamycin-induced dimerization of FRB and FKBP. A version of GRAMD1b with its N-terminal region, including the GRAM domain, replaced by a miRFP-tagged FKBP module (miRFP–FKBP–GRAMD1b) was expressed in GRAMD1 TKO cells together with an mCherry-tagged FRB module that is targeted to the PM (PM-FRB–mCherry). (D) Left: time course of normalized miRFP signal in response to rapamycin, as assessed by TIRF microscopy of GRAMD1 TKO cells expressing the indicated miRFP–FKBP–GRAMD1b constructs and PM-FRB–mCherry [wild-type (WT) and mutant versions with a StART-like domain that lacks cholesterol transport activity (5P or T469D)]. Rapamycin addition (200 nM) is indicated [mean ± SEM, n = 29 cells (WT), n = 29 cells (5P), n = 27 cells (T469D); all data are pooled from two independent experiments]. Right: values of ΔF/F0 at a time point corresponding to the end of the experiments (as shown by arrows). Dunnet’s multiple comparisons test, n.s. denotes not significant. (E) Confocal images of GRAMD1 TKO (TKO) HeLa cells expressing miRFP–FKBP–GRAMD1b and PM-FRB–mCherry with or without rapamycin (200 nM) treatment for 60 min at 37°C and then stained with recombinant EGFP–D4H proteins (15 µg/ml) for 15 min at room temperature. Insets show at higher magnification the regions indicated by white dashed boxes. Scale bars, 10 µm. (F) Values of EGFP–D4H signals at the PM after background subtraction, as assessed by confocal microscopy and line scan analysis, from GRAMD1 TKO HeLa cells expressing the indicated miRFP–FKBP–GRAMD1b constructs and PM-FRB–mCherry, which were stained with recombinant EGFP–D4H protein after rapamycin addition (200 nM) for either 30 min or 60 min, as shown in panel (E). Peak EGFP–D4H signals around the PM marked by peak PM-FRB–mCherry signals were quantified (see Materials and methods) [mean ± SEM, n = 40 cells for each condition; all data are pooled from two independent experiments; Tukey’s multiple comparisons test, **p<0.0001, n.s. denotes not significant].
-
Figure 7—source data 1
Dataset for Figure 7.
- https://cdn.elifesciences.org/articles/51401/elife-51401-fig7-data1-v2.xlsx
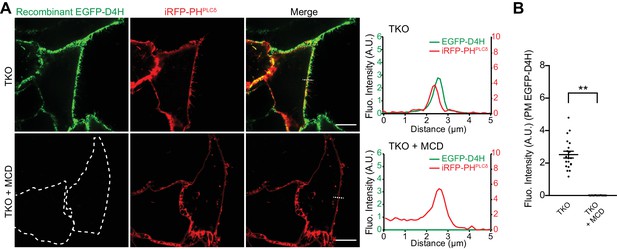
Increased D4H binding to the PM of GRAMD1 triple knockout (TKO) cells is dependent on the presence of cholesterol.
(A) Left: confocal images of live GRAMD1 TKO HeLa cells expressing a PI(4,5)P2 probe/PM marker (iRFP-PH-PLCδ) with or without methyl-β-cyclodextrin (MCD) treatment (10 mM for 30 min at 37°C), which were stained with recombinant EGFP–D4H proteins (15 µg/ml) for 15 min at room temperature. Scale bars, 10 µm. Right: line scan analysis of the regions indicated by white dotted lines in the images shown in confocal images. Note the loss of EGFP–D4H signals from the PM after MCD treatment. (B) Values of EGFP–D4H signals at the PM after background subtraction, as assessed by confocal microscopy and line scan analysis from GRAMD1 TKO HeLa cells that are stained with recombinant EGFP–D4H proteins as shown in panel (A) [mean ± SEM, n = 20 cells for each condition; data are pooled from two independent experiments; two-tailed unpaired Student’s t-test, **p<0.0001.].
-
Figure 7—figure supplement 1—source data 1
Dataset for Figure 7—figure supplement 1.
- https://cdn.elifesciences.org/articles/51401/elife-51401-fig7-figsupp1-data1-v2.xlsx
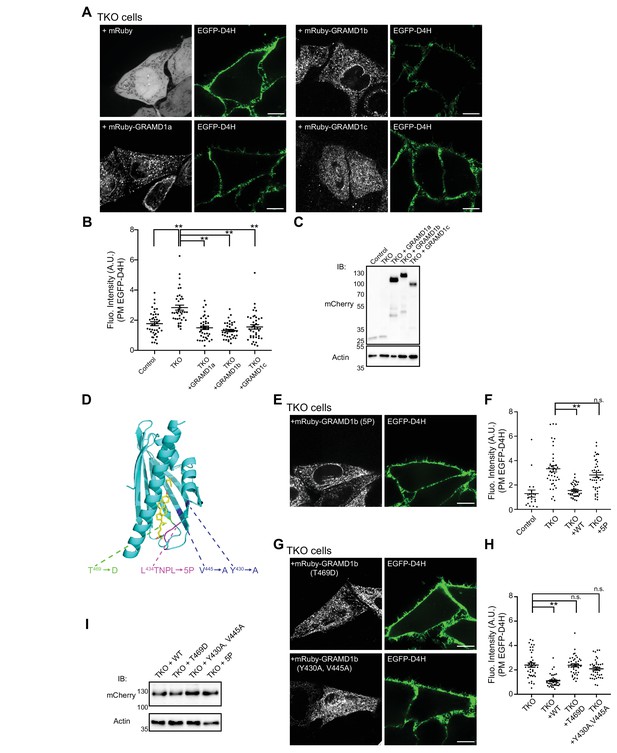
The cholesterol-transporting property of the StART-like domain is essential for removal of the expanded pool of D4H-accessible PM cholesterol in GRAMD1 knockout cells.
(A, E, G) Confocal images of live GRAMD1 TKO HeLa cells expressing either mRuby control or mRuby-tagged GRAMD1 constructs as indicated, together with a PI(4,5)P2 probe/PM marker (iRFP-PH-PLCδ), that are stained with recombinant EGFP–D4H proteins (15 µg/ml) for 15 min at room temperature. Note that re-expression of wild-type mRuby–GRAMD1s but not the mutant versions of mRuby–GRAMD1b in TKO cells reduced EGFP–D4H accessibility on the PM of TKO cells, rescuing the phenotype [see panels (B, F, H) for quantification]. Scale bars, 10 µm. (D) Design of the mutant StART-like domain that is defective in lipid harboring. Ribbon diagram of the modeled GRAMD1bStART (see Materials and methods) with designed mutations in the sterol-binding pocket (T469D, V445A and Y430A) and in the Ω1 loop of GRAMD1b that is predicted to open and close to capture sterol (5P). (B, F, H) Values of EGFP–D4H signals at the PM after background subtraction, as assessed by confocal microscopy and line scan analysis, from GRAMD1 TKO HeLa cells that are stained with recombinant EGFP–D4H protein as shown in panels (A, E, G) Peak EGFP–D4H signals around the PM, which are marked by peak iRFP-PH-PLCδ signals, were quantified (see Materials and methods) [mean ± SEM; (B) n = 40 cells for all the conditions; Tukey’s multiple comparisons test, **p<0.0001; (F) n = 40 cells for each condition except n = 20 for control; Tukey’s multiple comparisons test, **p<0.0001; n.s. denotes not significant; (H) n = 40 cells for all the conditions; Tukey’s multiple comparisons test, **p<0.0001; n.s. denotes not significant; 'Control' denotes wild-type HeLa cells; all data are pooled from two independent experiments]. (C, I) Lysates of wild-type (Control) and GRAMD1 TKO (TKO) HeLa cells expressing the indicated mRuby-tagged constructs were processed by SDS–PAGE and immunoblotting (IB) with anti-mCherry and anti-actin antibodies as indicated. Note that our anti-mCherry antibodies are able to detect mRuby-tagged proteins.
-
Figure 7—figure supplement 2—source data 1
Dataset for Figure 7—figure supplement 2.
- https://cdn.elifesciences.org/articles/51401/elife-51401-fig7-figsupp2-data1-v2.xlsx
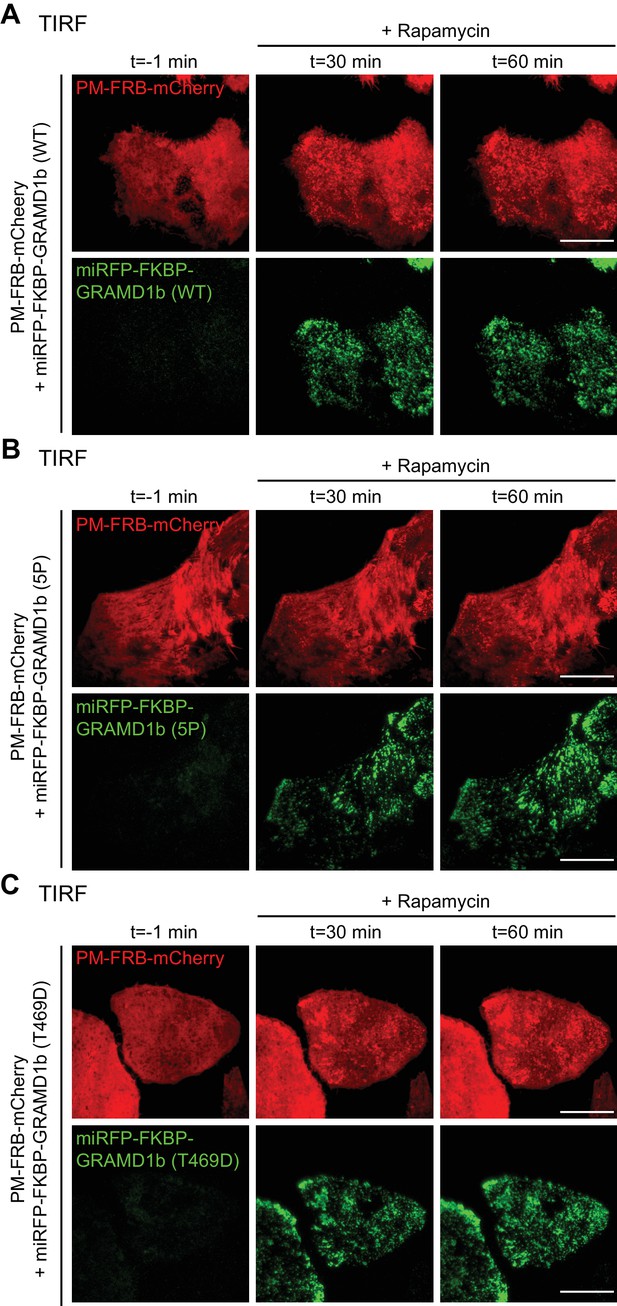
Rapamycin-induced acute recruitment of FKBP-tagged GRAMD1b to the PM in GRAMD1 triple knockout (TKO) cells.
(A–C). Snapshots of the cortical regions of the GRAMD1 TKO HeLa cells expressing PM-FRB–mCherry and the indicated miRFP-FKBP–GRAMD1b constructs, imaged under TIRF microscopy, at different times. Images were taken every 20 s. Rapamycin (200 nM) addition is indicated. Scale bars, 20 µm.
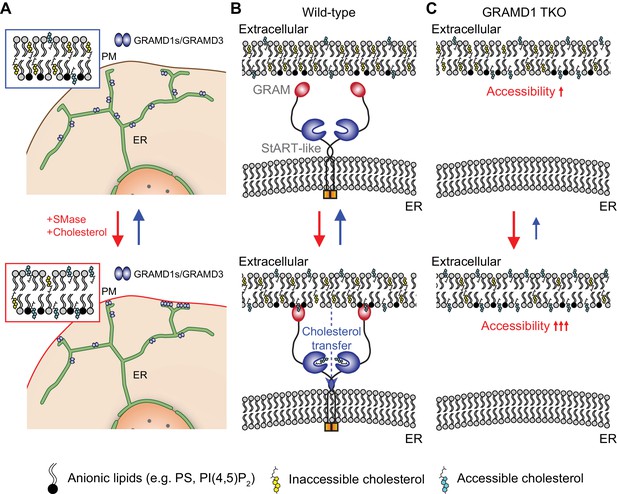
GRAMD1s facilitate the transport of the accessible pool of cholesterol from the PM to the ER, thereby contributing to PM cholesterol homeostasis.
(A) Distinct pools of cholesterol co-exist in PM bilayers at steady state: a major pool is inaccessible (i.e. sequestered or chemically inactive) and a smaller pool is accessible (i.e. unsequestered or chemically active). Sequestration is in part mediated by sphingomyelin. Compositions of phospholipids also influence the overall accessibility of PM cholesterol. Top: at rest, GRAMD1 complexes localize on the tubular ER network with little enrichment at ER–PM contact sites. Bottom: a transient expansion of the accessible pool of PM cholesterol (e.g. hydrolysis of sphingomyelin, cholesterol loading to the PM) induces the acute recruitment of GRAMD1 complexes to ER–PM contacts once the levels of accessible PM cholesterol exceed a certain threshold. The red arrow denotes the expansion of the accessible pool; the blue arrow denotes transport of the newly expanded pool of accessible PM cholesterol to the ER by GRAMD1s and other, yet to be identified, intracellular cholesterol transport systems. (B) Top: in wild-type cells, the GRAM domain of GRAMD1s only weakly interacts with the PM at rest due to sequestration of the majority of cholesterol in this bilayer. Bottom: upon reaching a certain threshold of the accessibility (i.e. a transient expansion in the accessible pool of PM cholesterol), the interaction of the GRAM domain with the PM is enhanced, and GRAMD1s are recruited to ER–PM contacts. Supporting this model, the binding of the purified GRAM domains of GRAMD1a and GRAMD1b to liposomes containing phosphatidylserine, a major anionic lipid in the PM, is dramatically enhanced by the presence of unsequestered/accessible cholesterol in a dose-dependent manner with a switch-like response (Figure 3C–F, and Figure 3—figure supplement 2B). Upon recruitment of GRAMD1s to ER–PM contacts, the StART-like domain initiates the extraction of accessible PM cholesterol and facilitates its transport to the ER, contributing to the suppression of SREBP-2 cleavage and to PM cholesterol homeostasis. (C) Top: in GRAMD1 TKO cells at rest, D4H binding to the PM increases, indicating chronic expansion of the accessible pool of PM cholesterol due to the absence of GRAMD1s-mediated transport of accessible cholesterol from the PM to the ER (Figure 7A,B). Bottom: upon a transient expansion of the accessible pool of PM cholesterol, GRAMD1 TKO cells show exaggerated accumulation of the accessible pool of PM cholesterol due to the absence of GRAMD1s-mediated transport of accessible PM cholesterol to the ER. Consistent with this model, sphingomyelinase treatment leads to enhanced recruitment of the GRAM domain of GRAMD1b, a novel biosensor for detecting acute expansion of the accessible pool of PM cholesterol that we identified in this study, to the PM in GRAMD1 TKO cells (Figure 4D,E). Reduced transport of accessible cholesterol from the PM to the ER in GRAMD1 TKO cells also results in less efficient suppression of SREBP-2 cleavage and continuous increase in the binding of recombinant D4 proteins to the PM (Figure 6B,C and Figure 6—figure supplement 2A,B). Other parallel cholesterol transport and regulatory systems, yet to be identified, may operate to maintain the total levels of PM cholesterol in the absence of GRAMD1s.
Videos
GRAMD1b is recruited to ER–PM contacts upon cholesterol loading.
HeLa cells expressing EGFP–GRAMD1b were imaged under TIRF microscopy. Images were taken every 20 s, and 200 μM cholesterol/MCD was added at the 5 min time point. Image size, 66.1 µm x 66.1 µm.
GRAMD1b ΔGRAM is not recruited to ER–PM contacts upon cholesterol loading.
HeLa cells expressing EGFP–GRAMD1b ΔGRAM were imaged under TIRF microscopy. Images were taken every 20 s, and 200 μM cholesterol/MCD was added at the 5 min time point. Image size, 66.1 µm x 66.1 µm.
Comparison of the recruitment to the PM of a wild-type GRAMD1b and of a mutant version of GRAMD1b that is defective in complex formation upon sphingomyelinase treatment.
HeLa cells expressing (left) EGFP–GRAMD1b or (right) EGFP–GRAMD1b TM swap were imaged under TIRF microscopy. Images were taken every 20 s, and 100 mU/ml of sphingomyelinase (SMase) was added at the 10 min time point. Image size, 66.1 µm x 66.1 µm.
Rapamycin-induced acute recruitment of GRAMD1b to the PM in GRAMD1 triple knockout (TKO) cells.
GRAMD1 TKO HeLa cells expressing PM-FRB–mCherry and miRFP-FKBP–GRAMD1b (WT) were imaged under TIRF microscopy. Images were taken every 20 s, and 200 nM rapamycin was added at the 5 min time point. Note the rapamycin-induced recruitment of miRFP-FKBP–GRAMD1b to the PM. Image size, 66.1 µm x 66.1 µm.
Rapamycin-induced acute recruitment of a mutant version of GRAMD1b (5P) to the PM in GRAMD1 triple knockout (TKO) cells.
GRAMD1 TKO HeLa cells expressing PM-FRB–mCherry and miRFP-FKBP–GRAMD1b (5P) were imaged under TIRF microscopy. Images were taken every 20 s, and 200 nM rapamycin was added at the 5 min time point. Note the rapamycin-induced recruitment of miRFP-FKBP–GRAMD1b (5P) to the PM. Image size, 66.1 µm x 66.1 µm.
Additional files
-
Supplementary file 1
Key resources table.
- https://cdn.elifesciences.org/articles/51401/elife-51401-supp1-v2.docx
-
Supplementary file 2
Table 1.
A list of sequence-based reagents. DNA sequences for oligos and primers used in this study are described.
- https://cdn.elifesciences.org/articles/51401/elife-51401-supp2-v2.docx
-
Supplementary file 3
Table 2.
Lipid compositions of liposomes used for lipid transfer assays. Moles% of lipids used for the acceptor and donor liposomes in FRET-based lipid transfer experiments are described.
- https://cdn.elifesciences.org/articles/51401/elife-51401-supp3-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/51401/elife-51401-transrepform-v2.docx





